Back to Journals » OncoTargets and Therapy » Volume 13
Long Noncoding RNA ADAMTS9-AS2 Inhibits the Proliferation, Migration, and Invasion in Bladder Tumor Cells
Authors Zhang Z, Jia JP, Zhang YJ, Liu G, Zhou F, Zhang BC
Received 13 January 2020
Accepted for publication 29 May 2020
Published 21 July 2020 Volume 2020:13 Pages 7089—7100
DOI https://doi.org/10.2147/OTT.S245826
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Sanjay Singh
Zhan Zhang,1,* Jin-Peng Jia,2,* Yin-Jiang Zhang,3,* Gang Liu,4,5 Fan Zhou,6 Bi-Cheng Zhang7
1Department of Rehabilitation Medicine, Changhai Hospital, Second Military Medical University, Shanghai 200433, People’s Republic of China; 2Department of Orthopaedics, General Hospital of Chinese People’s Liberation Army, Beijing 100853, People’s Republic of China; 3School of Pharmacy, Minzu University of China, Beijing 100081, People’s Republic of China; 4Department of Urology, Huangshi Central Hospital, Affiliated Hospital of Hubei Polytechnic University, Edong Healthcare Group, Huangshi 435000, Hubei, People’s Republic of China; 5Hubei Key Laboratory of Kidney Disease Pathogenesis and Intervention, Hubei Polytechnic University, Huangshi 435000, Hubei, People’s Republic of China; 6Department of Urology, Wuhan Fourth Hospital, Pu’ai Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430033, Hubei, People’s Republic of China; 7Cancer Center, Renmin Hospital of Wuhan University, Wuhan 430060, Hubei, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Bi-Cheng Zhang Email [email protected]
Background: Bladder tumor is the fifth most prevalent tumor in men, yet its pathogenesis remains to be fully identified. Albeit a host of long noncoding RNAs (lncRNA) are emerging as new players involved in bladder tumor, the functions of many lncRNAs are still enigmatic. Reports on the deluge of studies on lncRNA ADAMTS9-AS2 have been convincingly associated with various tumors, but without mention of its roles in bladder tumor. Therefore, the roles of ADAMTS9-AS2 in bladder tumor cells were explored in our study.
Materials and Methods: Quantitative real-time PCR assays and bioinformatic tools were applied in bladder tumor cells to identify the ADAMTS9-AS2 and ADAMTS9 expression. Western blot assays were performed to obtain the protein levels of bladder tumor related key molecules. CCK8, clonogenic assay, scratch wound healing, and transwell assays were separately applied to identify the functional roles of ADAMTS9-AS2 on proliferation, migration, and invasion in bladder tumor cells.
Results: First, ADAMTS9-AS2 downregulation in bladder tumor cells was identified. Overexpression and knockdown experiments showed that ADAMTS9-AS2 expression was positively related to ADAMTS9, which is in accordance with the results from GEO database. Second, ADAMTS9-AS2 contributed to the inhibition of proliferation, migration, and invasion in bladder tumor cells. Third, ADAMTS9-AS2 was linked with PI3K/AKT/mTOR pathway related-molecules, several key autophagy, and apoptotic proteins.
Conclusion: Conjointly, our findings suggested that ADAMTS9-AS2 might function as a tumor suppressor to restrain the proliferation, migration, and invasion in bladder tumor cells. The potential mechanism of ADAMTS9-AS2 related to PI3K/AKT/mTOR signal pathway was further identified. Of note, we found that ADAMTS9-AS2 has a significant effect on several key autophagy and apoptotic proteins. Therefore, these observations will provide supportive evidence to ADAMTS9-AS2 as a potential biomarker in patients with bladder tumor.
Keywords: lncRNA ADAMTS9-AS2, proliferation, PI3K/AKT/mTOR, bladder tumor
Introduction
Bladder cancer is the fourth most common cancer in male and it develops when abnormal cells grow in an uncontrolled way.1 Patients are widely categorized as non-muscle-invasive or muscle-invasive bladder tumor.2 Common treatment options are recommended, including surgery, chemotherapy, or combined with radiation therapy, but patients often end up with a relatively poor prognosis.3 Moreover, non-muscle-invasive bladder tumor constitutes the majority of patients, more often than not, which incurs a high cost and inconvenience due to frequent recurrence.4 Therefore, exploring more prognostic biomarkers that participate in the bladder tumor cell growth, migration, and invasion are necessary for the improvement of the patients’ prognosis and life quality.
