Back to Journals » OncoTargets and Therapy » Volume 12
Long Non-Coding RNA HOXA11-AS Promotes Non-Small Cell Lung Cancer Tumorigenesis Through microRNA-148a-3p/DNMT1 Regulatory Axis
Authors Bai Y, Lang L, Zhao W, Niu R
Received 15 December 2018
Accepted for publication 1 October 2019
Published 17 December 2019 Volume 2019:12 Pages 11195—11206
DOI https://doi.org/10.2147/OTT.S198367
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr XuYu Yang
Yue Bai,1 Lili Lang,2 Wentao Zhao,1 Rong Niu1
1Department One of Thoracic Surgery, Gansu Provincial Cancer Hospital, Gansu, People’s Republic of China; 2Department of Radiology, Gansu Provincial Cancer Hospital, Gansu, People’s Republic of China
Correspondence: Rong Niu
Department One of Thoracic Surgery, Gansu Provincial Cancer Hospital, No.2, Little West Lake, East Street, Qilihe District, Lanzhou City, Gansu Province, People’s Republic of China
Tel +86-13893604704
Email [email protected]
Objective: Our present study aimed to further investigate the molecular basis of long non-coding RNA homeobox A11 antisense (HOXA11-AS) in the tumorigenesis of non-small cell lung cancer (NSCLC).
Methods: HOXA11-AS, microRNA-148a-3p (miR-148a-3p), and DNA methyltransferase 1 (DNMT1) mRNA levels were measured by RT-qPCR assay. DNMT1 protein level was determined by Western blot assay. Cell proliferative capacity and apoptotic rate were determined by CCK-8 assay and flow cytometry analysis, respectively. The relationships of HOXA11-AS, miR-148a-3p, and DNMT1 were tested through bioinformatics analysis, luciferase assay, and RNA pull down assay. Mouse xenograft models of NSCLC were established to examine the biological function of HOXA11-AS in vivo.
Results: HOXA11-AS expression was notably upregulated and miR-148a-3p expression was conspicuously downregulated in NSCLC tissues and cells. HOXA11-AS knockdown curbed NSCLC cell proliferation and promoted cell apoptosis through directly increasing miR-148a-3p expression. Moreover, miR-148a-3p overexpression suppressed NSCLC cell proliferation and induced cell apoptosis. HOXA11-AS functioned as a competing endogenous RNA (ceRNA) of miR-148a-3p to increase DNMT1 expression in NSCLC cells. And, DNMT1 upregulation weakened the influence of HOXA11-AS1 loss on NSCLC cell proliferation and apoptosis. Additionally, HOXA11-AS knockdown suppressed NSCLC xenograft growth by upregulating miR-148a-3p and downregulating DNMT1 in vivo.
Conclusion: HOXA11-AS facilitated NSCLC tumorigenesis through miR-148a-3p/DNMT1 axis in vitro and in vivo, deepening our understanding of the molecular basis of HOXA11-AS in the development of NSCLC.
Keywords: non-small cell lung cancer, tumorigenesis, HOXA11-AS, miR-148a-3p, DNMT1
Introduction
Lung cancer is a huge threat for human health and life with an estimated 2.1 million new cases and 1.8 million deaths in 2018 alone worldwide.1 Moreover, the morbidity and mortality of lung cancer ranks first in all malignancies.1 Non-small cell lung cancer (NSCLC), a major histological subtype in lung cancer, accounts for approximately 85% of all cases.2,3 Despite the vast improvement in the management of NSCLC, most NSCLC patients are diagnosed with advanced or metastatic disease and the clinical outcomes of current therapeutic strategies are unsatisfactory.4–6 Therefore, it is of great importance to have a deep insight into the etiologies of NSCLC and seek potential biomarkers or targets for screening, diagnosis, prognosis, and treatment of NSCLC.
Long non-coding RNAs (lncRNAs) with a length of longer than 200 nucleotides (nt) and microRNAs (miRNAs) with a size of about 20 nt are a class of transcripts that lack protein-coding potential.7 Although the functions of lncRNAs and miRNAs are largely uncharacterized, growing evidence suggests that they are involved in the regulation of gene expression and fundamental biological processes.8,9 Moreover, accumulating lncRNAs and miRNAs have been found to be central players in the development and progression of many diseases including cancers.10 LncRNA homeobox A11 antisense (HOXA11-AS), located on chromosome 7p15.2, has been reported to be abnormally expressed in multiple cancers, either as a tumor suppressor or an oncogenic factor.11,12 For instance, HOXA11-AS functioned as a tumor accelerator in breast cancer,13 hepatocellular cancer,14 and gastric cancer,15 whereas it exerted anti-tumor effects in glioblastoma,16 epithelial ovarian cancer,17 and colorectal cancer.18 Furthermore, previous studies showed that HOXA11-AS could promote the development and progression of NSCLC.19–21
Bioinformatics examination showed that HOXA11-AS could possibly bind with miR-148a-3p. And, Sun et al demonstrated that HOXA11-AS could bind with enhancer of zeste homolog 2 (EZH2) and argonaute 2 (Ago2), and EZH2 could interact with DNA methyltransferase 1 (DNMT1) in GC cells.15 Ago2 is a core component of RNA-induced silencing complex (RISC), which serves as a crucial player in miRNAs-mediated gene silence.22 Hence, we supposed that HOXA11-AS could regulate DNMT1 expression by some miRNAs. DNMT1 has been demonstrated to be a target of miR-148a-3p in some cancers such as laryngeal squamous cell cancer,23 and bladder cancer.24 And, Chen et al disclosed that miR-148a-3p inhibited DNMT1 expression in NSCLC cells.25 MiR-148a, miR-148b, and miR-152 are members of the miR-148/miR-152 family, which have been reported as multi-faceted role players in the development of normal, non-tumor, and tumor tissues.26,27 And, miR-148a has been found to be a potential tumor suppressor in many malignancies including NSCLC.28 These data suggested the link of HOXA11-AS, miR-148a-3p, and DNMT1. Consequently, we further explored whether HOXA11-AS could exert its functions through miR-148a-3p/DNMT1 regulatory axis in NSCLC.
