Back to Journals » OncoTargets and Therapy » Volume 13
LncRNA CASC15 is Upregulated in Osteosarcoma Plasma Exosomes and CASC15 Knockdown Inhibits Osteosarcoma Progression by Regulating miR-338-3p/RAB14 Axis
Authors Zhang H , Wang J, Ren T, Huang Y, Yu Y, Chen C, Huang Q, Guo W
Received 16 September 2020
Accepted for publication 5 November 2020
Published 23 November 2020 Volume 2020:13 Pages 12055—12066
DOI https://doi.org/10.2147/OTT.S282053
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Leo Jen-Liang Su
Hongliang Zhang,1,2 Jun Wang,1,2 Tingting Ren,1,2 Yi Huang,1,2 Yiyang Yu,1,2 Chenglong Chen,1,2 Qingshan Huang,1,2 Wei Guo1,2
1Musculoskeletal Tumor Center, Peking University People’s Hospital, Beijing, People’s Republic of China; 2Beijing Key Laboratory of Musculoskeletal Tumor, Beijing, People’s Republic of China
Correspondence: Wei Guo Email [email protected]
Background: Currently, plenty of studies have demonstrated that lncRNAs can act as crucial roles during the progression of various tumors, including osteosarcoma (OS), and emerging evidences indicated that lncRNAs are abundant and stable in exosomes. The objective of this study is to reveal the dysregulated lncRNAs in OS plasma exosomes and explore their functions in OS.
Materials and Methods: Microarray was performed to analyze dysregulated exosomal lncRNAs. Western blot, qRT-PCR assays, and Dual-luciferase reporter assay were used to verify the interaction among cancer susceptibility 15 (CASC15), miR-338-3p, and RAB14. Cck-8, colony formation assay, and transwell assay were performed to explore and characterize the effects of CASC15 on OS cells. Animal experiments were used to verify the effects of CASC15 in vivo.
Results: Upregulated CASC15 was observed in OS plasma exosomes compared with control, and the same expression was observed in the OS tissues and cell lines. Further assays indicated that CASC15 knockdown could restrain the proliferation, migration, and invasion of OS cells, and inhibit the growth of OS in xenograft models. Furthermore, our results demonstrated CASC15 regulated OS progression via acting as miR-338-3p sponge, and RAB14 was a direct downstream target of miR-338-3p. Rescue experiments verified CASC15 promotes OS cell growth and metastasis by upregulating RAB14 expression.
Conclusion: Overall, our findings indicate that CASC15 plays a key role in OS progression by targeting the miR-338-3p/RAB14 axis and can serve as a possible therapeutic target for OS patients.
Keywords: bone tumor, exosome, lncRNA, CASC15, miR-338-3p
Introduction
Osteosarcoma (OS) is a common type of malignant bone tumor that often occurs in people under the age of 20.1 And in the past 30 years, the 5-year survival rate of OS patients has increased to 60–70% attributed to the improvement of the traditional treatments and the emergence of novel therapeutic strategies, including neoadjuvant chemotherapy, targeted therapies, and immunotherapy.2 However, the prognosis is still dissatisfactory because of the high rates of drug resistance, relapse, and metastasis.3 Thus, more researches are urgently needed to clarify the molecular mechanisms and highlight the potential therapeutic targets of OS.
Exosome, a type of extracellular vehicle ranging from 30 to 150 nm in diameter, has been reported that could be a key mediator between cellular communication.4 And emerging evidences indicated that lncRNAs are abundant and stable in exosomes.5 However, there are limited studies for the expression profile of plasma exosomal lncRNAs and their roles in OS.
LncRNA is a kind of non-coding RNA, and plenty of lncRNAs have been demonstrated to participate in the development of various tumors.6 Also, increasing evidences have shown the participation of lncRNAs in various biological processes of OS.7 CASC15 is a novel tumor-related lncRNA, and its aberrant expression has been reported in several tumors, like liver cancer, ovarian cancer, and melanoma.8–10 But in OS, the expression and function of CASC15 is still vague and needs to be explored.
