Back to Journals » Cancer Management and Research » Volume 12
Long Noncoding RNA LINC00173 Promotes the Malignancy of Melanoma by Promoting the Expression of IRS4 Through Competitive Binding to microRNA-493
Authors Yang F, Lei P, Zeng W, Gao J, Wu N
Received 27 December 2019
Accepted for publication 31 March 2020
Published 5 May 2020 Volume 2020:12 Pages 3131—3144
DOI https://doi.org/10.2147/CMAR.S243869
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Eileen O'Reilly
This paper has been retracted
Fan Yang, 1 Pengzhen Lei, 2 Weihui Zeng, 3 Jianwu Gao, 1 Na Wu 1
1Department of Dermatology, Shaanxi Provincial People’s Hospital, Xi’an, Shaanxi 710068, People’s Republic of China; 2Department of Orthopedics, Shaanxi Provincial People’s Hospital, Xi’an, Shaanxi 710068, People’s Republic of China; 3Department of Dermatology, The Second Affiliated Hospital, School of Medicine, Xi’an Jiaotong University, Xi’an, Shaanxi 710048, People’s Republic of China
Correspondence: Fan Yang
Department of Dermatology, Shaanxi Provincial People’s Hospital, No. 256 Youyi West Road, Xi’an, Shaanxi 710068, People’s Republic of China
Email [email protected]
Purpose: Long intergenic non-protein-coding RNA 173 (LINC00173) plays crucial roles in lung cancer. However, the expression and biological functions of LINC00173 in melanoma have not yet been investigated. In this study, we aimed to characterize the involvement of LINC00173 in melanoma and elucidate its mechanisms of action.
Materials and Methods: Reverse-transcription quantitative PCR was performed to measure LINC00173 expression in melanoma. A CCK-8 assay, flow cytometry, and migration and invasion assays were applied to examine melanoma cell proliferation, apoptosis, migration, and invasion, respectively. A xenograft tumor experiment was performed to determine the tumorous growth of melanoma cells in vivo.
Results: We found that LINC00173 was upregulated in melanoma tissues and cell lines. High LINC00173 expression was closely associated with TNM stage, lymph node metastasis, and shorter overall survival of patients with melanoma. Functional assays revealed that LINC00173 downregulation inhibited melanoma cell proliferation, migration, and invasion and induced apoptosis, suggesting that LINC00173 acts as an oncogenic RNA. LINC00173 knockdown retarded the tumorous growth of melanoma cells in vivo. Mechanistically, LINC00173 increased insulin receptor substrate 4 (IRS4) expression by sponging microRNA-493 (miR-493), thereby acting as a competing endogenous RNA. The effects of LINC00173 knockdown on the malignant phenotype of melanoma cells were reversed by overexpression of IRS4 or knockdown of miR-493.
Conclusion: The LINC00173–miR-493–IRS4 pathway regulates melanoma characteristics by increasing the expression of IRS4 via competitive binding of LINC00173 to miR-493, suggesting that this pathway is a potential target for the diagnosis, prognosis, and/or treatment of melanoma.
Keywords: LINC00173, miR-493, melanoma, therapeutic target
Introduction
Melanoma, resulting from malignant transformation of melanocytes located at the basement of the epidermis, is the most frequent and aggressive type of skin cancer.1,2 The global morbidity rate of melanoma has been increasing gradually in recent years.3 It is estimated that there would be over 150,000 novel cases and 50,000 deaths due to melanoma yearly worldwide.4,5 Even though some sophisticated therapeutic regimens have evolved from surgery, chemotherapy, targeted therapy, and immunotherapy,6–10 the prognosis of patients with melanoma has not improved significantly.11 The 5-year survival rate of these patients is lower than 15%, which is a consequence of the high metastatic potential and uncontrollable growth.12 Melanoma cases represent only 4% of patients with skin tumors but are responsible for ~74% of skin tumor–related deaths.13 UV damage has been validated as a major risk factor of melanoma; genetic and epigenetic changes also exert important actions during melanoma initiation and progression.14 Nonetheless, the exact events underlying the pathogenesis of melanoma remain largely unknown. Therefore, comprehensive elucidation of melanoma pathogenesis is urgently needed to identify novel and promising therapeutic techniques.
Long noncoding RNAs (lncRNAs) are a group of endogenous RNA transcripts (with a length of >200 bp) that cannot be translated into proteins.15 LncRNAs are implicated in nearly all physiological and pathological activities, especially in carcinogenesis and cancer progression.16 The actions of lncRNAs are mediated by diverse molecular mechanisms, such as transcriptional modulation, chromatin remodeling, histone modification, regulation of mRNA splicing, and the competing endogenous RNA (ceRNA) mechanism.17,18 Existing studies have identified a close relation between lncRNAs and human cancers.19–21 Much evidence suggests that numerous lncRNAs are differentially expressed in melanoma, and their dysregulation is involved in melanoma progression because lncRNAs can act as oncogenic factors or tumor suppressors.22–24 Accordingly, in-depth exploration of lncRNAs’ functions in the formation and progression of melanoma is essential for identifying new targets for the diagnosis and management of this fatal disease.
MicroRNAs (miRNAs), a family of single-stranded, highly conserved and noncoding RNAs, are identified as gene regulators by directly binding to the 3ʹ-untranslated region (3ʹ-UTRs) of their target mRNAs, and thereby resulting in mRNAs degradation and/or translation suppression.25,26 Interestingly, accumulating studies demonstrated the enrollment of miRNAs in the oncogenesis and progression of melanoma.27–29 MiRNAs may perform tumor-suppressing or tumor-promoting activities in melanoma, and are implicated in the regulation of multiple aggressive processes.30,31 Hence, studying the expression and roles of miRNAs in melanoma my highlight potential targets for managing melanoma.
LINC00173 is a crucial modulator of the malignancy of lung cancer.32,33 Nevertheless, the expression and biological roles of LINC00173 in melanoma have not yet been explored. Here, we attempted to analyze the expression of LINC00173 in melanoma and its clinical significance. Effects of LINC00173 on the malignancy characteristics of melanoma cells in vitro and in vivo were tested. The next step was investigation of the mechanism of LINC00173-mediated promotion of melanoma progression.