Our present study demonstrated that HOXA11-AS knockdown suppressed NSCLC cell proliferation and induced cell apoptosis in vitro and hampered NSCLC xenograft growth in vivo through upregulating miR-148a-3p and downregulating DNMT1.
Materials And Methods
Clinical Samples And Cell Culture
A total of 36 NSCLC patients who underwent surgical resection were enrolled in our project from Gansu Provincial Cancer Hospital during January 2017 to August 2017. These patients signed the written informed consents and did not receive any treatment prior to tissue collection. Also, our project got approval from Research Ethics Committee of Gansu Provincial Cancer Hospital. Once resected, these NSCLC tissues and adjacent normal lung tissues were immediately snap-frozen in liquid nitrogen and then stored at −80°C.
Normal human bronchial epithelial cell line 16HBE was obtained from Institute of Biochemistry and Cell Biology of the Chinese Academy of Sciences (Shanghai, China). Three NSCLC cell lines (95D, H460, H1299) were purchased from Cell Bank of Chinese Academy of Sciences (Shanghai, China). 16HBE cells were cultured in MEM medium (Thermo Scientific, Rockford, IL, USA) supplemented with 10% fetal bovine serum (FBS; Thermo Scientific). H460, 95D, and H1299 cells were grown in RPMI-1640 medium (Thermo Scientific) containing 10% FBS (Thermo Scientific).
Reagents And Cell Transfection
Small interference RNAs (siRNAs) targeting HOXA11-AS (siHOXA11-AS) and a scramble control (scrambled) were designed and synthesized from GenePharma Co., Ltd. (Shanghai, China). MiR-148a-3p mimic and its negative control miR-NC, miR-148a-3p inhibitor (anti-miR-148a-3p) and corresponding control anti-miR-NC were purchased from Thermo Scientific Co., ltd. HOXA11-AS and DNMT1 overexpression plasmids (HOXA11-AS, DNMT1) and their empty vectors were customized from Genomeditech Co., ltd. (Shanghai, China). These oligonucleotides or/and plasmids were transfected into NSCLC cells using Lipofectamine 3000 Reagent (Thermo Scientific) referring to the manufacturer’s instructions.
Reverse Transcription-Quantitative PCR (RT-qPCR) Assay
Total RNA was extracted and purified using miRNeasy Mini Kit (Qiagen, Dusseldorf, Germany) and RNase-free DNase I (Thermo Scientific). Then, TaqMan MicroRNA Reverse Transcription Kit and TaqMan MicroRNA Assay Kit (Thermo Scientific) were used to measure miR-148a-3p expression with U6 snRNA as the endogenous inference. For the detection of HOXA11-AS and DNMT1 expression levels, RNA was reverse transcribed into cDNA first strands using M-MLV Reverse Transcriptase (Thermo Scientific), and subsequent real time PCR analysis was performed using SYBR™ Green PCR Master Mix (Thermo Scientific) and specific primers. GAPDH acted as the house-keeping gene to normalize the expression of HOXA11-AS and DNMT1. The quantitative primers were listed as follows: 5ʹ-CGGCTAACAAGGAGATTTGG-3ʹ (sense) and 5ʹ-AGGCTCAGGGATGGTAGTCC-3ʹ (antisense) for HOXA11-AS, 5ʹ-GCACAAACTGACCTGCTTCA-3ʹ (sense) and 5ʹ-GCCTTTTCACCTCCATCAAA-3ʹ (antisense) for DNMT1, 5ʹ-TCCCTGAGCTGAACGGGAAG-3ʹ (sense) and 5ʹ-GGAGGAGTGGGTGTCGCTGT-3ʹ (antisense) for GAPDH.
