Back to Journals » International Journal of General Medicine » Volume 13
Long Non-Coding RNA LUCAT1 Promotes Progression of Thyroid Carcinoma by Reinforcing ADAM10 Expression Through Sequestering microRNA-493
Authors Xiong G, Chen J, Wu Z, He S, Lian M, Fang J
Received 22 July 2020
Accepted for publication 11 September 2020
Published 14 October 2020 Volume 2020:13 Pages 847—860
DOI https://doi.org/10.2147/IJGM.S273461
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Scott Fraser
Guofeng Xiong,1,2 Jiaming Chen,1 Zhen Wu,1 Shizhi He,1 Meng Lian,1 Jugao Fang1
1Department of Otorhinolaryngology, Head and Neck Surgery, Beijing Tongren Hospital, Capital Medical University, Beijing 100730, People’s Republic of China; 2Department of Otolaryngology, Head and Neck Surgery, Wenzhou Central Hospital, Wenzhou 325000, People’s Republic of China
Correspondence: Meng Lian; Jugao Fang Department of Otorhinolaryngology
Head and Neck Surgery, Beijing Tongren Hospital, Capital Medical University, No. 8 Chongwenmen Inner Street, Dongcheng District (East District), Beijing 100730, People’s Republic of China
Tel/Fax +86-010-58266699
Email [email protected]; [email protected]
Background: Long non-coding RNA (lncRNA) LUCAT1 has recently been recognized as an oncogene in several malignancies. This study was launched to probe its role in thyroid carcinoma (TC) development and the implicated molecules.
Methods: LUCAT1 expression in TC cell lines and in normal thyroid follicular epithelial cell line Nthy-ori3-1 was determined by RT-qPCR. Binding relationships between LUCAT1 and microRNA (miR)-493, and between miR-493 and a disintegrin and metalloproteinase-10 (ADAM10) were predicted on a bioinformatics system and then validated through luciferase reporter gene assays. Expression of miR-493 and ADAM10 in TC cells was determined. Gain- and loss-of functions of LUCAT1, miR-493 and ADAM10 were performed to explore their influences on the behaviors of TC cells. Xenograft tumors were induced in nude mice for in vivo studies.
Results: LUCAT1 and ADAM10 were highly expressed, while miR-493 was poorly expressed in TC cell lines. LUCAT1 served as a miR-493 sponge to upregulate ADAM10 expression. Silencing of LUCAT1 discouraged proliferation, invasion, and migration but triggered apoptosis of TC cells. By contrast, these changes were abrogated by further miR-493 inhibition or ADAM10 upregulation. The in vitro experiment results were reproduced in vivo. In addition, miR-493 inhibition or ADAM10 overexpression was found to increase the phosphorylation of STAT3 in cells.
Conclusion: This study evidenced that LUCAT1 increases ADAM10 expression through sequestering miR-493, leading to JAK-STAT activation and TC cell growth and metastasis. LUCAT1 and ADAM10 may serve as therapeutic targets for TC treatment.
Keywords: long non-coding RNA LUCAT1, microRNA-493, ADAM10, thyroid carcinoma, JAK-STAT signaling pathway
Introduction
Thyroid carcinoma (TC) ranks at the ninth place for incidence among all cancer cases worldwide, and though the mortality has remained stable, the incidence of this disease has been increasing greater than other cancer types with a higher tendency in females.1,2 Ionizing radiation is the only well-established risk factor for TC, especially when exposure is in childhood, although some other factors such as obesity, hormonal exposures smoking, and certain environmental causes are suggested to possibly play a role.1 Papillary thyroid carcinoma (PTC) is the most frequent TC type, accounting for approximately 85% of all cases, and the 5-year relative survival of patients with well-differentiated PTC after an indolent clinical course reached 97.5%; yet the aggressive poorly differentiated types and anaplastic carcinomas can be fatal.3 Once transferring to distant organs, the TC cells could be almost completely resistant to the current clinical regimens and consequently lead to an incurable disease and death.4 Considering the increasing incidence of this malignancy, developing novel therapeutic options for TC remains a major challenge and a focus issue in the medical field.
