Back to Journals » OncoTargets and Therapy » Volume 13
LncRNA LINC00665 Promotes Prostate Cancer Progression via miR-1224-5p/SND1 Axis
Authors Chen W, Yu Z, Huang W, Yang Y, Wang F, Huang H
Received 9 December 2019
Accepted for publication 11 March 2020
Published 26 March 2020 Volume 2020:13 Pages 2527—2535
DOI https://doi.org/10.2147/OTT.S241578
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Carlos E Vigil
Wei Chen,* Zhixian Yu,* Weiping Huang, Yu Yang, Feng Wang, Hang Huang
Department of Urology, First Affiliated Hospital of Wenzhou Medical University, Wenzhou 325000, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Hang Huang; Feng Wang
Department of Urology, First Affiliated Hospital of Wenzhou Medical University, Wenzhou 325000, People’s Republic of China
Email [email protected]; [email protected]
Background: Increasing researches have revealed a critical role of long noncoding RNAs (lncRNAs) in tumor progression. LINC00665 is a poorly investigated lncRNA. In this research, we sought to determine the potential role of LINC00665 in prostate cancer (PC) progression.
Methods: LINC00665 expression was analyzed by bioinformatics method and qRT-PCR. Proliferation was determined via CCK8 and colony formation assays. Transwell assay was conducted to analyze migration and invasion. Xenograft assay was used to test the roles of LINC00665 in vivo. Luciferase reporter assay, pulldown assay and RIP assay were utilized to confirm the interaction between LINC00665 and miR-1224-5p.
Results: LINC00665 expression was increased in PC samples in contrast to control tissues, according to bioinformatics analysis and qRT-PCR validation. LINC00665 high expression was related to a poor prognosis. LINC00665 knockdown markedly attenuated growth and metastasis of PC cells and impaired tumor propagation in vivo. Mechanistic investigation revealed that LINC00665 was the sponge for miR-1224-5p. By inhibiting miR-1224-5p level, LINC00665 dramatically promoted the expression of SND1 in PC cells. Ectopic expression of SND1 significantly rescued the effects of LINC00665 silencing.
Conclusion: LINC00665 is a novel oncogenic gene in PC by targeting miR-1224-5p/SND1 pathway and may be a therapeutic target.
Keywords: LINC00665, miR-1224-5p, SND1, prostate cancer, progression
Introduction
Prostate cancer (PC) is a very malignant and common cancer in men worldwide.1 This cancer has gradually become one of the leading causes, threatening men’s health.2 Although some improvement achieved on PC diagnosis, about 10% of new cases show advanced stages and about 5% ones display distant metastasis.3 Nowadays, the prognosis of PC patients is still unsatisfactory.4 Thus, understanding the molecular mechanism of PC pathogenesis is urgently required.
Long noncoding RNAs (lncRNAs) are characterized by no or limited protein-coding potential and have over 200 nucleotides in length.5 As the development of RNA sequencing, over 60,000 lncRNAs have been identified in human tissues.6 Although some reports have demonstrated the diverse functions of lncRNAs in biological processes, including tumor growth, metastasis and differentiation. Roles of most lncRNAs are still unclear.7,8 For example, LINC00460 upregulation in bladder urothelial cancer indicates poor prognosis.9 LncRNA H19 overexpression promotes epithelial–mesenchymal transition of papillary thyroid cancer.10 Aberrant downregulation of lncRNA STXBP5-AS1 in gastric cancer leads to accelerated growth and enhanced metastasis of tumor cells through activation of PI3K/AKT signal.11 Due to their important roles, several lncRNAs may be effective biomarkers for diagnosis or prognosis in cancer.12,13
LINC00665 has been reported to promote progression of several cancers, including lung cancer, gastric cancer and breast cancer.14–16 Nevertheless, the functions and mechanism of LINC00665 in PC are unclear. We found that LINC00665 was highly expressed in prostate cancer tissues through bioinformatics analysis and qRT-PCR method. Thus, we sought to determine the potential function and mechanism of LINC00665 in the regulation of PC progression. LINC00665 levels were elevated in PC tissues. LINC00665 knockdown suppressed PC cell proliferation, migration and invasion. Furthermore, LINC00665 was identified to be an effective sponge for miR-1224-5p, which further directly targeted SND1. Through regulating miR-1224-5p/SND1 axis, LINC00665 was demonstrated to act as oncogenic roles. Our results suggested that LINC00665 may serve as a novel target for PC therapy.
