Back to Journals » Cancer Management and Research » Volume 12
Long Non-Coding RNA GAS5 Targeting microRNA-21 to Suppress the Invasion and Epithelial–Mesenchymal Transition of Uveal Melanoma
Authors Qi Y, Cui Q , Zhang W , Yao R , Xu D, Zhang F
Received 1 May 2020
Accepted for publication 10 July 2020
Published 27 November 2020 Volume 2020:12 Pages 12259—12267
DOI https://doi.org/10.2147/CMAR.S260866
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Eileen O'Reilly
Ying Qi, Qingqing Cui, Wenjing Zhang, Renjie Yao, Dong Xu, Fengyan Zhang
Department of Ophthalmology, The First Affiliated Hospital of Zhengzhou University, The Laboratory for Ophthalmology and Vision Science, Henan Eye Hospital, Zhengzhou 450052, Henan, People’s Republic of China
Correspondence: Fengyan Zhang Department of Ophthalmology
The First Affiliated Hospital of Zhengzhou University, The Laboratory for Ophthalmology and Vision Science, Henan Eye Hospital, Zhengzhou 450052, Henan, People’s Republic of China
Email [email protected]
Objective: Human uveal melanoma (UM) is a common ocular malignant tumor with a high risk of metastasis. Emerging evidence indicates that long non-coding RNAs (lncRNAs) are correlated with the development of UM. Here, we aimed to determine the biological significance of lncRNA growth arrest-specific transcript 5 (GAS5) in UM.
Methods: The expression levels of GAS5 and microRNA-21 (miR-21) in UM tissues and cells were detected by qRT-PCR analysis. CCK-8 assay was performed to investigate the viability of UM cells after cell transfections, and the migration and invasion of UM cells were determined by transwell assay. The protein expression levels were detected by Western blot assay. The relationship between miR-21 and GAS5 in UM cells was confirmed by bioinformatics prediction and luciferase report assay.
Results: Our experiments demonstrated that GAS5 was markedly downregulated in UM cells and clinical specimens. Overexpression of GAS5 inhibited, whereas knockdown of GAS5 promoted the viability, migration, and invasion of UM cells. The epithelial-to-mesenchymal transition (EMT) process of UM cells was also suppressed by upregulating of GAS5 and enhanced by downregulating of GAS5. Additionally, as a competitive endogenous RNA (ceRNA), GAS5 directly binded to the oncogenic miR-21 in UM cells, and overexpression of miR-21 attenuated the EMT-suppressing effect of GAS5.
Conclusion: Taken together, our findings suggest that GAS5/miR-21 axis is implicated in the pathogenesis of UM and might serve as a potential therapeutic target.
Keywords: GAS5, uveal melanoma, invasion, epithelial–mesenchymal transition, microRNA-21
Introduction
Uveal melanoma (UM), arising from the iris, ciliary body, or choroid of the eye, is the most common primary intraocular tumor in adults.1 UM has a high ability to metastasize, most commonly to the liver initially.2 Despite advances in the diagnosis and therapy of UM, the prognosis of those diagnosed with metastatic UM remains disappointing, with the median overall survival of only 2–15 months.3 Accordingly, investigation of the underlying mechanisms involved in UM development and metastasis is urgently required.
Long non-coding RNAs (lncRNAs), a class of noncoding transcripts longer than 200 nucleotides with limited or no protein-coding capacity, are emerging as new players in various human cancers through acting as molecular sponges or competing endogenous RNAs (ceRNAs) to weaken the suppressive effect of microRNAs (miRNAs) on protein-coding genes.4 Growth arrest-specific transcript 5 (GAS5), a 651-bp lncRNA located on chromosome 1q25.1 (https://www.ncbi.nlm.nih.gov/nuccore/NR_002578.2), has previously been reported to be under-expressed and function as a tumor suppressor in various human cancers, including gastric cancer,5 ovarian cancer,6 non-small cell lung cancer7 and cervical cancer.8 In detail, GAS5 inhibits tumorigenesis and enhances radiosensitivity in non-small cell lung cancer cells via suppressing miR-135b expression.7 And, GAS5 functions as a ceRNA of miR-222-3p to increase PTEN level, leading to the inhibition of papillary thyroid carcinoma progression.9 However, the expression pattern, biological function and the molecular mechanisms of GAS5 in UM progression have not been fully elucidated.
In the present study, we report for the first time, the potential role and underlying mechanisms of GAS5 in UM. Our results demonstrate that GAS5 is downregulated in UM, and overexpression of GAS5 inhibits the viability, migration, and invasion of UM cells. Thus, we suggest that GAS5 might be a promising therapeutic target for UM.