Western Blot Assay
Cells or tumor tissues were lysed using RIPA buffer (Beyotime, Shanghai, China) supplemented with protease inhibitor (Thermo Scientific) and then centrifuged to obtain cell supernatants. Protein concentration in cell supernatants was determined through Pierce BCA Protein Assay Kit (Thermo Scientific). Subsequently, proteins (40 μg/lane) were separated through SDS-PAGE and transferred to nitrocellulose membranes (Millipore, Billerica, MA, USA). Then, the membranes were blocked for 1 h at room temperature using 5% non-fat milk, probed overnight at 4°C with anti-DNMT1 and anti-GAPDH primary antibodies (Abcam, Cambridge, UK) and incubated for 1 h at room temperature with horseradish peroxidase (HRP)-conjugated secondary antibody (Abcam). Finally, protein bands were visualized using Pierce™ ECL Western Blotting Substrate (Thermo Scientific) and the intensity of protein signals was estimated through Quantity One 4.1 (Bio-Rad Laboratories, Hercules, CA, USA).
Luciferase Reporter Assay
HOXA11-AS-Wt (wild type), DNMT1-Wt reporter (wild type), HOXA11-AS-Mut (mutant type) and DNMT1-Mut (mutant type) reporters containing wild or mutant miR-148a-3p binding sites were ordered from Hanbio Biotechnology Co., ltd. (Shanghai, China). Then, these reporters were respectively transfected into H460 and H1299 cells along with miR-NC or miR-148a-3p mimic. At 48 h after transfection, luciferase activities were determined using a dual luciferase reporter assay kit (Promega, Madison, WI, USA).
Cell Counting Kit-8 (CCK-8) Assay
Cell proliferative ability was assessed using the CCK-8 kit (Sigma-Aldrich, St. Louis, MO, USA). Briefly, transfected cells (100 μL/well) were inoculated into 96-well plates. Then, 10 μL of the CCK-8 solution was added into each well at 0, 24, 48, 72 h post-transfection. After another 3 h of incubation, cell absorbance was detected at 450 nm using a microplate reader.
Cell Apoptosis Analysis
Cell apoptosis rate was examined using an FITC AnnexinV Apoptosis Detection Kit (BD Biosciences, San Jose, CA, USA) according to the instructions of the manufacturer. Briefly, cells were collected at 48 h after transfection and resuspended in 1× Binding Buffer. Next, cells were incubated with FITC AnnexinV and propidium iodide solution for 15 min at room temperature in the dark. Finally, cell apoptosis patterns were analyzed using flow cytometry (BD Biosciences).
RNA Pull-Down Assay
RNA pull-down assay was carried out following a protocol as previously described.29 Briefly, biotinylated wild type miR-148a-3p (Bio-miR-148a-3p-Wt), biotinylated mutant type miR-148a-3p (Bio-miR-148a-3p-Mut), and their negative control Bio-miR-NC were ordered from Dharmacon Research Inc. (Lafayette, CO, USA) and transfected into H460 and H1299 cells, respectively. Forty-eight hours later, cells were collected and lysed using lysis buffer containing protease and RNase inhibitors. Next, cell supernatants were collected into new microcentrifuge tubes by centrifugation and then co-incubated overnight at 4°C with pre-treated Streptavidin-Dyna beads (Thermo Scientific). Next, RNA was isolated and purified from beads and HOXA11-AS level pulled down by these biotinylated miRNAs was measured through RT-qPCR assay.
Mouse Xenograft Experiments
BALB/c nude mice (n = 20, male, 6–8 weeks old) were purchased from Laboratory Animal Center of Zhengzhou University (Zhengzhou, China) and fed or treated following the national standards of the care and use of laboratory animals. Also, our animal experiments got the approval of Institutional Animal Care and Use Committee of Gansu Provincial Cancer Hospital. Mice were randomly divided into shNC and shHOXA11-AS groups with 10 mice in each group. Lentiviruses carrying HOXA11-AS knockdown fragment (shHOXA11-AS) and the control lentiviruses (shNC) were customized from Hanbio Biotechnology Co., ltd. For xenograft experiments, H460 cells (107 cells/mice) infected with shNC or shHOXA11-AS lentiviruses were subcutaneously injected into the flanks of mice in shNC or shHOXA11-AS group, respectively. Tumor volume was measured every 3 days for a total of 27 days using a caliper and estimated using the formula: volume = 0.5 × length × width.2 Twenty-seven days later, mice were killed and tumors were excised, weighed, and stored for the following RT-qPCR and Western blot assays.
Statistical Analysis
Data were obtained from more than three independent experiments and presented as means ± standard deviation (SD). Data were analyzed through GraphPad Prism software (La Jolla, CA, USA) and SPSS software (Chicago, IL, USA). The difference of groups was examined through Student’s t-test (for two group data) or one-way ANOVA (for over two groups data) with P < 0.05 as statistically significant.
Results
There Was High Expression Of HOXA11-AS And Low Expression Of miR-148a-3p In NSCLC Tissues And Cells
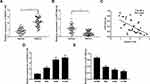
At first, RT-qPCR assay revealed that HOXA11-AS expression was markedly upregulated and miR-148a-3p expression was strikingly downregulated in 36 cases of NSCLC tissues compared to that in adjacent normal lung tissues (Figure 1A and B). Moreover, HOXA11-AS level was negatively associated with miR-148a-3p level in NSCLC tissues (n = 36) (Figure 1C). Also, as expected, higher HOXA11-AS expression and lower miR-148a-3p expression was observed in NSCLC cells (95D, H460 and H1299) compared with 16HBE cells (Figure 1D and E).