While no more than 2% of the human genome encodes proteins, biological roles for the so-called junk genome, termed non-coding (nc) RNAs, are being increasingly recognized to participate in the modification of the epigenetic profile of cancer cells and mediation in the expression of other RNA molecules.5 Long ncRNAs (lncRNAs) are a class of ncRNAs larger than 200 nucleotides, whose aberrant expression is frequently witnessed in multiple cancer types and closely related to the malignant phenotypical changes.6 Meanwhile, another major class of ncRNAs, microRNAs (miRNAs) are implicated in key processes in cancer onset and progression upon dysregulation, owing to their functions in primarily interacting to thousands of target mRNAs.7 Similar to protein-coding RNAs, lncRNAs can also subjected to post-transcriptional modifications.8 Therefore, a special RNA interaction network by which lncRNAs bind to miRNAs to mediate expression of specific mRNAs was proposed, termed competing endogenous RNA (ceRNA).9,10 LUCAT1 is a recently found lncRNA that has been documented to promote cancer progression through such ceRNA networks.11,12 It was suggested as a potential prognostic biomarker for patients with PTC.13 But its exact function on TC progression remains largely unknown.11 Importantly, data on a bioinformatics system (http://starbase.sysu.edu.cn/) suggested miR-493 as a sponge for LUCAT1, while a disintegrin and metalloproteinase-10 (ADAM10) was suggested as a target mRNA of miR-493. miR-493 was demonstrated as a tumor inhibitor in many malignancies14,15 with its role in TC unknown, while ADAM10 has been recognized as an oncogene whose suppression by miRNAs was reported to alleviate cancer development.16,17 Thereby, we hypothesized that LUCAT1 sponges miR-493 to mediate ADAM10 and to regulate TC progression. Dual-luciferase reporter gene assays were conducted to validate their binding relationships, and altered expression of these molecules was introduced in TC cells to evaluate their functions on cell growth in vitro and in vivo.
Materials and Methods
Clinical Sample Collection
TC tissues and the adjacent normal ones (over 2 cm away from the lesion sites) were collected from 47 TC patients who underwent surgical resection in Beijing Tongren Hospital, Capital Medical University from 2017 to 2019. All patients had complete clinical information and were free of other malignancies or a history of chemo-radiotherapy. The characteristics of tissues were confirmed by three pathologists. The tissues were collected from surgery and frozen at −80°C. The research was carried out with the approval and supervision of the Ethics Committee of Beijing Tongren Hospital, Capital Medical University and in line with the guidelines in Declaration of Helsinki. Written confirmed consent was acquired from each respondent.
Cell Culture and Treatment
Human normal thyroid follicular epithelial cell line Nthy-ori3-1 and TC cell lines 8305C, HTC-C3, B-CPAP and TPC-1 were acquired from Nanjing Cobioer Biotechnologies Co., Ltd. (Jiangsu, China). Cells were cultivated in Roswell Park Memorial Institute-1640 (Gibco, Thermo Fisher Scientific, Waltham, MA, USA) containing 10% fetal bovine serum (FBS, Gibco) and 100 mg/mL streptomycin and penicillin (100U/mL) at 37°C with 5% CO2.
Cells in good growth conditions were detached in trypsin and sorted on 24-well plates. Three batches of transfection were performed. In the first batch, the cells were transfected with short hairpin (sh)-negative control (NC), sh-LUCAT1-1, sh-LUCAT1-2 or sh-LUCAT1-3; in the second batch, the cells were transfected with overexpression (oe)-NC (empty vector), oe-LUCAT1, sh-NC, sh-LUCAT1, mimic NC, miR-493 mimic, inhibitor NC, and miR-493 inhibitor, respectively; as for the third batch, the cells were transfected with sh-NC. sh-LUCAT1, shLUCAT1 + inhibitor NC, sh-LUCAT1 + miR-493 inhibitor, sh-LUCAT1 + oe-NC, and sh-LUCAT1 + oe-ADAM10, respectively. All transfection procedures were performed using a Lipofectamine 2000 kit (Invitrogen, Thermo Fisher). Cells were cultivated in complete medium for 48 h post transfection for further use.
Dual-Luciferase Reporter Gene Assay
The binding sites between miR-493 and ADAM10-3ʹUTR or LUCAT1 were first predicted on the Starbase system (http://starbase.sysu.edu.cn/). Next, the ADAM10-3ʹUTR and LUCAT1 fragments containing the putative binding sites with miR-493 were synthetized and inserted into pGL3 vectors to construct pGL3-ADAM10-3ʹUTR wile-type (WT) and pGL3-LUCAT1-WT vectors. The mutant type (MUT) vectors were constructed based on the mutant binding sites and defined as pGL3-ADAM10-3ʹUTR-MUT and pGL3-LUCAT1-MUT vectors. Well-constructed vectors were co-transfected with miR-493 mimic or mimic NC into HEK293T cells (ATCC, Manassas, VA, USA). Forty-eight hours later, the cells were collected and lysed, and the relative luciferase activity was determined using a luciferase detection kit (K801-200, BioVision, Mountain View, CA, USA) and a Dual-Luciferase-Reporter-Gene System (Promega, Madison, WI, USA). Three independent experiments were conducted.
Cell Counting Kit-8 (CCK-8) Method
Exponentially growing cells were resuspended in 10% FBS-supplemented Dulbecco’s modified Eagle’s medium to 1 × 104 cells/mL. The cell suspension was sorted in 96-well plates at 100 μL per well with 8 duplicated wells set for each group. The plates were placed in a 37°C incubator, and each well was loaded with 10 µL CCK-8 solution (Sigma-Aldrich Chemical Company, St Louis, MO, USA) at 24, 48, and 72 h, respectively, for another 2 h of incubation. The optical density (OD) value of each well at 450 nm was determined using a microplate reader (NYW-96M, Noah Wei Instrument Co., Ltd., Beijing, China). The experiments were repeated three times.