Materials and Methods
Clinical Tissue Specimens
Forty-one pairs of PC samples and normal controls were collected from First Affiliated Hospital of Wenzhou Medical University. All samples were histopathologically confirmed by two independent pathologists. This study was approved by the Ethics Committee of First Affiliated Hospital of Wenzhou Medical University. Written informed consents were achieved from patients. This study was conducted in accordance with the Declaration of Helsinki.
Cell Lines and Culture
Human PC cell lines (LNCaP, PC-3, DU-145 and 22RV1) and normal prostate epithelial cell line RWPE-1 were obtained from the American Type Culture Collection (ATCC, Manassas, VA, USA). Cells were cultured as previously described.17
qRT-PCR
RNA was isolated with TRIzol (Beyotime, Shanghai, China) and then reversely transcribed into complementary DNA (cDNA) using RETROscriptTM Reverse Transcription Kit (Invitrogen). qPCR was carried out by using VeriQuest SYBR Green qPCR Master Mix (Thermo Fisher Scientific, Waltham, MA). U6 was the normalized control for miRNA and 18S was the normalized control for lncRNA. Expression was calculated according to the 2−ΔΔCt method.
Cell Transfection
To knock down LINC00665, siLINC00665 (#1: 5ʹ-TACAAAGAAACAGGTGAAATT-3ʹ; #2: 5ʹ-GGGATTACAGGTCCAGATTTT-3ʹ) or control siRNA (5ʹ-AATTCTCCGAACGTGTCACGT-3ʹ) was transfected into DU-145 or LNCaP cells. After 48 hrs, the transfection efficiency was validated by qRT-PCR. siRNAs were purchased from GenePharma (Shanghai, China). For SND1 overexpression, the coding sequence of SND1 was constructed into the pcDNA3 vector.
CCK8 Assay
CCK8 assay (Dojindo, Kumamoto, Japan) was used to detect cell proliferation. In brief, 2000 cells per well were seeded into 96-well plates. After cultured for 0 day, 1 day, 2 days and 3 days, 10 μL CCK8 solution per well was added and incubated for 2 hrs. Then, absorbance at 450 nm was determined using a VarioskanTM LUX microplate reader (Thermo Fisher Scientific).
Colony Formation Assay
For colony formation assay, 500 cells per well were seeded into the 6-well plates. After cultured for 14 days, the clones were fixed and stained using crystal violet. And the colony number was calculated.
EdU Assay
EdU assay was performed as described before.18 Briefly, tumor cells were incubated with 10 μmol/L EdU for 4 h, followed by Hoechst staining. EdU positive cells were then analyzed by FACS.
Transwell Assay
Transwell assays were performed as reported.19
Xenograft Assay
Four-week Balb/c nude female mice were from Shanghai Laboratory Animal Company (Shanghai, China). DU-145 cells were injected subcutaneously into the right flank of nude mice (n=5 for each group). And the tumor volumes were measured every week. After 5 weeks, the tumor tissues were removed and weighed. The animal experiments were approved by the Ethics Committee of First Affiliated Hospital of Wenzhou Medical University. All animal operations were performed in accordance with the Animal Policy and Welfare Committee of First Affiliated Hospital of Wenzhou Medical University.
Luciferase Reporter Assay
Luciferase reporters were designed and reporter assay was performed as previously described.20 The association between LINC00665 and miR-1224-5p or between miR-1224-5p and SND1 was tested by using the Dual-Luciferase Reporter Assay System (Promega, Madison, WI). In brief, 5000 DU-145 cells per well were seeded into 24-well plates and transfected with pmirGLO reporter (Promega) containing wild-type (wt) or mutant (mut) LINC00665 or SND1-3ʹ-UTR region and miR-1224-5p mimics using Lipofectamine 3000 reagent (Invitrogen). After 48h, luciferase activity was determined according to the manufacturer’s protocols.
