Back to Journals » Cancer Management and Research » Volume 13
LncRNA AGAP2-AS1 Promotes Cancer Cell Proliferation, Migration and Invasion in Colon Cancer by Forming a Negative Feedback Loop with LINC-PINT
Authors Ji L, Chen S, Gu L, Wang J, Zhang X
Received 28 April 2020
Accepted for publication 25 November 2020
Published 2 March 2021 Volume 2021:13 Pages 2153—2161
DOI https://doi.org/10.2147/CMAR.S260371
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Eileen O'Reilly
Liechen Ji,1,* Shuo Chen,1,* Liqiang Gu,1 Juan Wang,2 Xipeng Zhang1
1Department of Colorectal Surgery, Tianjin Union Medical Center, Nankai University, Tianjin, 300121, People’s Republic of China; 2Department of General Surgery, Tianjin Union Medical Center, Nankai University, Tianjin, 300121, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Xipeng Zhang
Department of Colorectal Surgery, Tianjin Union Medical Center, Nankai University, No. 190 Jieyuan Road, Hongqiao District, Tianjin, 300121, People’s Republic of China
Tel +862287729595
Email [email protected]
Introduction: It has been reported that lncRNA AGAP2-AS1 promotes the development of gastric cancer, lung cancer and breast cancer. This study aimed to investigate the role of AGAP2-AS1 in colon cancer.
Methods: A total of 66 patients with colon cancer were enrolled. RT-qPCR was performed to detect the differential expression of AGAP2-AS1 in tumor tissues and adjacent normal tissues. To test the interaction between AGAP2-AS1 and LINC-PINT in colon cancer, overexpression vector or inhibitor of AGAP2-AS1 and LINC-PINT were transfected into RKO and HCT 116 cells. CCK-8 assay was used to detect cell proliferation. Transwell assays were performed to evaluate cell migration and invasion. The expression of p-LATS1, p-YAP and nuclear YAP were detected by Western blot and immunofluorescence.
Results: The expression of AGAP2-AS1 was upregulated in colon cancer tissues compared with that in adjacent normal tissues, and the expression of AGAP2-AS1 in colon cancer tissues was not significantly affected by tumor stages. In addition, we found that the expression of LINC-PINT was downregulated in colon cancer tissues compared with that in adjacent normal tissues and had a reverse correlation with the expression of AGAP2-AS1 in colon cancer tissues. Moreover, overexpression of AGAP2-AS1 downregulated the expression of LINC-PINT, and overexpression of LINC-PINT also altered the expression of AGAP2-AS1 in colon cancer cells. Inhibition of AGAP2-AS1 upregulated the expression of LINC-PINT, and inhibition of LINC-PINT promoted the expression levels of AGAP2-AS1 in colon cancer cells. Furthermore, overexpression of AGAP2-AS1 could increase the proliferation, invasion and migration of colon cancer cells, while overexpression of LINC-PINT could attenuate the effects of overexpression of AGAP2-AS1 on the proliferation, migration and invasion of colon cancer cells. We also found that AGAP2-AS1 promoted colon cancer cell proliferation, migration and invasion through the Hippo signaling.
Conclusion: Upregulated expression of AGAP2-AS1 promoted proliferation, invasion and migration in colon cancer by forming a negative feedback loop with LINC-PINT.