Colony Formation Assay
Exponentially growing cells were sorted in 75 mm-culture dishes at 800 cells per dish for a 9-d incubation. After that, the cells were washed by phosphate buffer saline (PBS, pH = 6.8), fixed in methanol for 20 min, stained with 10% Giemsa for 20 min, and then rinsed by deionized water and dried. The number of cell colonies (over 20 cells) was counted under the microscope and the colony formation rate was calculated. Three independent experiments were performed.
Transwell Assays
Cell migration and invasion were determined by Transwell assays. As for invasion assay, Matrigel (YB356234, Yu Bo Biotech Co., Ltd., Shanghai, China) was mixed and diluted in an equal volume of serum-free medium at 4°C. Each apical chamber was pre-loaded with 50 µL Matrigel. Cells were resuspended in serum-free medium to 1 × 105 cells/mL and loaded to the apical chambers, while the basolateral chambers were loaded with 10% FBS-supplemented medium. The chambers were warm-incubated at 37°C for 24 h and taken out. The cells in the inner membrane were wiped out, and the invaded cells were fixed in 5% glutaraldehyde at 4°C, stained with 0.1% crystal violet for 30 min and then observed under an inverted microscope. Cell migration was performed in a similar manner but without precoating Matrigel in the apical chambers. At least four fields of views were included in microscope observation. Three independent experiments were performed.
Wound-Healing Assay
Migration of cells was further determined by a wound-healing assay. In brief, B-CPAP and TPC-1 were seeded in 6-well plates. When cells reached a 90% cell confluence, a sterile 200-μL pipette tip was used to produce scratches on the cell monolayer at an equal interval. The cells were then cultured in for 24 h in serum-free medium (Gibco), scratches, and the width of the scratches was observed under an inverted microscope (Carl Zeiss, Carl Zeiss MicroImaging, Inc., Thornwood, NY, USA)
Flow Cytometry
An Annexin V-fluorescein isothiocyanate (FITC)/propidium iodide staining kit (ab14085, Abcam Inc., Cambridge, MA, USA) was used for apoptosis detection. After transfection, cells were washed, centrifuged, and resuspended in 200 μL binding buffer, and then treated with 10 μL Annexin V-FITC and 5 μL PI for 15 min of reaction avoiding light exposure. Then, the apoptosis of cells was examined on a flow cytometer (Attune NxT, Thermo Fisher, USA) at an excitation wavelength of 488 nm, and the data were analyzed by the FlowJo software.
The cell cycle distribution was further determined. In short, the cells were detached in trypsin and fixed in 75% pre-chilled ethanol. Then, cells were washed and resuspended in 200 μL PBS, and then added with 10 μL PI and 20 μL RNaseA on ice for 35 min of incubation. Next, the cells were run on the flow cytometer and the data were analyzed by the FlowJo again.
Reverse Transcription Quantitative Polymerase Chain Reaction (RT-qPCR)
Total RNA was extracted using the TRIzol reagent (15,596,026, Invitrogen, Car, Cal, USA) and then reversely transcribed into cDNA using a PrimeScript RT Reagent Kit (RR047A, Takara, Japan). The synthetized cDNA was used for real-time qPCR using a Fast SYBR Green PCR Kit (Applied Biosystems, Foster City, CA, USA) and a qPCR System (7500, ABI Company, Oyster Bay, NY, USA). The primer sequences are presented in Table 1. U6 was served as an internal reference for miR-493 while GAPDH for other genes. Relative gene expression was evaluated by using the 2−ΔΔCt.
 |
Table 1 Primer Sequences for RT-qPCR |
Nuclear-Cytoplasmic RNA Separation
Nuclear and cytoplasmic RNA from B-CPAP and TPC-1 cells was separated using a PARIS kit (Invitrogen) as per the kit’s instructions. Then, the expression of RNA sample was determined by RT-qPCR, and GAPDH and U6 were used as the control for cytoplasmic RNA and nuclear RNA.
Western Blot Analysis
Cells were lysed in radio-immunoprecipitation assay cell lysis buffer (Boster Biological Technology Co., Ltd., Wuhan, Hubei, China) to collect the total protein of cells. The protein concentration was measured using a bicinchoninic acid method. Then, the protein was run on SDS-PAGE and transferred onto polyvinylidene fluoride membranes. Following 2 h of 5% bovine serum albumin treatment to block non-specific binding, the membranes were incubated with the primary antibodies to B-cell lymphoma-2 (Bcl-2, 1:1000, ab32124, Abcam) and Bcl-2-associated X (Bax, 1:1000, ab32503, Abcam), ADAM10 (1:1000, ab1997, Abcam), N-cadherin (1:1000, ab18203, Abcam), matrix metalloproteinase-9 (MMP-9, #13,667, Cell Signaling Technology, Boston, UK), Signal transducer and activator of transcription 3 (STAT3, 1:1000, #30,835, CST), p-STAT3 (1:2000, #9145, CST), GAPDH (1:2000, ab9485, Abcam) at 4°C overnight, and then incubated with goat-anti rabbit secondary antibody (ab205719, 1:2000, Abcam) at 20°C for 1 h. After that, the protein blots were developed using enhanced chemiluminescence reagent (EMD Millipore, Corp. Billerica, MA, USA), and the grey value was determined using an Image J software. Three independent experiments were performed.
