Back to Journals » OncoTargets and Therapy » Volume 13
LncRNA CTBP1-AS2 Facilitates Gastric Cancer Progression via Regulating the miR-139-3p/MMP11 Axis
Authors Yang Y , Gao M, Li Y, Li M, Ma Q
Received 25 May 2020
Accepted for publication 29 September 2020
Published 10 November 2020 Volume 2020:13 Pages 11537—11547
DOI https://doi.org/10.2147/OTT.S264394
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Federico Perche
Yudan Yang,1 Ming Gao,1 Yunpeng Li,2 Mengyi Li,1 Qingqing Ma1
1Department of Oncology, The First Affiliated Hospital of Zhengzhou University, Zhengzhou 450052, Henan, People’s Republic of China; 2Department of Cardiology, The First Affiliated Hospital of Zhengzhou University, Zhengzhou 450052, Henan, People’s Republic of China
Correspondence: Yudan Yang Email [email protected]
Background: This study aimed at probing into the effect of long non-coding RNA (lncRNA) C-terminal binding protein 1 antisense RNA 2 (CTBP1-AS2) on gastric cancer (GC) cell proliferation and apoptosis, and its regulatory function on miR-139-3p and MMP11.
Methods: Quantitative real-time polymerase chain reaction (qRT-PCR) was employed to examine the expressions of CTBP1-AS2, miR-139-3p and MMP11 mRNA in GC cell lines and clinical specimens. Cell counting kit-8 (CCK-8) assay, flow cytometry and EdU assay were conducted to examine the effects of CTBP1-AS2 and miR-139-3p on GC cell proliferation and apoptosis. Western blot was applied for detecting the expressions of Bax, Bcl-2 and MMP11. A lung metastasis mouse model was used to evaluate metastasis of GC cells in vivo. Bioinformatics, dual-luciferase report assay, RIP and RNA pull-down assays were utilized to validate the targeted relationship between CTBP1-AS2 and miR-139-3p as well as the targeting relationship between miR-139-3p and MMP11.
Results: CTBP1-AS2 was highly expressed in GC, and its high expression was strongly associated with increased TNM stage, increased tumor size and low degree of differentiation of the tumor tissues. Meanwhile, CTBP1-AS2 promoted GC cell proliferation, metastasis and suppressed apoptosis, while miR-139-3p could weaken these effects. In addition, CTBP1-AS2 was identified as a molecular sponge for miR-139-3p, and MMP11 was verified as a target gene of CTBP1-AS2. CTBP1-AS2 could increase the expression of MMP11 via repressing miR-139-3p.
Conclusion: CTBP1-AS2 promotes GC cells and inhibits apoptosis by regulating the miR-139-3p/MMP11 molecular axis.
Keywords: CTBP1-AS2, GC, proliferation, apoptosis
Introduction
Ranking fifth in morbidity and third in mortality globally, gastric cancer (GC) is defined as one of the most threatening malignancies.1,2 Currently, surgery remains the primary treatment for GC, and the five-year overall survival rate among GC patients is only approximately 35%.3 Exploring the mechanism of GC pathogenesis is of great scientific and clinical significance, which is expected to open up new paths for the clinical prevention and treatment of GC.
Recognized as a class of RNAs with a length of over 200 nucleotides, long non-coding RNAs (lncRNAs) are mainly located in the nucleus or cytoplasm and cannot encode proteins.4 Reportedly, lncRNAs can markedly regulate the progression of multiple cancers including GC.5–7 For instance, as an oncogenic lncRNA, lncRNA AFDN-DT can inhibit GC cell growth through transcriptional regulation;8 as a tumor-suppressive factor, LINC00460 is significantly up-regulated in GC tissues, and promotes the malignant biological behaviors of cancer cells via epigenetically silencing CCNG2.9 LncRNA C-terminal binding protein 1 antisense RNA 2 (CTBP1-AS2) is not only related to the susceptibility to type 2 diabetes10 but also closely connected with the recurrence of papillary thyroid cancer.11 Nevertheless, the role of CTBP1-AS2 in cancer biology, especially in GC, is still unclear.