Long noncoding RNAs (lncRNAs) are a kind of noncoding RNA with more than transcripts >200 nt in length.5 Ample evidence has indicated their molecular roles in tumor cell growth, metastasis, invasion, and apoptosis.6 For example, TUG1 is involved in cell growth of small cell lung cancer;7 knockdown of GHET1 inhibits cell proliferation and invasion of colorectal cancer;8 upregulated FGF13-AS1 in breast cancer promotes invasion and metastasis.9 These studies contribute to a better understanding of the importance of dysregulated lncRNAs in various tumors. Many functions of lncRNAs in tumor biology have been discovered and investigated, but the molecular mechanisms of most lncRNAs in bladder tumors remain largely unknown.
In our study, lncRNA ADAMTS9-AS2 has drawn our attention due to recent literature reports. ADAMTS9-AS2 is known to be the antisense transcript of ADAMTS910,12 and has been identified as a tumor suppressor gene if mutated.13,14 Thus, to get in-depth knowledge of ADAMTS9-AS2, extensive researches have been reported regarding its diverse roles in tumors including clear cell renal cell carcinoma,15 esophageal cancer,16 and ovarian cancer.17 Besides, accumulating researches find that PI3K/AKT is partially accountable for tumorigenesis and metastasis in bladder cancer.18,20 It is well-known that many lncRNAs display abnormal expression in parallel with phosphorylated AKT1 (p-AKT1) activation of PI3K/AKT pathway.21,27 Therefore, we proposed a hypothesis that ADAMTS9-AS2 cooperated with ADAMTS9 and may participate in the bladder tumor progression by regulating the PI3K/AKT signal pathway.
In our study, we concentrated on the potential roles of ADAMTS9-AS2 by investigating its influence on proliferation, migration, and invasion, which may contribute to the further understanding of bladder tumor and offer new insights for its clinical application in patients.
Materials and Methods
Cell Culture and Treatment
We purchased five bladder tumor cell lines (UC3, 5637, J82, T24, and EJ) and one normal cell line SV-HUC-1 from the American Type Culture Collection (ATCC, Manassas, VA, USA). Cells were maintained in DMEM (Thermo Fisher Scientific, Waltham, MA, USA) supplemented with 10% FBS (Thermo Fisher Scientific), with a mixture of 100 U/mL of penicillin/streptomycin. Cells were cultivated at 37°C in a humidified atmosphere containing 5% CO2 in air.
Quantitative PCR (q-PCR) Assays
ADAMTS9-AS2 and ADAMTS9 expression were detected by q-PCR. Total RNA was first extracted from treated cells using the TRIzol reagent (Thermo Fisher Scientific) based on the manufacturer’s instructions. One microgram of the total RNA was purified and used as the template for cDNA synthesis using a PrimeScript RT Reagent Kit with cDNA Eraser (Takara Biotech, Dalian, China). q-PCR was conducted using SYBR Premix Ex Taq (Takara Biotech, Dalian, China) on an ABI 7900 system (Thermo Fisher Scientific). The GAPDH was used as an internal control in comparisons of gene expression. Analysis of relative gene expression data was performed using 2−ΔΔCt. The PCR primers are shown as below: ADAMTS9-AS2, forward 5ʹ-AAGAAACCCTGATGTCTGGCTGAA-3ʹ and reverse 5ʹ-GTGTTACTTGAGGAGAAAGCGAAA-3ʹ; ADAMTS9, forward: 5ʹ-TGGGTTTTCCAGTTTTCAG-3ʹ and reverse 5ʹ-GTTGATGCTAAAACGACCC-3ʹ; GAPDH, forward 5ʹ- ACGGATTTGGTCGTATTGGGCG-3ʹ and reverse 5ʹ-GCTCCTGGAAGATGGTGATGGG-3ʹ.
Plasmid Construction and Cell Transfection
The pcDNA3.1 vector was purchased from Invitrogen (Thermo Fisher Scientific) and small interfering RNAs (siRNAs) were obtained from Ribobio (Guangzhou, China). pcDNA 3.1-ADAMTS9-AS2 (pcDNA 3.1-AS2) was constructed by cloning the fragment of ADAMTS9-AS2 into a pcDNA3.1 vector at the BamHI-EcoRI sites. si-ADAMTS9-AS2-1 and si-ADAMTS9-AS2-2 (si-AS2-1 and si-AS2-2) and the negative control siRNA (si-NC) were provided by Ribobio (Guangzhou, China). Cells were plated to the 12-well plates at a density of 2×103 cells per well and cultured at least 24 h before transfection. Cells were transfected with pcDNA3.1 or pcDNA3.1-AS2, and si-NC, si-AS2-1 or si-AS2-2 by using LipofectamineTM 2000 (Thermo Fisher Scientific) following the manufacturer’s instructions. Cells were incubated for 24 h and then collected for further assays after transfection. The target sequences were listed as follows: pcDNA3.1-AS2, forward 5ʹ-GGGGTACCAAACTTGACGTACACACG-3ʹ and reverse 5ʹ-AGCCGGAATTCTTTTCTGTTTTTATAATGTAC-3ʹ; si-AS2-1 5ʹ-GCATGACGCAACTTTGCTA-3ʹ (sense), si-AS2-2 5ʹ-CCTGTCTACAGGCTGATAT-3ʹ (sense).
