Back to Journals » Cancer Management and Research » Volume 13
LINC00908 Promotes Diffuse Large B-Cell Lymphoma Development by Down-Regulating miR-671-5p
Authors Zeng H, Wei Y, Wei X, Feng R
Received 9 January 2021
Accepted for publication 15 April 2021
Published 30 April 2021 Volume 2021:13 Pages 3589—3599
DOI https://doi.org/10.2147/CMAR.S299715
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Professor Harikrishna Nakshatri
This paper has been retracted.
Hong Zeng, Yongqiang Wei, Xiaolei Wei, Ru Feng
Department of Hematology, Nanfang Hospital, Southern Medical University, Guangzhou, Guangdong, People’s Republic of China
Correspondence: Ru Feng
Department of Hematology, Nanfang Hospital, Southern Medical University, 1838 Guanzghou Avenue, Guagzhou, Guangdong, 510515, People’s Republic of China
Tel +86-13922865798
Email [email protected]
Introduction: Emerging evidence has revealed that long noncoding RNA (lncRNA) play important role in almost all kinds of human cancers. LINC00908 has been reported to be involved in the development of prostate cancer, colorectal cancer and gastric cancer which was functioned as an oncogene. However, the potential biology role and molecular mechanism of LINC00908 in diffuse large B-cell lymphoma are still unclear.
Methods: LINC00908 and miR-671-5p expression were evaluated in DLBCL tissues and cell lines using RT-qPCR. CCK-8 and transwell assay were used to analyze the in vitro role of LINC00908 in DLBCL progression. The xenograft model was used to explore the in vivo role of LINC00908 in DLBCL growth. The physical interaction between LINC00908 and miR-671-5p was confirmed using bioinformatics analysis and a dual luciferase assay, RIP and RNA pull down.
Results: The expression of LINC00908 was markedly up-regulated in diffuse large B-cell lymphoma tissues and cell lines, and the decreased expression of LINC00908 significantly inhibited diffuse large B-cell lymphoma cell proliferation and invasion. Then, we revealed that LINC00908 directly interacted with miR-671-5p, which was down-regulated in diffuse large B-cell lymphoma cells and highly expressed with LINC00908 knockdown. Moreover, luciferase reporter assays and RNA immunoprecipitation (RIP) assay further proved that miR-671-5p is a direct target of LINC00908 in diffuse large B-cell lymphoma cells. Rescue experiments were also performed, and we confirmed that LINC00908 acts as an oncogene role in diffuse large B-cell lymphoma through miR-671-5p. Finally, the influence of LINC00908 silence significantly inhibited diffuse large B-cell lymphoma growth in vivo.
Conclusion: LINC00908 promotes malignancy of diffuse large B-cell lymphoma through regulating miR-671-5p.
Keywords: LINC00908, diffuse large B-cell lymphoma, miR-671-5p, proliferation, invasion
Introduction
Worldwide, diffuse large B cell lymphoma (DLBCL) accounts for more than one-third cases of all non-Hodgkin lymphoma (NHL), is the commonest subtype of NHL. DLBCL is a rather aggressive hyperplastic disease in lymphatic system, moreover, approximately 40% of DLBCL patients showing resistance to the clinical therapeutic protocols available now.1 Despite a large number of diagnosis and therapy methods of DLBCL have been made, the prognosis DLBCL patients remain rather unsatisfactory. Thus, it is imperative to explore underlying molecular mechanisms and identify novel biomarker of DLBCL.
Long non-coding RNAs (lncRNAs) are defined as transcripts with a length of >200 nucleotides. Emerging researches have suggested that lncRNAs serve crucial functions in the initiation and development of DLBCL.2–4
LINC00908 was a recently identified lncRNA. It was reported that LINC00908 inhibited prostate cancer progression.5 Nevertheless, other studies reported the oncogenic role of LINC00908 in hepatocellular carcinoma and colorectal cancer.6,7 Obviously, the biological effects and underlying mechanism of LINC00908 in the development of main human malignant diseases, including DLBCL, remain elusive.
The present study assessed the expression of LINC00908 in DLBCL tissues and paired normal lymph nodes. Furthermore, the biological functions of LINC00908 in DLBCL were assessed in vitro and in vivo. Moreover, the current study also examined the association between LINC00908 and miR-671-5p to reveal the molecular mechanism of LINC00908 in DLBCL development. To the best of our knowledge, this study was the first to reveal that LINC00908 exerts oncogenic roles in the progression of DLBCL.