MicroRNAs (miRNAs), also belonging to non-coding RNAs (ncRNAs), emerge as a type of endogenous ncRNAs with 19–24 nucleotides, and can bind to the 3ʹ-UTR of target mRNA to further modulate the target gene expression.12 In addition, miRNAs take part in regulating the malignant phenotypes of cancer cells in many different tumors including GC, such as cell proliferation, differentiation and apoptosis.13–15 For example, miR-638 is down-regulated in GC, and its overexpression suppresses cell growth via inhibiting MACCI expression.16 MiR-139-3p is lowly expressed in ovarian cancer tissues and it impedes tumor cell growth, and ELAVL1 is its downstream target which can be negatively regulated by it.17 Besides, miR-139-3p can negatively regulate matrix metalloproteinase 11 (MMP11), and via this mechanism, miR-139-3p can repress GC cell growth and promote apoptosis.18 However, the upstream mechanism for regulating miR-139-3p in GC is still unclear.
With the data from the cancer genome atlas (TCGA), we noticed that CTBP1-AS2 was up-regulated in GC tissues, suggesting its relationship with the tumorigenesis of GC. In this work, we studied the expression characteristic, prognostic value, and biological function of CTBP1-AS2 in GC. Furthermore, we established a competitive endogenous RNA (ceRNA) network with CTBP1-AS1, miR-139-3p and MMP11, which participates in the progression of GC.
Materials and Methods
Clinical Samples
For the present study, tissue samples, including GC tissues and adjacent tissues (3 cm away from GC tissue), of 37 patients were collected during the gastrectomy in the First Affiliated Hospital of Zhengzhou University, from June 2017 to August 2019. All samples were immediately stored in liquid nitrogen after collection until RNA extraction. The diagnosis of the patients was confirmed by pathological examination. The informed consent from the patients was obtained before the gastrectomy. The study protocol was endorsed by the Medical Research Ethics Committee of the First Affiliated Hospital of Zhengzhou University.
Cell Culture
Human normal gastric mucosal cells (GES-1) and GC cell lines (SNU-1, NCI-N87, HGC-27 and AGS) were purchased from the American Type Culture Collection (ATCC; Rockville, MD, USA). All the above cells were cultured in Dulbecco’s Modified Eagle Medium/Ham’s Nutrient mixture F12 (DMEM/F12, Gibco, Grand Island, NY, USA) with 100 U/mL penicillin and 0.1 mg/mL streptomycin (Sigma, St. Louis, MO, USA) and 10% fetal bovine serum (FBS; HyClone, Logan, UT, USA) in 5% CO2 at 37°C. When reaching 70–80% confluence, the cells were treated with 0.25% trypsin and subcultured.
Cell Transfection
SNU-1 and AGS cells were used for transfection when their confluency was around 80%. LipofectaminTM2000 (Thermo Fisher Scientific, Waltham, MA, USA) was utilized to transfect AGS cells with CTBP1-AS2 overexpression plasmids and SNU-1 cells with siRNAs targeting CTBP1-AS2.
Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR)
After AGS and SNU-1 cells during logarithmic growth were collected, the cell lysis was performed using TRIzol reagent (Invitrogen, Carlsbad, CA, USA). Total RNA extraction was conducted following the instruction of the kit, and Nanodrop-spectrophotometer was employed to measure the RNA purity and concentration. Then, RNAs were kept at –80°C for further use. Based on the manufacturer’s protocol, complementary DNA (cDNA) was synthesized from 1 µg of total RNA with the PrimeScript-RT Kit (Madison, WI, USA), and then the SYBR® Premix-Ex-Taq™ (Takara, TX, USA) and ABI7300 system were used to perform PCR. U6 and GAPDH were used as the internal references for miRNA, and mRNA (lncRNA), respectively. All experiments were conducted in triplicate, and Table 1 shows the specific primer sequences.
 |
Table 1 Sequences Used for qRT-PCR |
Databases and Bioinformatics Analysis
GEPIA (http://gepia.cancer-pku.cn/) was utilized to analyze CTBP1-AS2 expression in GC samples from TCGA; Kaplan–Meier plotter (http://kmplot.com/analysis/index.php) was employed to relationship between CTBP1-AS2 and the prognosis of GC patients; LncBase Predicted v2.0 (http://carolina.imis.athena-innovation.gr/diana_tools/web/index.php) was employed to predict the candidate miRNAs binding to CTBP1-AS2, and TargetScan (http://www.targetscan.org/) was used for predicting downstream mRNAs of the miRNA.
Cell Counting Kit-8 (CCK-8) Assay
After the transfection of SNU-1 cells and AGS cells were resuspended in 10% serum-containing DMEM/F12 medium, and the cell concentration was modulated to 1 × 104 cells/mL. The cells were subsequently inoculated into 96-well culture plates (100 μL/well) with 6 replicate wells in each group. Ten microliters of CCK-8 solution (Thermo Fisher Scientific, Waltham, MA, USA) was added into each well of the plate. After 1 h of culture, the optical density (OD) value of each well was measured at 450 nm. The OD values at 0, 24, 48 and 72 h were measured, respectively. Then, the growth curve was plotted.
