Back to Journals » OncoTargets and Therapy » Volume 13
Long Noncoding RNA HEIH Promotes Proliferation, Migration and Invasion of Retinoblastoma Cells Through miR-194-5p/WEE1 Axis
Authors Gao S, Chu Q, Liu X, Zhao X, Qin L, Li G, Liu Q
Received 30 June 2020
Accepted for publication 12 October 2020
Published 23 November 2020 Volume 2020:13 Pages 12033—12041
DOI https://doi.org/10.2147/OTT.S268942
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr William C. Cho
Sheng Gao,1 Qingxia Chu,1 Xia Liu,1 Xia Zhao,2 Libao Qin,1 Guoliang Li,1 Qinghuai Liu3
1Department of Ophthalmology, Nanjing Pukou Central Hospital, Nanjing 211800, People’s Republic of China; 2Department of Ophthalmology, Tangshan Eye Hospital, Tangshan 063000, People’s Republic of China; 3Department of Ophthalmology, The First Affiliated Hospital of Nanjing Medical University, Nanjing 210029, People’s Republic of China
Correspondence: Qinghuai Liu
Department of Ophthalmology, The First Affiliated Hospital of Nanjing Medical University, 300# Guangzhou Road, Gulou District, Nanjing 210029, People’s Republic of China
Email [email protected]
Background: Abnormally expressed long noncoding RNA (lncRNA) high expression in hepatocellular carcinoma (HEIH) has been implicated in many types of human cancer, and plays crucial roles in tumor development and progression. However, little is known about its function in retinoblastoma.
Methods: qRT-PCR was used to determine the expression levels of HEIH, miR-194-5p and WEE1 in retinoblastoma tissues and cell lines. The trypan blue exclusion method, colony formation assay, wound-healing assay and transwell invasion assay were performed to evaluate the effects of HEIH, miR-194-5p and WEE1 on cell proliferation, migration and invasion. Bioinformatics analysis, dual-luciferase reporter assay and Western blot were employed to investigate the regulatory relationship among HEIH, miR-194-5p and WEE1.
Results: We found that HEIH was up-regulated in retinoblastoma tissues and cell lines. Furthermore, high level of HEIH was associated with TNM stage, optic nerve invasion and choroidal invasion of patients with retinoblastoma. Functional studies showed that HEIH knockdown significantly suppressed retinoblastoma cell proliferation, migration and invasion. Mechanistically, HEIH promoted retinoblastoma progression by serving as a sponge of miR-194-5p to regulate WEE1 expression.
Conclusion: Our work suggests that HEIH acts as an oncogenic lncRNA to promote retinoblastoma proliferation and metastasis, providing a new insight into the retinoblastoma treatment.
Keywords: retinoblastoma, HEIH, miR-194-5p, WEE1, metastasis
Introduction
Retinoblastoma is the most common aggressive childhood intraocular cancer, with approximately 9000 newly established cases annually.1 The typical clinical feature of retinoblastoma patients is leukocoria and strabismus. According to a study conducted by Epstein et al from a cohort of 1196 retinoblastoma, median age at presentation is 15 months with 51% male, 49% female and 53% unilateral, 47% bilateral.2 Indeed, the mutation of the Rb1 gene in retinal cells has been characterized as a risk factor for retinoblastoma. However, evidence now support the tent that retinoblastoma initiation is involved in genetic and epigenetic alterations as well as dysregulated noncoding RNA.3 Hence, the molecular mechanisms underlying retinoblastoma pathogenesis is complex and more efforts are needed to tackle this issue.
Over the past few decades, the survival of retinoblastoma patients has been significantly improved, due to advancements in the enucleation, radiotherapy and neoadjuvant chemotherapy. But the prognosis between high-income and low-income countries is very different. In high-income countries, the disease-free survival rate is over 95%.4,5 In low-income countries, the survival rate is still very poor, such as 62.1% at 5 years in Jordan, 26.6% at 3 years in Kenyatta and 23.1% at 5 years in Harare.6–8 Additionally, retinoblastoma can rapidly fill the eye, extend into the optic nerve and the central nervous system, and easily metastasizes.9 Nevertheless, it is urgent to decipher the molecular mechanisms involved in the metastasis of retinoblastoma to discover new biomarkers for diagnosis and treatment.