Cell Proliferation and Clonogenic Assays
The CCK8 assays (Dojindo Molecular Technologies, Inc., Kumamoto, Japan) and clonogenic assays were conducted to evaluate cell proliferation ability. As for the CCK8 assays, cells were cultured at a density of 4×104 cells/well in 96-well culture plates for 18 h prior to transfection. Given the transfection efficiency of si-AS2-2 was higher than si-AS2-1, si-AS2-2 was selected in the experiments below. Cells were transfected with pcDNA 3.1 or pcDNA 3.1-AS2, and si-NC or si-AS2-2. Cells were treated with 10 mL/well of CCK8 solution during the last four hours of culture. The OD value was plotted using the 450 nm absorbance by Multiskan FC (Thermo Fisher Scientific) at each time point. In terms of clonogenic assays, in brief, cells at a density of 500 cells/well were cultured in six-well plates and cultured for two weeks under 37°C containing 5% CO2. The colonies were fixed with 4% paraformaldehyde for 10 min, stained with 0.1% crystal violet for 15 min and washed twice with PBS. The number of clones of each well were calculated with ImageJ software (National Institutes of Health, Bethesda, Maryland, USA). The clone formation rate of each well was counted. All experiments were performed in triplicate.
Cell Migration Assays
The scratch wound healing assay was applied to observe cell migration ability. Cells were cultured at a density of 1×106 cells/well in six-well culture plates for 18 h. Uniform wounds were generated prior to transfection (pcDNA 3.1 or pcDNA 3.1-AS2; si-NC or si-AS2-2). Microphotographs were taken at 0, 24, 48, and 72 h after transfection, and images were obtained under a magnification of 40× for each group. All experiments were performed in triplicate.
Cell Invasion Assays
The invasive ability of transfected cells was determined by transwell assay. Cells were cultured in 100 μL serum-free DMEM after transfection for 48 h. Transwell insert chamber coated with Matrigel (BD Biosciences, San Jose, CA, USA) was used according to manufacturer’s instruction. By contrast, the migrated cells on the bottom side were fixed with 4% polyoxymethylene and stained with 0.1% crystal violet for 20 min at 4°C. The invading cells at 100× magnification were counted using the Adobe Photoshop CS5. All experiments were performed in triplicate.
Western Blot Assays
Total cell lysates were prepared with sample buffer and boiled at 95°C for five minutes. Protein samples were transferred to 10% polyacrylamide gel electrophoresis (SDS-PAGE) for three hours, and then the samples were moved to polyvinylidene fluoride (PVDF). After incubation with primary antibodies (1:10,000) for ADAMTS9, AKT, p-AKT1, PIK3CB, mTOR, p-mTOR, LC3, BECN1, SQSTM1, Bax and Bcl-2 at 4°C overnight. All these antibodies were purchased from ABclonal (China). Peroxidase-conjugated goat anti-mouse (1:10,000) was used as a secondary antibody for GAPDH and BECN1 at 37°C for one hour. Peroxidase-conjugated goat anti-rabbit IgG (1:10,000) were used as a secondary antibody for other proteins at 37°C for one hour. The PVDF membrane was developed using ECL chemiluminescent reagent (EMD Millipore, Billerica, MA, USA). GAPDH was used as an internal control. The images were obtained using Bio-Rad Gel Doc XR+ system (Bio-Rad Laboratories Inc., Hercules, CA, USA). Each sample was examined in triplicate.
Gene Expression Omnibus (GEO) Data Extraction
Using GEO database (https://www.ncbi.nlm.nih. gov/geo/), the clinical samples of bladder cancer were obtained. First, the gene expression data sets of bladder cancer and adjacent tissues were processed by R language. Second, the matrix of gene expressions was generated and uploaded to Gene Set Enrichment Analysis (GSEA) platform (http://software.broadinstitute.org/gsea/index.jsp) for enrichment analysis. Further, the expression profiles, pathway enrichment and the gene enrichment maps were obtained, separately. Finally, the correlation coefficient map between of ADAMTS9-AS2 and ADAMTS9 was calculated by Cor function in the R-language.
Statistical Analysis
All data were analyzed with GraphPad Prism 7.0 software (GraphPad Software Inc., La Jolla, CA, USA) and the data were expressed as mean ±SD. Student’s t-test was used to analyze differences between two groups. All experiments were performed in triplicate. P <0.05 was considered to indicate a statistically significant difference.