Materials and Methods
Clinical Tissue Sample Collection
This study was performed with the approval of the Research Ethics Committee of Shaanxi Provincial People’s Hospital. In addition, written informed consent forms were signed by all the patients who participated in this research. Melanoma tissue samples and adjacent normal tissues were obtained from 45 patients with melanoma undergoing a surgical procedure at the Shaanxi Provincial People’s Hospital. Patients who had received chemotherapy, radiotherapy, targeted therapy, or immunotherapy were excluded from this study. All tissue samples were immediately frozen and stored in liquid nitrogen.
Cell Culture
Human epidermal melanocytes (HEMs) were bought from ScienCell Research Laboratories, Inc. (San Diego, CA, USA) and grown in a melanocyte medium (ScienCell Research Laboratories, Inc.). Four human melanoma cell lines, ie, A375, A2058, SKMEL1, and HT144, were purchased from the American Type Culture Collection (Manassas, VA, USA). Dulbecco’s modified Eagle’s medium (DMEM; Gibco; Thermo Fisher Scientific, Inc., Waltham, MA, USA) containing 10% fetal bovine serum (FBS; Gibco; Thermo Fisher Scientific), 100 U/mL penicillin, and 100 µg/mL streptomycin was utilized for culturing melanoma cells. All cell lines were maintained at 37°C in a humidified atmosphere supplied with 5% CO2.
Cell Transfection
An miR-493 mimic, negative control (NC) miRNA mimic (miR-NC), miR-493 inhibitor, and NC inhibitor were acquired from GenePharma Technology (Shanghai, China). Small interfering RNAs (siRNA) specific to LINC00173 (si-LINC00173) and NC siRNA (si-NC) were synthesized by RiboBio (Guangzhou, China). IRS4-overexpressing plasmid pcDNA3.1-IRS4 was bought from Sangon Biotech (Shanghai, China). Cells were grown up to 60% confluence and transfected with the miRNA mimic (100 pmol), miRNA inhibitor (100 pmol), siRNA (100 pmol) or plasmid (4 µg) using Lipofectamine 2000® (Invitrogen; Thermo Fisher Scientific).
Reverse-Transcription Quantitative PCR (RT-qPCR)
The TRIzol reagent (Invitrogen; Thermo Fisher Scientific) was applied for total-RNA isolation. An absorbance ratio (A260/A280), which was determined using Nanodrop 2000 (Invitrogen; Thermo Fisher Scientific) was used to analyze the quality of the isolated total RNA. To quantitate the expression of miR-493, first-strand cDNAs were produced from the total RNA using the miScript Reverse Transcription Kit (Qiagen GmbH, Hilden, Germany). qPCR was performed using an miScript SYBR Green PCR Kit (Qiagen GmbH). The expression of miR-493 was normalized to that of U6 small nuclear RNA. To quantify IRS4 mRNA and LINC00173 expression, the total RNA was reverse-transcribed into cDNA with a PrimeScript RT Reagent Kit (Takara Bio, Dalian, China). The synthesized cDNA was analyzed by qPCR with the SYBR Premix Ex Taq™ Kit (Takara Bio). GAPDH was regarded as an endogenous control for IRS4 mRNA and LINC00173 normalization. All reactions were performed on a 7500 Sequence Detection System (Applied Biosystems; Thermo Fisher Scientific). All data were analyzed by the 2−ΔΔCq method.
The primers were designed as follows: miR-493, 5′-TGTGATTGGAATGGAAATTTAATTT-3′ (forward) and 5′-ACTATCCTACACTCCCCTACCCTAC-3′ (reverse); U6, 5′-CTCGCTTCGGCAGCACA-3′ (forward) and 5′- AACGCTTCACGAATTTGCGT-3′ (reverse); LINC00173, 5′-GGAATGTTGCGATCCTCTGG-3′ (forward) and 5′-CAGCCATGTCTCAGAGGTGA-3′ (reverse); IRS4, 5′- CCGACACCTCATTGCTCTTTTC-3′ (forward) and 5′-TTTCCTGCTCCGACTCGTTCTC-3′ (reverse); and GAPDH, 5′-CAGCCTCAAGATCATCAGCA-3′ (forward) and 5′- TGTGGTCATGAGTCCTTCCA-3′ (reverse).
Cell Counting Kit-8 (CCK-8) Assay
Suspensions of transfected cells were diluted to a certain concentration and then seeded in 96-well plates at an initial density of 2000 cells/well. The cells were incubated at 37°C in a humidified atmosphere containing 5% CO2 for four periods after inoculation: 0, 24, 48, or 72 h. A total of 10 µL of the CCK-8 solution (Beyotime Institute of Biotechnology, Shanghai, China) was added into each well at each time point. Subsequent to additional 2 h incubation, the absorbance value of every well at a wavelength of 450 nm was measured on a microplate reader.
Apoptosis and Cell Cycle Assessment via Flow-Cytometric Analysis
Apoptotic cells were quantified using an Annexin V-Fluorescein Isothiocyanate (FITC) Apoptosis Detection Kit (BioLegend, San Diego, CA, USA). After 48 h culture, transfected cells were collected via treatment with trypsin without EDTA, and precooled phosphate-buffered saline was utilized to wash the transfected cells thrice. The transfected cells were centrifuged and resuspended in 100 μL of flow cytometry binding buffer, after which the cells were labeled with 5 μL of Annexin-V-FITC and 5 μL of a propidium iodide (PI) solution at room temperature in the dark for 15 min. The apoptotic cells were quantitated on a flow cytometer (FACScan™, BD Biosciences, Franklin Lakes, NJ, USA).
Transfected cells were fixed in 70% ethanol at 4°C overnight, followed by centrifugation at 4°C for 5 min. The supernate was discarded and the transfected cells were probed with 50 µL RNase (100 µg/mL) at 37°C for 20 min. Following incubation at room temperature with 25 µL PI (Biolegend) diluted in 500 µL cell-staining buffer.Finally, a flow cytometry was utilized to analyze the cell cycle status.