HOXA11-AS Knockdown Inhibited NSCLC Cell Proliferation And Induced Cell Apoptosis By Directly Increasing miR-148a-3p Expression
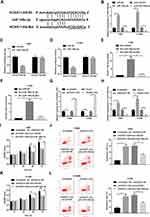
Bioinformatics analysis by StarBase online website presented that HOXA11-AS could possibly interact with miR-148a-3p (Figure 2A). Also, transfection efficiency revealed that the transfection of miR-148a-3p mimic was efficient to increase miR-148a-3p expression, and the introduction of miR-148a-3p inhibitor led to the notable reduction of miR-148a-3p expression in H460 and H1299 cells (Figure 2B). Next, luciferase reporter assay and RNA pull down assay were carried out to further validate the interaction of HOXA11-AS and miR-148a-3p. Luciferase reporter assay showed that miR-148a-3p overexpression could remarkably reduce the luciferase activity of HOXA11-AS-Wt reporter in H460 and H1299 cells, but did not have much influence on luciferase activity of HOXA11-AS-Mut reporter (Figure 2C and D). Moreover, RNA pull down assay disclosed that HOXA11-AS was substantially enriched by biotin-labeled wild type miR-148a-3p (Bio-miR-148a-3p-Wt) in H460 and H1299 cells, but not by mutant type miR-148a-3p (Bio-miR-148a-3p-Mut) (Figure 2E and F). That was to say, HOXA11-AS could interact with miR-148a-3p by putative complementary sites in NSCLC cells. To have a deep insight into the functions of HOXA11-AS in the tumorigenesis of NSCLC, siHOXA11-AS1 and HOXA11-AS overexpression plasmids were transfected into H460 and H1299 cells. As displayed in Figure 2G, HOXA11-AS expression was notably reduced in H460 and H1299 cells transfected with siHOXA11-AS1, but was conspicuously increased in cells transfected with HOXA11-AS overexpression plasmid, suggesting the practical values of siHOXA11-AS1 and HOXA11-AS overexpression plasmid in ensuing functional experiments. Moreover, RT-qPCR assay further unveiled that HOXA11-AS loss led to the obvious increase of miR-148a-3p level, whereas HOXA11-AS overexpression triggered the noticeable reduction of miR-148a-3p level in H460 and H1299 cells (Figure 2H). Next, functional analysis revealed that HOXA11-AS knockdown inhibited cell proliferation and promoted cell apoptosis in H460 and H1299 cells, while these effects were abrogated by miR-148a-3p inhibitor (Figure 2I–L). In a word, these data evinced that HOXA11-AS facilitated proliferation and suppressed apoptosis by reducing miR-148a-3p expression in NSCLC cells.
miR-148a-3p Overexpression Inhibited NSCLC Cell Proliferation And Promoted Cell Apoptosis
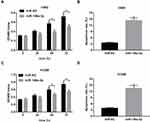
Then, CCK-8 assay revealed that cell proliferative ability was remarkably reduced in H460 and H1299 cells following the transfection of miR-148a-3p mimic at 48 h or 72 h post-transfection (Figure 3A and C). And, ectopic expression of miR-148a-3p led to the prominent elevation of cell apoptosis rate in H460 and H1299 cells (Figure 3B and D). That was to say, miR-148a-3p curbed cell proliferation and facilitated cell apoptosis in NSCLC.
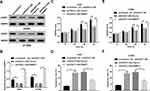
HOXA11-AS Promoted DNMT1 Expression By Downregulating miR-148a-3p In NSCLC Cells
Next, prediction analysis by TargetScan online website revealed that DNMT1 is a potential target of miR-148a-3p (Figure 4A). Then luciferase reporter assay further demonstrated that miR-148a-3p overexpression induced the notable downregulation of luciferase activity of DNMT1-Wt reporter, but did not affect the luciferase activity of DNMT1-Mut reporter in H460 and H1299 cells (Figure 4B and C), suggesting that miR-148a-3p could bind with DNMT1 3ʹ UTR through predicted binding sites. Moreover, we further demonstrated that enforced expression of miR-148a-3p led to the obvious downregulation of DNMT1 mRNA and protein levels in H460 and H1299 cells (Figure 4D–F). Inversely, miR-148a-3p depletion induced the increase of DNMT1 expression at mRNA and protein levels in H460 and H1299 cells (Figure 4D–F). These data manifested that miR-148a-3p inhibited DNMT1 expression through direct interaction in NSCLC cells. Further analysis revealed that DNMT1 mRNA and protein expression was remarkably elevated in HOXA11-AS-overexpressed H460 and H1299 cells, but was dramatically reduced in HOXA11-AS-depleted cells (Figure 4G–L). Moreover, miR-148a-3p overexpression inhibited the increase of DNMT1 expression induced by HOXA11-AS in H460 and H1299 cells (Figure 4G–I). And, the depletion of miR-148a-3p abrogated the inhibitory effect of HOXA11-AS knockdown on DNMT1 expression in H460 and H1299 cells (Figure 4J–L). That was to say, HOXA11-AS could function as a ceRNA of miR-148a-3p to sequester miR-148a-3p from its target DNMT1, leading to the elevation of DNMT1 level in NSCLC cells.