Results
Downregulation of ADAMTS9-AS2 in Bladder Tumor Cells
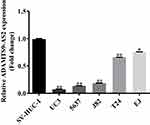
ADAMTS9-AS2 expression was determined in one normal cell line SV-HUC-1 and five bladder tumor cell lines (UC3, 5637, J82, T24, and EJ) using qRCR. Compared to the normal cell line SV-HUC-1, ADAMTS9-AS2 expression was downregulated in UC3, 5637, J82, T24, and EJ cells, apparently in UC3 and 5637 (Figure 1, P <0.001). Thus, bladder tumor cell lines UC3 and 5637 were chosen for the functional experiments. Besides, the ADAMTS9 expression was also identified using qRCR. It suggested that ADAMTS9 was downregulated in UC3, 5637, J82, T24, and EJ cell lines (Figure S1). The results suggested that ADAMTS9-AS2 and ADAMTS9 were downregulated in bladder tumor cells.
ADAMTS9-AS2 was Positively Related to ADAMTS9 in 5637 and UC3 Cells
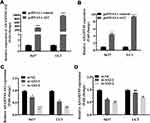
Using the GEO database, a weak positive correlation with 0.42 has been observed between ADAMTS9-AS2 and ADAMTS9 (Figure S2). Importantly, to pinpoint the relationship between ADAMTS9-AS2 and ADAMTS9, ADAMTS9-AS2 knockdown and overexpression experiments were conducted separately. ADAMTS9-AS2 overexpression experiments were conducted in the first place. After pcDNA3.1 control and pcDNA3.1-AS2 were transfected into 5637 and UC3 cell lines, respectively, ADAMTS9-AS2 expression was rising in parallel with increasing ADAMTS9 expression (Figure 2A and B; P <0.01). For ADAMTS9-AS2 knockdown experiments, si-NC, si-AS2-1 and si-AS2-2 were transfected into 5637 and UC3 cell lines, respectively, which showed that downregulated ADAMTS9-AS2 resulted in the reduction of ADAMTS9 expression and no significant changes in UC3 cells transfected with si-AS2-1 group (Figure 2C and D; P <0.05). The results suggested that ADAMTS9-AS2 was positively related to ADAMTS9 in bladder tumor cells.
ADAMTS9-AS2 Inhibits the Proliferation Ability of 5637 and UC3 Cells
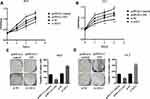
To further explore the effect of ADAMTS9-AS2 on proliferation ability, ADAMTS9-AS2 overexpression and knockdown experiments in 5637 and UC3 cell lines were performed. The CCK8 and clonogenic assays demonstrated that in cells transfected with pcDNA3.1-AS2 group, the proliferation ability was curbed compared with those transfected with pcDNA3.1 control group (Figure 3A–D; P <0.05), while it was promoted in cells transfected with si-AS2-2 as compared to si-NC group (Figure 3A–D; P <0.05). The results suggested that ADAMTS9-AS2 inhibits the proliferation ability of bladder tumor cells.
ADAMTS9-AS2 Suppresses the Migration Ability of 5637 and UC3 Cells
The scratch wound healing assays showed that the migration ability in 5637 and UC3 cells transfected with the pcDNA 3.1-AS2 group was restrained in comparison with those in pcDNA 3.1 group (Figure 4A–D). On the flip side, cells were transfected with si-NC or si-AS2-2 to further confirm the influence of ADAMTS9-AS2, which showed that cell migration ability in the si-AS2-2 group was strengthened compared to si-NC (Figure 4A–D). Concisely, ADAMTS9-AS2 suppresses the migration ability of bladder tumor cells.
ADAMTS9-AS2 Restrains the Invasion Ability of 5637 and UC3 Cells
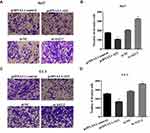
The transwell assays found that the invasion ability of 5637 and UC3 in cells transfected with the pcDNA 3.1-AS2 group had abated compared to the corresponding NC group (Figure 5A–D; P <0.01). Cells were transfected with si-NC or si-AS2-2 to further identify the roles of ADAMTS9-AS2. The knockdown experiments in 5637 and UC3 cell lines suggested that cell invasion ability in si-AS2-2 groups were enhanced compared to their corresponding si-NC group (Figure 5A–D; P <0.01). Collectively, these findings suggested that ADAMTS9-AS2 inhibits the proliferation, migration, and invasion ability of bladder tumor cells.