EdU Assay
Twenty-four hours after the transfection, medium was discarded and cells were rinsed once with PBS. Subsequently, cells were trypsinized to prepare single-cell suspension, which was then seeded into 96-well plates and incubated for 12 h. The EdU working solution (Beyotime Biotechnology, Shanghai, China) was added, with which the cells were incubated for 4 h. Next, 100 μL of paraformaldehyde fixative was added before cells were incubated on a decolorizing shaker for 30 min at room temperature. 2 mg/mL glycine was added, followed by incubation for 5 min on a decolorizing shaker. After that, 100 μL 1 × Apollo® staining solution was added prior to the incubation for 30 min. Then, 100 μL of penetrant was added, with which the cells were incubated for 10 min. Afterwards, DAPI reaction liquid was added and the cells were incubated at room temperature for 30 min. After the reaction liquid was washed off with PBS, the pictures of cells were taken under the fluorescence microscope. The blue fluorescence represented the total number of cells detected, and the red fluorescence symbolized proliferated cells. Cell proliferation rate = number of red fluorescence cells/number of blue fluorescence cells × 100%.
Flow Cytometry to Detect Apoptosis
AnnexinV/7-AAD Apoptosis Detection Kit (Southern Biotechnology, Birmingham, Al, USA) was used. Cells during logarithmic growth were taken. The cells of each group were centrifuged for 5 min at 2000 r/min, and medium was discarded. Five microliters of 7-AAD staining solution was added into 500 μL of binding buffer, then the cells were resuspended. Next, the cells in each group were added with 1 μL of AnnexinV-PE staining solution and incubated in the dark at room temperature for 15 min. After that, the cells were immediately detected by flow cytometry, the result of which was expressed by the apoptosis rate.
Dual-Luciferase Reporter Assay
The dual-luciferase activity detection kit was purchased from Promega Corporation (Madison, WI, USA), and luciferase reporter vectors were also synthesized by Promega. Reporter gene plasmids for wild-type (WT) and mutant-type (MUT) CTBP1-AS2 and MMP11-3ʹUTR were co-transfected into HEK293T cells with miR-139-3p and miR-con, respectively. After 48 h of transfection, for each group, firefly luciferase and renilla luciferase activities were detected with a microplate reader. Firefly luciferase activity was then normalized by renilla luciferase activity.
RNA Pull-Down Assay
AGS cells were lysed using RIPA lysis buffer (Biossci, Wuhan, China), and then incubated with Biotinylated (Bio)-NC, Bio-CTBP1-AS2-WT or Bio-CTBP1-AS2-MUT (GenePharma, Shanghai, China) for 1 h. Next, Dynabeads M-280 streptavidin (Invitrogen, CA, USA) was employed to isolate biotin-labeled RNA, and qRT-PCR was utilized for detecting the enrichment of miR-139-3p in the pull-down complex.
RNA Immunoprecipitation (RIP)
RIP analysis was conducted with EZ-Magna RIP™ RNA-Binding Protein Immunoprecipitation Kit (Merck Millipore, Billerica, MA, USA). Briefly, based on the manufacturer’s instructions, AGS and SNU-1 cells were lysed in RIP lysis buffer, and incubated with anti-IgG or anti-Ago2 antibodies conjugated with magnetic beads. After that, the lysis was incubated with proteinase K to remove the proteins. After that, RNA was extracted with TRIzol method, and then qRT-PCR was performed for detecting the expression of co-precipitated RNAs.
Western Blot
RIPA buffer (Biossci, Wuhan, China) was added to the cells in each group. After fully mixed, the lysis was incubated on ice for 30 min. Then, the lysis was centrifuged for 15 min at 12,000 r/min at 4°C, and the supernatant was collected. After SDS-PAGE, the proteins were transferred to PVDF membranes (Merck Millipore, Billerica, MA, USA) and afterwards the membranes were blocked with 5% skimmed milk. Then, the membranes were incubated at 4°C overnight with primary antibodies anti-Bcl-2, Bax, MMP11 and β-actin. After washing the membranes with TBST, they were incubated with HRP-labeled secondary antibodies for 1 h at room temperature. Subsequently, equivoluminal liquid A and liquid B of the hypersensitive ECL chemiluminescence kit (Millipore, Bedford, MA, USA) was mixed and cover the PVDF membranes to react with antigen-antibody complex. Then, the protein bands were scanned. The antibodies used in this study included anti-MMP11 antibody (ab53143, 1:2000), anti-Bcl-2 antibody (ab185002, 1:2000), anti-Bax antibody (ab32503, 1:2000) and anti-β-actin antibody (ab179467, 1:2000), which were all available from Abcam (Shanghai, China).