Statistical Analysis
GraphPad Prism Software was used for statistical analyses. Student’s t-test and one-way ANOVA were used for analysis of statistical significance as appropriately. Kaplan-Meier analysis and log-rank test were used for analysis of survival rate. P < 0.05 was considered as statistically significant.
Results
Expression Patterns of LINC00665 in PC Tissues and Cell Lines
To analyze important lncRNAs in PC progression, we used bioinformatics approach and identified LINC00665 as a significantly upregulated lncRNA in PC tissues (Figure 1A), which was further confirmed via TCGA data (Figure 1B). Through qRT-PCR analysis, we validated LINC00665 upregulation in 41 PC tissues compared to adjacent normal tissues (Figure 1C). Kaplan-Meier analysis show overexpression of LINC00665 predicted decreased survival rate in PC patients (Figure 1D). Then, LINC00665 level in PC cell lines was examined. We found that LINC00665 levels were also upregulated in PC cell lines (Figure 1E).
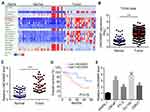 |
Figure 1 Expression patterns of LINC00665 in PC tissues and cell lines. (A) Differentially expressed lncRNAs in PC tissues and normal tissues according to TCGA database were analyzed through bioinformatics method (http://lncrnator.ewha.ac.kr/index.html). The heatmap data were directly available from the lncRNAtor database. TCGA: the cancer genome atlas. (B) Expression patterns of LINC00665 in PC tissues and normal controls based on TCGA data according to the TANRIC tool (https://ibl.mdanderson.org/tanric/_design/basic/query.html). (C) 41 pairs of PC tissues and normal tissues were used for the analysis of LINC00665 expression by qRT-PCR. 18S was as the normalized control. (D) Survival rate analysis of PC patients based on LINC00665 expression levels. The 41 patients were divided into LINC00665 high or low expression groups. (E) LINC00665 expression was determined in PC cell lines by qRT-PCR. **P<0.01 and ***P<0.001. |
Effects of LINC00665 Knockdown on PC Cell Proliferation
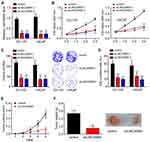
Then, we chose DU-145 and LNCaP cells for experiments. We knocked down LINC00665 expression in DU-145 and LNCaP cells by two independent siRNAs (Figure 2A). CCK8 assay was then performed. LINC00665 knockdown significantly inhibited proliferation of DU-145 and LNCaP cells (Figure 2B). Similarly, LINC00665 silencing led to reduced colony number and EdU incorporation in DU-145 and LNCaP cells (Figure 2C and D), suggesting LINC00665 regulates PC cell proliferation. Moreover, we performed xenograft experiments. At indicated time points, we measured tumor volumes. And 5 weeks later post-injection, we analyzed tumor weights. The results showed that LINC00665 knockdown suppressed tumor volumes and weights significantly (Figure 2E and F). Thus, LINC00665 promotes PC cell growth in vitro and in vivo.
LINC00665 Knockdown Suppressed PC Cell Migration and Invasion
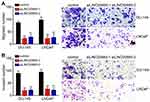
Afterwards, Transwell assay was carried out. We found that LINC00665 knockdown markedly reduced numbers of migrated and invaded DU-145 and LNCaP cells (Figure 3A and B), indicating that LINC00665 regulates PC cell metastasis.