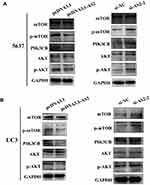
ADAMTS9-AS2 Blocked PI3K/AKT/mTOR Signal Pathway in 5637 and UC3 Cells
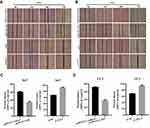
As mentioned earlier, the activation of PI3K/AKT was closely related to abnormal expression of a large number of lncRNAs.21,27 To investigate the potential mechanism of ADAMTS9-AS2 in bladder cancer, therefore, the associations between ADAMTS9-AS2 and PI3K/AKT/mTOR pathway related markers were analyzed. Western blot assays were first conducted to analyze the expression of p-AKT, PIK3CB and p-mTOR at phosphorylation level. ADAMTS9-AS2 overexpression experiment or knockdown experiment were conducted in 5637 and UC3 cell lines. It showed that the expressions of p-AKT, PIK3CB and p-mTOR were inhibited in cells transfected with pcDNA 3.1-AS2 group compared to pcDNA 3.1 group (Figure 6A and B). ADAMTS9-AS2 knockdown experiment suggested that the expression of p-AKT, PIK3CB and p-mTOR was enhanced in the si-AS2-2 group compared to si-NC group (Figure 6A and B). ADAMTS9-AS2 overexpression or knockdown had no significant effects on AKT1 and mTOR (Figure 6A and B). These results suggested that ADAMTS9-AS2 might block the PI3K/AKT/mTOR signal pathway in bladder cancer cells.
ADAMTS9-AS2 was Essential to Autophagy and Apoptosis in 5637 and UC3 Cells
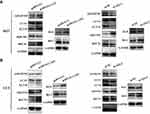
The PI3K/AKT/mTOR pathway has been widely explored to reflect the apoptosis process.28,30 In fact, the mechanistic target of rapamycin (mTOR) serves as a central controller of cell growth signaling to autophagy.31 To further reveal the functional roles of ADAMTS9-AS2, its effect on autophagy and apoptosis was investigated. Similarly, ADAMTS9-AS2 overexpression and knockdown experiments were conducted in 5637 and UC3 cell lines. Expressions of several key autophagy (LC3-I, LC3-II, BECN1, and SQSTM1) and apoptotic proteins (Bax and Bcl-2) were observed by Western blot assays.
For ADAMTS9-AS2 overexpression experiment, cells were transfected with pcDNA 3.1 or pcDNA 3.1-AS2. We found that ADAMTS9, LC3-II, BECN1, and proapoptotic Bax expression were increased, but SQSTM1 and antiapoptotic Bcl-2 expression were decreased in pcDNA 3.1-AS2 group compared to their corresponding pcDNA 3.1 group (Figure 7A and B). For the ADAMTS9-AS2 knockdown experiment, we noticed that ADAMTS9, LC3-II, BECN1, and proapoptotic Bax expression were reduced in cells transfected with si-AS2-2 group compared with si-NC, while SQSTM1 and antiapoptotic Bcl-2 expression were elevated in si-AS2-2 group compared to si-NC group (Figure 7A and B). We did not detect obvious impact on LC3-I in ADAMTS9-AS2 overexpression or knockdown experiment (Figure 7A and B). All these results indicate a potential regulatory mechanism of ADAMTS9-AS2 involved in autophagy and apoptosis in bladder tumor cells.
Discussion
It has been well-recognized that a host of lncRNAs were closely related to the pathogenesis of bladder cancer.32 Nevertheless, the roles of ADAMTS9-AS2 in the bladder tumor remain largely unknown. It is well known that ADAMTS9-AS2 is an antisense transcript of ADAMTS9.11,33 A growing number of research has shown that a pair of sense and antisense transcripts commonly plays a pivotal role in the tumors, like SPINT1-AS1 and SPINT1 in colorectal cancer,34 FOXF1-AS1 and FOXF1 in osteosarcoma cells;35 AFAP1-AS1 and AFAP1 in breast cancer;36 MAPT-AS1 and MAPT in breast cancer.37 From the GEO analysis results, red represents positive correlation, which indicated that ADAMTS9-AS2 and ADAMTS9 had a weak positive correlation. Therefore, the potential association of ADAMTS9-AS2 and ADAMTS9 gave rise to the insights into the investigation of bladder cancer. Using the qPCR assays, ADAMTS9-AS2 expressions in one normal cell line SV-HUC-1 and five bladder cancer cell lines (UC3, 5637, J82, T24, and EJ) were first identified. Because ADAMTS9-AS2 was apparently downregulated in the 5637 and UC3 cell lines, the two cell lines were applied in following assays. From the ADAMTS9-AS2 overexpression and knockdown experiments, the association between ADAMTS9-AS2 and ADAMTS9 were identified. We found that ADAMTS9-AS2 was positively related to ADAMTS9 in 5637 and UC3 cells, which is in line with the previous study.38 Albeit the diverse roles of ADAMTS9-AS2 has been reported in a variety of tumors,15,17 it still remains largely unclear whether ADAMTS9-AS2 could bring out the effects on bladder tumor cells. To identify the functional roles of ADAMTS9-AS2 in bladder tumor