Metastasis Assays in vivo
All procedures in animal experiments were approved by the Animal Care and Use Committee of The First Affiliated Hospital of Zhengzhou University and performed strictly in accordance with institutional policies and approved guidelines of experiment operations. Briefly, nude mice, 4–6 weeks old, were purchased from Zhengzhou University Experimental Animal Center (Henan, China). 2 × 106 AGS cells (suspended in 0.2 mL PBS) were injected into the lateral tail vein of each mouse (n=4 in each group). After 30 days, the mice were sacrificed, and their lungs were examined for tumor metastases after hematoxylin and eosin staining.
Statistical Analysis
The statistical software SPSS22.0 (SPSS Inc., Chicago, IL, USA) was employed for the analysis of experimental data, and GraphPad Prism 8.0 (La Jolla, CA, USA) was utilized to draw the graphs. Mean ± standard deviation was the expression form of all data. One-way ANOVA was performed for comparing the difference among 3 or more groups, and t-test was utilized for comparing the difference between two groups. P < 0.05 represented that differences were of statistical significance.
Results
CTBP1-AS2 Was Up-Regulated in GC and Related to Patients’ Poor Prognosis
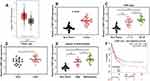
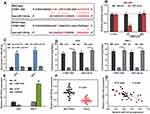
At first, we found through GEPIA database that in GC samples of TCGA, CTBP1-AS2 was up-regulated (Figure 1A). Subsequently, qRT-PCR was carried out to examine CTBP1-AS2 expression in the adjacent tissue and GC tissue samples of 37 patients, and consistent with the result of GEPIA, compared to adjacent tissues, CTBP1-AS2 expression level in GC tissues was significantly higher (Figure 1B). Next, the relationship between CTBP1-AS2 expression and pathological indicators of patients was analyzed, and the results demonstrated that high CTBP1-AS2 expression was markedly linked to the increase in TNM stage, larger tumor size and low degree of differentiation (Figure 1C–E). In addition, Kaplan–Meier plotter displayed that CTBP1-AS2 expression was negatively correlated with the patient’s post-progression survival (Figure 1F). These data indicate that in GC, CTBP1-AS2 was highly expressed and was significantly associated with the patient’s poor prognosis.
CTBP1-AS2 Promoted GC Cell Proliferation, Metastasis and Inhibited Apoptosis
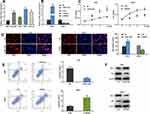
To further decipher the biological effects of CTBP1-AS2 in GC, we detected its expression in GC cells by qRT-PCR and found that in comparison to GES-1 cells, GC cell lines (SNU-1, NCI-N87, HGC-27 and AGS) expressed higher level of CTBP1-AS2; among the four GC cell lines, CTBP1-AS2 expression level was the highest in SNU-1 cells and was the lowest in AGS cells (Figure 2A). Therefore, AGS cells were transfected with CTBP1-AS2 overexpression plasmids, and siRNAs targeting CTBP1-AS2 were transfected into SNU-1 cells. qRT-PCR showed that the transfection was successful; additionally, the knockdown effect of si-RNA#2 was more significant than that of si-RNA#1, so it was used in the subsequent experiments (Figure 2B). CCK-8 assay, EdU assay and flow cytometry experiments were carried out to examine cell proliferation and apoptosis, and it was revealed that in contrast to the NC group, overexpression of CTBP1-AS2 observably promoted AGS cell proliferation and suppressed apoptosis; as against the si-NC group, si-RNA#2 significantly inhibited SNU-1 cell proliferation and facilitated apoptosis (Figure 2C–E). Western blot results showed that in the CTBP1-AS2 overexpression group, Bcl-2 expression was higher and Bax expression was lower; conversely, in the si-RNA#2 group, opposite phenomena could be found (Figure 2F). Then, we used a lung metastasis mouse model to evaluate the potential of GC cells to metastasize, and it indicated that over-expression of CTBP1-AS2 enhanced pulmonary metastasis of AGS cells in vivo (Supplementary Figure 1A). Collectively, these results indicated that CTBP1-AS2 could promote GC cell proliferation, and metastasis, and inhibit apoptosis.