Therefore, our objective is to reveal the expression of CASC15 both in plasma exosomes and tissues of OS patients and elucidate the function and mechanisms of CASC15 in tumor progression.
Materials and Methods
Clinical Specimens
Thirty osteosarcoma specimens, thirty normal bone tissues, and five OS plasma were collected at the Musculoskeletal Tumor Center, Peking University People’s Hospital (Beijing, China). The health plasma was donated by five health donators (HD). Written informed consent was acquired from all patients, and the study was approved by the Ethics Committee of Peking University People’s Hospital.
Cell Culture and Transfection
OS cell lines were obtained from the American Type Culture Collection (ATCC, Manassas, VA, USA). All cells were cultured in DMEM medium (Hyclone, Logan, UT, USA), and maintained in a moist incubator with 5% CO2 at 37°C. Agents like sh-CASC15/sh-NC, siRAB14 #1, siRAB14 #2, miR-338-3p inhibitor/mimic, and NC inhibitor/mimic were synthesized by GenePharma (Shanghai, China), and transfected through Lipofectamine 3000 (Invitrogen, Waltham, MA, USA).
2.3.qRT-PCR
TRIzol (Invitrogen) was used to isolate total RNAs, and the PrimeScriptRT reagent Kit (Takara, Kusatsu, Japan) was used to synthesize cDNAs. Then, the qRT-PCR was performed on a CFX96 Real-Time System (Bio-Rad, Hercules, CA, USA) using iTaq Universal One-Step RT-qPCR Kits (Bio-Rad). U6/GAPDH was used as endogenous control.
Western Blot
The assay was performed as previously described.11 And anti-CD9 (1:1000, sc-13,118, Santa Cruz, USA), anti-CD63 (1:1000, sc-5275), anti-N-cadherin (1:1000, Cell Signaling Technology (CST), 13,116), anti-E-cadherin (1:1000, CST, 3195), anti-RAB14 (1:1000, sc-271,401) and anti-Actin (1:1000, sc-8432) were used in this study.
Colony Formation and EdU Assay
Eight hundred cells were added in 6-well plates and cultured for 120 hours. Then, the residual medium was discarded, and cells were washed two times. Finally, methanol was used to fix cells, and the 0.1% crystalline violet solution was used to stain cells. The cell colonies were observed and photographed for further analysis. EdU assay was performed as previously described.12
Transwell and Wound-Healing Assays
For transwell assay, Matrigel-coated or noncoated Transwell plate chambers (BD Biosciences, USA) was used. In short, bottom chambers were added with complete medium, and top chambers were added with a serum-free medium. Thirty thousand cells were planted in top chambers, and cultured for 24 hours. Then, the residual medium was discarded, and the chambers were washed three times. Methanol was used to fix cells, and the 0.1% crystalline violet solution was used to stain cells. Finally, five fields were observed and photographed using an inverted microscope. For the wound-healing assay, cells in complete medium were added in 6-well plates, and when cell confluence reached 80–90%, a wound was scratched by a pipette tip. Then, the residual medium was discarded, and replaced with a medium with 0.2% FBS. The status of the wound was observed and photographed at 0h and 24h separately using an inverted microscope.
Exosome Extraction and Microarray Analysis
Exosupur Kit (Echo Biotech, Beijing, China) was used to extract the exosomes from OS or HD plasma based on the principle of Size Exclusion Chromatography (SEC). Then, 10µL of exosome solution was pipetted onto the copper mesh, and stained with 2% uranyl acetate. Finally, the exosomes were observed and photographed under a transmission electron microscope (TEM) at 80 kV. Besides, after exosomes were diluted, its size distribution was analyzed by nanoparticle tracking analysis (NTA). For Microarray analysis, the plasma was collected from five OS patients and five health donators, and the exosome was isolated by the method described above. Then, TRIzol was used to isolate total RNAs, and analysis was performed by the commercial corporation Echo Biotech (Beijing, China). Besides, p-value <0.05 and |log2 Fold change|≧1 were used as the threshold for dysregulated lncRNAs.