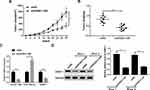
HOXA11-AS1 Exerted Its Functions By Upregulating DNMT1 In NSCLC Cells
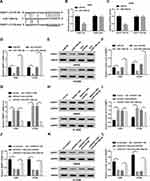
Next, Western blot assay confirmed that the introduction of DNMT1 overexpression plasmid reversed the detrimental effect of HOXA11-AS1 knockdown on DNMT1 expression in H460 and H1299 cells (Figure 5A and B). Restoration experiments showed that DNMT1 upregulation weakened the effects of HOXA11-AS loss on the proliferation and apoptosis of H460 and H1299 cells (Figure 5C–F). In summary, these results manifested that HOXA11-AS1 knockdown suppressed NSCLC cell proliferation and promoted cell apoptosis by downregulating DNMT1.
HOXA11-AS Knockdown Suppressed NSCLC Xenograft Growth By Upregulating miR-148a-3p And Downregulating DNMT1 In Vivo
Next, in vivo experiments further showed that HOXA11-AS knockdown resulted in the notable reduction of NSCLC xenograft tumor volume and weight (Figure 6A and B). Moreover, RT-qPCR assay demonstrated that HOXA11-AS level was markedly reduced in shHOXA11-AS-infected NSCLC xenograft tumors compared with control groups (Figure 6C). Also, a noticeable increase of miR-148a-3p level and an obvious reduction of DNMT1 mRNA level were observed in HOXA11-AS-depleted NSCLC xenograft tumors (Figure 6C). Moreover, we further demonstrated that HOXA11-AS knockdown led to the notable downregulation of DNMT1 protein level in two randomly selected NSCLC xenograft tumors (Figure 6D).
Discussion
Over the past decades, mounting experimental evidence has suggested that lncRNAs and miRNAs can serve as signaling transducers and potential biomarkers in plenty of cancers including NSCLC.30,31 Moreover, lncRNAs can regulate the expression of miRNAs and downstream targets by acting as ceRNAs in tumor biology.32 Also, some lncRNAs were found to regulate tumorigenesis and progression of cancers by affecting mRNA stability. For instance, Gou et al showed that enforced expression of lncRNA AB074169 suppressed papillary thyroid cancer (PTC) cell proliferation in vitro and hindered PTC xenograft tumor growth in vivo by reducing KH-type splicing regulatory protein (KHSRP) expression through destabilizing KHSRP mRNA transcripts.33 An in-depth understanding of these non-coding RNAs and their mechanisms of action will help us to manage cancer better.
In this document, we demonstrated that HOXA11-AS expression was strikingly upregulated in NSCLC tissues and cells, which was in line with previous reports.19,20 Also, The Cancer Genome Atlas (TCGA) database analysis showed that HOXA11-AS was highly expressed in NSCLC tissues compared to that in normal lung tissues.20,34 Functional analysis showed that HOXA11-AS knockdown inhibited NSCLC cell proliferation and promoted cell apoptosis in vitro and impeded NSCLC xenograft tumor growth in vivo. Also, the oncogenic effects of HOXA11-AS have been presented in previous documents.19–21 For instance, Chen et al demonstrated that the depletion of HOXA11-AS suppressed cell invasion and epithelial-mesenchymal transition (EMT) in NSCLC.19 Zhang et al showed that HOXA11-AS knockdown weakened proliferative, migratory, and invasive capacities of NSCLC cells and facilitated NSCLC cell apoptosis in vitro as well as curbing NSCLC tumorigenesis and angiogenesis in vivo.20
MiR-148a has been reported to be aberrantly expressed in multiple malignancies, which was closely correlated with tumor size, development, and prognosis.24 Although some reports pointed out that miR-148a exerted oncogenic effects in glioblastoma and osteosarcoma,35,36 miR-148a has been widely reported as a tumor suppressor in plenty of cancers such as breast cancer and papillary thyroid cancer.37,38 Moreover, previous reports disclosed that miR-148a level was markedly reduced in the serum of NSCLC patients, accompanied with higher possibility as a promising biomarker in NSCLC screening.39,40 Also, some studies demonstrated that miR-148a expression was remarkably downregulated in NSCLC tissues and cell lines and miR-148a exerted tumor-suppressive effects in NSCLC.41–43 For instance, the loss of miR-148a expression was associated with tumor progression and poor clinical outcomes, and enforced expression of miR-148a repressed cell migration and invasion via targeting Wnt1 in NSCLC.43 And, miR-148a overexpression impaired migratory, invasive, and proliferative abilities of NSCLC cells in vitro and impeded NSCLC tumorigenesis in vivo through downregulating matrix metallopeptidase 15 and Rho associated coiled-coil containing protein kinase 1.44
Consistent with previous reports,41–43 our present study demonstrated that miR-148a-3p expression was dramatically reduced in NSCLC tissues and cells, and miR-148a-3p overexpression inhibited NSCLC cell proliferation and induced cell apoptosis. Moreover, our data revealed that miR-148a-3p expression was negatively associated with HOXA11-AS expression in NSCLC tissues. Additionally, miR-148a-3p could bind with HOXA11-AS and miR-148a-3p downregulation attenuated the effects of HOXA11-AS loss on NSCLC cell proliferation and apoptosis.