Long noncoding RNAs (lncRNAs) are a class of regulatory RNAs with more than 200 nucleotides in length. Several studies have demonstrated that lncRNAs exert pivotal functions in retinoblastoma via acting as tumor suppressors or oncogenes.10–12 High expression in hepatocellular carcinoma (HEIH), also known as hepatocellular carcinoma up-regulated EZH2-associated long non-coding RNA, is a recently identified intergenic lncRNA located in chromosome chr5:180256954–180258618. HEIH was firstly characterized to be overexpressed in HBV-related hepatocellular carcinoma tissues compared with paired peritumoral tissues.13 Subsequently, researchers reported that HEIH was frequently up-regulated in various types of human cancer, including colorectal cancer, melanoma, esophageal squamous cell carcinoma, breast cancer and cholangiocarcinoma, and promoted cancer cell proliferation, migration and invasion through sponging tumor suppressor miRNAs.14–18 Taken together, the above mentioned findings suggest that HEIH is a functional lncRNA in tumor progression, yet its biological role in retinoblastoma remains largely unknown.
This study aimed to investigate the expression profiling and role of HEIH in retinoblastoma. qRT-PCR was first used to determine HEIH level in retinoblastoma tissues and normal retinal tissues. To explore the effects of abnormally expressed HEIH on the cellular biological behavior of retinoblastoma cells, we performed loss-of-function assays through transfection with specific siRNA for HEIH. We further attempted to reveal the possible underlying mechanism through which HEIH exerted its biological functions in retinoblastoma.
Materials and Methods
Clinical Samples
Thirty-five retinoblastoma tissues and seven normal retinal tissues were collected from the enucleation in Nanjing Pukou Central Hospital between June 2014 and August 2016. No patients had received any chemotherapy or radiotherapy prior to the surgery. Collections and use of human tissue samples were approved by the Ethics Committee of Nanjing Pukou Central Hospital (No.20170812) and written informed consent was obtained from each participant. All protocols followed the principles outlined in the Declaration of Helsinki. The clinicopathological characteristics are summarized in Table 1.
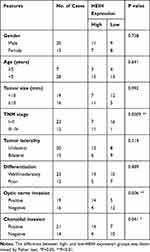 |
Table 1 The Relationship Between HEIH Expression and Clinicopathologic Features of Retinoblastoma Patients |
Cell Culture and Transfection
Human normal retinal epithelial cell line ARPE-19 and retinoblastoma cell lines (Y79 and SO-Rb50) were purchased from ATCC and maintained in RPMI-1640 medium supplemented with 10% FBS at 37°C in 5% CO2. si-RNA targeting HEIH, and corresponding negative control si-NC, miR-194-5p mimics, inhibitor and negative control, WEE1 overexpressing vector pcDNA3.1-WEE1 and negative control pcDNA3.1 were supplied by RiboBio (Guangzhou, China). Cells were transfected using Lipofectamine 2000 (Invitrogen) according to the manufacturer’s protocol. At 48 h transfection, the cells were harvested for functional analysis.
Cell Proliferation Assay
After 24 h transfection, cells were plated in 6-well plates and then cultured for 48h. The number of viable cells was counted with the trypan blue exclusion method on a Coulter counter (Beckman Coulter, Miami, FL, USA). For colony formation assay, 500 transfected cells were plated in 6-well plates for 14 days. Then the cells were washed two times with PBS, fixed with methanol and stained with 0.1% crystal violet (Sigma, St. Louis, MO, USA). The colonies were photographed and counted using a microscope (Olympus, Tokyo, Japan).
qRT-PCR Assay
Total RNA was isolated from tissues or cultured cells with Trizol reagent (Thermo Fisher Scientific, Waltham, MA, USA). cDNA was reverse-transcribed with the All-in-One First-Strand cDNA Synthesis kit (GeneCopoeia, Rockville, MD, USA). All reactions were conducted using Platinum SYBR Green qPCR SuperMix-UDG reagents (Thermo Fisher Scientific) in triplicate for each sample. Gene expression was calculated using the 2-ΔΔCt method relative to U6 or GAPDH. The primers sequences are as follows:
HEIH-Forward: 5ʹ-CCTCTTGTGCCCCTTTCT-3ʹ,
HEIH-Reverse: 5ʹ- AGGTCTCATGGCTTCTCG-3ʹ;
GAPDH-Forward: 5ʹ-CTCTGCTCCTCCTGTTCGAC-3ʹ,
GAPDH-Reverse: 5ʹ- GACTCCGACCTTCACCTTCC-3ʹ;
miR-194-5p-Forward: 3ʹ-CTAGTACCTAGAGGAACCTTTGAAGACTGTTACAGCTCAGCA-5ʹ,
miR-194-5p-Reverse: 5ʹ- AGCTTGCTGAGCTGTAACAGTCTTCAAAGGTTCCTCTAGGTA-3ʹ;
U6-Forward: 5ʹ-GTGGACCGCACAAGCTCGCT-3ʹ,
U6-Reverse:5ʹ-TTGTTGAACGGCACTGTGTATAGCA-3ʹ.