Materials and Methods
Clinical Samples
Eight enucleated eyes were collected from the patients with large UM lesions and stored in a liquid nitrogen tank (−196°C). Normal uveal samples were obtained from the Beijing Tongren Eye Bank (Beijing, China). The use of human tissue samples was approved by the Ethic Committee of the First Affiliated Hospital of Zhengzhou University. The informed written consents were obtained from patients or their relatives as the eyesight for some patients was too low.
Cell Culture and Transfection
Human UM cell lines MUM-2B and C918 were purchased from Chinese Academy of Sciences Cell Bank (Shanghai, China) and cultured in RPMI 1640 medium (KeyGene, Nanjing, China) with 10% fetal bovine serum (FBS; HyClone, Logan, UT, USA) and 50 µg/mL penicillin/streptomycin in humidified air at 37°C with 5% CO2. Human melanocytes (D78) obtained from Chinese Academy of Sciences Cell Bank were culture in Dulbecco’s modified Eagle’s medium (DMEM; KeyGene).
The GAS5 sequence was synthesized according to the full-length GAS5 sequence and then sub-cloned into a pcDNA3.1 vector (Invitrogen, Shanghai, China). GAS5 siRNAs negative control siRNA (si-NC) were specifically synthesized by Shanghai GenePharma Co, Ltd. (Shanghai, China). The sequences of the GAS5 siRNAs were as follows: si-GAS5 (#1), AGTGTGGCTCTGGATAGCACCTTAT; si-GAS5 (#2), AGGAAGGATGAGAATAGCTACTGAA; si-GAS5 (#3), CAGTGTGGCTCTGGATAGCACCTTA. miR-21 mimics and mimics negative control were purchased from Genecopoeia (Guangzhou, China). Cells at 70~80% confluence were selected for transfection using Lipofectamine 2000 reagent (Invitrogen). Forty-eight hours after transfection, the cells were collected for further analysis.
RNA Isolation and qRT-PCR Analysis
Total RNA was extracted from the tissues and cells using TRIZOL reagent (Invitrogen). For mRNA and lncRNA, RNA was reverse transcribed to cDNA using the PrimeScript™ RT reagent kit (Takara, Dalian, China). For miRNA, reverse transcription was performed using the TaqMan MicroRNA RT kit (Applied Biosystems, Foster City, CA, USA). qPCR was then performed using SYBR® Premix Ex Taq™ II (Takara) on the ABI PRISM 7000 Fluorescent Quantitative PCR System (Applied Biosystems). U6 or GAPDH was used as an internal control. The relative expression of genes is presented as fold change and calculated using the 2−ΔΔCt method.10 The primer sequences are listed in Table 1.
 |
Table 1 Sequences of PCR Primers |
Western Blot Assay
Proteins were extracted from the cells using RIPA protein extraction reagent (Sigma, St. Louis, MO, USA) containing the protease inhibitor cocktail (Roche, Mannheim, Germany). Equal quantity of proteins was then separated by SDS-PAGE gels, and transferred to PVDF membranes (Millipore, Billerica, MA, USA). Then, the membranes were blocked with 5% non-fat milk and incubated with specific primary antibodies against E-cadherin (1:1000, Cell Signaling, Beverly, MA, USA), Vimentin (1:1000, Cell Signaling), N-cadherin (1:1000, Cell Signaling), and β-actin (1:500, Santa Cruz, Dallas, TX, USA) at 4 °C overnight. Thereafter, the membranes were incubated with HRP-conjugated secondary antibodies (1:10,000, Santa Cruz) at room temperature for 1 hour. The bands were visualized using the enhanced chemiluminescence kit (Amersham, Little Chalfont, UK). Protein expression was quantified by densitometry (Quantity One software; Bio-Rad, CA, USA), with β-actin used as a control.
CCK-8 Assay
Cell viability was determined over the next 24–96 hours following cell transfections using the Cell Counting Kit-8 (CCK-8; Beyotime, Haimen, China). Cells were plated at 3000 cells per well in 96-well plates. At the indicated time after transfection, 10 μL CCK-8 solution was added into each well, and wells were incubated for another 2 hours. The absorbance was measured at 450 nm using a microtiter plate reader.
Transwell Assay
Cell migration and invasion were determined by using transwell chambers (24-well insert; 8-μm pore size; BD Biosciences, Franklin Lakes, NJ, USA). In brief, after transfection, 1×105 cells in 200 μL serum-free medium were seeded in the uncoated or Matrigel (30 μL; BD Biosciences)-coated upper chambers, and the lower chambers were filled with medium containing 10% FBS. After 48 hours of incubation, the cells on the bottom surface of the membrane were fixed with methanol, stained with 0.1% crystal violet (Solarbio, Beijing, China) for 10 min at room temperature, and then counted using a light microscope (Olympus, Tokyo, Japan).