LINC00665 Was a Sponge for miR-1224-5p
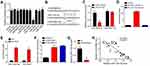
To further explore the mechanism, we analyzed the possible targets of LINC00665 through bioinformatics analysis (miRDB tool). We chose the ten most potential target miRNAs for validation. Through luciferase reporter assay, we found that only miR-1224-5p mimics inhibited the luciferase activity of LINC00665 reporter (Figure 4A). Then, we further constructed LINC00665-wt and mut luciferase reporter vectors (Figure 4B). Luciferase reporter assays demonstrated miR-1224-5p mimics suppressed the activity of LINC00665-wt vector but not the mutant one (Figure 4C). RNA pulldown assay also showed that LINC00665 was enriched by biotin-labeled miR-1224-5p-wt but not the mutant one or control (Figure 4D), suggesting a direct interaction between miR-1224-5p and LINC00665. RIP assay using anti-Ago2 also confirmed the direct interaction between LINC00665 and miR-1224-5p (Figure 4E). Importantly, LINC00665 knockdown also increased the level of miR-1224-5p and vice versa (Figure 4F and G). We also found a negative correlation between miR-1224-5p and LINC00665 in PC tissues (Figure 4H). Thus, LINC00665 was a sponge for miR-1224-5p.
SND1 Was Targeted by miR-1224-5p
We also analyzed the targets of miR-1224-5p through TargetScan7 tool. Many potential candidates were identified. And SND1 ranked top among all candidates. Thus, we chose it to validate. Firstly, the wt and mut SND1 luciferase reporter vectors were cloned (Figure 5A). Luciferase reporter assay showed that SND1-WT reporter activity was decreased as miR-1224-5p overexpression (Figure 5B). MiR-1224-5p mimics also inhibited SND1 level (Figure 5C). Moreover, qRT-PCR analysis and TCGA data indicated that SND1 expression was upregulated in PC tissues (Figure 5D and E), suggesting it may be an oncogene. We also observed that miR-1224-5p expression was negatively correlated with SND1 in PC tissues (Figure 5F).
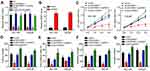
Ectopic Expression of SND1 Reversed the Effects of LINC00665 Knockdown
Through qRT-PCR analysis, we found that LINC00665 silencing suppressed SND1 expression (Figure 6A). However, inhibition of miR-1224-5p reversed it (Figure 6A). Thus, LINC00665 promoted SND1 expression by inhibiting miR-1224-5p in PC. To determine whether LINC00665 regulates PC progression through SND1, we overexpressed SND1 and confirmed its upregulation by qRT-PCR (Figure 6B). Then, we performed rescue assays. CCK8, colony formation and EdU incorporation assays indicated that ectopic expression of SND1 significantly promoted proliferation of DU-145 and LNCaP cells after LINC00665 knockdown (Figure 6C–E). Similarly, the abilities of migration and invasion were also rescued by SND1 overexpression in LINC00665-depleted DU-145 and LNCaP cells (Figure 6F and G). More importantly, we found that SND1 overexpression further promoted PC progression compared to the control group (Figure 6C–G). Thus, LINC00665 promotes PC progression through upregulating SND1 expression.
Discussion
As increasing numbers of lncRNAs are identified by RNA-sequencing, more and more attention are attracted to their functions in tumorigenesis.21 In our work, we revealed LINC00665 expression was upregulated in PC tissues. Furthermore, LINC00665 upregulation indicated unsatisfactory prognosis. We revealed that LINC00665 could activate SND1 expression via sponging miR-1224-5p. Then, upregulated SND1 expression led to enhanced proliferation, migration and invasion of PC cells.
Previously, several lncRNAs were reported to modulate various biological processes of PC cells.22,23 For example, lncRNA FOXC2-AS1 promotes growth and metastasis of PC cells through regulating miR-1253/EZH2 axis.24 LncRNA LOXL1-AS1 promotes CCND1 expression to facilitate the growth of PC cells via sponging miR-541-3p.25 PVT1 promotes PC cell proliferation and migration through modulating P38 expression.26 As a poorly researched lncRNA, LINC00665 was reported to promote lung cancer progression and tumor cell resistance to Gefitinib.14,27 Whether LINC00665 participates in PC progression remains largely unclear. We firstly identified LINC00665 as a significantly upregulated lncRNA in PC tissues by bioinformatics analysis. We also validated it by qRT-PCR. Moreover, we revealed that LINC00665 expression was closely linked to patients’ prognosis. Thus, LINC00665 might be a biomarker for prognosis in PC patients. Then, through loss-of-function assay, we demonstrated that LINC00665 knockdown suppressed growth and metastasis of PC cells.