Xenograft Tumors in Nude Mice
Thirty-six BALB/c female nude mice (3–4 weeks old, 14 ± 2 g) purchased from SLAC Laboratory Animal Co., Ltd. (Shanghai, China) were acclimated in a suitable condition (25–27°C, 45–50% humidity) for one week and allowed standard chow pellets and water ad libitum. The mice were allocated into 6 groups: sh-NC, sh-LUCAT1, sh-LUCAT1 + inhibitor NC, sh-LUCAT1 + miR-493 inhibitor, sh-LUCAT1 + oe-NC, and sh-LUCAT1 + oe-ADAM10 groups. Well-transfected cells were adjusted to 1 × 107 cells/mL, and each mouse was implanted with 20 μL cell suspension. Then, the growth of xenograft tumors was monitored and recorded. The volume (V) of tumors was determined every week as the following formula: V = a × b2/2, where ‘a’ indicates the length while ‘b’ indicates the width. On the 42nd d, the mice were euthanized through an intraperitoneal injection of overdose of pentobarbital sodium (150 mg/kg) to collect and weigh the tumors. The study was ratified by the Ethics Committee of Beijing Tongren Hospital, Capital Medical University. Animal experiments were performed in compliance with the Ethical Guidelines for The Study of Experimental Pain in Conscious Animals. Great efforts were made to minimize the number and pain of animals.
Statistical Analysis
Data were analyzed using the SPSS 21.0 system (IBM Corp. Armonk, NY, USA). Data were exhibited as mean ± standard deviation (SD) from at least three independent experiments. Differences were compared by t-test (for two groups) and one-way or two-way analysis of variance (ANOVA, for multiple groups) followed by Tukey’s multiple comparisons test. The clinical information of patients was analyzed by Fisher’s exact test. p was obtained from two-tailed tests, and p < 0.05 was considered statistically significant.
Results
Silencing of LUCAT1 Inhibits Proliferation but Promotes Apoptosis of TC Cells
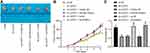
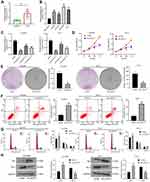
LUCAT1 (Ensembl: ENSG00000248323) was suggested as a potential prognostic biomarker for PTC patients.13 But its exact functions on TC cell behaviors and the molecular mechanisms are largely unknown. Here, we first determined expression of LUCAT1 in the collected tumor and normal samples from 47 patients. It was found that the LUCAT1 expression was notably higher in the tumor tissues than that in normal ones (Figure 1A). Then, according to the average value of LUCAT1 expression (3.07), the samples were allocated into high-LUCAT1 (n = 23) and low-LUCAT1 expression (n = 24) groups. Then, the correlation between LUCAT1 expression and the clinical parameters was analyzed (Table 2). It was found that high expression of LUCAT1 was linked to lymph node metastasis, increased tumor node metastasis stage, and poor tumor differentiation. Then, we determined LUCAT1 expression in TC cell lines (8305C, HTC-C3, B-CPAP and TPC-1) and in Nthy-ori3-1 cells. The RT-qPCR results suggested that the expression of this lncRNA was notably higher in the TC cell lines than that in Nthy-ori3-1 cells, especially in B-CPAP and TPC-1 cells, which were used for the subsequent experiments (Figure 1B).
 |
Table 2 Correlation Between LUCAT1 Expression and Clinical Parameters of TC Patients |
Next, artificial downregulation of LUCAT1 was introduced in B-CPAP and TPC1 cells lines through administrating shRNAs. Three shRNAs including sh-LUCAT1-1, sh-LUCAT1-2 and sh-LUCAT1-3 were used. It was found that sh-LUCAT1-1, sh-LUCAT1-2 and sh-LUCAT1-3 inhibited the LUCAT1 expression in both cell lines (Figure 1C). The sh-LUCAT1-1, which presented the highest interfering efficiency, was selected for further use.