Luciferase Report Assay
The GAS5 fragment containing the putative miR-21 binding site was amplified by PCR and cloned into a pGL3 promoter vector (Promega, Madison, WI, USA). The QuickChange® Site-Directed Mutagenesis Kit (Stratagene, La Jolla, CA, USA) was used to mutate the binding site of GAS5 and miR-21. C918 cells were seeded into 24-well plates and co-transfected with luciferase reporter vectors (WT-GAS5 or MUT-GAS5) and miR-21 mimics or mimics negative control using Lipofectamine 2000 reagent (Invitrogen). Forty-eight hours post transfection, the cells were harvested, and the relative luciferase activities were measured using the Dual Luciferase Reporter Assay System (Promega). Renilla luciferase activity was normalized to firefly luciferase activity.
Statistical Analysis
Data are expressed as the mean ± standard deviation (SD) from at least three separate experiments. Data analysis, graph generation, and statistical analysis were performed using Graphpad Prism (version 6.01) software (GraphPad Software, Inc., La Jolla, CA, USA). Differences between groups were assessed using either Student’s t-test or one-way analysis of variance (ANOVA). The correlation between GAS5 and miR-21 expression was evaluated by Spearman correlation analysis. P-values of less than 0.05 were considered to be statistically significant.
Results
GAS5 is Downregulated in UM and Predicts a Poor Prognosis
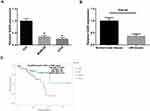
To determine whether GAS5 plays a role in the development of UM, qRT-PCR analysis was performed to measure the expression of GAS5 in UM cells. As shown in Figure 1A, the expression of GAS5 was significantly decreased in UM cell lines compared to that in D78 cells. Furthermore, GAS5 was also downregulated in UM tissues compared with that in normal uveal samples (Figure 1B). In addition, UM patients with GAS5 low expression presented with lower overall survival rates than patients with GAS5 high expression (Figure 1C).
GAS5 Inhibits UM Cell Viability and Invasion
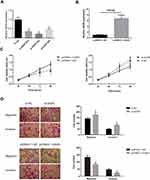
To further explore the biological role of GAS5 in UM progression, GAS5 was knocked-down in MUM-2B cells by using GAS5 siRNAs (Figure 2A), and overexpressed by transfecting pcDNA3.1-GAS5 into C918 cells (Figure 2B) as C918 cells showed the lower expression level of GAS5 than MUM-2B. si-GAS5 (#2) was used for all the RNAi experiments in this study. UM cell viability was then detected by CCK-8 assay at 24, 48, 72, and 96 hour time points and found that increased GAS5 expression significantly inhibited the viability of C918 cells, whereas knockdown of GAS5 in MUM-2B cells resulted in a significant increase in cell viability (Figure 2C). GAS5 effects on migratory and invasive abilities of UM cells were then assayed. After knockdown of GAS5, the migratory and invasive abilities were remarkably enhanced in MUM-2B cells, but was suppressed by ectopic expression of GAS5 in C918 cells (Figure 2D).
GAS5 Suppresses EMT Process in UM Cells
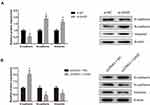
EMT is vital for cell invasion, therefore we determined the effects of GAS5 on EMT in UM cells. The results show that the expression levels of Vimentin and N-cadherin were significantly increased whereas E-cadherin expression was reduced when GAS5 was knocked down in MUM-2B cells (Figure 3A). Ectopic expression of GAS5 in C918 cells inhibited the expression levels of N-cadherin and Vimentin, but promoting the E-cadherin expression (Figure 3B).
GAS5 Functions as a ceRNA for miR-21 in UM
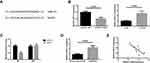
Using an online database (http://cbcsrv.watson.ibm.com/rna22.html), we found that GAS5 has one predicted miR-21 target binding site (Figure 4A). Additionally, as shown in Figure 4B, GAS5 could negatively regulate the expression of miR-21 in UM cells. The luciferase reporter assay manifested that miR-21 significantly suppressed luciferase activity that carried WT but not MUT 3′-UTR of GAS5 (Figure 4C). Further analysis revealed that the expression of miR-21 was increased in UM tissues (Figure 4D), and the increased miR-21 expression was closely correlated with the decreased GAS5 expression in UM tissues (Figure 4E).
miR-21 Attenuates the Function of GAS5 in UM
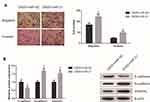
To further determine the role of miR-21 in the GAS5-mediated effects on UM cells, rescue experiments were subsequently performed. We found that upregulation of miR-21 abolished GAS5-induced inhibition of the migration and invasion of C918 cells (Figure 5A). Furthermore, miR-21 rescued repression of EMT process caused by GAS5 overexpression in C918 cells (Figure 5B).