LncRNA was shown to function as ceRNA for miRNAs.28,29 For instance, lncRNA LINC00460 sponges miR-939-5p to promote colorectal cancer invasion.30 LncRNA SLCO4A1-AS1 contributes to bladder cancer progression by inhibiting miR-335-5p.31 LncRNA PVT1 facilitates bladder cancer development by sponging miR-143.32 Thus, we analyzed the potential targets of LINC00665. We identified miR-1224-5p and demonstrated miR-1224-5p bound to LINC00665. MiR-1224-5p was found to suppress the development of several cancers, including gliomas and melanoma.33,34 Its roles in PC have not been reported. In our research, our data indicated that miR-1224-5p may be a tumor suppressor in PC. Furthermore, we also found the direct target of miR-1224-5p by bioinformatics analysis and identified SND1 (Staphylococcal nuclease domain-containing 1). Importantly, we found that LINC00665 promotes SND1 expression by inhibiting miR-1224-5p in PC cells.
SND1 has been shown to be overexpressed in several human tumor tissues. A recent study reported that SND1 overexpression promotes growth and metastasis of glioma cells.35 Besides, other works also indicated that SND1 upregulation contributes to tumorigenesis, such as osteosarcoma, hepatocellular carcinoma and breast cancer.36–38 Our data indicated that SND1 expression was upregulated in PC tissues. Moreover, we found that SND1 significantly enhanced proliferation, migration and invasion of PC cells transfected with LINC00665 siRNA, indicating the oncogenic roles of SND1 in PC.
Summarily, we demonstrated LINC00665 promoted PC progression through targeting miR-1224-5p/SND1.
Data Sharing Statement
All data were included in the manuscript.
Funding
Our work was granted by Wenzhou Science and Technology Project (Y20170231) and Zhejiang Provincial Natural Science Foundation of China under Grant (LQ20H050001 and LY20H160013).
Disclosure
The authors report no conflicts of interest in this work.
References
1. Martignano F, Rossi L, Maugeri A, et al. Urinary RNA-based biomarkers for prostate cancer detection. Clin Chim Acta. 2017;473:96–105. doi:10.1016/j.cca.2017.08.009
2. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2015. CA Cancer J Clin. 2015;65(1):5–29. doi:10.3322/caac.21254
3. Liu T, Chi H, Chen J, et al. Curcumin suppresses proliferation and in vitro invasion of human prostate cancer stem cells by ceRNA effect of miR-145 and lncRNA-ROR. Gene. 2017;631:29–38. doi:10.1016/j.gene.2017.08.008
4. Lin C, Wang J, Wang Y, et al. GRP78 participates in PCA3-regulated prostate cancer progression. Anticancer Res. 2017;37(8):4303–4310. doi:10.21873/anticanres.11823
5. Yabushita S, Fukamachi K, Tanaka H, et al. Metabolomic and transcriptomic profiling of human K-ras oncogene transgenic rats with pancreatic ductal adenocarcinomas. Carcinogenesis. 2013;34(6):1251–1259. doi:10.1093/carcin/bgt053
6. Iyer MK, Niknafs YS, Malik R, et al. The landscape of long noncoding RNAs in the human transcriptome. Nat Genet. 2015;47(3):199–208. doi:10.1038/ng.3192
7. Zhu P, Wang Y, Wu J, et al. LncBRM initiates YAP1 signalling activation to drive self-renewal of liver cancer stem cells. Nat Commun. 2016;7:13608. doi:10.1038/ncomms13608
8. Schmitt AM, Chang HY. Long noncoding RNAs in cancer pathways. Cancer Cell. 2016;29(4):452–463. doi:10.1016/j.ccell.2016.03.010