Keywords: colon cancer, lncRNA AGAP2-AS1, lncRNA LINC-PINT, proliferation, migration, invasion
Introduction
Colon cancer affects more than 1 million new cases every year and it is one of the major causes of cancer deaths worldwide.1 Extensive efforts have been made to improve cancer diagnosis and treatment. However, survival of colon cancer patients is generally poor, especially for those at advanced stages.2,3 In clinical practice, about 1 out of 5 colon patients is diagnosed with the existence of cancer metastasis, in which radical therapy is applicable.4 Biological target therapy, radiotherapy and chemotherapy can be used to treat advanced colon cancer patients, however, more than half of the patients die of tumor recurrence in the second year of diagnosis.4,5
Long (200 nt) non-coding RNAs (lncRNAs) were considered as “dark matter” or “noise” of the transcriptome without biological function.6,7 However, an increasing number of studies have shown that lncRNAs participate in almost all biological processes, such as cell proliferation and growth, cell differentiation and cell apoptosis through regulation of target genes.8 Altered expression of lncRNAs may result in abnormal expression of disease-related genes, such as oncogenes or tumor suppressors,9 indicating the potential role of lncRNAs in cancer diagnosis and therapy.10 However, the functions of most lncRNAs remain unknown. It has been reported that lncRNA AGAP2-AS1 promotes the development of gastric cancer, lung cancer and breast cancer.11–13 However, the role of AGAP2-AS1 in colon cancer is unclear. This study was carried out to investigate the function of AGAP2-AS1 in colon cancer.
Materials and Methods
Research Subjects
A total of 66 patients with colon cancer (39 males and 27 females, 31 to 67 years old, mean age 49.8 ± 6.1 years old, Table 1) admitted at Cancer Hospital of China Medical University from July 2015 to January 2018 were enrolled in this study. All patients were diagnosed by histopathological examinations. Inclusion criteria: 1) newly diagnosed cases; 2) no therapy was carried out before admission; 3) patients signed the informed consent. Exclusion criteria: 1) patients transferred from other institutes; 2) patients complicated with other clinical conditions besides colon cancer; 3) therapies were initiated. All patients were staged according to AJCC criteria, and there were 16, 14, 20 and 18 cases at stage I–IV, respectively. This study was approved by the Ethics Committee of the aforementioned hospital. Experiments were conducted in accordance with the Declaration of Helsinki. All patients signed the written informed consent.
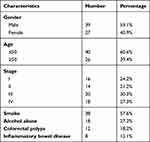 |
Table 1 The Clinical Characteristics of Colorectal Cancer |
Tissue Specimens and Cell Lines
Prior to therapy, colon biopsy was carried out under the guidance of MRI to collect tumor and adjacent (3 cm around tumors) non-tumor tissues from all patients. All tissue samples were confirmed by histopathological exams. Colon carcinoma cell lines (ATCC, USA) RKO and HCT 116 were used. Cells were cultivated in a cell culture medium composed of 10% FBS and 90% Eagle’s Minimum Essential Medium at 37 °C with 95% humidity and 5% CO2.
Cell Transfections
Full length of the genomic DNA AGAP2-AS1 or LINC-PINT was inserted into pcDNA 3.1 vector (Sangon, Shanghai, China) to construct AGAP2-AS1 or LINC-PINT expression vector. Lipofectamine 2000 (Thermo Fisher Scientific) was used to transfect 10 nM vector (empty vector was negative control, NC) into 106 cells. Untransfected cells were used as the Control (C) cells. Cells were collected at 24 h post transfection for the subsequent experiments.
Real-Time Quantitative PCR (RT-qPCR)
Total RNAs were extracted from tissues or RKO and HCT 116 cells using RNAzol reagent (Sigma-Aldrich, St. Louis, MO, USA). Total RNAs were reverse transcribed into cDNA using GoScript™ Reverse Transcription System (Promega). To detect the expression of AGAP2-AS1 and LINC-PINT, SYBR Green Master Mix (Bio-Rad) was used to prepare qPCR reaction mixtures. The expression levels of AGAP2-AS1 and LINC-PINT were normalized to 18S RNA using 2−ΔΔCT method.
Cell Proliferation Assay
Cell proliferation assay was carried out using Cell Counting Kit-8 (CCK-8), (ab228554, Abcam, Cambridge, UK) at 24 h post transfection. Briefly, cell density was adjusted to 5 × 104 cells/mL. Cells were cultivated in a 96-well plate with 100 μL cell suspension per well at 37 °C with 5% CO2. CCK-8 solution (10 μL) was added at 24, 48, 72 and 96 h after the beginning of cell culture. After that, cells were cultivated for an additional 4 h and 10 μL dimethyl sulfoxide (DMSO) was added. OD values at 450 nm were measured to reflect cell proliferation abilities.