Discussion
In addition to microRNAs, lncRNAs have emerged as critical players in UM progression.11–13 Altered GAS5 expression is related to many human cancers. This study might be the first direct investigation of the relationship between GAS5 expression and UM. Here, we observed that GAS5 is downregulated in UM tissue samples and cell lines, and overexpression of GAS5 markedly inhibited the viability, migration and invasion of UM cells in vitro. These findings indicate that GAS5 might function as a tumor suppressor in UM.
Cancer metastasis is a complex process involving multiple cellular steps. Featured by the loss of intracellular junctions and acquirement of mesenchymal characteristics, EMT is a key process for the dissemination of malignant cells to adjacent tissues and distant sites.14,15 EMT-associated factors could promote invasive properties of UM cells.16 A recent study reported that overexpression of GAS5 could inhibited the EMT of osteosarcoma cells17 In this study, we observed that the expression levels of EMT-associated proteins, including N-cadherin and vimentin, were reduced in GAS5-overexpressed UM cells, indicating that GAS5 inhibited human UM metastasis by suppressing EMT process.
Recently, a large body of experimental evidence has shown that lncRNAs can act as either ceRNAs which bind to miRNAs to influence miRNAs-induced gene silencing or natural miRNA sponges to regulate gene expression, playing critical roles in human diseases.18,19 The direct regulatory association between GAS5 and miR-21 was previously reported.20,21 As a well-known oncogenic miRNA, miR-21 was also found to be upregulated in formalin-fixed, paraffin-embedded UM samples.22 In the present study, through bioinformatics prediction and functional assays, we lend credence to the previous study, that lncRNA GAS5 could directly bind to miR-21 in UM, and that overexpression of miR-21 could attenuate the effects of GAS5 on UM cells.
PTEN is a well-known tumor suppressor, and plays a tumor-suppressive role during UM pathogenesis.23 Loss of PTEN expression is associated with shortened disease-free survival of UM patients.23 It has been reported that miR-21 induced EMT in lung cancer cells via targeting PTEN.24 Therefore, we conjecture that GAS5 may function as a ceRNA of miR-21 to induce PTEN expression, resulting in the suppressions of invasion and EMT in UM. We intend to clarify it in following assays.
In summary, this study identifies GAS5 serves as a novel potential cancer suppressor in UM. GAS5 might act as a ceRNA to inhibit the EMT process by sponging miR-21. Our findings provide a more adequate theoretical basis for understanding the molecular pathology of UM, and describe a novel GAS5/miR-21/EMT regulatory network involved in UM development.
Disclosure
The authors report no funding and no conflicts of interest for this work.
References
1. Singh AD, Turell ME, Topham AK. Uveal melanoma: trends in incidence, treatment, and survival. Ophthalmology. 2011;118(9):1881–1885. doi:10.1016/j.ophtha.2011.01.040
2. Coupland SE, Lake SL, Zeschnigk M, Damato BE. Molecular pathology of uveal melanoma. Eye (Lond). 2013;27(2):230–242. doi:10.1038/eye.2012.255
3. Augsburger JJ, Correa ZM, Shaikh AH. Effectiveness of treatments for metastatic uveal melanoma. Am J Ophthalmol. 2009;148(1):119–127. doi:10.1016/j.ajo.2009.01.023
4. Gibb EA, Brown CJ, Lam WL. The functional role of long non-coding RNA in human carcinomas. Mol Cancer. 2011;10:38. doi:10.1186/1476-4598-10-38
5. Sun M, Jin FY, Xia R, et al. Decreased expression of long noncoding RNA GAS5 indicates a poor prognosis and promotes cell proliferation in gastric cancer. BMC Cancer. 2014;14:319. doi:10.1186/1471-2407-14-319