Migration and Invasion Assays
The transfected cells were resuspended in FBS-free DMEM. For the invasion assay, 200 μL of a cell suspension containing 5 × 104 transfected cells was seeded on the upper insert of a 24-well Transwell plate (8 μm pore size; Corning Inc., Corning, NY, USA) that was precoated with Matrigel (BD Biosciences). The basolateral inserts were covered with 600 μL DMEM that was supplemented with 10% FBS functioning as a chemoattractant. After 24 h incubation, noninvasive cells (remaining on the top layer of the membrane) were gently wiped away with a cotton swab, and the invasive cells were fixed with 4% paraformaldehyde and stained with 0.1% crystal violet. After extensive washing and air drying, the invasive cells were counted under an inverted microscope (Olympus Corporation, Tokyo, Japan) at 200× magnification to evaluate the invasiveness of cancer cell lines. A migration assay was performed following the same experimental steps as in the invasion assay except that the membranes were not coated with Matrigel.
Xenograft Tumor Experiment and Terminal Deoxynucleotidyl Transferase dUTP Nick End Labeling (TUNEL) Assay
Short hairpin RNA (shRNA) specific to LINC00173 (sh-LINC00173) and NC shRNA (sh-NC) was acquired from GenePharma Technology and subsequently inserted into the lentiviral pLKO vector, thus yielding lentiviruses pLKO-sh-LINC00173 and pLKO-sh-NC, respectively. To stably silence LINC00173 in A375 cells, either the pLKO-sh-LINC00173 or pLKO-sh-NC lentivirus was transduced into A375 cells. A375 cells stably expressing sh-LINC00173 were selected with puromycin.
All animal experiments were performed with the approval of the animal ethics committee of Shaanxi Provincial People’s Hospital and in conformity with the Animal Protection Law of the People’s Republic of China-2009 for experimental animals. Male, BALB/c nude mice at 4 weeks of age were bought from Huafukang Bioscience Co., Inc. (Beijing, China). A375 cells stably expressing either sh-LINC00173 or sh-NC were collected, resuspended in phosphate-buffered saline, and subcutaneously injected into the flank of the mice. Each group contained three mice. Starting at 7 days postinjection, the length (L) and width (W) of the tumor xenografts were measured every 2 days; the tumor volume was computed as L × W2 × 0.5. All mice were euthanized through cervical dislocation at 4 weeks after the tumor xenografting. Tumor xenografts were collected, weighed, and analyzed with RT-qPCR and Western blotting.
After fixation in 4% formalin, the tumor xenografts were embedded in paraffin and subjected to Anin situ terminal deoxynucleotidyl transferase dUTP nick end labeling (TUNEL) kit (Roche Applied Science) for the detection of cell apoptosis. The number of apoptotic cells was counted in both sh-LINC00173 and sh-NC groups.
Nuclear-and-Cytoplasmic Separation Assay
RNA located in the cytoplasm or nucleus of melanoma cells was separated using a Cytoplasmic and Nuclear RNA Purification Kit (Norgen, Thorold, ON, Canada). Cytoplasmic RNA and nuclear RNA samples were analyzed by RT-qPCR to determine LINC00173 expression distribution within the melanoma cell.
Bioinformatic Analysis and the Luciferase Reporter Assay
LINC00173–miRNA interactions were predicted using starBase 3.0 (http://starbase.sysu.edu.cn/). Fragments of LINC00173 carrying either the putative wild-type (WT) or mutant (MUT) miR-493–binding sequence were generated by GenePharma Technology, followed by insertion into the psi-CHECK2 luciferase reporter vector (Promega Corporation, Madison, WI, USA). The luciferase reporter plasmids were named as LINC00173-WT and LINC00173-MUT, respectively. Lipofectamine 2000® was employed to transfect melanoma cells with either the miR-493 mimic or miR-NC plus either LINC00173-WT or LINC00173-MUT. The cells were harvested at 48 h post-transfection and subjected to the measurement of luciferase activity via a Dual Luciferase Reporter Assay System (Promega Corporation). The activity of firefly luciferase was normalized to that of Renilla luciferase.
RNA Immunoprecipitation (RIP) Assay
This assay was performed for testing the interaction between LINC00173 and miR-493 in melanoma cells. We used the Magna RIP RNA‐Binding Protein Immunoprecipitation Kit (Millipore, Burlington, MA, USA). In short, cells were lysed in RIP lysis buffer. An anti-Argonaute 2 (AGO2) antibody or control IgG (both from Millipore) was conjugated to magnetic beads and was incubated with the cell extract. After overnight incubation at 4°C, the magnetic beads were collected, washed, and digested with Proteinase K. The immunoprecipitated RNA was extracted and analyzed by RT-qPCR to assess the enrichment of LINC00173 and miR-493 on the AGO2-containing beads.
Western Blotting
Total protein was extracted using radioimmunoprecipitation assay buffer (Beyotime, Shanghai, China). A BCA Protein Assay Kit (Beyotime) was employed to quantity the total-protein samples. Equivalent amounts of protein were separated by SDS 10% polyacrylamide gel electrophoresis and transferred onto polyvinylidene difluoride membranes. After 2 h blocking with 5% fat-free milk, primary antibodies were incubated with the membranes overnight at 4°C. The membranes were probed with a horseradish peroxidase–conjugated antibody (ab6721; 1:5,000 dilution; Abcam) at room temperature for 2 h. The protein signals were developed using the ECL Detection Kit (GE Healthcare Life Sciences, Chalfont, UK). The primary antibodies included IRS4 (cat. # ab52622; 1:500; Abcam, Cambridge, MA, USA), Bcl-2 (cat. # ab182858; 1:500; Abcam), BAX (cat. # ab32503; 1:500; Abcam), Bcl-XL (cat. # ab178844; 1:500; Abcam), CDK-2 (cat. # ab32147; 1:500; Abcam), CDK-4 (cat. # ab199728; 1:500; Abcam), cyclin D1 (cat. # ab16663; 1:500; Abcam), and GAPDH (ab128915; 1:500; Abcam).