Dual-Luciferase Reporter Assay
The assay was performed as previously described.13
Immunohistochemistry (IHC)
IHC staining was performed as previously described [13]. And anti-RAB14 (1:80, sc-271,401, Santa Cruz), anti-N-cadherin (1:100, CST, 13,116), anti-E-cadherin (1:100, CST, 3195) and anti- Ki-67 (1:100, sc-23,900, Santa Cruz) were used in this study.
Tumor Xenografts
In this study, 8 BALB/c nude mice were used in this research, and 5×106 transfected 143B cells were subcutaneously injected in the right flanks of mice. All mice were fed in the SPF environment, and tumor volume (width2 × length/2) was measured every four days for sixteen days. Then, mice were sacrificed and samples were processed for further research. All animal experiments were performed with written confirmation authorized by the Animal Care and Use Committee of Peking University People’s Hospital. Animal experiments complied with the ARRIVE guidelines and followed the National Institutes of Health Guide for the Care and Use of Laboratory Animals.
Statistical Analysis
The data of this study were analyzed by GraphPad Prism 8 and expressed as mean ± SD. The significant differences between groups were analyzed using Student’s t-test or Fisher’s exact test, and indicated by *p < 0.05 and **p < 0.01.
Results
Expression Profiles of Plasma Exosomal lncRNAs in OS
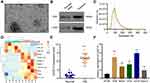
Plasma exosomes were extracted by SEC, and TEM results showed that exosomes have a bowl-shaped structure with a diameter of 100–150 nm (Figure 1A). Exosome marker proteins CD9 and CD63 were detected by Western blot (Figure 1B). And the result of NTA revealed that the diameter of plasma exosomes ranged around 100 nm (Figure 1C). Furthermore, Microarray was performed to analyze dysregulated exosomal lncRNAs, and the results demonstrated that 3 lncRNAs were downregulated and 8 lncRNAs were upregulated in plasma exosomes of OS patients compared with that of health donators (Figure 1D). Among which, we noticed that CASC15 was significantly upregulated in OS plasma exosomes, whereas there have no reports about its expression and function in OS.
CASC15 is Upregulated in OS Tissues and Associated with Tumor Metastasis
We performed qRT-PCR in thirty OS specimens and thirty normal bone tissues to further explore the expression of CASC15, and the results revealed that CASC15 was upregulated in OS tissues (Figure 1E). Consistently, the expression of CASC15 was higher in OS cell lines than that in normal osteoblast hFOB (Figure 1F). Furthermore, the result of the correlation analysis indicated that CASC15 upregulation was associated with tumor metastasis (p=0.004, cutoff value=mean) (Table 1).
 |
Table 1 Relationship Between CASC15 Expression Level and Clinicopathological Features in Osteosarcoma |
CASC15 Knockdown Restrains the Proliferation, Migration, and Invasion of OS Cells
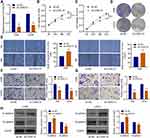
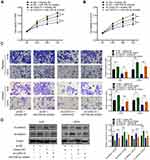
After the OS cells were transfected with sh-CASC15, qRT-PCR was performed to verify the knockdown of CASC15 (Figure 2A). CCK8 assay revealed that CASC15 knockdown could restrain the proliferation of 143B and U2OS cells (Figure 2B and C). Consistently, further assays demonstrated that CASC15 knockdown could restrain the colony formation ability of 143B and U2OS cells (Figure 2D). Furthermore, wound-healing assays revealed that CASC15 knockdown could suppress the migration ability of 143B and U2OS cells (Figure 2E). And transwell assays revealed that CASC15 knockdown could restrain the ability of migration (Figure 2F) and invasion of OS cells (Figure 2G). Western blot results showed that CASC15 knockdown could significantly upregulate the expression of E-cadherin and downregulated the expression of N-cadherin in OS cells (Figure 2H).