Transwell Assay
Cell migration and invasion abilities were measured by Transwell migration and invasion assays at 24 h post transfection. Briefly, 1 mL serum-free Eagle’s Minimum Essential Medium was used to resuspend cell pellets containing 3 × 104 cells to make cell suspensions. The upper Transwell chamber was used to cultivate cells (0.1 mL per well) and the lower chamber was filled with a mixture of 80% Eagle’s Minimum Essential Medium and 20% FBS. Matrigel (356,234, Millipore, USA) was used to coat the membrane before the invasion assay. Cells were cultivated under the aforementioned conditions for 12 h, followed by staining the lower surface of membranes using 0.1% crystal violet (Sigma-Aldrich). Stained cells were observed under a light microscope. All groups were normalized to the C group.
Western Blot
Total proteins were isolated from cells using RIPA buffer reagent (Thermo Fisher Scientific, Inc., Waltham, MA, USA) containing protease inhibitor (Cocktail, Roche, Basel, Switzerland). The same amount (30 μg) of protein samples was separated by 10% SDS-PAGE (Mlbio, Shanghai, China) and transferred to nitrocellulose membranes (GE Healthcare, Westborough, MA). The membranes were blocked with 5% BSA at room temperature for 2 h, followed by incubation with primary antibodies: anti-p-LATS1 (1:1000, ab111344, Abcam), ANTI-LATS1 (1:1000, ab243656, Abcam), anti-p-YAP (1:1000, ab76252, Abcam), anti-YAP (1:1000, ab52771, Abcam) and anti-GAPDH (1:1000, ab8245, Abcam) at 4 °C overnight. TBST buffer was used to elute the membrane for two times. The membranes were then incubated with the secondary antibody for 2 h at 37 °C. The ECL detection system (Thermo Fisher Scientific, Waltham, MA, USA) was used to image the protein bands.
Immunofluorescence Staining
RKO and HCT 116 cells were fixed in 4% paraformaldehyde for 15 min and permeabilized in 0.1% Triton X-100 at room temperature for 20 min. Cardiomyocytes were then blocked in PBS containing 0.5% bovine serum albumin for 60 min and incubated with primary rabbit antibody against YAP (1:200, abcam) at 4 °C overnight, followed by incubation with Alexa Fluor 488-conjugated anti-rabbit IgG secondary antibody. Finally, the nuclei were stained with DAPI. Images were visualized and captured with a Leica SP8 confocal microscopy.
Nuclear/Cytoplasmic Fractionation
For nuclear YAP accumulation assay, RKO and HCT 116 cells were harvested and lysed to obtain cytoplasmic and nuclear lysates using the Keygen Nuclear-Cytosol Protein Extraction Kit from Nanjing KeyGen Biotech. Co., Ltd. (Nanjing, China). The cytoplasmic and nuclear lysates were performed with Western blotting.
Statistical Analyses
Three biological replicates were included in each experiment and mean values were used in all data analyses. Paired t-test was used to explore differences between two types of tissue. ANOVA Tukey’s test was used to compare differences among multiple patients and cell groups. Correlations were analyzed by linear regression. P < 0.05 was considered as statistically significant.
Results
AGAP2-AS1 Was Upregulated in Colon Cancer Patients but Not Affect by Clinical Stage
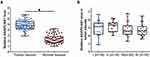
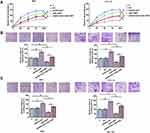
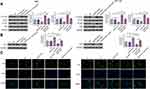
RT-qPCR was performed to detect the expression of AGAP2-AS1 in tumor and adjacent normal tissue specimens. It was observed that AGAP2-AS1 was upregulated in tumor tissues compared to that in adjacent normal tissues (Figure 1A, p < 0.05), indicating the upregulation of AGAP2-AS1 in colon cancer. In addition, no significant difference was observed in the expression levels of AGAP2-AS1 in tumor tissue specimens among four clinical stages (Figure 1B).