9. Wen L, Zhang X, Bian J, et al. The long non-coding RNA LINC00460 predicts the prognosis and promotes the proliferation and migration of cells in bladder urothelial carcinoma. Oncol Lett. 2019;17(4):3874–3880. doi:10.3892/ol.2019.10023
10. Liang WQ, Zeng CCF, Sun SM, Lu XF, Peng CY, Lin HY. Long noncoding RNA H19 is a critical oncogenic driver and contributes to epithelial-mesenchymal transition in papillary thyroid carcinoma. Cancer Manag Res. 2019;11:2059–2072. doi:10.2147/CMAR.S195906
11. Cen D, Huang H, Yang L, Guo K, Zhang J. Long noncoding RNA STXBP5-AS1 inhibits cell proliferation, migration, and invasion through inhibiting the PI3K/AKT signaling pathway in gastric cancer cells. Oncol Targets Ther. 2019;12:1929–1936. doi:10.2147/OTT.S194463
12. Chen X, Dai M, Zhu H, et al. Evaluation on the diagnostic and prognostic values of long non-coding RNA BLACAT1 in common types of human cancer. Mol Cancer. 2017;16(1):160. doi:10.1186/s12943-017-0728-2
13. Li JK, Chen C, Liu JY, et al. Long noncoding RNA MRCCAT1 promotes metastasis of clear cell renal cell carcinoma via inhibiting NPR3 and activating p38-MAPK signaling. Mol Cancer. 2017;16(1):111. doi:10.1186/s12943-017-0681-0
14. Cong Z, Diao Y, Xu Y, et al. Long non-coding RNA linc00665 promotes lung adenocarcinoma progression and functions as ceRNA to regulate AKR1B10-ERK signaling by sponging miR-98. Cell Death Dis. 2019;10(2):84. doi:10.1038/s41419-019-1361-3
15. Qi H, Xiao Z, Wang Y. Long non-coding RNA LINC00665 gastric cancer tumorigenesis by regulation miR-149-3p/RNF2 axis. Oncol Targets Ther. 2019;12:6981–6990. doi:10.2147/OTT.S214588
16. Guo B, Wu S, Zhu X, et al. CIP2A-BP encoded by LINC00665 inhibits triple-negative breast cancer progression. EMBO J. 2019;e102190.
17. Qin X, Zhu W, Lu A, Wang G, Ye X, Weng G. Long non-coding RNA SAP30L-AS1 promotes prostate cancer growth through repressing SAP30L. Gene. 2019;690:120–128. doi:10.1016/j.gene.2018.12.047
18. Qiu X, Dong J, Zhao Z, Li J, Cai X. LncRNA LINC00668 promotes the progression of breast cancer by inhibiting apoptosis and accelerating cell cycle. Oncol Targets Ther. 2019;12:5615–5625. doi:10.2147/OTT.S188933
19. Zhang G, He X, Ren C, Lin J, Wang Q. Long noncoding RNA PCA3 regulates prostate cancer through sponging miR-218-5p and modulating high mobility group box 1. J Cell Physiol. 2018.
20. Sun W, Zhang L, Yan R, Yang Y, Meng X. LncRNA DLX6-AS1 promotes the proliferation, invasion, and migration of non-small cell lung cancer cells by targeting the miR-27b-3p/GSPT1 axis. Oncol Targets Ther. 2019;12:3945–3954. doi:10.2147/OTT.S196865
21. Hua JT, Ahmed M, Guo H, et al. Risk SNP-mediated promoter-enhancer switching drives prostate cancer through lncRNA PCAT19. Cell. 2018;174(3):564–575 e18. doi:10.1016/j.cell.2018.06.014
22. Wu M, Huang Y, Chen T, et al. LncRNA MEG3 inhibits the progression of prostate cancer by modulating miR-9-5p/QKI-5 axis. J Cell Mol Med. 2019;23(1):29–38. doi:10.1111/jcmm.2019.23.issue-1
23. Chen Q, Sun T, Wang F, et al. Long noncoding RNA IGF2AS is acting as an epigenetic tumor suppressor in human prostate cancer. Urology. 2019;124:310 e1–310 e8. doi:10.1016/j.urology.2018.11.002