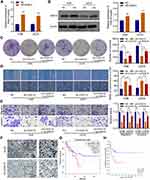
CASC15 Knockdown Inhibits the Growth of OS in vivo
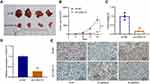
To explore whether CASC15 knockdown could inhibit the growth of OS in vivo, xenograft models were established using 143B cells. Compared with the control, CASC15 knockdown significantly reduced the growth rate, tumor volume, and tumor weight of OS (Figure 3A-C). qRT-PCR result verified that CASC15 was downregulated in the sh-CASC15 group compared with the sh-NC group (Figure 3D). Besides, IHC was performed in xenograft samples, and downregulated Ki67, N-cadherin, but upregulated E-cadherin was observed in CASC15 knockdown group (Figure 3E). All these findings suggested that CASC15 knockdown inhibited the growth of OS in vivo.
miR-338-3p is a Target Gene of CASC15
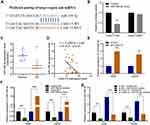
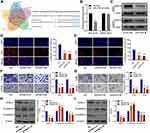
Using DIANA-LncBase v2, miR-338-3p was predicted as a potential target miRNA that could be sponged by CASC15 (Figure 4A). And dual-luciferase reporter assay verified their direct interaction (Figure 4B). Then, further analysis in Gene Expression Omnibus (GEO) database (GSE28423) demonstrated the downregulation of miR-338-3p in OS (Figure 4C), and qRT-PCR results revealed that miR-338-3p expression was negatively correlated with CASC15 expression in OS tissues (Figure 4D). Furthermore, CASC15 knockdown significantly upregulated miR-338-3p in OS cells, whereas the miR-338-3p inhibitor could reverse its effect (Figure 4E-G).
CASC15 Regulated OS Progression via Acting as miR-338-3p Sponge
Consistently, functional assays demonstrated that sh-CASC15-mediated inhibition of proliferation, migration, and invasion could be reversed by miR-338-3p inhibitor in OS cells (Figure 5A-C). Furthermore, Western blot results showed that miR-338-3p inhibitor could reverse sh-CASC15-mediated regulation of epithelial-mesenchymal transition (EMT) markers E-cadherin and N-cadherin in OS cells (Figure 5D). All these results collectively indicated that CASC15 regulated OS progression via acting as a miR-338-3p sponge.
RAB14 Was a Direct Target of miR-338-3p
Five bioinformatics tools were used to predict the target genes of miR-338-3p, and RAB14 was found as one of two intersecting genes in all five databases (Figure 6A). Subsequent dual-luciferase reporter assay verified their direct interaction (Figure 6B), and Western blot indicated that miR-338-3p could downregulate RAB14 protein expression (Figure 6C). Besides, siRAB14 sequences were used to knock down the expression of RAB14 in OS cells, and the results of the EdU assay indicated that the knockdown of RAB14 significantly inhibited cell proliferation (Figure 6D and E). And transwell assays revealed that RAB14 knockdown could suppress the ability of migration (Figure 6F) and invasion of OS cells (Figure 6G). Consistently, Western blot results showed that RAB14 knockdown could significantly upregulate the expression of E-cadherin and downregulated the expression of N-cadherin in OS cells (Figure 6H and I).
CASC15 Promotes OS Cell Growth and Metastasis by Upregulating RAB14 Expression
We further performed rescue experiments to prove that CASC15 promotes OS cell growth and metastasis by upregulating RAB14 expression. As shown in Figure 7A and B, the overexpression vector targeting RAB14 was transfected into OS cells, and overexpressed RAB14 was observed at both mRNA and protein levels. Then, the colony formation assays showed that RAB14 overexpression could significantly reverse sh-CASC15-induced proliferation inhibition (Figure 7C). And the results of wound-healing and transwell indicated that RAB14 overexpression could also partially reverse sh-CASC15- mediated metastasis inhibition in OS cells (Figure 7D and E). Furthermore, IHC staining of RAB14 was performed in xenograft samples, and downregulated RAB14 was observed in the CASC15 knockdown group (Figure 7F). Kaplan-Meier assays were analyzed using the GEPIA database and R2 database. GEPIA result showed that patients with highly expressed RAB14 displayed poorer overall survival rates (p=0.048) (Figure 7G), and the R2 result indicated that metastasis-free survival probability of OS patients was negatively associated with RAB14 expression (p=0.007) (Figure 7H). The aforementioned findings indicated that RAB14 was released in OS cells due to CASC15 sponges miR-338-3p, and participated in the regulatory network of OS progression.