6. Li J, Huang H, Li Y, Li L, Hou W, You Z. Decreased expression of long non-coding RNA GAS5 promotes cell proliferation, migration and invasion, and indicates a poor prognosis in ovarian cancer. Oncol Rep. 2016;36(6):3241–3250. doi:10.3892/or.2016.5200
7. Xue Y, Ni T, Jiang Y, Li Y. Long noncoding RNA GAS5 inhibits tumorigenesis and enhances radiosensitivity by suppressing miR-135b expression in non-small cell lung cancer. Oncol Res. 2017;25(8):1305–1316. doi:10.3727/096504017X14850182723737
8. Yang W, Hong L, Xu X, Wang Q, Huang J, Jiang L. LncRNA GAS5 suppresses the tumorigenesis of cervical cancer by downregulating miR-196a and miR-205. Tumour Biol. 2017;39(7):1010428317711315. doi:10.1177/1010428317711315
9. Zhang XF, Ye Y, Zhao SJ. LncRNA Gas5 acts as a ceRNA to regulate PTEN expression by sponging miR-222-3p in papillary thyroid carcinoma. Oncotarget. 2018;9(3):3519–3530. doi:10.18632/oncotarget.23336
10. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods. 2001;25(4):402–408. doi:10.1006/meth.2001.1262
11. Lu L, Yu X, Zhang L, et al. The long non-coding RNA RHPN1-AS1 promotes uveal melanoma progression. Int J Mol Sci. 2017;18(1):226. doi:10.3390/ijms18010226
12. Lu Q, Zhao N, Zha G, Wang H, Tong Q, Xin S. LncRNA HOXA11-AS exerts oncogenic functions By repressing p21 and miR-124 in uveal melanoma. DNA Cell Biol. 2017;36(10):837–844.
13. Zheng X, Tang H, Zhao X, Sun Y, Jiang Y, Liu Y. Long non-coding RNA FTH1P3 facilitates uveal melanoma cell growth and invasion through miR-224-5p. PLoS One. 2017;12(11):e0184746. doi:10.1371/journal.pone.0184746
14. Yang J, Weinberg RA. Epithelial-mesenchymal transition: at the crossroads of development and tumor metastasis. Dev Cell. 2008;14(6):818–829. doi:10.1016/j.devcel.2008.05.009
15. Savagner P. The epithelial-mesenchymal transition (EMT) phenomenon. Ann Oncol. 2010;21 Suppl 7:vii89–vii92. doi:10.1093/annonc/mdq292
16. Asnaghi L, Gezgin G, Tripathy A, et al. EMT-associated factors promote invasive properties of uveal melanoma cells. Mol Vis. 2015;21:919–929.
17. Ye K, Wang S, Zhang H, Han H, Ma B, Nan W. Long noncoding RNA GAS5 suppresses cell growth and epithelial-mesenchymal transition in osteosarcoma by regulating the miR-221/ARHI pathway. J Cell Biochem. 2017;118(12):4772–4781. doi:10.1002/jcb.26145
18. Tay Y, Rinn J, Pandolfi PP. The multilayered complexity of ceRNA crosstalk and competition. Nature. 2014;505(7483):344–352. doi:10.1038/nature12986
19. Paraskevopoulou MD, Hatzigeorgiou AG. Analyzing MiRNA-LncRNA Interactions. Methods Mol Biol. 2016;1402:271–286.
20. Wen Q, Liu Y, Lyu H, et al. Long noncoding RNA GAS5, which acts as a tumor suppressor via microRNA 21, regulates cisplatin resistance expression in cervical cancer. Int J Gynecol Cancer. 2017;27(6):1096–1108. doi:10.1097/IGC.0000000000001028
21. Cao L, Chen J, Ou B, Liu C, Zou Y, Chen Q. GAS5 knockdown reduces the chemo-sensitivity of non-small cell lung cancer (NSCLC) cell to cisplatin (DDP) through regulating miR-21/PTEN axis. Biomed Pharmacother. 2017;93:570–579. doi:10.1016/j.biopha.2017.06.089
22. Ragusa M, Barbagallo C, Statello L, et al. miRNA profiling in vitreous humor, vitreal exosomes and serum from uveal melanoma patients: pathological and diagnostic implications. Cancer Biol Ther. 2015;16(9):1387–1396. doi:10.1080/15384047.2015.1046021
23. Abdel-Rahman MH, Yang Y, Zhou XP, Craig EL, Davidorf FH, Eng C. High frequency of submicroscopic hemizygous deletion is a major mechanism of loss of expression of PTEN in uveal melanoma. J Clin Oncol. 2006;24(2):288–295. doi:10.1200/JCO.2005.02.2418
24. Dai L, Chen F, Zheng Y, et al. miR-21 regulates growth and EMT in lung cancer cells via PTEN/Akt/GSK3beta signaling. Front Biosci (Landmark Ed). 2019;24:1426–1439. doi:10.2741/4788
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.