cells, CCK8, clonogenic assay, scratch wound healing, and transwell assay were performed, respectively. We found that ADAMTS9-AS2 overexpression inhibits the proliferation, migration, and invasion ability in bladder cancer lines 5637 and UC3. Conversely, its knockdown had positive results. Given the information from GEO database and several pairs of sense and antisense related studies,34,37 we speculated that ADAMTS9-AS2 might play a role in bladder tumor by regulating the ADAMTS9 expression. Besides, ADAMTS9 functions as a critical player in the process of AKT/mTOR signaling pathway as evidenced by Chen et al.33,39 Noticeably, the activation of the PI3K/AKT/mTOR signaling pathway is widely recognized as an essential part of regulatory mechanism in tumors including bladder cancer.40,42 As previously mentioned, ADAMTS9-AS2 expression was correspondent with ADAMTS9. We thus proceeded with deep investigation concerning the association of ADAMTS9-AS2 and PI3K/Akt/mTOR pathway related markers. Results based on Western blot assays indicated that ADAMTS9-AS2 inhibited p-AKT, PIK3CB, and p-mTOR. Previous study noticed that PI3K/Akt/mTOR was closely related to autophagy and apoptosis.31,43 The autophagy related several key markers (LC3-I, LC3-II, BECN1, and SQSTM1) and apoptotic proteins (Bax and Bcl-2) were further explored. It suggested that ADAMTS9-AS2 promoted LC3-II, BECN1, and proapoptotic Bax, but inhibited SQSTM1 and antiapoptotic Bcl-2 expression. The data we present might highlight the biological functions of ADAMTS9-AS2 in bladder tumor. In addition, we further explored the reasonable mechanism that ADAMTS9-AS2 inhibited the proliferation, migration, and invasion via PI3K/Akt/mTOR signaling pathway. Nevertheless, our study exists some limitations, such as, functional experiments were performed only in two cell lines; the precise molecular mechanism underlying the ADAMTS9-AS2 and ADAMTS9 require further investigation. Therefore, further studies will be considered to reveal the specific mechanisms of ADAMTS9-AS2 in bladder tumor.
Conclusion
Our results found the functional roles and potential mechanisms of ADAMTS9-AS2 in bladder tumor. It suggested that ADAMTS9-AS2 could inhibit cell proliferation, migration, and invasion ability in bladder tumor. Our work might provide valuable insights into a possible mechanism that ADAMTS9-AS2 is a tumor suppressor involved in the PI3K/AKT/mTOR signal pathway causing cell autophagy and apoptosis in bladder tumor.
Abbreviations
lncRNA, long non-coding RNA; siRNA, small interfering RNA; q-PCR, quantitative PCR.
Disclosure
The authors confirmed no conflicts of interest in this work.
References
1. Tyagi A, Agarwal C, Glode LM, Agarwal R. Silibinin efficacy in human bladder transitional cell carcinoma: modulation of cell cycle and apoptosis cascades. Cancer Res. 2004;64:730.
2. Tinay I, Mungan NA. Management of Non-muscle-Invasive and Muscle-Invasive Bladder Cancers. Principles and Practice of Urology. 2017;59-74.
3. Kanady KE, Shipley WU, Zietman AL, Kaufman DS, Althausen AF, Heney NM. Treatment strategies using transurethral surgery, chemotherapy, and radiation therapy with selection that safely allows bladder conservation for invasive bladder cancer. Semin Surg Oncol. 1997;13(5):359–364. doi:10.1002/(SICI)1098-2388(199709/10)13:5<359::AID-SSU10>3.0.CO;2-I
4. Barocas DA, Globe DR, Colayco DC, et al. Surveillance and treatment of non-muscle-invasive bladder cancer in the USA. Adv Urol. 2012;2012:1–8. doi:10.1155/2012/421709
5. Yang Q, Jia L, Li X, et al. Long noncoding RNAs: new players in the osteogenic differentiation of bone marrow- and adipose-derived mesenchymal stem cells. Stem Cell Rev Rep. 2018;14(7414):1–12. doi:10.1007/s12015-018-9801-5
6. Sültmann H, Diederichs S. Long noncoding RNA: “LNCs” to cancer. Eur Urol. 2014;65(6):1152–1153. doi:10.1016/j.eururo.2014.01.032
7. Niu Y, Ma F, Huang W, et al. Long non-coding RNA TUG1 is involved in cell growth and chemoresistance of small cell lung cancer by regulating LIMK2b via EZH2. Mol Cancer. 2017;16(1):5. doi:10.1186/s12943-016-0575-6
8. Zhou J, Li X, Wu M, Lin C, Guo Y, Tian B. Knockdown of long noncoding RNA GHET1 inhibits cell proliferation and invasion of colorectal cancer. Oncol Res. 2016;23(6):303–309. doi:10.3727/096504016X14567549091305
9. Ma F, Liu X, Zhou S, et al. Long non-coding RNA FGF13-AS1 inhibits glycolysis and stemness properties of breast cancer cells through FGF13-AS1/IGF2BPs/Myc feedback loop. Cancer Lett. 2019;450:63–75.