Cell Migration and Invasion Assays
For wound-healing assay, transfected cells were seeded in the 6-well plates. The monolayer cells were scraped with a 200 μL sterile pipette tip. The wound healing was then captured at 0 and 24 h under a microscope. For Transwell invasion assay, transfected cells were seeded into the upper chamber pre-coated with Matrigel (BD Biosciences, San Jose, CA, USA). RPMI-1640 medium containing 10% FBS was added to the bottom chamber. After a 24-h incubation, the invasive cells on the lower surface were fixed and stained with 0.1% crystal violet, and counted in six randomly fields under a microscope.
Western Blot Analysis
Cultured cells were harvested and lysed with RIPA buffer. The equivalent protein samples (30 μg) were loaded into 12% SDS-PAGE and electroblotted onto PVDF membranes (EMD Millipore, Billerica, MA, USA). Membranes were blocked by 5% non-fat milk for 2 h at room temperature and then incubated with primary antibodies as follows: WEE1 (#13084, Cell Signaling Technology) and GAPDH (ab181602). After incubation overnight at 4 °C, the membranes were incubated with HRP-conjugated goat anti-rabbit secondary antibodies for 1 h at room temperature. The blot signals were visualized with ECL reagents (Pierce, Waltham, MA, USA).
Dual-Luciferase Reporter Assay
The luciferase reporters of wild-type and mutated HEIH and 3ʹUTR of WEE1 containing predicted binding site of miR-194-5p were constructed into the psiCHECK-2 vector (Promega). Then, reporter plasmids wt-HEIH, mut-HEIH, wt-WEE1 and mut-WEE1 with miR-194-5p mimics or negative control were co-transfected into retinoblastoma cells using Lipofectamine 2000. Forty-eight hours after transfection, 100 μL PLB buffer was added to each well and cell lysates were collected after 15 min shake at room temperature. The cell lysates were used to evaluate the luciferase activities using the dual luciferase reporter assay system (Promega Corporation, Fitchburg, WI, USA).
Statistical Analysis
All data are presented as the means ± SD from at least 3 independent experiments. The difference was determined by Student’s t-test and the one-way analysis of variance (ANOVA). Overall survival rate was analyzed used Kaplan-Meier method and Log rank test according to the median value of HEIH expression. The associations between HEIH expression and clinicopathologic characteristics were determined by Fisher test. Results were analyzed using GraphPad Prism v6.0 (GraphPad Software, Inc., La Jolla, CA, USA). Differences were considered statistically significant at P < 0.05.
Results
HEIH Up-Regulation Was Associated with TNM Stage, Optic Nerve Invasion and Choroidal Invasion in Retinoblastoma
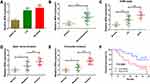
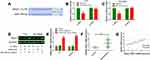
We first determined the expression level of HEIH in normal retinal epithelial cell line ARPE-19 and retinoblastoma cell lines (Y79 and SO-Rb50). As shown in Figure 1A, HEIH was up-regulated 2.2-fold and 2.5-fold in Y79 and SO-Rb50 relative to ARPE-19. Then, we validated HEIH expression in 35 retinoblastoma tissues and 7 normal retinal tissues. Consistently, HEIH exhibited a mean 2.3-fold increase in retinoblastoma tissues (Figure 1B). We also evaluated the correlation between HEIH expression and patients’ characteristics (Table 1). Notably, a higher level of HEIH was observed in advanced tumor ICRB stage patients than that of early tumor stage (Figure 1C). Interestingly, HEIH expression was significantly related to retinoblastoma optic nerve invasion (Figure 1D) as well as choroidal invasion (Figure 1E). Furthermore, up-regulated HEIH predicted an unfavorable survival of retinoblastoma patients (P=0.0255; Figure 1F). Thus, HEIH might play an important regulatory role in retinoblastoma carcinogenesis.