The Expression of LINC-PINT Was Inversely Correlated with the Expression of AGAP2-AS1 in Colon Cancer
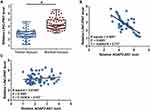
The expression levels of LINC-PINT were also measured by qPCR. LINC-PINT was significantly downregulated in tumor tissues compared to that in adjacent normal tissues (Figure 2A, p < 0.05). In addition, correlation analysis results showed that the expression of LINC-PINT was significantly and inversely correlated with the expression of AGAP2-AS1 across tumor tissues (Figure 2B). However, the expression of AGAP2-AS1 and LINC-PINT were not correlated with each other in adjacent normal tissues (Figure 2C). Moreover, we found that the expression of AGAP2-AS1 (Supplementary Figure 1A, p = 0.26) and LINC-PINT (Supplementary Figure 1B, p = 0.046) were not correlated with the overall survival of colon cancer patients.
AGAP2-AS1 and LINC-PINT Downregulated Each Other in Colon Cancer Cell Lines
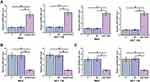
To analyze the interaction between AGAP2-AS1 and LINC-PINT in colon cancer, a vector expressing AGAP2-AS1 or LINC-PINT was transfected into RKO and HCT 116 cells. It was observed that, compared to the C and NC groups, AGAP2-AS1 and LINC-PINT were overexpressed at 24 h after transfection (Figure 3A, p < 0.05). Overexpression of AGAP2-AS1 resulted in significantly downregulated LINC-PINT (Figure 3B, p < 0.05). In addition, overexpression of LINC-PINT resulted in significantly downregulated AGAP2-AS1 (Figure 3C, p < 0.05).
AGAP2-AS1 and LINC-PINT Inhibitor Upregulated Each Other in Cells of Colon Cancer Cell Lines
To further test the interaction between AGAP2-AS1 and LINC-PINT in colon cancer, AGAP2-AS1 or LINC-PINT inhibitor was transfected into RKO and HCT 116 cells. Compared to the C and NC groups, the expression of AGAP2-AS1 and LINC-PINT were inhibited at 24 h after transfection (Figure 4A, p < 0.05). Inhibition of AGAP2-AS1 significantly upregulated LINC-PINT (Figure 4B, p < 0.05). In addition, inhibition of LINC-PINT significantly upregulated AGAP2-AS1 (Figure 4C, p < 0.05).
AGAP2-AS1 and LINC-PINT Regulate Colon Cancer Cell Proliferation, Migration and Invasion
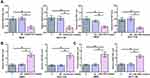
Next, the role of AGAP2-AS1 and LINC-PINT in colon cancer cell proliferation, migration and invasion was investigated. Overexpression of AGAP2-AS1 and LINC-PINT were verified (Figure 3A, p < 0.05). Cell proliferation was detected by CCK-8 assay. Overexpression of AGAP2-AS1 increased the cell proliferation rate, while overexpression of AGAP2-AS1 and LINC-PINT together decreased the cell proliferation rate compared with the overexpression of AGAP2-AS1 alone, and increased the cell proliferation rate compared with the overexpression of LINC-PINT alone in RKO and HCT 116 cells (Figure 5A, p < 0.05). In addition, Transwell assay showed that AGAP2-AS1 promoted cell migration and invasion, while LINC-PINT had the opposite effect on RKO and HCT 116 cells. Moreover, cells with the overexpression of both AGAP2-AS1 and LINC-PINT showed significantly inhibited migration and invasion compared with the overexpression of AGAP2-AS1 alone, and the co-transfection also significantly promoted migration and invasion compared to cells with the overexpression of LINC-PINT alone in both RKO and HCT 116 cells (Figure 5B and C, p < 0.05).