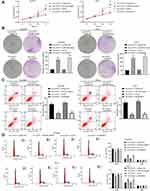
Then, the biological behaviors of cells were determined. The CCK-8 method results suggested that proliferation of B-CPAP and TPC-1 cells was decreased after LUCAT1 downregulation (Figure 1D). A similar trend was found in the colony formation ability of both cells (Figure 1E). In addition, the flow cytometry results showed that after sh-LUCAT1 administration, the apoptosis rate of B-CPAP and TPC-1 cells was increased (Figure 1F). Accordingly, the cell cycle was increasingly arrested in the G0/G1 phase (Figure 1G). The Western blot assay further evidenced that the expression of apoptotic Bax in cells was increased while the expression of anti-apoptotic Bcl-2 was decreased upon LUCAT1 downregulation (Figure 1H).
Silencing of LUCAT1 Inhibits Migration and Invasion of TC Cells
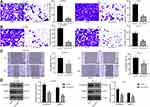
The Transwell assay results showed that silencing of LUCAT1 led to a decline in number of migrated and invaded cells in both B-CPAP and TPC-1 cells (Figure 2A and B). In concert with this, the wound-healing assay also suggested that silencing of LUCAT1 reduced the migration ability of TC cells (Figure 2C). In a cytokine perspective, the Western blot analysis found that the levels of metastasis-related proteins N-cadherin and MMP in cells were reduced after sh-LUCAT1 transfection (Figure 2D).
Overexpression of LUCAT1 Promoted the Malignant Behaviors of TC Cells
The above findings suggested that sh-LUCAT1 suppressed the malignancy of TC cells. Then, overexpression of LUCAT1 was also introduced in B-CPAP and TPC-1 cells using oe-LUCAT1 vectors. The transfection efficacy was confirmed by RT-qPCR (Figure 3A). Then, the CCK-8 and colony formation assays suggested that the proliferation and viability of B-CPAP and TPC-1 cells were increased (Figure 3B and C). The flow cytometry results showed that the apoptosis and cell cycle arrest at the G0/G1 phase were reduced upon LUCAT1 overexpression (Figure 3D and E). In addition, the wound-healing and Transwell assays suggested that overexpression of LUCAT1 increased the migratory and invasive potentials of TC cells (Figure 3F and G).
LUCAT1 Upregulates ADAM10 Through Sequestering miR-493
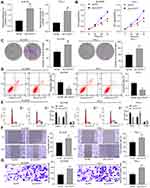
To further explore the potential molecules involved, we first determined the sub-cellular localization of LUCAT1 in B-CPAP and TPC-1 cells. The nuclear-cytoplasmic RNA separation assay suggested that LUCAT1 was mainly located in cytoplasm in cells (Figure 4A). The expression of miR-493 and ADAM10 in the collected tumor and adjacent normal ones was determined by RT-qPCR, and miR-493 was found to be poorly expressed while ADAM10 was highly expressed in the tumor tissues (Figure 4B). Further, we measured expression of miR-493 and ADAM10 mRNA in TC cell lines and in Nthy-ori3-1. It was found that miR-493 was highly expressed in 8305C, HTC-C3, B-CPAP and TPC-1 cells compared to that in Nthy-ori3-1 cells (Figure 4C).
Binding sites between LUCAT1 and miR-493, and between miR-493 and ADAM10 were first predicted on Starbase, and the luciferase vectors containing the WT and MT sequences were constructed (Figure 4D and E). Then, these binding relationships were further confirmed through luciferase assays. Compared to mimic NC transfection, cells co-transfected with miR-493 mimic and ADAM10-3ʹUTR-WT or LUCAT1-WT vectors presented significant reduced luciferase activity, while those transfected with MUT vectors showed little changes (Figure 4F and G). Then, altered expression of LUCAT1 or miR-493 was introduced in B-CPAP and TPC-1 cells. It was found that overexpression of LUCAT1 led to a decline in miR-493 expression but an increase in ADAM10 mRNA expression, while silencing of LUCAT1 led to converse results (Figure 4H). After miR-493 mimic transfection, LUCAT1 expression showed little changes, miR-493 was increased, while the mRNA expression of ADAM10 was notably decreased. Again, miR-493 inhibitor treatment resulted in opposite trends (Figure 4H).
LUCAT1 Mediates Proliferation and Colony Formation of TC Cells Through the miR-493/ADAM10 Axis
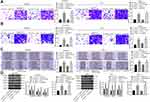
Downregulation of miR-493 or upregulation of ADAM10 was further introduced in B-CPAP and TPC-1 cells with stable sh-LUCAT1 transfection. According to the CCK-8, colony formation and flow cytometry results, it was found the inhibition in cell proliferation and colony formation and the increase in cell apoptosis by sh-LUCAT1 were counteracted by miR-493 inhibitor or oe-ADAM10 (Figure 5A–C). In addition, the cell cycle arrest in G0/G1 phase induced by sh-LUCAT1 was then blocked by miR-493 inhibitor or oe-ADAM10 with an increased number of cells entered the S phase (Figure 5D). These results indicate that miR-493 upregulation and ADAM10 downregulation are possibly responsible for the tumor-suppressing events of LUCAT1 silencing.