Statistical Analysis
All experiments were performed at least in triplicate, and repeated three times. All measurement data are presented as mean ± standard deviations. Comparisons between two groups were evaluated with Student’s t test. One-way analysis of variance (ANOVA) together with Tukey’s post hoc test was performed to compare the data among multiple groups. The chi-square test was performed to determine the association between LINC00173 expression and clinical features of the patients with melanoma. Overall-survival curves were constructed by the Kaplan–Meier method, and the logrank test was performed to analyze the differences. Spearman correlation analysis was performed on some parameters. All statistical analyses were performed using SPSS 20.0 software (IBM Corp., Armonk, NY, USA), and P less than 0.05 was considered indicative of a statistically significant difference.
Results
LINC00173 Is Upregulated in Melanoma Tissues and Cell Lines
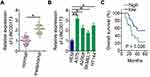
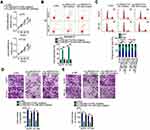
To analyze the expression status of LINC00173 in melanoma, RT-qPCR analysis was performed to measure the amounts of LINC00173 in the 45 pairs of melanoma tissue samples and adjacent normal tissues. LINC00173 was overexpressed in the melanoma tissues compared to the adjacent normal tissues (Figure 1A). LINC00173 expression in HEMs and four human melanoma cell lines (A375, A2058, SKMEL1, and HT144) was determined by RT-qPCR. LINC00173 level was higher in the four tested melanoma cell lines than in HEMs (Figure 1B).
Evaluation of the relation between LINC00173 expression and clinical features indicated that higher LINC00173 expression was significantly associated with TNM stage (P = 0.007) and lymph node metastasis (P = 0.002) in the 45 patients with melanoma (Table 1). Patients with melanoma featuring high LINC00173 expression showed shorter overall survival than did the patients with low LINC00173 expression (Figure 1C; P = 0.026). Therefore, LINC00173 was upregulated in melanoma, and this upregulation negatively correlated with the patients’ overall survival, implying that LINC00173 may perform crucial functions in melanoma progression.
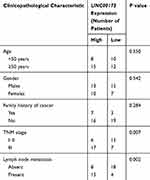 |
Table 1 Correlation Between LINC00173 Expression and Clinical Characteristics of Patients with Melanoma (n = 45 Patients) |
Downregulation of LINC00173 Restrains Melanoma Cell Proliferation, Migration, and Invasion and Induces Apoptosis in vitro
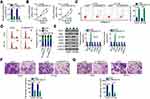
Given that LINC00173 was more highly expressed in melanoma cell lines A375 and HT144 compared to the other two melanoma cell lines, these two cell lines were chosen for further experiments. To investigate whether dysregulation of LINC00173 is functionally implicated in melanoma tumorigenesis, knockdown experiments were performed on A375 and HT144 cells using the siRNA specific to LINC00173 (si-LINC00173). RT-qPCR analysis confirmed good knockdown efficiency of si-LINC00173. Transfection with si-LINC00173 resulted in a marked decrease of LINC00173 expression in both A375 and HT144 cells (Figure 2A). LINC00173 knockdown obviously restrained the proliferation of A375 and HT144 cells (Figure 2B), as determined by the CCK-8 assay. The effect of LINC00173 silencing on melanoma cell apoptosis and cell cycle was tested by flow cytometry. LINC00173 depletion notably promoted the apoptosis (Figure 2C) and induced the G0/G1 cycle arrest (Figure 2D) of A375 and HT144 cells.
In addition, expression of apoptotic-associated proteins (Bcl-2, BAX and Bcl-XL) and cell cycle-associated proteins (CDK-2, CDK-4 and cyclin D1) in the LINC00173-deficiency A375 and HT144 cells was measured via Western blotting. The results (Figure 2E) were in accord with the respective cell apoptotic rate and cell cycle progression data.
To test whether LINC00173 could affect the migration and invasiveness of melanoma cells, the migratory and invasive abilities of LINC00173-deficient A375 and HT144 cells were determined in the migration and invasion assays. Treatment with si-LINC00173 drastically impaired the migration (Figure 2F) and invasiveness (Figure 2G) of A375 and HT144 cells. Thus, LINC00173 enhanced the malignancy of melanoma cells.
LINC00173 Serves as a ceRNA in Melanoma Cells by Sponging miR-493
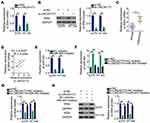
Mechanistically, lncRNAs can function as ceRNAs that sponge miRNAs and thereby increase the expression of miRNAs’ target mRNAs.34 To explore the mechanisms behind the strong involvement of LINC00173 in melanoma, we first evaluated LINC00173 expression distribution in the melanoma cell. According to the nuclear-and-cytoplasmic separation assay, LINC00173 was enriched in the cytoplasm of A375 and HT144 cells (Figure 3A), suggesting that LINC00173 may act as a ceRNA in melanoma. In online database starBase, version 3.0, LINC00173 was predicted to harbor a potential binding site for miR-493 (Figure 3B).
A luciferase reporter assay was performed to validate the direct binding between LINC00173 and miR-493 in melanoma cells. The miR-493 mimic–mediated upregulation of miR-493 (Figure 3C) effectively reduced the luciferase activity of plasmid LINC00173-WT in A375 and HT144 cells. In contrast, the luciferase activity of LINC00173-MUT was unaffected by the miR-493 mimic introduction (Figure 3D). The RIP assay revealed that LINC00173 and miR-493 were both enriched on the anti-AGO2 antibody–containing magnetic beads (Figure 3E). These observations confirmed the interaction between LINC00173 and miR-493 in melanoma cells.
To test whether miR-493 can be sponged by LINC00173 in melanoma cells, RT-qPCR was performed to measure miR-493 expression in A375 and HT144 cells after transfection with either si-LINC00173 or si-NC. The results showed that miR-493 was obviously induced in si-LINC00173–transfected A375 and HT144 cells relative to cells transfected with si-NC (Figure 3F). Clinical tissue test results indicated that miR-493 was downregulated in melanoma tissues (Figure 3G), consistently with the results of another study.35 An inverse association between LINC00173 and miR-493 expression levels in melanoma tissue samples was identified by Spearman correlation analysis (Figure 3H; r = –0.6006, P < 0.0001). Therefore, LINC00173 as a ceRNA is capable of sponging miR-493 in melanoma cells.