10. Liu C, Yang Z, Deng Z, et al. Upregulated lncRNA ADAMTS9-AS2 suppresses progression of lung cancer through inhibition of miR-223-3p and promotion of\r TGFBR3. IUBMB Life. 2018;70(6):536–546.
11. Yao J, Zhou B, Zhang J, et al. A new tumor suppressor LncRNA ADAMTS9-AS2 is regulated by DNMT1 and inhibits migration of glioma cells. Tumor Biol. 2014;35(8):7935–7944. doi:10.1007/s13277-014-1949-2
12. Barreto-Luis A, Pino-Yanes M, Corrales A, et al. Genome-wide association study in Spanish identifies ADAM metallopeptidase with thrombospondin type 1 motif, 9 (ADAMTS9), as a novel asthma susceptibility gene. J Allergy Clin Immunol. 2016;137(3):964–966. doi:10.1016/j.jaci.2015.09.051
13. Zhu N, Hou J, Wu Y, et al. Integrated analysis of a competing endogenous RNA network reveals key lncRNAs as potential prognostic biomarkers for human bladder cancer. Medicine. 2018;97(35):e11887. doi:10.1097/MD.0000000000011887
14. LI Q, Dai Y, Wang F, Hou S. Differentially expressed long non-coding RNAs and the prognostic potential in colorectal cancer. Neoplasma. 2016;63(06):977–983. doi:10.4149/neo_2016_617
15. EL S, L X, L W, et al. LncRNA ADAMTS9-AS2 inhibits cell proliferation and decreases chemoresistance in clear cell renal cell carcinoma via the miR-27a-3p/FOXO1 axis. Aging. 2019;11(15):5705.
16. Liu D, Wu K, Yang Y, Zhu D, Zhang C, Zhao S. Long noncoding RNA ADAMTS9-AS2 suppresses the progression of esophageal cancer by mediating CDH3 promoter methylation. Mol Carcinog. 2020;59(1):32–44.
17. Wang A, Jin C, Li H, Qin Q, Li L. LncRNA ADAMTS9-AS2 regulates ovarian cancer progression by targeting miR-182-5p/FOXF2 signaling pathway. Int J Biol Macromol. 2018;120:1705–1713.
18. Meng C, Jian G, Delclos GL, et al. Genetic variations of the PI3K–AKT–mTOR pathway and clinical outcome in muscle invasive and metastatic bladder cancer patients. Carcinogenesis. 2010;8:8.
19. Meng C, Jian G, Grossman H, et al. Abstract #1983: genetic variations in PI3K-AKT-mTOR pathway and bladder cancer clinical outcome. Cancer Res. 2009;69(4):1553–1554. doi:10.1158/0008-5472.CAN-08-1921
20. Calderaro J, Rebouissou S, Koning LD, Masmoudi A, Hérault A. PI3K/AKT pathway activation in bladder carcinogenesis. Int J Cancer. 2014;134(8):1776–1784. doi:10.1002/ijc.28518
21. Huang Y, Zhang J, Hou L, et al. LncRNA AK023391 promotes tumorigenesis and invasion of gastric cancer through activation of the PI3K/Akt signaling pathway. J Exp Clin Cancer Res. 2017;36(1):194. doi:10.1186/s13046-017-0666-2
22. Koirala P, Huang J, Ho -T-T, Wu F, Ding X, Mo -Y-Y. LncRNA AK023948 is a positive regulator of AKT. Nat Commun. 2017;8(1):14422. doi:10.1038/ncomms14422
23. Yan J, Dang Y, Liu S, Zhang Y, Zhang G. LncRNA HOTAIR promotes cisplatin resistance in gastric cancer by targeting miR-126 to activate the PI3K/AKT/MRP1 genes. Tumor Biol. 2016;37(12):16345–16355.
24. Wang Y, Kuang H, Xue J, Liao L, Yin F, Zhou X. LncRNA AB073614 regulates proliferation and metastasis of colorectal cancer cells via the PI3K/AKT signaling pathway. Biomed Pharmacother. 2017;93:1230–1237. doi:10.1016/j.biopha.2017.07.024
25. Wei S, Mei JZ, Zhang M. Long noncoding RNA PlncRNA-1 promotes colorectal cancer cell progression by regulating the PI3K/Akt signaling pathway. Oncol Res. 2018;26(2):261–268.