DNMT1, a central DNA methyltransferase in mammalian cells, can regulate gene expression by epigenetic modification in a variety of biological processes such as cell apoptosis, cell cycle regulation, and chromatin organization.45 In addition, DNMT1 plays a significant role in the initiation, maintenance, and progression of cancers including NSCLC.46,47 For instance, the inhibition of DNMT1 by 5-aza-2ʹ-deoxycytidine resulted in the demethylation of some anti-tumor genes, the downregulation of NSCLC cell proliferative ability, and the inhibition of NSCLC xenograft tumor growth.48 And, DNMT1 knockdown suppressed cell migration, invasion, and EMT by inactivating Wnt/β-catenin signaling pathway in NSCLC.49
In this text, we demonstrated that DNMT1 was a target of miR-148a-3p, which was in line with previous reports.25,26 Moreover, our outcomes further presented that HOXA11-AS could function as a ceRNA of miR-148a-3p, giving rise to the downregulation of miR-148a-3p level and the upregulation of DNMT1 expression in NSCLC cells. Additionally, DNMT1 overexpression weakened the effects of HOXA11-AS depletion on proliferation and apoptosis of NSCLC cells. Furthermore, HOXA11-AS knockdown suppressed NSCLC xenograft tumor growth by upregulating miR-148a-3p and downregulating DNMT1 in vivo.
Conclusion
Taken together, these data revealed that HOXA11-AS facilitated NSCLC tumorigenesis in vitro and in vivo through regulating miR-148a-3p/DNMT1 axis, deepening our understanding of the regulatory mechanisms of HOXA11-AS and DNMT1, and providing some promising biomarkers and targets in the screening and treatment of NSCLC. However, the downstream targets of DNMT1 need to be further investigated. Also, it is requisite to explore the effects of HOXA11-AS/miR-148a-3p/DNMT1 on cell migration, invasion, and EMT in NSCLC cell and mouse models. Also, immunohistochemistry analysis and in situ hybridization experiments were indispensable to examine the expression patterns of HOXA11-AS, miR-148a-3p, and DNMT1 in NSCLC tissues.
Ethical Statement
A total of 36 NSCLC patients who underwent surgical resection were enrolled in our project from Gansu Provincial Cancer Hospital during January 2017 to August 2017. These patients signed the written informed consents and did not receive any therapy prior to tissue collection. Also, our project got the approval of Research Ethics Committee of Gansu Provincial Cancer Hospital. Our animal experiments got the approval of Institutional Animal Care and Use Committee of Gansu Provincial Cancer Hospital.
Disclosure
The authors declare that there are no financial and non-financial conflicts of interest existing in this study.
References
1. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424.
2. Ettinger DS, Akerley W, Borghaei H, et al. Non–small cell lung cancer. J Natl Comp Cancer Netw. 2012;10(10):1236–1271. doi:10.6004/jnccn.2012.0130
3. Ettinger DS, Wood DE, Akerley W, et al. Non–small cell lung cancer, version 1.2015. J Natl Comp Cancer Netw. 2014;12(12):1738–1761. doi:10.6004/jnccn.2014.0176
4. Wakelee H, Kelly K, Edelman MJ. 50 years of progress in the systemic therapy of non-small cell lung cancer. Am Soc Clin Oncol Educ Book. 2014;177–189. doi:10.14694/EdBook_AM.2014.34.177
5. Liu P, Wu Y, Zhou L, et al. [Prognostic analysis of patients with advanced non-small cell lung cancer in different genotypes]. Zhongguo Fei Ai Za Zhi. 2017;20(11):741–750. doi:10.3779/j.issn.1009-3419.2017.11.04
6. Ben Amar J, Ben Safta B, Zaibi H, Dhahri B, Baccar MA, Azzabi S. Prognostic factors of advanced stage non-small-cell lung cancer. Tunis Med. 2016;94(5):360–367.