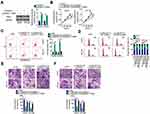
LINC00173 Positively Modulates IRS4 Expression in Melanoma Cells
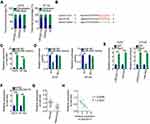
IRS4 mRNA is a direct target of miR-493 in melanoma cells.35 To examine the regulatory relation between LINC00173 and IRS4 in melanoma, the expression of IRS4 was evaluated in A375 and HT144 cells after knockdown of LINC00173. RT-qPCR and Western blotting results showed that mRNA (Figure 4A) and protein (Figure 4B) levels of IRS4 in A375 and HT144 cells were obviously downregulated by si-LINC00173 transfection. We found that IRS4 mRNA was overexpressed in the melanoma tissue samples (Figure 4C), showing a positive correlation with LINC00173 expression (Figure 4D; r = 0.5027, P = 0.0004). Rescue assays were then performed to determine whether the positive influence of LINC00173 on IRS4 expression is dependent on the sponging of miR-493. For this purpose, the miR-493 inhibitor was cotransfected with either si-LINC00173 or si-NC into A375 and HT144 cells. The transfection efficiency of the miR-493 inhibitor was validated by RT-qPCR (Figure 4E). Silencing of LINC00173 expression increased the expression of miR-493 in A375 and HT144 cells, and this phenomenon was reversed by cotransfection with the miR-493 inhibitor (Figure 4F). Downregulation of IRS4 mRNA (Figure 4G) and protein (Figure 4H) by si-LINC00173 in A375 and HT144 cells was attenuated by miR-493 inhibitor cotransfection. Consequently, LINC00173 acts as a ceRNA of miR-493 and thereby positively modulates IRS4 expression in melanoma cells.
Cancer-Promoting Activities of LINC00173 in Melanoma Cells are Dependent on Upregulation of miR-493–IRS4 Axis Output
Two other rescue assays were performed to test whether LINC00173 exerts its oncogenic actions via regulation of the miR-493–IRS4 axis. First, LINC00173-deficient A375 and HT144 cells were transfected with the miR-493 inhibitor. The downregulation of LINC00173 significantly inhibited proliferation (Figure 5A), promoted the apoptosis (Figure 5B) and induced the G0/G1 cycle arrest (Figure 5C) of A375 and HT144 cells, and these effects were abrogated by the miR-493 inhibitor cotransfection. Similarly, the effect of the LINC00173 knockdown on the migration (Figure 5D) and invasiveness (Figure 5E) of A375 and HT144 cells was reversed by miR-493 inhibition.
A rescue assay was performed on A375 and HT144 cells after cotransfection with si-LINC00173 and either IRS4-overexpressing plasmid pcDNA3.1-IRS4 or the empty pcDNA3.1 vector. Transfection with pcDNA3.1-IRS4 notably raised IRS4 protein (Figure 6A) levels in A375 and HT144 cells, as evidenced Western blotting. Functional experiments suggested that overexpression of IRS4 attenuated the LINC00173 depletion–induced effects on cell proliferation (Figure 6B), apoptosis (Figure 6C), cell cycle (Figure 6D), migration (Figure 6E), and invasiveness (Figure 6F) of A375 and HT144 cells. In brief, the oncogenic roles of LINC00173 in melanoma cells were found to be dependent on upregulation of miR-493–IRS4 axis output.
Knockdown of LINC00173 Inhibits the Tumorous Growth of Melanoma Cells in vivo by Enhancing miR-493 Expression and Reducing IRS4 Expression
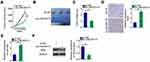
A xenograft tumor experiment was performed to examine the effect of LINC00173 on the growth of melanoma cells in vivo. A375 cells stably transfected with either sh-LINC00173 or sh-NC were subcutaneously inoculated into the flank of nude mice. The tumorous growth of the transplanted tumor cells was dramatically slower in the sh-LINC00173 group than in the sh-NC group (Figure 7A and B). At 4 weeks after tumor xenografting, all the mice were euthanized, and the tumor xenografts were resected. The weight of tumor xenografts was significantly lower in the sh-LINC00173 group than in the sh-NC group (Figure 7C). TUNEL assay manifested that cell apoptosis was obviously promoted by low expression of LINC00173 in the nude mice (Figure 7D). RT-qPCR and Western blotting analyses of these tumor xenografts indicated that miR-493 expression was higher (Figure 7E) but the IRS4 protein amount was lower (Figure 7F) in the tumor xenografts derived from stably sh-LINC00173–transfected A375 cells. Thus, LINC00173 knockdown impeded the tumorous growth of melanoma cells in vivo by decreasing the output of the miR-493–IRS4 axis.
Discussion
In recent years, lncRNAs were identified as novel regulators of tumorigenesis and tumor progression.36,37 Approximately one fifth of lncRNAs are predicted to control the aggressive phenotype of human cancers.38 Regarding melanoma, a variety of lncRNAs are abnormally expressed and are closely related to the patients’ prognosis.39–41 They play an important part in the malignant characteristics of melanoma in vitro and in vivo because these lncRNAs exert oncogenic or tumor-suppressive effects.42–44 Therefore, studying the specific roles of lncRNAs in melanoma may uncover effective therapeutic targets in this disease. Here, we first assessed the expression of LINC00173 in melanoma tissues and cell lines. Second, we applied siRNA to silence endogenous LINC00173 expression in melanoma cells in order to evaluate the influence of the LINC00173 knockdown on the malignant phenotype of melanoma cells in vitro and in vivo. Third, we explored the events behind the oncogenic activities of LINC00173 in melanoma cells.
LINC00173 is upregulated in non-small cell lung cancer32 and small cell lung cancer.33 Upregulation of LINC00173 is closely related to chemoresistance and a more advanced stage in patients with small cell lung cancer.33 Patients with small cell lung cancer overexpressing LINC00173 manifest shorter overall survival compared with patients with low LINC00173 expression.33 Functionally, LINC00173 has been confirmed as an oncogenic lncRNA in small cell lung cancer and is known to promote cancer cell chemoresistance, proliferation, and metastasis in vitro and tumor chemoresistance and growth in vivo.33 However, the expression pattern, clinical value, and detailed participation of LINC00173 in melanoma are poorly understood. In this work, our results indicate that LINC00173 expression is high in melanoma tissues and cell lines. High LINC00173 expression was associated with adverse clinical features and shorter overall survival of patients with melanoma. LINC00173 knockdown suppressed melanoma cell proliferation, migration, and invasion in vitro; increased apoptosis in vitro; and restricted tumorous growth in vivo.