The LUCAT1/miR-493/ADAM10 Network Regulates Migration and Invasion of TC Cells and the Activation of the JAK-STAT Signaling Pathway
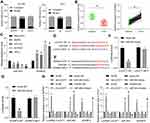
Likewise, migration and invasion of B-CPAP and TPC-1 cells were measured as well. The Transwell assay and scratch test results suggested that further inhibition of miR-493 or upregulation of ADAM10 elevated the number of migratory and invasive cells that were first reduced by sh-LUCAT1 (Figure 6A–C). Again, the Western blot analysis showed that the levels of N-cadherin and MMP-9 in cells were recovered by miR-493 inhibitor or oe-ADAM10 (Figure 6D). ADAM10 was recently noted to be linked to the activation of the JAK-STAT signaling pathway and could increase the growth and metastasis of colorectal cancer.18 Here, we noticed that the ADAM10 expression and phosphorylation of STAT3 was decreased by sh-LUCAT1 but then recovered following further administration of miR-493 inhibitor or oe-ADAM10 (Figure 6D), indicates that the JAK-STAT3 signaling is possibly involved in the LUCAT1-mediated events.
Silencing of LUCAT1 Inhibits Growth of Xenograft Tumors in Mice
B-CPAP cells with stable transfection were implanted into nude mice for in vivo experiments. It was found that silencing of LUCAT1 in cells reduced the volume and weight of xenograft tumors (Figure 7A–C). Still, this inhibition was blocked by further administration of miR-493 inhibitor or oe-ADAM10 in B-CPAP cells. These results suggest that LUCAT1 also inhibits growth of xenograft tumors in mice through the miR-493/ADAM10 axis.
Discussion
Though the prognosis is better than many other malignancies, TC remains a high health concern due to the increasing incidence and the possibility to death.3 During the conventional radioiodine treatment, patients are exposed to higher risks of developing radioiodine-refractory disease or being overtreated, therefore additional biomarkers may be beneficial and helpful for making decisions.19 In this research, we evidenced a novel ceRNA network involving aberrant expression of LUCAT1/miR-493/ADAM10 which is possibly participating in TC progression, during which the JAK-STAT activation is activated.
Emerging evidence has noted the implications of lncRNAs in pathogenesis, diagnosis and therapy in TC because of their mediation in angiogenesis, metastasis, proliferation apoptosis, and differentiation of cancer cells.19,20 For example, lncRNA ZFAS121 and lncRNA H1922 were recently found as potential biomarkers, reversely relating to the prognosis of TC patients. Similarly, downregulation of lncRNA AFAP1-AS1 was recently evidenced to promote apoptosis while inhibiting tumor growth and metastasis in TC.23 As aforementioned, LUCAT1 has been suggested as a prognostic biomarker for PTC. Here, we first identified a high-expression profile of LUCAT1 in TC cell lines. Then, artificial downregulation of LUCAT1 was introduced in TC cell lines, after which the proliferation, colony formation, migration and invasion, and resistance to apoptosis of TC cells were decreased. In a cytokine perspective, the expression of metastasis-related factors N-cadherin and MMP-9 in cells was decreased after LUCAT1 downregulation. The oncogenic functions of LUCAT1 have been well recognized. It was reported to regulate the stemness and trigger the progression of breast cancer with the involvement of the miR-5582-3p/TCF7L2 axis24 Likewise, LUCAT1 was found to trigger malignant behaviors from proliferation and invasion to chemo-resistance of cells in different cancer types.25,26 Here, we confirmed the implication of aberrant LUCAT1 expression in TC pathogenesis as well. In addition, the animal experiments where silencing of LUCAT1 inhibited growth of xenograft tumors in nude mice further validated its tumorigenic role in vivo.
Importantly, the data on a Starbase suggested a possible interaction among LUCAT1, miR-493 and ADAM10. Then, the binding relationships between LUCAT1 and miR-493, and between miR-493 and ADAM10 were validated through luciferase assays. And as expected, miR-493 presented a converse correlation while ADAM10, yet showed a positive correlation with LUCAT1 expression in TC cells. Further, inhibition of miR-493 or upregulation of ADAM10 was found to block the inhibitory roles of LUCAT1 silencing in cells. Though there is limited evidence concerning the correlation between miR-493 and TC, studies have noted that it serves as a tumor suppressor. For instance, silencing of miR-493 was found to increase the resistance of lung cancer cells to cisplatin by targeting TCRP1.27 On the contrary, high expression of miR-493-5p was positively associated with the treating outcome of lung cancer patients.28 miR-493-3p was found to directly target five different mRNAs to trigger apoptosis of ovarian cancer cells.29 Here, we identified a similar function of miR-493 in TC, indicating miR-493 inhibition is involved in the events mediated by LUCAT1. As for ADAM10, it is a member of the ADAMs family that modulates a wide array of cellular functions including cell adhesion, migration, proteolysis and other cell-signaling events.30 ADAM10 itself has been implicated in multiple human diseases from neurodegeneration to cancer, thus serving as a promising therapeutic target.31,32 Silencing of ADAM10 by miRNAs has been evidenced to reduce malignancy of cancer cells.17,33 Here, a similar trend was found in TC. Intriguingly, activation of the JAK-STAT signaling was found to be responsible for the oncogenic effect of ADAM10 in colorectal cancer.18 This signaling pathway has disclosed highly conserved programs correlating cytokine signaling with the mediation of key cellular mechanisms including proliferation, invasion, survival, inflammation and immunity, leaving its aberrant activation leading to cancer progression34 including that of TC.35 In this research, we noted that silencing of miR-493 or upregulation of ADAM10 increased phosphorylation of STAT3, implying that this signaling is possibly accountable for the oncogenic role of LUCAT1.