HEIH Knockdown Inhibited Retinoblastoma Cell Proliferation, Migration and Invasion
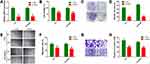
Loss-of-function assays were performed to establish the functional role of HEIH by transfecting si-HEIH into Y79 and SO-Rb50 cells. qRT-PCR results showed that HEIH expression was significantly decreased after si-HEIH transfection (Figure 2A). As shown in Figure 2B, HEIH knockdown obviously reduced the number of viable cells. In addition, a similar effect was observed in colony formation assays (Figure 2C and D). We also evaluated the effects of HEIH on the migration and invasion of retinoblastoma cells. Wound-healing experiment proved that the migratory distance was markedly reduced by HEIH silence (Figure 2E and F). Transwell assay result showed that HEIH knockdown resulted in a smaller number of invasive cells than negative control (Figure 2G and H). Therefore, HEIH down-regulation suppressed proliferation and metastasis of retinoblastoma cells.
HEIH Negatively Regulated miR-194-5p Expression in Retinoblastoma
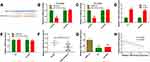
To explore the underlying mechanism of HEIH in retinoblastoma, bioinformatics analysis was performed to search for potential targets. Based on starBase v3.0, HEIH had a putative binding site for miR-194-5p (Figure 3A). We then constructed mutant and wild type luciferase reporter of HEIH for miR-194-5p. The results showed that wt-HEIH overexpression remarkably reduced the luciferase activity when co-transfection with miR-194-5p mimics compared with miRNA negative, but no obvious change for mutant HEIH (Figure 3B and C). We also assessed the regulatory relationship between HEIH and miR-194-5p. HEIH knockdown could up-regulate miR-194-5p expression (Figure 3D). Meanwhile, miR-194-5p overexpression had no significant influence on HEIH level (Figure 3E). Furthermore, miR-194-5p expression was detected using qRT-PCR in retinoblastoma tissues and cell lines. As shown in Figure 3F and G, miR-194-5p level was lower in retinoblastoma tissues and cell lines than normal retinal tissues and ARPE-19 cells, respectively. More importantly, there was a significant negative correlation between miR-194-5p and HEIH in retinoblastoma tissues (R=−0.5461; P=0.0007; Figure 3H). These data suggest that miR-194-5p was a downstream target of HEIH in retinoblastoma.
HEIH Positively Regulated WEE1 Expression via miR-194-5p
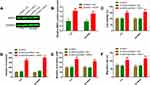
Utilizing TargetScan v7.2, we further predicted the possible target genes of miR-194-5p. As shown in Figure 4A, we identified that the conserved 3ʹ UTR of WEE1 contained complementary sequence of miR-194-5p at the position of 225–231. Luciferase reporter assay was performed to validate whether miR-194-5p could directly interact with WEE1 3ʹUTR. In both Y79 and SO-Rb50 cells, luciferase activity of wt-WEE1+miR-194-5p mimics group was evidently decreased compared to mut-WEE1+miR-194 mimics (Figure 4B and C). We also performed Western blot to test whether HEIH could regulate WEE1 expression through interacting with miR-194-5p. We found that HEIH knockdown could down-regulate WEE1 expression, but miR-194-5p inhibitor transfection reversed the effects of HEIH knockdown on WEE1 level (Figure 4D and E). In retinoblastoma tissues, a higher expression level of WEE1 was observed than normal retinal tissues (Figure 4F). Additionally, WEE1 level was positively associated with HEIH (Figure 4G). We conclude that HEIH serves as a sponge of miR-194-5p to up-regulate WEE1 expression in retinoblastoma.
Overexpression of WEE1 Reversed Malignant Phenotypes Inhibition of Retinoblastoma Cells Induced by HEIH Knockdown
To test whether HEIH exerted its oncogenic roles through regulating WEE1 expression, we first constructed WEE1 overexpressing vector pcDNA 3.1-WEE1 and its control vector pcDNA 3.1. Western blot analysis validated the up-regulation of WEE1 by pcDNA3.1-WEE1 transfection (Figure 5A and B). The proliferation of si-HEIH+pcDNA3.1-WEE1 was significantly increased compared with si-HEIH+pcDNA3.1-NC group (Figure 5D and E). Besides, ectopic expression of WEE1 also rescued the migrative and invasive capability of retinoblastoma cells (Figure 5F). Taken together, these findings indicate that HEIH promotes retinoblastoma cell proliferation, migration and invasion through up-regulating WEE1 expression.