Subcellular distribution of lncRNAs determines the functions of lncRNAs. The ceRNA theory indicates that when an lncRNA is mainly enriched in the cytoplasm, this RNA acts as a molecular sponge sequestering target miRNAs consequently de-repressing the miRNAs’ targets at the post-transcriptional level.34,45 Here, LINC00173 was demonstrated to be predominantly localized in the cytoplasm of melanoma cells, suggesting that LINC00173 may work as a ceRNA. Bioinformatic analysis then revealed that LINC00173 contains a binding site for miR-493. This prediction was validated by luciferase reporter and RIP assays. Knockdown of LINC00173 was found to decrease the expression of IRS4 (the target of miR-493) in melanoma cells, whereas this regulatory impact was abrogated by inhibition of miR-493 expression. A positive correlation between LINC00173 and IRS4 levels was confirmed in the melanoma tissues. Our rescue assays indicate that the oncogenic actions of LINC00173 on the progression of melanoma are dependent on enhancement of miR-493–IRS4 axis output. Therefore, the influence of LINC00173 on the aggressiveness of melanoma can be partly explained by the ceRNA mechanism involving LINC00173, miR-493, and IRS4 mRNA.
MiR-493 is dysregulated in multiple types of human cancer,46–51 including melanoma.35 Functionally, exogenous miR-493 expression attenuates cell proliferation and induces cell cycle arrest in melanoma.35 Mechanistically, IRS4 is a direct target gene of miR-493 in melanoma cells.35 IRS4 is a part of a family of cytoplasmic docking proteins mediating signals from cell surface receptors to downstream effectors.52 In humans, IRS4 is increasingly implicated in cancer initiation and progression. Overexpression of IRS4 is seen in hepatocellular carcinoma,53 breast cancer,54 colorectal cancer,55 and lung cancer.56 Our results indicate that IRS4 is upregulated by the LINC00173–miR-493 axis in melanoma. LINC00173 can interact with miR-493 directly to raise the expression of IRS4. Hence, a new LINC00173–miR-493–IRS4 pathway was identified here in melanoma cells and seems to control tumorigenesis and tumor progression. This study may offer novel ideas for the discovery of antimelanoma therapies.
Conclusions
Our study identified an oncogenic lncRNA, LINC00173, involved in melanoma. LINC00173 promotes the malignant characteristics of melanoma by increasing IRS4 expression via competitive binding to miR-493. Validating the participation of the LINC00173–miR-493–IRS4 pathway (Figure 8) in melanoma pathogenesis may be useful in the identification of novel therapeutic targets.
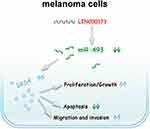 |
Figure 8 Schematic diagram of proposed mechanism. LINC00173 promotes the malignant characteristics of melanoma by increasing IRS4 expression via competitive binding to miR-493. |
Abbreviations
3′-UTR, 3′-untranslated region; CCK-8, Cell Counting Kit-8; DMEM, Dulbecco’s modified Eagle’s medium; FBS, fetal bovine serum; FITC, fluorescein isothiocyanate; lncRNA, long noncoding RNA; miRNA, miR, microRNA; MUT, mutant; NC, negative control; RIP, RNA immunoprecipitation; RT-qPCR, reverse-transcription quantitative PCR; shRNA, short hairpin RNA; siRNA, small interfering RNA; WT, wild-type.
Ethics and Consent Statement
This study was performed with the approval of the Research Ethics Committee of Shaanxi Provincial People’s Hospital. In addition, written informed consent forms were signed by all the patients who participated in this research. All animal experiments were performed with the approval of the animal ethics committee of Shaanxi Provincial People’s Hospital and in conformity with the Animal Protection Law of the People’s Republic of China-2009 for experimental animals.
Disclosure
The authors declare that they have no competing interests.
References
1. Owens B. Melanoma. Nature. 2014;515:S109. doi:10.1038/515S109a
2. Mohammadpour A, Derakhshan M, Darabi H, Hedayat P, Momeni M. Melanoma: where we are and where we go. J Cell Physiol. 2019;234:3307–3320. doi:10.1002/jcp.27286
3. Little EG, Eide MJ. Update on the current state of melanoma incidence. Dermatol Clin. 2012;30:355–361. doi:10.1016/j.det.2012.04.001
4. Slipicevic A, Herlyn M. Narrowing the knowledge gaps for melanoma. Ups J Med Sci. 2012;117:237–243. doi:10.3109/03009734.2012.658977
5. Gershenwald JE, Scolyer RA, Hess KR, et al.; P. for members of the American Joint Committee on Cancer Melanoma Expert, D. the International Melanoma, P. Discovery. Melanoma staging: evidence-based changes in the American Joint Committee on Cancer eighth edition cancer staging manual. CA Cancer J Clin. 2017;67:472–492. doi:10.3322/caac.21409
6. Malvi P, Chaube B, Singh SV, et al. Elevated circulatory levels of leptin and resistin impair therapeutic efficacy of dacarbazine in melanoma under obese state. Cancer Metab. 2018;6:2. doi:10.1186/s40170-018-0176-5
7. Mohammad N, Malvi P, Meena AS, et al. Cholesterol depletion by methyl-beta-cyclodextrin augments tamoxifen induced cell death by enhancing its uptake in melanoma. Mol Cancer. 2014;13:204. doi:10.1186/1476-4598-13-204
8. Chaube B, Malvi P, Singh SV, Mohammad N, Meena AS, Bhat MK. Targeting metabolic flexibility by simultaneously inhibiting respiratory complex I and lactate generation retards melanoma progression. Oncotarget. 2015;6:37281–37299. doi:10.18632/oncotarget.6134
9. Malvi P, Chaube B, Singh SV, et al. Weight control interventions improve therapeutic efficacy of dacarbazine in melanoma by reversing obesity-induced drug resistance. Cancer Metab. 2016;4:21. doi:10.1186/s40170-016-0162-8
10. Malvi P, Chaube B, Pandey V, et al. Obesity induced rapid melanoma progression is reversed by orlistat treatment and dietary intervention: role of adipokines. Mol Oncol. 2015;9:689–703. doi:10.1016/j.molonc.2014.11.006
11. Tsao H, Chin L, Garraway LA, Fisher DE. Melanoma: from mutations to medicine. Gene Dev. 2012;26:1131–1155. doi:10.1101/gad.191999.112
12. Pacheco I, Buzea C, Tron V. Towards new therapeutic approaches for malignant melanoma. Expert Rev Mol Med. 2011;13:e33. doi:10.1017/S146239941100202X
13. Arnold J, Engelmann JC, Schneider N, Bosserhoff AK, Kuphal S. miR-488-5p and its role in melanoma. Exp Mol Pathol. 2020;112:104348. doi:10.1016/j.yexmp.2019.104348
14. Lugovic-Mihic L, Cesic D, Vukovic P, Bilic GN, Situm M, Spoljar S. Melanoma development: current knowledge on melanoma pathogenesis. Acta Dermatovener Cr. 2019;27:163–168.