7. Beermann J, Piccoli MT, Viereck J, Thum T. Non-coding RNAs in development and disease: background, mechanisms, and therapeutic approaches. Physiol Rev. 2016;96(4):1297–1325. doi:10.1152/physrev.00041.2015
8. Khorkova O, Hsiao J, Wahlestedt C. Basic biology and therapeutic implications of lncRNA. Adv Drug Deliv Rev. 2015;87:15–24. doi:10.1016/j.addr.2015.05.012
9. Mohr AM, Mott JL. Overview of microRNA biology. Semin Liver Dis. 2015;35(1):3–11. doi:10.1055/s-00000069
10. Adams BD, Parsons C, Walker L, Zhang WC, Slack FJ. Targeting noncoding RNAs in disease. J Clin Invest. 2017;127(3):761–771. doi:10.1172/JCI84424
11. Lu CW, Zhou DD, Xie T, et al. HOXA 11 antisense long noncoding RNA (HOXA 11‐AS): a promising lnc RNA in human cancers. Cancer Med. 2018;7(8):3792–3799. doi:10.1002/cam4.1571
12. Xue J-Y, Huang C, Wang W, Li H-B, Sun M, Xie M. HOXA11-AS: a novel regulator in human cancer proliferation and metastasis. OncoTargets Ther. 2018;11:4387–4393. doi:10.2147/OTT.S166961
13. Li W, Jia G, Qu Y, Du Q, Liu B, Liu B. Long non-coding RNA (LncRNA) HOXA11-AS promotes breast cancer invasion and metastasis by regulating epithelial-mesenchymal transition. MedSci Monit. 2017;23:3393–3403. doi:10.12659/MSM.904892
14. Yu J, Hong J, Kang J, Liao L, Li C. Promotion of LncRNA HOXA11-AS on the proliferation of hepatocellular carcinoma by regulating the expression of LATS1. Eur Rev Med Pharmacol Sci. 2017;21(15):3402–3411.
15. Sun M, Nie F, Wang Y, et al. LncRNA HOXA11-AS promotes proliferation and invasion of gastric cancer by scaffolding the chromatin modification factors PRC2, LSD1, and DNMT1. Cancer Res. 2016;76(21):6299–6310. doi:10.1158/0008-5472.CAN-16-0356
16. Se Y-B, Kim SH, Kim JY, et al. Underexpression of HOXA11 is associated with treatment resistance and poor prognosis in glioblastoma. Cancer Res Treat. 2017;49(2):387–398. doi:10.4143/crt.2016.106
17. Richards EJ, Permuth-Wey J, Li Y, et al. A functional variant in HOXA11-AS, a novel long non-coding RNA, inhibits the oncogenic phenotype of epithelial ovarian cancer. Oncotarget. 2015;6(33):34745–34757. doi:10.18632/oncotarget.5784
18. Li T, Xu C, Cai B, Zhang M, Gao F, Gan J. Expression and clinicopathological significance of the lncRNA HOXA11-AS in colorectal cancer. Oncol Lett. 2016;12(5):4155–4160. doi:10.3892/ol.2016.5129
19. Chen J-H, Zhou L-Y, Xu S, Zheng Y-L, Wan Y-F, Hu C-P. Overexpression of lncRNA HOXA11-AS promotes cell epithelial–mesenchymal transition by repressing miR-200b in non-small cell lung cancer. Cancer Cell Int. 2017;17(1):64. doi:10.1186/s12935-017-0433-7
20. Zhang Y, Chen W-J, Gan T-Q, et al. Clinical significance and effect of lncRNA HOXA11-AS in NSCLC: a study based on bioinformatics, in vitro and in vivo verification. Sci Rep. 2017;7(1):5567.
21. Yu W, Peng W, Jiang H, Sha H, Li J. LncRNA HOXA11-AS promotes proliferation and invasion by targeting miR-124 in human non–small cell lung cancer cells. Tumor Biol. 2017;39(10):1010428317721440. doi:10.1177/1010428317721440
22. Gregory RI, Chendrimada TP, Cooch N, Shiekhattar R. Human RISC couples microRNA biogenesis and posttranscriptional gene silencing. Cell. 2005;123(4):631–640. doi:10.1016/j.cell.2005.10.022
23. Wu T, Qu L, He G, et al. Regulation of laryngeal squamous cell cancer progression by the lncRNA H19/miR-148a-3p/DNMT1 axis. Oncotarget. 2016;7(10):11553.