AGAP2-AS1 Promoted Colon Cancer Cell Proliferation, Migration and Invasion Through the Hippo Signaling
The Hippo signaling pathway plays a key role in cancer progression. The involvement of the Hippo signaling pathway was tested in colon cancer cell lines. Immunoblotting exhibited that overexpression of AGAP2-AS1 inhibited LAST1 phosphorylation and YAP reduction in RKO and HCT 116 cells (Figure 6A, p < 0.05). In addition, overexpression of AGAP2-AS1 resulted in decreased nuclear expression of YAP, indicated by immunofluorescence and nuclei-cytoplasm fractionation in RKO and HCT 116 cells. Moreover, all the effects of AGAP2-AS1 on the Hippo signaling pathway were almost abolished by overexpression of LINC-PINT (Figure 6B and C, p < 0.05).
Discussion
AGAP2-AS1 has been reported to promote the development and progression of several cancers.11–13 The role of AGAP2-AS1 in colon cancer is unknown. Our study is the first to report the oncogenic function of AGAP2-AS1 in colon cancer and we also proved that the function of AGAP2-AS1 in colon cancer is likely achieved through the interaction with tumor suppressor LINC-PINT.
A recent large-scale expressional and functional study has revealed that LINC-PINT, a p53-regulated lncRNA, is downregulated in many types of cancer including colon cancer,14 and LINC-PINT may inhibit tumor cell invasion to suppress cancer development.14 Another study reported that reduced expression levels of LINC-PINT could serve as potential prognostic and diagnostic biomarkers for pancreatic cancer.15 We also observed downregulated expression of LINC-PINT in colon cancer tissues compared to that in adjacent normal tissues. In addition, overexpression of LINC-PINT resulted in decreased proliferation, migration and invasion of colon cancer cells. Therefore, our study confirmed the oncogenic role of LINC-PINT in colon cancer.
A conserved sequence of LINC-PINT mediates its interaction with PRC2, which is a key player in cancer cell invasion, to suppress the invasive phenotype of cancer cells.16 It is known that lncRNAs cancer interacts with tumor-suppressive or oncogenic pathways to participate in cancer biology.17 Interestingly, our study proved that LINC-PINT and AGAP2-AS1 can form a negative feedback regulation loop to participate in the regulation of colon cancer cell proliferation, migration and invasion.
AGAP2-AS1 and LINC-PINT interact with multiple oncogenic or tumor-suppressive factors. AGAP2-AS1 is upregulated in lung cancer and can interact with EZH2 and LSD1 to suppress the expression of LATS2 and KLF2, thereby promoting tumor growth.11 In gastric cancer, SP1 activates AGAP2-AS1 to increase cancer cell migration and proliferation.12 In breast cancer, overexpression of AGAP2-AS1 regulates the methylation of MyD88 to promote the development of chemoresistance.13 In contrast, LINC-PINT is downregulated in many cancers and interacts with PRC2 through a highly conserved sequence element to inhibit tumor cell invasion.14 Therefore, AGAP2-AS1 and LINC-PINT may form a complex gene expression regulation network to regulate the development and progression of colon cancer. More studies are needed to identify other players in this process.
It was reported that Hippo signaling pathway exerts an important role in cancer. Yes-associated protein (YAP), a downstream regulator of the Hippo pathway, is upregulated in cancers and is involved in proliferation, apoptosis, migration, invasion.18 The decreased phosphorylation level of LATS1, YAP was found after the overexpression of AGAP2-AS1, while LINC-PINT has the opposite effect. Furthermore, overexpression of AGAP2-AS1 improved the YAP translocation, while LINC-PINT decreased the nuclear level of YAP. These data demonstrated that the Hippo signaling pathway is required for the proliferation, migration and invasion effects of AGAP2-AS1.
Conclusion
In conclusion, AGAP2-AS1 is an oncogene, and LINC-PINT is a tumor suppressor in colon cancer. AGAP2-AS1 and LINC-PINT may form a negative feedback regulation loop to participate in colon cancer.
Disclosure
The authors declare that they have no conflicts of interest.