15. Castro-Oropeza R, Melendez-Zajgla J, Maldonado V, Vazquez-Santillan K. The emerging role of lncRNAs in the regulation of cancer stem cells. Cell Oncol. 2018;41:585–603. doi:10.1007/s13402-018-0406-4
16. Liz J, Esteller M. lncRNAs and microRNAs with a role in cancer development. Biochim Biophys Acta. 2016;1859:169–176. doi:10.1016/j.bbagrm.2015.06.015
17. Wei YG, Liu ZY, Fang JH. H19 functions as a competing endogenous RNA to regulate human epidermal growth factor receptor expression by sequestering let-7c in gastric cancer. Mol Med Rep. 2018;17:2600–2606. doi:10.3892/mmr.2017.8184
18. Shang W, Adzika GK, Li Y, et al. Molecular mechanisms of circular RNAs, transforming growth factor-beta, and long noncoding RNAs in hepatocellular carcinoma. Cancer Med. 2019;8:6684–6699. doi:10.1002/cam4.2553
19. Zong W, Ju SQ, Jing RR, Cui M. Long non-coding RNA-mediated regulation of signaling pathways in gastric cancer. Clin Chem Lab Med. 2018;56:1828–1837. doi:10.1515/cclm-2017-1139
20. Ng M, Santer L, Emmrich S, Schwarzer A, Heckl D, Klusmann JH. Crispri/a screening to identify functional long noncoding RNAs in pediatric acute myeloid leukemia. Ann Hematol. 2017;96:S78.
21. Yao F, Wang Q, Wu QM. The prognostic value and mechanisms of lncRNA UCA1 in human cancer. Cancer Manag Res. 2019;11:7685–7696. doi:10.2147/CMAR.S200436
22. Luan WK, Ding YT, Ma SJ, Ruan HR, Wang JL, Lu F. Long noncoding RNA LINC00518 acts as a competing endogenous RNA to promote the metastasis of malignant melanoma via miR-204-5p/AP1S2 axis. Cell Death Dis. 2019;10. doi:10.1038/s41419-019-2090-3
23. Gao H, Liu R, Sun X. STAT3-induced upregulation of lncRNA SNHG17 predicts a poor prognosis of melanoma and promotes cell proliferation and metastasis through regulating PI3K-AKT pathway. Eur Rev Med Pharmaco. 2019;23:8000–8010.
24. Wei XH, Gu XL, Ma M, Lou CX. Long noncoding RNA HCP5 suppresses skin cutaneous melanoma development by regulating RARRES3 gene expression via sponging miR-12. Once targets Ther. 2019;12:6323–6335. doi:10.2147/OTT.S195796
25. Ameis D, Khoshgoo N, Iwasiow BM, Snarr P, Keijzer R. MicroRNAs in lung development and disease. Paediatr Respir Rev. 2017;22:38–43. doi:10.1016/j.prrv.2016.12.002
26. Muhammad N, Bhattacharya S, Steele R, Ray RB. Anti-miR-203 suppresses ER-positive breast cancer growth and stemness by targeting SOCS3. Oncotarget. 2016;7:58595–58605. doi:10.18632/oncotarget.11193
27. Varrone F, Caputo E. The miRNAs role in melanoma and in its resistance to therapy. Int J Mol Sci. 2020;21:878. doi:10.3390/ijms21030878
28. Lu NH, Wei CY, Qi FZ, Gu JY. Hsa-let-7b suppresses cell proliferation by targeting UHRF1 in melanoma. Cancer Invest. 2020;38:52–60. doi:10.1080/07357907.2019.1709482
29. Chen L, Karisma VW, Liu H, Zhong L. MicroRNA-300: A transcellular mediator in exosome regulates melanoma progression. Front Oncol. 2019;9:1005. doi:10.3389/fonc.2019.01005
30. Xu Y, Wang L, Jiang L, Zhang X. Novel microRNA biomarkers, miR-142-5p, miR-550a, miR-1826, and miR-1201, were identified for primary melanoma. J Comput Biol. 2019. doi:10.1089/cmb.2019.0198
31. Zhao G, Yin Y, Zhao B. miR-140-5p is negatively correlated with proliferation, invasion, and tumorigenesis in malignant melanoma by targeting SOX4 via the Wnt/beta-catenin and NF-kappaB cascades. J Cell Physiol. 2020;235:2161–2170. doi:10.1002/jcp.29122
32. Yang Q, Tang Y, Tang C, et al. Diminished LINC00173 expression induced miR-182-5p accumulation promotes cell proliferation, migration and apoptosis inhibition via AGER/NF-kappaB pathway in non-small-cell lung cancer. Am J Transl Res. 2019;11:4248–4262.
33. Zeng F, Wang Q, Wang S, et al. Linc00173 promotes chemoresistance and progression of small cell lung cancer by sponging miR-218 to regulate Etk expression. Oncogene. 2019;39:293–307.