Conclusion
To sum up, this study evidenced that LUCAT1 increases ADAM10 expression through sequestering miR-493, leading to JAK-STAT activation and growth and metastasis of TC cells. These molecules may serve as novel prognostic biomarkers or novel therapeutic targets for TC. We hope more studies in this field will be performed to offer more ideas for TC control.
Funding
This work was supported by Zhejiang Provincial Medicine and Health Technology Project (2019KY660).
Disclosure
All authors declare that they have no conflicts of interest in this study.
References
1. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. doi:10.3322/caac.21492
2. Lubitz CC, Zhan T, Gunda V, et al. Circulating BRAF(V600E) levels correlate with treatment in patients with thyroid carcinoma. Thyroid. 2018;28(3):328–339. doi:10.1089/thy.2017.0322
3. Baloch ZW, LiVolsi VA. Special types of thyroid carcinoma. Histopathology. 2018;72(1):40–52.
4. Wang Q, Shang J, Zhang Y, Zhou Y, Tang L. MiR-451a restrains the growth and metastatic phenotypes of papillary thyroid carcinoma cells via inhibiting ZEB1. Biomed Pharmacother. 2020;127:109901. doi:10.1016/j.biopha.2020.109901
5. Ferreira HJ, Esteller M. Non-coding RNAs, epigenetics, and cancer: tying it all together. Cancer Metastasis Rev. 2018;37(1):55–73.
6. Li CH, Chen Y. Insight into the role of long noncoding RNA in cancer development and progression. Int Rev Cell Mol Biol. 2016;326:33–65.
7. Harrandah AM, Mora RA, Chan EKL. Emerging microRNAs in cancer diagnosis, progression, and immune surveillance. Cancer Lett. 2018;438:126–132. doi:10.1016/j.canlet.2018.09.019
8. Salehi S, Taheri MN, Azarpira N, Zare A, Behzad-Behbahani A. State of the art technologies to explore long non-coding RNAs in cancer. J Cell Mol Med. 2017;21(12):3120–3140.
9. Ergun S, Oztuzcu S. Oncocers: ceRNA-mediated cross-talk by sponging miRNAs in oncogenic pathways. Tumour Biol. 2015;36(5):3129–3136. doi:10.1007/s13277-015-3346-x
10. Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP. A ceRNA hypothesis: the rosetta stone of a hidden RNA language? Cell. 2011;146(3):353–358. doi:10.1016/j.cell.2011.07.014
11. Gao YS, Liu XZ, Zhang YG, Liu XJ, Li LZ. Knockdown of long noncoding RNA LUCAT1 inhibits cell viability and invasion by regulating miR-375 in glioma. Oncol Res. 2018;26(2):307–313. doi:10.3727/096504017X15088061795756
12. Nai Y, Pan C, Hu X, LncRNA MY. LUCAT1 contributes to cell proliferation and migration in human pancreatic ductal adenocarcinoma via sponging miR-539. Cancer Med. 2020;9(2):757–767. doi:10.1002/cam4.2724
13. Luzon-Toro B, Fernandez RM, Martos-Martinez JM, Rubio-Manzanares-Dorado M, Antinolo G, Borrego S. LncRNA LUCAT1 as a novel prognostic biomarker for patients with papillary thyroid cancer. Sci Rep. 2019;9(1):14374.
14. Deng J, Ma M, Jiang W, Zheng L, Cui S. MiR 493 induces cytotoxic autophagy in prostate cancer cells through regulation on PHLPP2. Curr Pharm Biotechnol. 2020;21. doi:10.2174/1389201021666200318120733
15. Sakai H, Sato A, Aihara Y, et al. MKK7 mediates miR-493-dependent suppression of liver metastasis of colon cancer cells. Cancer Sci. 2014;105(4):425–430. doi:10.1111/cas.12380
16. Liu F, Zhuang L, Wu R, Li D. miR-365 inhibits cell invasion and migration of triple negative breast cancer through ADAM10. J BUON. 2019;24(5):1905–1912.