Discussion
Recent genome and transcriptome studies have shown that only about 1–2% of the human genome is responsible for encoding proteins, and most transcripts are a large set of non-coding RNAs.19,20 Previous studies have implicated that abnormally expressed lncRNAs were involved in many human cancers, including retinoblastoma. For example, highly expressed lncRNA SNHG16 predicted poor overall survival of patients with retinoblastoma.21 Silencing of lncRNA LINC00152 has been shown to inhibit cell proliferation, colony formation, migration, and invasion and promote cell apoptosis in retinoblastoma.22 In addition, lncRNA UCA1 promoted multidrug resistance of retinoblastoma cells by sponging miR-513a-5p.23 Taken together, these findings suggested that targeting lncRNAs could interfere with retinoblastoma progression, and identification of functional lncRNAs might provide potential therapeutic targets for retinoblastoma treatment.
In this study, we observed that HEIH was significantly overexpressed in retinoblastoma cell lines and tissues. Furthermore, up-regulated HEIH indicated advanced tumor stage, optic nerve invasion and choroidal invasion. Loss-of-function assays showed that HEIH knockdown obviously suppressed retinoblastoma cell proliferation, migration and invasion. Our data suggest that HEIH in retinoblastoma serves as an oncogene, which was consistent with previous studies of HEIH in several types of human cancer.14–18
Accumulating data demonstrated that lncRNA transcripts can act as endogenous decoys for miRNAs to influence cancer-related gene expression through their miRNA binding sites. For example, lncRNA H19 increased retinoblastoma cell viability, migration and invasion via miR-143/RUNX2 axis.24 HEIH promoted breast cancer progression through down-regulating microRNA-200b.17 To investigate the underlying mechanism through which HEIH exerted its malignant phenotypes in retinoblastoma, we performed bioinformatics analysis using starBase. The results showed that HEIH contained putative binding site for miR-194-5p. Further investigations revealed that HEIH could directly bind miR-194-5p and reduced miR-194-5p expression in retinoblastoma. These data indicate that miR-194-5p is a downstream target of HEIH in retinoblastoma.
Growing studies have suggested that miR-194-5p plays key roles in tumor initiation and progression. For example, miR-194-5p overexpression was reported to suppress proliferation, migration and invasion of hypopharyngeal carcinoma cells by regulating SMURF1.25 In bladder cancer, miR-194-5p was down-regulated and associated with node metastasis and differentiation.26 Restoration of miR-194-5p could inhibit cell migration and invasion through targeting E2F3. Besides, Zhu et al found that miR-194 inhibited the proliferation, invasion, migration, and enhanced the cisplatin sensitivity by targeting FOXA1 in non-small cell lung cancer.27 These mentioned findings indicate that miR-194-5p functions as a tumor suppressor. In breast cancer, however, miR-194-5p acted as an oncogenic miRNA to promote cell proliferation, migration and invasion via regulating the Wnt/β-catenin signaling pathway.28 These findings indicate that miR-194-5p has a complex role in tumorigenesis.
WEE1 is a protein kinase that plays an important role in cell cycle and DNA replication by regulating the G2 checkpoint and preventing entry into mitosis in response to DNA damage.29 Recently, several studies demonstrated that WEE1 were implicated in tumor growth and metastasis in laryngeal carcinoma, glioma, non-small cell lung cancer, bladder cancer and hepatocellular carcinoma.30–34 In this study, we observed that miR-194-5p could directly target WEE1 3ʹUTR and regulating its expression, which was consistent with Li’s report of miR-194 targeting WEE1 in laryngeal squamous cell carcinoma.35 Additionally, HEIH knockdown significantly reduced WEE1 expression via miR-194-5p. More importantly, WEE1 overexpression rescued the inhibition of HEIH knockdown on retinoblastoma cell proliferation, migration and invasion.
Conclusion
Taken together, our data revealed that HEIH functions as an oncogene to promote retinoblastoma cell proliferation, migration and invasion through miR-194-5p/WEE1 axis. These findings demonstrated that HEIH is a useful prognostic biomarker in retinoblastoma detection and has the potential to be a novel promising therapeutic target for retinoblastoma treatment, especially for those patients who have developed optic nerve and choroidal invasion.
Disclosure
The authors declare that they have no competing interests.