24. Wang X, Liang Z, Xu X, et al. miR-148a-3p represses proliferation and EMT by establishing regulatory circuits between ERBB3/AKT2/c-myc and DNMT1 in bladder cancer. Cell Death Dis. 2016;7(12):e2503. doi:10.1038/cddis.2016.373
25. Chen Y, Min L, Zhang X, et al. Decreased miRNA-148a is associated with lymph node metastasis and poor clinical outcomes and functions as a suppressor of tumor metastasis in non-small cell lung cancer. Oncol Rep. 2013;30(4):1832–1840. doi:10.3892/or.2013.2611
26. Chen Y, Song Y-X, Wang Z-N. The microRNA-148/152 family: multi-faceted players. Mol Cancer. 2013;12(1):43. doi:10.1186/1476-4598-12-43
27. Friedrich M, Pracht K, Mashreghi MF, Jäck HM, Radbruch A, Seliger B. The role of the miR‐148/‐152 family in physiology and disease. Eur J Immunol. 2017;47(12):2026–2038. doi:10.1002/eji.201747132
28. Li Y, Deng X, Zeng X, Peng X. The role of Mir-148a in cancer. J Cancer. 2016;7(10):
29. Phatak P, Donahue JM. Biotinylated micro-RNA pull down assay for identifying miRNA targets. RNA Biol. 2018;15(1):55–61. doi:10.1080/15476286.2017.1391441
30. Ricciuti B, Mencaroni C, Paglialunga L, et al. Long noncoding RNAs: new insights into non-small cell lung cancer biology, diagnosis and therapy. Med Oncol. 2016;33(2):18. doi:10.1007/s12032-016-0731-2
31. Slaby O, Laga R, Sedlacek O. Therapeutic targeting of non-coding RNAs in cancer. Biochem J. 2017;474(24):4219–4251. doi:10.1042/BCJ20170079
32. Sui J, Li YH, Zhang YQ, et al. Integrated analysis of long non-coding RNAassociated ceRNA network reveals potential lncRNA biomarkers in human lung adenocarcinoma. Int J Oncol. 2016;49(5):2023–2036. doi:10.3892/ijo.2016.3716
33. Gou Q, Gao L, Nie X, et al. Long noncoding RNA AB074169 inhibits cell proliferation via modulation of KHSRP-mediated CDKN1a expression in papillary thyroid carcinoma. Cancer Res. 2018;78(15):4163–4174. doi:10.1158/0008-5472.CAN-17-3766
34. Zhang Y, He R-Q, Dang Y-W, et al. Comprehensive analysis of the long noncoding RNA HOXA11-AS gene interaction regulatory network in NSCLC cells. Cancer Cell Int. 2016;16(1):89. doi:10.1186/s12935-016-0366-6
35. Kim J, Zhang Y, Skalski M, et al. microRNA-148a is a prognostic oncomiR that targets MIG6 and BIM to regulate EGFR and apoptosis in glioblastoma. Cancer Res. 2014;74:1541–1553. doi:10.1158/0008-5472.CAN-13-1449
36. Ma W, Zhang X, Chai J, Chen P, Ren P, Gong M. Circulating miR-148a is a significant diagnostic and prognostic biomarker for patients with osteosarcoma. Tumor Biol. 2014;35(12):12467–12472. doi:10.1007/s13277-014-2565-x
37. Xu X, Zhang Y, Jasper J, et al. MiR-148a functions to suppress metastasis and serves as a prognostic indicator in triple-negative breast cancer. Oncotarget. 2016;7(15):20381–20394. doi:10.18632/oncotarget.7953
38. Han C, Zheng W, Ge M, Wang K, Xiang Y, Wang P. Downregulation of cyclin-dependent kinase 8 by microRNA-148a suppresses proliferation and invasiveness of papillary thyroid carcinomas. Am J Cancer Res. 2017;7(10):2081–2090.
39. Yang J-S, Li B-J, Lu H-W, et al. Serum miR-152, miR-148a, miR-148b, and miR-21 as novel biomarkers in non-small cell lung cancer screening. Tumor Biol. 2015;36(4):3035–3042. doi:10.1007/s13277-014-2938-1
40. Li L, Chen Y-Y, S-q L, Huang C, Y-z Q. Expression of miR-148/152 family as potential biomarkers in non-small-cell lung cancer. Med Sci Monitor. 2015;21:1155–1161. doi:10.12659/MSM.892940
41. Li J, Song Y, Wang Y, Luo J, Yu W. MicroRNA-148a suppresses epithelial-to-mesenchymal transition by targeting ROCK1 in non-small cell lung cancer cells. Mol Cell Biochem. 2013;380(1–2):277–282. doi:10.1007/s11010-013-1682-y
42. Li J, Yu T, Cao J, et al. MicroRNA-148a suppresses invasion and metastasis of human non-small-cell lung cancer. Cell Physiol Biochem. 2015;37(5):1847–1856. doi:10.1159/000438546
43. Chen Y, Min L, Ren C, et al. miRNA-148a serves as a prognostic factor and suppresses migration and invasion through Wnt1 in non-small cell lung cancer. PloS ONE. 2017;12(2):e0171751. doi:10.1371/journal.pone.0171751
44. Joshi P, Jeon Y-J, Laganà A, et al. MicroRNA-148a reduces tumorigenesis and increases TRAIL-induced apoptosis in NSCLC. Proc Nat Acad Sci. 2015;112(28):8650–8655. doi:10.1073/pnas.1500886112
45. Svedruzic ZM. Dnmt1 structure and function. Prog Mol Biol Transl Sci. 2011;101:221–254. doi:10.1016/B978-0-12-387685-0.00006-8
46. Lakshminarasimhan R, Liang G. The role of DNA methylation in cancer. Adv Exp Med Biol. 2016;945:151–172.
47. Singh V, Sharma P, Capalash N. DNA methyltransferase-1 inhibitors as epigenetic therapy for cancer. Current Cancer Drug Targets. 2013;13(4):379–399.
48. Liu B, Song J, Luan J, et al. Promoter methylation status of tumor suppressor genes and inhibition of expression of DNA methyltransferase 1 in non-small cell lung cancer. Exp Biol Med (Maywood). 2016;241(14):1531–1539. doi:10.1177/1535370216645211
49. Bu X, Zhang X, Xu J, et al. Inhibition of DNA methyltransferase 1 by RNA interference reverses epithelial-mesenchymal transition in highly metastatic 95D lung cancer cells by inhibiting the Wnt signaling pathway. Oncol Lett. 2018;15(6):9242–9250. doi:10.3892/ol.2018.8449
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.