CTBP1-AS2 Regulated miR-139-3p
With LncBase Predicted v2.0, it was found that CTBP1-AS2 had the binding site for miR-139-3p (Figure 3A). Dual-luciferase reporter gene assay indicated that compared to miR-con, miR-139-3p mimics could inhibit the luciferase activity of CTBP1-AS2 WT, while it did not significantly affect the luciferase activity of CTBP1-AS2 MUT (Figure 3B). To further verify the direct interaction between CTBP1-AS2 and miR-139-3p, RNA pull-down and Ago2-RIP assays were performed. RNA pull-down assay illustrated that Bio-CTBP1-AS2 WT greatly increased the enrichment of miR-139-3p compared to Bio-probe NC in AGS cells (Figure 3C). Additionally, RIP assay suggested that, endogenous CTBP1-AS2 and miR-139-3p in Ago2-expressed cell lines were enriched in Ago2 pellet in comparison to the input control (Figure 3D). Moreover, qRT-PCR showed that CTBP1-AS2 overexpression could inhibit miR-139-3p expression, while si-RNA#2 promoted its expression (Figure 3E). Furthermore, miR-139-3p expression was down-regulated in GC tissues, and CTBP1-AS2 expression was in negative correlation with miR-139-3p expression (Figure 3F–G). To sum up, CTBP1-AS2 directly targeted miR-139-3p and negatively regulated its expression in GC.
CTBP1-AS2 Promoted GC Cell Proliferation and Suppressed Apoptosis by Adsorbing miR-139-3p
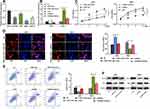
The foregoing experiments showed that CTBP1-AS2 could adsorb miR-139-3p, thereby inhibiting its expression. Then, rescue experiments were performed. CTBP1-AS2 + miR-139-3p mimics were transfected into AGS cells, and si-RNA#2 + miR-139-3p inhibitors were transfected into SNU-1 cells. Moreover, qRT-PCR verified the success of the transfection (Figure 4A and B). Subsequently, CCK-8 assay, EdU assay, flow cytometry and Western blot manifested that miR-139-3p mimics could weaken the promotional effect of CTBP1-AS2 on GC cell proliferation and the inhibitory effect of CTBP1-AS2 on GC cell apoptosis, while miR-139-3p inhibitors could reverse the inhibiting effect of si-RNA#2 on GC cell proliferation and the promotional effect of CTBP1-AS2 of on GC cell apoptosis (Figure 4C–F). These indicated that CTBP1-AS2 could promote GC cell proliferation and inhibit apoptosis by regulating miR-139-3p.
CTBP1-AS2 Up-Regulated MMP11 Expression by Adsorbing miR-139-3p
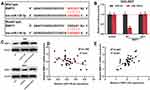
It is reported that miR-139-3p targetedly regulates MMP11 expression.18 In addition, Kaplan–Meier plotter displayed that MMP11 expression was negatively correlated with the patient’s survival time (Supplementary Figure 1B). TargetScan was utilized to predict the binding site between miR-139-3p and MMP11 (Figure 5A). Consistently, dual-luciferase reporter gene assay suggested that the luciferase activity of the cells was significantly decreased after co-transfection of miR-139-3p and MMP11 WT, while there was no significant change in the luciferase activity of the cells after co-transfection of miR-139-3p and MMP11 MUT (Figure 5B). Subsequently, Western blot indicated that miR-139-3p mimics could suppress MMP11 expression, while miR-139-3p inhibitors promoted its expression; CTBP1-AS2 enhanced MMP11 expression, while CTBP1-AS2 knockdown inhibited its expression (Figure 5C). In addition, miR-139-3p was negatively correlated with MMP11 expression in GC samples (Figure 5D); CTBP1-AS2 was positively correlated with MMP11 expression in GC samples (Figure 5E). All the above-mentioned findings supported that CTBP1-AS2 could up-regulate MMP11 expression by adsorbing miR-139-3p.