References
1. Dimaras H, Kimani K, Dimba EA, et al. Retinoblastoma. Lancet. 2012;379(9824):1436–1446. doi:10.1016/S0140-6736(11)61137-9
2. Epstein JA, Shields CL, Shields JA. Trends in the management of retinoblastoma: evaluation of 1,196 consecutive eyes during 1974 to 2001. J Pediatr Ophthalmol Strabismus. 2003;40(4):196–203. doi:10.3928/0191-3913-20030701-05
3. Benavente CA, Dyer MA. Genetics and epigenetics of human retinoblastoma. Annu Rev Pathol. 2015;10:547–562. doi:10.1146/annurev-pathol-012414-040259
4. MacCarthy A, Birch JM, Draper GJ, et al. Retinoblastoma: treatment and survival in Great Britain 1963 to 2002. Br J Ophthalmol. 2009;93(1):38–39. doi:10.1136/bjo.2008.139626
5. Fernandes AG, Pollock BD, Rabito FA. Retinoblastoma in the United States: a 40-year incidence and survival analysis. J Pediatr Ophthalmol Strabismus. 2018;55(3):182–188. doi:10.3928/01913913-20171116-03
6. Al-Nawaiseh I, Jammal HM, Khader YS, et al. Retinoblastoma in Jordan, 2003–2013: ocular survival and associated factors. Ophthalmic Epidemiol. 2014;21(6):406–411. doi:10.3109/09286586.2014.967781
7. Gichigo EN, Kariuki-Wanyoike MM, Kimani K, et al. Retinoblastom in Kenia: Überleben und prognostische Faktoren [Retinoblastoma in Kenya: survival and prognostic factors]. Ophthalmologe. 2015;112(3):255–260. doi:10.1007/s00347-014-3123-z
8. Joko-Fru WY, Parkin DM, Borok M, et al. Survival from childhood cancers in Eastern Africa: a population-based registry study. Int J Cancer. 2018;143(10):2409–2415. doi:10.1002/ijc.31723
9. Narang S, Mashayekhi A, Rudich D, Shields CL. Predictors of long-term visual outcome after chemoreduction for management of intraocular retinoblastoma. Clin Exp Ophthalmol. 2012;40(7):736–742. doi:10.1111/j.1442-9071.2012.02757.x
10. He H, Qin M. Long non-coding RNA LEF1-AS1 is involved in the progression of retinoblastoma through regulating the Wnt/β-catenin pathway. Clin Exp Pharmacol Physiol. 2020;47(5):886–891. doi:10.1111/1440-1681.13263
11. Ding F, Jiang K, Sheng Y, et al. LncRNA MIR7-3HG executes a positive role in retinoblastoma progression via modulating miR-27a-3p/PEG10 axis. Exp Eye Res. 2020;193:107960. doi:10.1016/j.exer.2020.107960
12. Wang L, Wang C, Wu T, et al. Long non-coding RNA TP73-AS1 promotes TFAP2B-mediated proliferation, metastasis and invasion in retinoblastoma via decoying of miRNA-874-3p. J Cell Commun Signal. 2020. doi:10.1007/s12079-020-00550-x
13. Yang F, Zhang L, Huo XS, et al. Long noncoding RNA high expression in hepatocellular carcinoma facilitates tumor growth through enhancer of zeste homolog 2 in humans. Hepatology. 2011;54(5):1679–1689. doi:10.1002/hep.24563
14. Cui C, Zhai D, Cai L, et al. Long noncoding RNA HEIH promotes colorectal cancer tumorigenesis via counteracting miR-939‒mediated transcriptional repression of Bcl-xL. Cancer Res Treat. 2018;50(3):992–1008. doi:10.4143/crt.2017.226
15. Meng H. Long noncoding RNA HEIH promotes melanoma cell proliferation, migration and invasion via inhibition of miR-200b/a/429. Biosci Rep. 2017;37(3):BSR20170682. doi:10.1042/BSR20170682
16. Wang D, You D, Pan Y, et al. Downregulation of lncRNA-HEIH curbs esophageal squamous cell carcinoma progression by modulating miR-4458/PBX3. Thorac Cancer. 2020.
17. Zhao J, Meng R, Yao Q, et al. Long non-coding RNA HEIH promotes breast cancer development via negative modulation of microRNA-200b. Pharmazie. 2019;74(8):471–476.