34. Abdollahzadeh R, Daraei A, Mansoori Y, Sepahvand M, Amoli MM, Tavakkoly-Bazzaz J. Competing endogenous RNA (ceRNA) cross talk and language in ceRNA regulatory networks: A new look at hallmarks of breast cancer. J Cell Physiol. 2019;234:10080–10100. doi:10.1002/jcp.27941
35. Cui A, Jin Z, Gao Z, et al. Downregulation of miR-493 promoted melanoma proliferation by suppressing IRS4 expression. Tumour Biol. 2017;39:1010428317701640. doi:10.1177/1010428317701640
36. Ghafouri-Fard S, Mohammad-Rahimi H, Taheri M. The role of long non-coding RNAs in the pathogenesis of thyroid cancer. Exp Mol Pathol. 2019;112:104332. doi:10.1016/j.yexmp.2019.104332
37. Gourvest M, Brousset P, Bousquet M. Long noncoding RNAs in acute myeloid leukemia: functional characterization and clinical relevance. Cancers. 2019;11:1638. doi:10.3390/cancers11111638
38. Zhao M, Zhu N, Hao F, et al. The regulatory role of non-coding RNAs on programmed cell death four in inflammation and cancer. Front Oncol. 2019;9:919. doi:10.3389/fonc.2019.00919
39. Yang S, Xu J, Zeng X. A six-long non-coding RNA signature predicts prognosis in melanoma patients. Int J Oncol. 2018;52:1178–1188. doi:10.3892/ijo.2018.4268
40. Yu X, Zheng H, Tse G, Chan MT, Wu WK. Long non-coding RNAs in melanoma. Cell Prolif. 2018;51:e12457.
41. Aftab MN, Dinger ME, Perera RJ. The role of microRNAs and long non-coding RNAs in the pathology, diagnosis, and management of melanoma. Arch Biochem Biophys. 2014;563:60–70.
42. Liao Z, Zhao J, Yang Y. Downregulation of lncRNA H19 inhibits the migration and invasion of melanoma cells by inactivating the NFkappaB and PI3K/Akt signaling pathways. Mol Med Rep. 2018;17:7313–7318. doi:10.3892/mmr.2018.8782
43. Li P, Gao Y, Li J, et al. LncRNA MEG3 repressed malignant melanoma progression via inactivating Wnt signaling pathway. J Cell Biochem. 2018;119:7498–7505. doi:10.1002/jcb.27061
44. Shi G, Li H, Gao F, Tan Q. lncRNA H19 predicts poor prognosis in patients with melanoma and regulates cell growth, invasion, migration and epithelial-mesenchymal transition in melanoma cells. Onco Targets Ther. 2018;11:3583–3595. doi:10.2147/OTT.S160143
45. Dong BS, Shi MJ, Su SB, Zhang H. Insight into long noncoding competing endogenous RNA networks in hepatic fibrosis: the potential implications for mechanism and therapy. Gene. 2019;687:255–260. doi:10.1016/j.gene.2018.11.063
46. Zhou W, Zhang C, Jiang H, Zhang Z, Xie L, He X. MiR-493 suppresses the proliferation and invasion of gastric cancer cells by targeting RhoC. Iran J Basic Med Sci. 2015;18(10):1027–1033.
47. Xu Y, Ge K, Lu J, Huang J, Wei W, Huang Q. MicroRNA-493 suppresses hepatocellular carcinoma tumorigenesis through down-regulation of anthrax toxin receptor 1 (ANTXR1) and R-Spondin 2 (RSPO2). Biomed Pharmacother. 2017;93:334–343. doi:10.1016/j.biopha.2017.06.047
48. Zhi D, Zhao X, Dong M, Yan C. miR-493 inhibits proliferation and invasion in pancreatic cancer cells and inversely regulated hERG1 expression. Oncol Lett. 2017;14:7398–7404. doi:10.3892/ol.2017.7178
49. Li J, Meng S, Xu M, et al. Downregulation of N(6)-methyladenosine binding YTHDF2 protein mediated by miR-493-3p suppresses prostate cancer by elevating N(6)-methyladenosine levels. Oncotarget. 2018;9:3752–3764. doi:10.18632/oncotarget.23365
50. Ueno K, Hirata H, Majid S, et al. Tumor suppressor microRNA-493 decreases cell motility and migration ability in human bladder cancer cells by downregulating RhoC and FZD4. Mol Cancer Ther. 2012;11:244–253. doi:10.1158/1535-7163.MCT-11-0592
51. Gu Y, Cheng Y, Song Y, et al. MicroRNA-493 suppresses tumor growth, invasion and metastasis of lung cancer by regulating E2F1. PLoS One. 2014;9:e102602.
52. Ikink GJ, Hilkens J. Insulin receptor substrate 4 (IRS4) is a constitutive active oncogenic driver collaborating with HER2 and causing therapeutic resistance. Mol Cell Oncol. 2017;4:e1279722. doi:10.1080/23723556.2017.1279722
53. Cantarini MC, de la Monte SM, Pang M, et al. Aspartyl-asparagyl beta hydroxylase over-expression in human hepatoma is linked to activation of insulin-like growth factor and notch signaling mechanisms. Hepatology. 2006;44:446–457.
54. Ikink GJ, Boer M, Bakker ER, Hilkens J. IRS4 induces mammary tumorigenesis and confers resistance to HER2-targeted therapy through constitutive PI3K/AKT-pathway hyperactivation. Nat Commun. 2016;7:13567. doi:10.1038/ncomms13567
55. Sanmartin-Salinas P, Toledo-Lobo MV, Noguerales-Fraguas F, Fernandez-Contreras ME, Guijarro LG. Overexpression of insulin receptor substrate-4 is correlated with clinical staging in colorectal cancer patients. J Mol Histol. 2018;49:39–49. doi:10.1007/s10735-017-9745-0
56. Weischenfeldt J, Dubash T, Drainas AP, et al. Pan-cancer analysis of somatic copy-number alterations implicates IRS4 and IGF2 in enhancer hijacking. Nat Genet. 2017;49:65–74. doi:10.1038/ng.3722
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.