Discussion
LncRNAs are involved in regulating multiple biological processes, and many of them are dysregulated in tumors, causing enhanced cell proliferation, metastasis and inhibited apoptosis.19 For example, lncRNA DRAIC is down-regulated in GC, and it can interact with UCHL5 to mediate NFRKB ubiquitination, in turn suppress the tumorigenesis of GC;20 LINC00460 is overexpressed in GC tissues, and it facilitate the proliferation, migration and invasion of GC cells by epigenetically silencing tumor suppressor CCNG2.9 LncRNA CTPB1-AS2 is a newly discovered lncRNA, and its function and mechanism in cancer biology is rarely investigated. A recent study reports that CTBP1-AS2 expression is significantly related to recurrence-free survival time of patients with papillary thyroid cancer, showing its potential to be used as a cancer-related biomarker.11 In this study, for the first time, we explored the role of CTBP1-AS2 in GC. We observed that CTBP1-AS2 expression was up-regulated in GC and markedly linked to the post-progression survival time of GC patients; high CTBP1-AS2 expression was also significantly associated with the increase in TNM stage, the increase of tumor size and low degree of differentiation of the tumor tissues. Further in vitro and in vivo experiments indicated that lncRNA CTBP1-AS2 could promote GC cell proliferation and metastasis, and inhibit apoptosis. These results proved that CTBP1-AS2 exerted a tumor-promoting function in GC.
Accumulating researches illustrate that in GC, lncRNAs competitively decoy miRNAs via miRNA response elements (MREs), and this process reduces the availability of miRNAs, and indirectly up-regulates the level of downstream mRNAs.21–23 For example, the expression of lncRNA DDX11-AS1 is up-regulated in GC tissues, and functionally, DDX11-AS1 facilitates GC progression by inhibiting miR-873-5p expression.24 LncRNA FTX expression in GC is also up-regulated, and FTX can promote GC proliferation and invasion by adsorbing miR-144 and up-regulating ZFX;25 conversely, as a tumor suppressor, lncRNA HAND2-AS1 inhibits the proliferation and migration of GC cells through modulating miR-590-3p/KCNT2 axis.26 In this study, we verified that CTBP1-AS2 was a molecular sponge for miR-139-3p. Furthermore, functional experiments illustrated that CTBP1-AS2 promoted cell proliferation and suppressed apoptosis by inhibiting miR-139-3p expression.
The tumor-suppressive characteristics of miR-139-3p have been well described in the previous studies. Its target gene includes a series of oncogenes including ELAVL1, Bcl-6, Wnt5A, Rab1A and so on.17,27–29 Matrix metalloproteinase (MMPs) are zinc-dependent endopeptidases and key regulators of extracellular matrix (ECM) degradation. As a member of the MMP family, matrix metalloproteinase 11 (MMP11) is associated with embryonic tissue remodeling, ECM remodeling, epithelial growth and wound healing.30 MMP11 expression in pancreatic cancer tissues is up-regulated, and high MMP11 expression is associated with the poor prognosis of pancreatic cancer patients.31 In papillary thyroid carcinoma, MMP11 expression is up-regulated; besides, MMP11 promotes colony formation and metastasis of cancer cells and is negatively regulated by miR-125a-3p.32 In GC, cancer-associated fibroblasts (CAFs) are the primary components of tumor stroma, and MMP11 is highly expressed in gastric CAFs, which is negatively regulated by miR-139-3p.18 Another study reports that in bladder cancer, miR-139-3p can significantly inhibit migration and invasion of cancer cells via repressing MMP11.33 In this work, consistently, we proved that miR-139-3p could suppress the expression of MMP11 via targeting it, and CTBP1-AS2 could positively regulate MMP11, probably via its regulation on miR-139-3p. These demonstrations partly explained the mechanism by which CTBP1-AS1/miR-139-3p participated in GC progression.
Altogether, this study shows that CTBP1-AS2 can promote GC cell proliferation, metastasis and inhibit apoptosis through targeted regulation of the miR-139-3p/MMP11 molecular axis, and high CTBP1-AS2 expression is significantly associated with poor prognosis of patients. Therefore, CTBP1-AS2 can serve as a promising therapeutic target and biomarkers for GC.
Data Sharing Statement
The data used to support the findings of this study are available from the corresponding author upon request.
Ethics Statement
Our study was approved by the Ethics Review Board of The First Affiliated Hospital of Zhengzhou University.
Author Contributions
All authors made a significant contribution to the work reported, whether that is in the conception, study design, execution, acquisition of data, analysis and interpretation, or in all these areas; took part in drafting, revising or critically reviewing the article; gave final approval of the version to be published; have agreed on the OncoTargets and Therapy; and agree to be accountable for all aspects of the work.
Funding
This work was supported by grants from Funded by Science and Technology Project of Henan Province (NO. 182102310346).
Disclosure
The authors declare that they have no competing interests.