18. Wan T, Wang H, Gou M, et al. LncRNA HEIH promotes cell proliferation, migration and invasion in cholangiocarcinoma by modulating miR-98-5p/HECTD4. Biomed Pharmacother. 2020;125:109916. doi:10.1016/j.biopha.2020.109916
19. ENCODE Project Consortium. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature. 2007;447(7146):799–816.
20. Guttman M, Amit I, Garber M, et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature. 2009;458(7235):223–227. doi:10.1038/nature07672
21. Yang L, Zhang L, Lu L, et al. Long noncoding RNA SNHG16 sponges miR-182-5p and miR-128-3p to promote retinoblastoma cell migration and invasion by targeting LASP1. Onco Targets Ther. 2019;12:8653–8662. doi:10.2147/OTT.S212352
22. Li S, Wen D, Che S, et al. Knockdown of long noncoding RNA 00152 (LINC00152) inhibits human retinoblastoma progression. Onco Targets Ther. 2018;11:3215–3223. doi:10.2147/OTT.S160428
23. Yang L, Zhang L, Lu L, et al. lncRNA UCA1 increases proliferation and multidrug resistance of retinoblastoma cells through downregulating miR-513a-5p. DNA Cell Biol. 2020;39(1):69–77. doi:10.1089/dna.2019.5063
24. Qi D, Wang M, Yu F. Knockdown of lncRNA-H19 inhibits cell viability, migration and invasion while promotes apoptosis via microRNA-143/RUNX2 axis in retinoblastoma. Biomed Pharmacother. 2019;109:798–805. doi:10.1016/j.biopha.2018.10.096
25. Xu S, Hui L, Yang N, et al. Upregulation of microRNA-194-5p inhibits hypopharyngeal carcinoma cell proliferation, migration and invasion by targeting SMURF1 via the mTOR signaling pathway. Int J Oncol. 2019;54(4):1245–1255.
26. Wang Y, Sun G, Wang C, et al. MiR-194-5p inhibits cell migration and invasion in bladder cancer by targeting E2F3. J BUON. 2018;23(5):1492–1499.
27. Zhu X, Li D, Yu F, et al. miR-194 inhibits the proliferation, invasion, migration, and enhances the chemosensitivity of non-small cell lung cancer cells by targeting forkhead box A1 protein. Oncotarget. 2016;7(11):13139–13152. doi:10.18632/oncotarget.7545
28. Yang F, Xiao Z, Zhang S. Knockdown of miR-194-5p inhibits cell proliferation, migration and invasion in breast cancer by regulating the Wnt/β-catenin signaling pathway. Int J Mol Med. 2018;42(6):3355–3363.
29. Do K, Doroshow JH, Wee KS. kinase as a target for cancer therapy. Cell Cycle. 2013;12(19):3159–3164. doi:10.4161/cc.26062
30. Wang L, Su K, Wu H, et al. LncRNA SNHG3 regulates laryngeal carcinoma proliferation and migration by modulating the miR-384/WEE1 axis. Life Sci. 2019;232:116597. doi:10.1016/j.lfs.2019.116597
31. Wu M, Li X, Liu Q, et al. miR-526b-3p serves as a prognostic factor and regulates the proliferation, invasion, and migration of glioma through targeting WEE1. Cancer Manag Res. 2019;11:3099–3110. doi:10.2147/CMAR.S192361
32. Fu J, Cai H, Wu Y, et al. Elevation of FGD5-AS1 contributes to cell progression by improving cisplatin resistance against non-small cell lung cancer cells through regulating miR-140-5p/WEE1 axis. Gene. 2020:144886. doi:10.1016/j.gene.2020.144886
33. Shin SS, Hwang B, Muhammad K, et al. Nimbolide represses the proliferation, migration, and invasion of bladder carcinoma cells via Chk2-mediated G2/M phase cell cycle arrest, altered signaling pathways, and reduced transcription factors-associated MMP-9 expression. Evid Based Complement Alternat Med. 2019;2019:3753587. doi:10.1155/2019/3753587
34. Li D, Tang X, Li M, et al. Long noncoding RNA DLX6-AS1 promotes liver cancer by increasing the expression of WEE1 via targeting miR-424-5p. J Cell Biochem. 2019;120(8):12290–12299. doi:10.1002/jcb.28493
35. Li P, Yang Y, Liu H, et al. MiR-194 functions as a tumor suppressor in laryngeal squamous cell carcinoma by targeting Wee1. J Hematol Oncol. 2017;10(1):32. doi:10.1186/s13045-017-0402-6
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.