17. Wu G, Zheng K, Xia S, et al. MicroRNA-655-3p functions as a tumor suppressor by regulating ADAM10 and beta-catenin pathway in Hepatocellular Carcinoma. J Exp Clin Cancer Res. 2016;35(1):89. doi:10.1186/s13046-016-0368-1
18. Hong YG, Xin C, Zheng H, et al. miR-365a-3p regulates ADAM10-JAK-STAT signaling to suppress the growth and metastasis of colorectal cancer cells. J Cancer. 2020;11(12):3634–3644. doi:10.7150/jca.42731
19. Mahmoudian-Sani MR, Jalali A, Jamshidi M, et al. Long non-coding RNAs in thyroid cancer: implications for pathogenesis, diagnosis, and therapy. Oncol Res Treat. 2019;42(3):136–142. doi:10.1159/000495151
20. Sui F, Ji M, Hou P. Long non-coding RNAs in thyroid cancer: biological functions and clinical significance. Mol Cell Endocrinol. 2018;469:11–22. doi:10.1016/j.mce.2017.07.020
21. Han CG, Huang Y, Qin L. Long non-coding RNA ZFAS1 as a novel potential biomarker for predicting the prognosis of thyroid cancer. Med Sci Monit. 2019;25:2984–2992. doi:10.12659/MSM.912921
22. Jiao X, Lu J, Huang Y, Zhang J, Zhang H, Zhang K. Long non-coding RNA H19 may be a marker for prediction of prognosis in the follow-up of patients with papillary thyroid cancer. Cancer Biomark. 2019;26(2):203–207. doi:10.3233/CBM-190273
23. Dai W, Tian Y, Jiang B, Chen W. Down-regulation of long non-coding RNA AFAP1-AS1 inhibits tumor growth, promotes apoptosis and decreases metastasis in thyroid cancer. Biomed Pharmacother. 2018;99:191–197. doi:10.1016/j.biopha.2017.12.105
24. Zheng A, Song X, Zhang L, et al. Long non-coding RNA LUCAT1/miR-5582-3p/TCF7L2 axis regulates breast cancer stemness via Wnt/beta-catenin pathway. J Exp Clin Cancer Res. 2019;38(1):305. doi:10.1186/s13046-019-1315-8
25. Wang W, Dong ML, Zhang W, Liu T. Long noncoding LUCAT1 promotes cisplatin resistance of non-small cell lung cancer by promoting IGF-2. Eur Rev Med Pharmacol Sci. 2019;23(12):5229–5234.
26. Yu H, Xu Y, Zhang D, Liu G. Long noncoding RNA LUCAT1 promotes malignancy of ovarian cancer through regulation of miR-612/HOXA13 pathway. Biochem Biophys Res Commun. 2018;503(3):2095–2100. doi:10.1016/j.bbrc.2018.07.165
27. Gu Y, Zhang Z, Yin J, et al. Epigenetic silencing of miR-493 increases the resistance to cisplatin in lung cancer by targeting tongue cancer resistance-related protein 1(TCRP1). J Exp Clin Cancer Res. 2017;36(1):114. doi:10.1186/s13046-017-0582-5
28. Liang Z, Kong R, He Z, et al. High expression of miR-493-5p positively correlates with clinical prognosis of non small cell lung cancer by targeting oncogene ITGB1. Oncotarget. 2017;8(29):47389–47399. doi:10.18632/oncotarget.17650
29. Kleemann M, Schneider H, Unger K, et al. Induction of apoptosis in ovarian cancer cells by miR-493-3p directly targeting AKT2, STK38L, HMGA2, ETS1 and E2F5. Cell Mol Life Sci. 2019;76(3):539–559. doi:10.1007/s00018-018-2958-x
30. Dempsey PJ. Role of ADAM10 in intestinal crypt homeostasis and tumorigenesis. Biochim Biophys Acta Mol Cell Res. 2017;1864(11 Pt B):2228–2239. doi:10.1016/j.bbamcr.2017.07.011
31. Smith TM
32. Wetzel S, Seipold L, Saftig P. The metalloproteinase ADAM10: A useful therapeutic target? Biochim Biophys Acta Mol Cell Res. 2017;1864(11 Pt B):2071–2081. doi:10.1016/j.bbamcr.2017.06.005
33. Guo W, Huang J, Lei P, Guo L, Li X. LncRNA SNHG1 promoted HGC-27 cell growth and migration via the miR-140/ADAM10 axis. Int J Biol Macromol. 2019;122:817–823. doi:10.1016/j.ijbiomac.2018.10.214
34. Pencik J, Pham HT, Schmoellerl J, et al. JAK-STAT signaling in cancer: from cytokines to non-coding genome. Cytokine. 2016;87:26–36. doi:10.1016/j.cyto.2016.06.017
35. Cletzer E, Klahn S, Dervisis N, LeRoith T. Identification of the JAK-STAT pathway in canine splenic hemangiosarcoma, thyroid carcinoma, mast cell tumor, and anal sac adenocarcinoma. Vet Immunol Immunopathol. 2020;220:109996. doi:10.1016/j.vetimm.2019.109996
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.