References
1. Wang N, Lu K, Qu H, et al. CircRBM33 regulates IL-6 to promote gastric cancer progression through targeting miR-149. Biomed Pharmacother. 2020;125:109876. doi:10.1016/j.biopha.2020.109876
2. Tian H, Wang W, Meng X, et al. ERas enhances resistance to cisplatin-induced apoptosis by suppressing autophagy in gastric cancer cell. Front Cell Dev Biol. 2019;7:375. doi:10.3389/fcell.2019.00375
3. Ma H-W, Xi D-Y, Ma J-Z, et al. Long noncoding RNA AFAP1-AS1 promotes cell proliferation and metastasis via the miR-155-5p/FGF7 axis and predicts poor prognosis in gastric cancer. Dis Markers. 2020;2020:8140989. doi:10.1155/2020/8140989
4. Luo Y, Yang J, Yu J, et al. Long non-coding RNAs: emerging roles in the immunosuppressive tumor microenvironment. Front Oncol. 2020;10:48. doi:10.3389/fonc.2020.00048
5. Yang C, Cai X, Yu M, et al. Long noncoding RNA OR3A4 promotes the proliferation and invasion of osteosarcoma cells by sponging miR-1227-5p. J Bone Oncol. 2020;21:100278. doi:10.1016/j.jbo.2020.100278
6. Jia W, Zhang J, Ma F, et al. Long noncoding RNA THAP9-AS1 is induced by and promotes cell growth and migration of gastric cancer. Onco Targets Ther. 2019;12:6653–6663. doi:10.2147/OTT.S201832
7. Wang Z, Zhang Q, Sun Y, Shao F. Long non-coding RNA PVT1 regulates BAMBI to promote tumor progression in non-small cell lung cancer by sponging miR-17-5p. Onco Targets Ther. 2020;13:131–142. doi:10.2147/OTT.S217335
8. Lai Y, Xu P, Wang J, Xu K, Wang L, Meng Y. Tumour suppressive long non-coding RNA AFDN-DT inhibits gastric cancer invasion via transcriptional regulation. J Cell Mol Med. 2020;24(5):3157–3166. doi:10.1111/jcmm.14988
9. Yang J, Lian Y, Yang R, et al. Upregulation of lncRNA LINC00460 facilitates GC progression through epigenetically silencing CCNG2 by EZH2/LSD1 and indicates poor outcomes. Mol Ther Nucleic Acids. 2020;19:1164–1175. doi:10.1016/j.omtn.2019.12.041
10. Erfanian Omidvar M, Ghaedi H, Kazerouni F, et al. Clinical significance of long noncoding RNA VIM-AS1 and CTBP1-AS2 expression in type 2 diabetes. J Cell Biochem. 2019;120(6):9315–9323. doi:10.1002/jcb.28206
11. Ma B, Liao T, Wen D, et al. Long intergenic non-coding RNA 271 is predictive of a poorer prognosis of papillary thyroid cancer. Sci Rep. 2016;6:36973. doi:10.1038/srep36973
12. Yao J, Zhang H, Liu C, Chen S, Qian R, Zhao K. miR-450b-3p inhibited the proliferation of gastric cancer via regulating KLF7. Cancer Cell Int. 2020;20:47. doi:10.1186/s12935-020-1133-2
13. Ogawa H, Nakashiro K-I, Tokuzen N, Kuribayashi N, Goda H, Uchida D. MicroRNA-361-3p is a potent therapeutic target for oral squamous cell carcinoma. Cancer Sci. 2020;111(5):1645–1651. doi:10.1111/cas.14359
14. Lin X, Qiu W, Xiao Y, et al. MiR-199b-5p suppresses tumor angiogenesis mediated by vascular endothelial cells in breast cancer by targeting ALK1. Front Genet. 2019;10:1397. doi:10.3389/fgene.2019.01397
15. Chen F-R, Sha S-M, Wang S-H, et al. RP11-81H3.2 promotes gastric cancer progression through miR-339-HNRNPA1 interaction network. Cancer Med. 2020;9(7):2524–2534. doi:10.1002/cam4.2867
16. Wei CJ, Zhang ZW, Lu JH, Mao YM. MiR-638 regulates gastric cardia adenocarcinoma cell proliferation, apoptosis, migration and invasion by targeting MACC1. Neoplasma. 2020;67(3):537–546. doi:10.4149/neo_2020_190719N651
17. Xue F, Li QR, Xu YH, Zhou HB. MicroRNA-139-3p inhibits the growth and metastasis of ovarian cancer by inhibiting ELAVL1. OncoTargets Ther. 2019;12:8935–8945. doi:10.2147/OTT.S210739