Discussion
LncRNAs have been reported as pivotal regulators in the development of various tumors.14 Mounting evidences revealed that lncRNAs can be abundant and stable in exosome, which is recently regarded as an important mediator of intercellular communication.15 Inspired by these evidences, we wanted to screen dysregulated lncRNAs in OS plasma exosomes and investigate their function in OS development. We performed microarray analysis and the result indicated that CASC15 was significantly upregulated in OS plasma exosomes compared with control.
CASC15 is a lncRNA locus in chromosome 6p22 and has been reported as a tumor promoter in several tumors. For example, in gastric cancer, Wu et al found that CASC15 was enriched and could promote tumor progression by targeting CDKN1A and ZEB1.16 In lung cancer, Bai et al reported that upregulated CASC15 could regulate tumor progression by targeting the miR-766-5p/KLK12 axis.17 Similarly, we verified that CASC15 was upregulated, and could serve as an oncogene in OS.
Plenty of recent evidences have revealed that lncRNAs could work as miRNA sponges to downregulate the expression of target miRNAs.18 Accordingly, we used bioinformatics prediction tools to predict the downstream target of CASC15, and miR-338-3p was predicted as one of the target genes. Given that aberrantly expressed miR-338-3p has been widely researched either as an oncogene or antioncogene in various tumors,19–21 and several recent articles also reported that miR-338-3p was lowly expressed in OS tissues,22,23 miR-338-3p was chosen as a candidate target gene of CASC15 for further verification and research. For example, Jia et al reported that miR-338-3p was downregulated in OS and could inhibit OS progression via regulating RUNX2 and CDK4.22 And Cao et al demonstrated that poorly expressed miR-338-3p could be an antioncogene by targeting AHSA1 in OS.23 As shown in results, our findings also indicated that miR-338-3p was lowly expressed in OS tissues and CASC15 regulated OS development via targeting miR-338-3p.
To further explore the downstream target gene of miR-338-3p, we used five bioinformatics prediction tools to predict, and RAB14 and LARP4 were found as two intersecting genes in all five databases. Recently, RAB14 has been reported as an oncogene in several tumors, including oral squamous cell carcinoma, pancreatic cancer, and bladder cancer.24–26 And several researches have revealed that miR-338-3p can directly bind to the 3ʹ-UTR of RAB14.27,28 Thus, we chose RAB14 for further research and confirmed that RAB14 was a direct target gene of miR-338-3p in OS cells. All these results indicated that RAB14 can be released in OS cells due to CASC15 sponges miR-338-3p, and participated in the regulatory network of OS progression.
Furthermore, our results demonstrated that knockdown of RAB14 significantly inhibited the proliferation, migration, and invasion of OS cells, and overexpression of RAB14 could partially reverse CASC15-knockdown mediated proliferation inhibition and metastasis inhibition. Besides, Kaplan-Meier assays revealed patients with highly expressed RAB14 displayed poorer prognosis. Taken together, in OS, RAB14 was confirmed as a key member in the regulatory network of CASC15.
EMT is a crucial process that participated in the metastasis of OS.27,29 And in the present study, we found that EMT markers E-cadherin and N-cadherin were associated with the expression of CASC15/miR-338-3p/RAB14 axis, indicating that CASC15/miR-338-3p/RAB14 axis can regulate the invasion and metastasis of OS via EMT.