24. Chen Y, Gu M, Liu C, et al. Long noncoding RNA FOXC2-AS1 facilitates the proliferation and progression of prostate cancer via targeting miR-1253/EZH2. Gene. 2019;686:37–42. doi:10.1016/j.gene.2018.10.085
25. Long B, Li N, Xu XX, et al. Long noncoding RNA LOXL1-AS1 regulates prostate cancer cell proliferation and cell cycle progression through miR-541-3p and CCND1. Biochem Biophys Res Commun. 2018;505(2):561–568. doi:10.1016/j.bbrc.2018.09.160
26. Wan B, Wu HY, Lv DJ, et al. Downregulation of lncRNA PVT1 expression inhibits proliferation and migration by regulating p38 expression in prostate cancer. Oncol Lett. 2018;16(4):5160–5166. doi:10.3892/ol.2018.9305
27. Liu X, Lu X, Zhen F, et al. LINC00665 induces acquired resistance to gefitinib through recruiting EZH2 and activating PI3K/AKT pathway in NSCLC. Mol Ther Nucleic Acids. 2019;16:155–161. doi:10.1016/j.omtn.2019.02.010
28. Ji D, Wang Y, Li H, Sun B, Luo X. Long non-coding RNA LINC00461/miR-149-5p/LRIG2 axis regulates hepatocellular carcinoma progression. Biochem Biophys Res Commun. 2019. doi:10.1016/j.bbrc.2019.03.049
29. Yu J, Zhang B, Zhang H, et al. E2F1-induced upregulation of long non-coding RNA LMCD1-AS1 facilitates cholangiocarcinoma cell progression by regulating miR-345-5p/COL6A3 pathway. Biochem Biophys Res Commun. 2019. doi:10.1016/j.bbrc.2019.03.054
30. Zhang Y, Liu X, Li Q, Zhang Y. lncRNA LINC00460 promoted colorectal cancer cells metastasis via miR-939-5p sponging. Cancer Manag Res. 2019;11:1779–1789. doi:10.2147/CMAR.S192452
31. Yang Y, Wang F, Huang H, Zhang Y, Xie H, Men T. lncRNA SLCO4A1-AS1 promotes growth and invasion of bladder cancer through sponging miR-335-5p to upregulate OCT4. Oncol Targets Ther. 2019;12:1351–1358. doi:10.2147/OTT.S191740
32. Chen J, Yu Y, Li H, et al. Long non-coding RNA PVT1 promotes tumor progression by regulating the miR-143/HK2 axis in gallbladder cancer. Mol Cancer. 2019;18(1):33. doi:10.1186/s12943-019-0947-9
33. Qian J, Li R, Wang YY, et al. MiR-1224-5p acts as a tumor suppressor by targeting CREB1 in malignant gliomas. Mol Cell Biochem. 2015;403(1–2):33–41. doi:10.1007/s11010-015-2334-1
34. Wang Q, Zhang R, Liu D. Long non-coding RNA ZEB1-AS1 indicates poor prognosis and promotes melanoma progression through targeting miR-1224-5p. Exp Ther Med. 2019;17(1):857–862. doi:10.3892/etm.2018.7005
35. Yu L, Xu J, Liu J, et al. The novel chromatin architectural regulator SND1 promotes glioma proliferation and invasion and predicts the prognosis of patients. Neuro Oncol. 2019. doi:10.1093/neuonc/noz038
36. Zhou M, Wang A, Yin B, et al. SND1 promotes the proliferation of osteosarcoma cells by upregulating COX2/PGE2 expression via activation of NFkappaB. Oncol Rep. 2019;41(1):579–589. doi:10.3892/or.2018.6822
37. Cui X, Zhao C, Yao X, et al. SND1 acts as an anti-apoptotic factor via regulating the expression of lncRNA UCA1 in hepatocellular carcinoma. RNA Biol. 2018;15(10):1364–1375. doi:10.1080/15476286.2018.1534525
38. Chen L, Yuan Y, Kar S, et al. PPARgamma ligand-induced Annexin A1 Expression determines chemotherapy response via deubiquitination of death domain kinase rip in triple-negative breast cancers. Mol Cancer Ther. 2017;16(11):2528–2542. doi:10.1158/1535-7163.MCT-16-0739
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.