18. Xu G, Zhang B, Ye J, et al. Exosomal miRNA-139 in cancer-associated fibroblasts inhibits gastric cancer progression by repressing MMP11 expression. Int J Biol Sci. 2019;15(11):2320–2329. doi:10.7150/ijbs.33750
19. Martínez-Barriocanal Á, Arango D, Dopeso H. PVT1 long non-coding RNA in gastrointestinal cancer. Front Oncol. 2020;10:38. doi:10.3389/fonc.2020.00038
20. Zhang Z, Hu X, Kuang J, Liao J, Yuan Q. LncRNA DRAIC inhibits proliferation and metastasis of gastric cancer cells through interfering with NFRKB deubiquitination mediated by UCHL5. Cell Mol Biol Lett. 2020;25:29. doi:10.1186/s11658-020-00221-0
21. Liu K, Wang B-J, Han W, et al. CFIm25-regulated lncRNA acv3UTR promotes gastric tumorigenesis via miR-590-5p/YAP1 axis. Oncogene. 2020;39(15):3075–3088. doi:10.1038/s41388-020-1213-8
22. Zhang W, Zhan F, Li D, Wang T, Huang H. RGMB-AS1/miR-22-3p/NFIB axis contributes to the progression of gastric cancer. Neoplasma. 2020;67(3):484–491. doi:10.4149/neo_2020_190418N350
23. Dong D, Lun Y, Sun B, et al. Silencing of long non-coding RNA PCAT6 restrains gastric cancer cell proliferation and epithelial-mesenchymal transition by targeting microRNA-15a. Gen Physiol Biophys. 2020;39(1):1–12. doi:10.4149/gpb_2019044
24. Ren Z, Liu X, Si Y, Yang D. Long non-coding RNA DDX11-AS1 facilitates gastric cancer progression by regulating miR-873-5p/SPC18 axis. Artif Cells, Nanomed Biotechnol. 2020;48(1):572–583. doi:10.1080/21691401.2020.1726937
25. Li H, Yao G, Zhai J, Hu D, Fan Y. LncRNA FTX promotes proliferation and invasion of gastric cancer via miR-144/ZFX axis. Onco Targets Ther. 2019;12:11701–11713. doi:10.2147/OTT.S220998
26. Yu L, Li H, Li Z, et al. Long non-coding RNA HAND2-AS1 inhibits growth and migration of gastric cancer cells through regulating the miR-590-3p/KCNT2 axis. Onco Targets Ther. 2020;13:3187–3196. doi:10.2147/OTT.S233256
27. Xu Y, Yu H, Liu G. Hsa_circ_0031288/hsa-miR-139-3p/Bcl-6 regulatory feedback circuit influences the invasion and migration of cervical cancer HeLa cells. J Cell Biochem. 2020;121(10):4251–4260. doi:10.1002/jcb.29650
28. Huo LW, Wang YF, Bai XB, Zheng HL, Wang MD. circKIF4A promotes tumorogenesis of glioma by targeting miR-139-3p to activate Wnt5a signaling. Mol Med. 2020;26(1):29. doi:10.1186/s10020-020-00159-1
29. Pei FL, Cao MZ, Li YF. Circ_0000218 plays a carcinogenic role in colorectal cancer progression by regulating miR-139-3p/RAB1A axis. J Biochem. 2020;167(1):55–65. doi:10.1093/jb/mvz078
30. Li -C-C, Hsieh M-J, Wang -S-S, et al. Impact of matrix metalloproteinases 11 gene variants on urothelial cell carcinoma development and clinical characteristics. Int J Environ Res Public Health. 2020;17(2):475. doi:10.3390/ijerph17020475
31. Lee J, Lee J, Kim JH. Identification of matrix metalloproteinase 11 as a prognostic biomarker in pancreatic cancer. Anticancer Res. 2019;39(11):5963–5971. doi:10.21873/anticanres.13801
32. Song M, Wang N, Li Z, et al. miR-125a-3p suppresses the growth and progression of papillary thyroid carcinoma cell by targeting MMP11. J Cell Biochem. 2020;121(2):984–995. doi:10.1002/jcb.29333
33. Yonemori M, Seki N, Yoshino H, et al. Dual tumor-suppressors miR-139-5p and miR-139-3p targeting matrix metalloprotease 11 in bladder cancer. Cancer Sci. 2016;107(9):1233–1242. doi:10.1111/cas.13002
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.