In conclusion, this research demonstrates that CASC15 is upregulated in OS plasma exosomes, and CASC15 knockdown can inhibit osteosarcoma progression by targeting the miR-338-3p/RAB14 axis. Which uncovers a new insight into molecular mechanisms of OS, and highlights the possibility of CASC15 as a biomarker and therapeutic target for OS patients.
Ethics Statement
This experiment was approved by the ethics committee of Peking University People’s Hospital. Animal experiments complied with the ARRIVE guidelines and followed the National Institutes of Health Guide for the Care and Use of Laboratory Animals.
Funding
This work was supported by the National Natural Science Foundation of China (No. 81572633, NO.81702657 and No.81972509) and Beijing Municipal Natural Science Foundation (No. 7182170).
Disclosure
The authors declare no conflicts of interests.
References
1. Isakoff MS, Bielack SS, Meltzer P, Gorlick R. Osteosarcoma: current treatment and a collaborative pathway to success. J Clin Oncol. 2015;33(27):3029–3035. doi:10.1200/JCO.2014.59.4895
2. Dean DC, Shen S, Hornicek FJ, Duan Z. From genomics to metabolomics: emerging metastatic biomarkers in osteosarcoma. Cancer Metastasis Rev. 2018;37:719–731. doi:10.1007/s10555-018-9763-8
3. Niu J, Yan T, Guo W, Wang W, Zhao Z. Insight into the role of autophagy in osteosarcoma and its therapeutic implication. Front Oncol. 2019;9:1232. doi:10.3389/fonc.2019.01232
4. Wortzel I, Dror S, Kenific CM, Lyden D. Exosome-mediated metastasis: communication from a distance. Dev Cell. 2019;49:347–360. doi:10.1016/j.devcel.2019.04.011
5. Fan Q, Yang L, Zhang X, et al. The emerging role of exosome-derived non-coding RNAs in cancer biology. Cancer Lett. 2018;414:107–115. doi:10.1016/j.canlet.2017.10.040
6. Chi Y, Wang J, Wang J, Yu W, Yang J. Long non-coding RNA in the pathogenesis of cancers. Cells. 2019;8(9):1015. doi:10.3390/cells8091015
7. Yang Z, et al. Long noncoding RNAs in the progression, metastasis, and prognosis of osteosarcoma. Cell Death Dis. 2016;7:e2389. doi:10.1038/cddis.2016.272
8. Wang C, Zi H, Wang Y, Li B, Ge Z, Ren X. LncRNA CASC15 promotes tumour progression through SOX4/Wnt/beta-catenin signalling pathway in hepatocellular carcinoma. Artif Cells Nanomed Biotechnol. 2020;48:763–769. doi:10.1080/21691401.2019.1576713
9. Shi Y, Gao S, Zheng Y, Yao M, Ruan F. LncRNA CASC15 functions as an unfavorable predictor of ovarian cancer prognosis and inhibits tumor progression through regulation of miR-221/ARID1A axis. Onco Targets Ther. 2019;12:8725–8736. doi:10.2147/OTT.S219900
10. Yin Y, Zhao B, Li D, Yin G. Long non-coding RNA CASC15 promotes melanoma progression by epigenetically regulating PDCD4. Cell Biosci. 2018;8:42. doi:10.1186/s13578-018-0240-4
11. Zhang H, Yang K, Ren T, et al. miR-100-5p inhibits malignant behavior of chordoma cells by targeting IGF1R. Cancer Manag Res. 2020;12:4129–4137. doi:10.2147/CMAR.S252185
12. Zhang H, Yang K, Ren T, et al. Bone marrow mesenchymal stem cell-derived exosomal miR-206 inhibits osteosarcoma progression by targeting TRA2B. Cancer Lett. 2020. doi:10.1016/j.canlet.2020.07.008
13. Zhang H, et al. miR-16-5p inhibits chordoma cell proliferation, invasion and metastasis by targeting Smad3. Cell Death Dis. 2018;9:680. doi:10.1038/s41419-018-0738-z