26. Guo HM, Yang SH, Zhao SZ, Li L, Yan MT, Fan MC. LncRNA NEAT1 regulates cervical carcinoma proliferation and invasion by targeting AKT/PI3K. Eur Rev Med Pharmacol Sci. 2018;22(13):4090–4097.
27. Sun L, Wang L, Chen T, Yao B, Wang Y. microRNA-1914, which is regulated by lncRNA DUXAP10, inhibits cell proliferation by targeting the GPR39-mediated PI3K/AKT/mTOR pathway in HCC. J Cell Mol Med. 2019;23(12):8292–8304. doi:10.1111/jcmm.14705
28. Gao N, Zhang Z, Jiang BH, Shi X. Role of PI3K/AKT/mTOR signaling in the cell cycle progression of human prostate cancer. Biochem Biophys Res Commun. 2003;310(4):1132.
29. Liu S, Wang X-J, Liu Y, Cui Y-F. PI3K/AKT/mTOR signaling is involved in (-)-epigallocatechin-3-gallate-induced apoptosis of human pancreatic carcinoma cells. Am J Chin Med. 2013;41(03):629–642. doi:10.1142/S0192415X13500444
30. Pan ST, Qin Y, Zhou ZW. Plumbagin induces G2/M arrest, apoptosis, and autophagy via p38 MAPK- and PI3K/Akt/mTOR-mediated pathways in human tongue squamous cell carcinoma cells. Drug Des Devel Ther. 2015;9(default):1601–1626.
31. Li -T-T, Zhu D, Mou T, et al. IL-37 induces autophagy in hepatocellular carcinoma cells by inhibiting the PI3K/AKT/mTOR pathway. Mol Immunol. 2017;87:132–140. doi:10.1016/j.molimm.2017.04.010
32. Qiu L, Tang Q, Li G, Chen K. Long non-coding RNAs as biomarkers and therapeutic targets: recent insights into hepatocellular carcinoma. Life Sci. 2017;191:273–82.
33. Du W, Wang S, Zhou Q, et al. ADAMTS9 is a functional tumor suppressor through inhibiting AKT/mTOR pathway and associated with poor survival in gastric cancer. Oncogene. 2013;32(28):3319–3328. doi:10.1038/onc.2012.359
34. Chen L, Wei L, Yanli Z, et al. Increased expression of antisense lncRNA SPINT1-AS1 predicts a poor prognosis in colorectal cancer and is negatively correlated with its sense transcript. Onco Targets Ther. 2018;11:3969–3978.
35. Kun-Peng Z, Chun-Lin Z, Xiao-Long M. Antisense lncRNA FOXF1-AS1 promotes migration and invasion of osteosarcoma cells through the FOXF1/MMP-2/-9 pathway. Int J Biol Sci. 2017;13(9):1180–1191. doi:10.7150/ijbs.21722
36. Dianatpour A, Faramarzi S, Geranpayeh L, Mirfakhraie R, Motevaseli E, Ghafouri-Fard S. Expression analysis of AFAP1-AS1 and AFAP1 in breast cancer. Cancer Biomarkers. 2018;22(1):49–54. doi:10.3233/CBM-170831
37. Pan Y, Pan Y, Cheng Y, Yang F, Yao Z, Wang O. Knockdown of LncRNA MAPT-AS1 inhibites proliferation and migration and sensitizes cancer cells to paclitaxel by regulating MAPT expression in ER-negative breast cancers. Cell Biosci. 2018;8(1):7.
38. Wenric S, ElGuendi S, Caberg JH, et al. Transcriptome-wide analysis of natural antisense transcripts shows their potential role in breast cancer. Sci Rep. 2017;7(1):17452.
39. Chen QY, Costa M. PI3K/Akt/mTOR signaling pathway and the biphasic effect of arsenic in carcinogenesis. Mol Pharmacol. 2018;94(1):784–792. doi:10.1124/mol.118.112268
40. S A, R N. Targeting the PI3K/AKT/mTOR pathway in bladder cancer. Methods Mol Biol. 2018;1655:335–350.
41. Cidado J, Park BH. Targeting the PI3K/Akt/mTOR pathway for breast cancer therapy. J Mammary Gland Biol Neoplasia. 2012;17(3–4):205–216. doi:10.1007/s10911-012-9264-2
42. YG G, WT W, CI L, CA N, YJ W. Involvement of mitochondrial dysfunction, endoplasmic reticulum stress, and the PI3K/AKT/mTOR pathway in nobiletin-induced apoptosis of human bladder cancer cells. Molecules. 2019;24(16):2881.
43. Chen B, Li D, Li M, Li S, Zhu Q. Induction of mitochondria-mediated apoptosis and PI3K/Akt/mTOR-mediated autophagy by aflatoxin B2 in hepatocytes of broilers. Oncotarget. 2016;7(51):84989–84998. doi:10.18632/oncotarget.13356
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.