References
1. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2017. CA Cancer J Clin. 2017;67(1):7–30. doi:10.3322/caac.21387
2. Lemmens V, van Steenbergen L, Janssen-Heijnen M, Martijn H, Rutten H, Coebergh JW. Trends in colorectal cancer in the south of the Netherlands 1975–2007: rectal cancer survival levels with colon cancer survival. Acta Oncol. 2010;49(6):784–796. doi:10.3109/02841861003733713
3. De Angelis R, Sant M, Coleman MP, et al. Cancer survival in Europe 1999–2007 by country and age: results of EUROCARE–5-a population-based study. Lancet Oncol. 2014;15(1):23–34. doi:10.1016/S1470-2045(13)70546-1
4. Hurwitz H, Fehrenbacher L, Novotny W, et al. Bevacizumab plus irinotecan, fluorouracil, and leucovorin for metastatic colorectal cancer. N Engl J Med. 2004;350(23):2335–2342. doi:10.1056/NEJMoa032691
5. Knoop K, Schwenk N, Schmohl K, et al. Mesenchymal stem cell-mediated, tumor stroma-targeted radioiodine therapy of metastatic colon cancer using the sodium iodide symporter as theranostic gene. J Nucl Med. 2015;56(4):600–606. doi:10.2967/jnumed.114.146662
6. Mercer TR, Dinger ME, Mattick JS. Long non-coding RNAs: insights into functions. Nat Rev Genet. 2009;10(3):155–159. doi:10.1038/nrg2521
7. Hauptman N, Glavac D. Long non-coding RNA in cancer. Int J Mol Sci. 2013;14:4655–4669. doi:10.3390/ijms14034655
8. Gibb E, Brown C, Lam W. The functional role of long non-coding RNA in human carcinomas. Mol Cancer. 2011;10. doi:10.1186/1476-4598-10-38
9. Gutschner T, Diederichs S. The hallmarks of cancer. RNA Biol. 2012;9(6):703–719. doi:10.4161/rna.20481
10. Qi P, Du X. The long non-coding RNAs, a new cancer diagnostic and therapeutic gold mine. Mod Pathol. 2012;26. doi:10.1038/modpathol.2012.160
11. Li W, Sun M, Zang C, et al. Upregulated long non-coding RNA AGAP2-AS1 represses LATS2 and KLF2 expression through interacting with EZH2 and LSD1 in non-small-cell lung cancer cells. Cell Death Dis. 2016;7:e2225. doi:10.1038/cddis.2016.126
12. Qi F, Liu X, Wu H, et al. Long noncoding AGAP2-AS1 is activated by SP1 and promotes cell proliferation and invasion in gastric cancer. J Hematol Oncol. 2017;10:1–4.
13. Dong H, Wang W, Mo S, et al. SP1-induced lncRNA AGAP2-AS1 expression promotes chemoresistance of breast cancer by epigenetic regulation of MyD88. J Exp Clin Cancer Res. 2018;37:1–5.
14. Marín-Béjar O, Mas A, González J, et al. The human lncRNA LINC-PINT inhibits tumor cell invasion through a highly conserved sequence element. Genome Biol. 2017;18. doi:10.1186/s13059-017-1331-y
15. Li L, Zhang G-Q, Chen H, et al. Plasma and tumor levels of Linc-pint are diagnostic and prognostic biomarkers for pancreatic cancer. Oncotarget. 2016;7:71773.
16. Hu P, Chu J, Wu Y, et al. NBAT1 suppresses breast cancer metastasis by regulating DKK1 via PRC2. Oncotarget. 2015;6(32):32410–32425. doi:10.18632/oncotarget.5609
17. Schmitt Adam M, Chang Howard Y. Long noncoding RNAs in cancer pathways. Cancer Cell. 2016;29(4):452–463. doi:10.1016/j.ccell.2016.03.010
18. Ma J, Fan Z, Tang Q, Xia H, Zhang T, Bi F. Aspirin attenuates YAP and β-catenin expression by promoting β-TrCP to overcome docetaxel and vinorelbine resistance in triple-negative breast cancer. Cell Death Dis. 2020;11(7):530. doi:10.1038/s41419-020-2719-2
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.