14. Bhan A, Soleimani M, Mandal SS. Long noncoding RNA and cancer: a new paradigm. Cancer Res. 2017;77:3965–3981. doi:10.1158/0008-5472.CAN-16-2634
15. Sun Z, Yang S, Zhou Q, et al. Emerging role of exosome-derived long non-coding RNAs in tumor microenvironment. Mol Cancer. 2018;17:1. doi:10.1186/s12943-018-0831-z
16. Wu Q, Xiang S, Ma J, et al. Long non-coding RNA CASC 15 regulates gastric cancer cell proliferation, migration and epithelial mesenchymal transition by targeting CDKN1A and ZEB1. Mol Oncol. 2018;12:799–813. doi:10.1002/1878-0261.12187
17. Bai Y, Zhang G, Cheng R, Yang R, Chu H. CASC15 contributes to proliferation and invasion through regulating miR-766-5p/KLK12 axis in lung cancer. Cell Cycle. 2019;18:2323–2331. doi:10.1080/15384101.2019.1646562
18. Thomson DW, Dinger ME. Endogenous microRNA sponges: evidence and controversy. Nat Rev Genet. 2016;17:272–283. doi:10.1038/nrg.2016.20
19. Wang Y, Qin H. miR-338-3p targets RAB23 and suppresses tumorigenicity of prostate cancer cells. Am J Cancer Res. 2018;8:2564–2574.
20. Zhang R, Shi H, Ren F, et al. MicroRNA-338-3p suppresses ovarian cancer cells growth and metastasis: implication of Wnt/catenin beta and MEK/ERK signaling pathways. J Exp Clin Cancer Res. 2019;38:494. doi:10.1186/s13046-019-1494-3
21. Sun F, Yu M, Yu J, et al. miR-338-3p functions as a tumor suppressor in gastric cancer by targeting PTP1B. Cell Death Dis. 2018;9:522. doi:10.1038/s41419-018-0611-0
22. Jia F, Zhang Z, Zhang X. MicroRNA-338-3p inhibits tumor growth and metastasis in osteosarcoma cells by targeting RUNX2/CDK4 and inhibition of MAPK pathway. J Cell Biochem. 2019;120:6420–6430. doi:10.1002/jcb.27929
23. Cao R, Shao J, Hu Y, et al. microRNA-338-3p inhibits proliferation, migration, invasion, and EMT in osteosarcoma cells by targeting activator of 90 kDa heat shock protein ATPase homolog 1. Cancer Cell Int. 2018;18:1. doi:10.1186/s12935-018-0551-x
24. Lian Q, Ma DM, Chen MG, Chen K, Li XJ. Silencing Rab14 represses the proliferation and migration of oral squamous cell carcinoma, and enhances cisplatin sensitivity. Am J Transl Res. 2017;9:4195–4205.
25. Ge J, Ge C. Rab14 overexpression regulates gemcitabine sensitivity through regulation of Bcl-2 and mitochondrial function in pancreatic cancer. Virchows Arch. 2019;474:59–69. doi:10.1007/s00428-018-2455-5
26. Chao H, Deng L, Xu F, et al. RAB14 activates MAPK signaling to promote bladder tumorigenesis. Carcinogenesis. 2019;40:1341–1351. doi:10.1093/carcin/bgz039
27. Xu Y, Yu J, Huang Z, et al. Circular RNA hsa_circ_0000326 acts as a miR-338-3p sponge to facilitate lung adenocarcinoma progression. J Exp Clin Cancer Res. 2020;39:57. doi:10.1186/s13046-020-01556-4
28. Li M, Bian Z, Jin G, et al. LncRNA SNHG 15 enhances cell proliferation in colorectal cancer by inhibiting miR-338-3p. Cancer Med. 2019;8:2404–2413. doi:10.1002/cam4.2105
29. Zhu ST, Wang X, Wang JY, Xi GH, Liu Y. Downregulation of miR-22 contributes to epithelial-mesenchymal transition in osteosarcoma by targeting Twist1. Front Oncol. 2020;10:406. doi:10.3389/fonc.2020.00406
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.