Materials and Methods
Clinical Samples
Totally, 28 DLBCL tissues and normal lymph nodes were obtained from DLBCL patients at the Nanfang Hospital, Southern Medical University (Guangzhou, China) from Feb 2016 to Oct 2018. All participants signed informed consent forms prior to sample collection. The samples collected during operation were rapidly frozen in liquid nitrogen until further use. This study was approved by the Ethic and Research Committees of Nanfang Hospital, Southern Medical University (Guangzhou, China). All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Cell Culture
Human lymphoblastoid B cell (GM12878) and human DLBCL cells (OCI-LY7, DB, U2932, and FARAGE) were purchased from American Type Culture Collection (ATCC). GM12878 cells were cultured in RPMI 1640 with 15% fetal bovine serum and 1% pen/strep. DLBCL cells (OCI-LY7, DB, U2932, and FARAGE) were grown in RPMI 1640 with 10% FBS and 0.05 mg/mL gentamicin. All cell cultures were maintained at 37°C in a humidified incubator with 5% CO2.
Constructs, Synthesized Oligos and Transfection
The short hairpin RNA (shRNA/sh) targeting LINC00908 (sh- LINC00908), miR-671-5p mimics, inhibitors and their corresponding negative controls (NCs) were purchased from Shanghai GeneChem Co., Ltd. All DNAs were inserted into pcDNA3.1. Finally, Lipofectamine 3000 was utilized to transfer the oligonucleotides and constructs into the U2932 and FARAGE cells.
RNA Extraction and qRT-qPCR
The total RNA was obtained from tissues and cells adopting TRIzol reagent. A total of 2 µg RNA was reverse transcribed into cDNA adopting the PrimeScript RT reagent kit. RT-qPCR was conducted using SYBR Green Master Mix on an ABI PRISM 7500 PCR system. GAPDH and U6 were used as controls and for the normalization of the expression levels of mRNA and miRNA, respectively. The specific primers were showed in Table 1.
 |
Table 1 Primer List |
Cell Proliferation Assay
Cell Counting Kit-8 (CCK-8) and colony formation assays were performed to evaluate the proliferation ability. For the CCK-8 assay, cells were seeded in 96-well plates (3000 cells/well). Then 10 μL CCK-8 solution was added after 24, 48, 72 and 96 h of culture. After 2 h, the plates were washed using PBS and finally the absorbance at 450 nm was measured using a microplate reader. For colony formation analysis, 1000 cells were seeded into a 6-well plate and continuously incubated for 12 days. The colonies were fixed using 4% paraformaldehyde and stained with 1% crystal violet. Finally, the colonies were counted and imaged.
Transwell Invasion Assay
The transwell invasion assays were conducted to determine the cell invasion potential using transwell plates coated with 50 µL Matrigel (BD Biosciences). Briefly, 1×105 cells were suspended in 300 μL serum-free medium and added to the upper chamber, while 800 μL complete medium was placed in the lower chamber. 24 h later, cells on the upper surface of the membrane were scraped off, while the cells on the lower side of the chamber were fixed and stained. The invaded cells were counted in more than 5 fields under a light microscope.
Subcellular Fractionation Assay
The PARIS Kit (Life Technologies) was used to isolate nuclear and cytoplasmic RNAs according to the manufacturer’s protocol. Reverse transcription of extracted RNAs and RT-PCR was conducted as described before.
Luciferase Reporter Assay
Mut (mutant-type) or wt (wild-type) fragments of LINC00908 containing the miR-671-5p targeting site were synthesized and cloned into a dual-luciferase reporter vector (pmirGLO; Shanghai GenePharma Co., Ltd.). Similarly, luciferase vectors and miR-671-5p mimics or miR-671-5p NC and Renilla plasmid were co-transfected into FARAGE cells using Lipofectamine 3000. 48 hours after transfection, a dual-luciferase assay was used to examine the Renilla and firefly luciferase activity according to the manufacturer’s protocol, and the levels of firefly luciferase activity were normalized to that of Renilla luciferase activity.
RNA-Binding Protein Immunoprecipitation (RIP) Assay
A Magna RIP RNA-Binding Protein Immunoprecipitation Kit (EMD Millipore) was adopted for the RIP assay. Cells were harvested and lysed. Lysis buffer containing magnetic beads was incubated with human Ago2 (anti-argonaute RISC catalytic component 2) antibody (cat. no. ab32381; 1:1000; Abcam) to conjugate the antibody to the magnetic beads. Subsequently, proteinase K was added to digest the protein, and the immunoprecipitated RNAs were isolated using TRIzol reagent and measured.
RNA Pull-Down Assay
FARAGE cells were transfected with biotin-labeled miR-671-5p mimic or NC. After 24 h, cells were collected and cultured with M-280 streptavidin magnetic beads (Invitrogen; Thermo Fisher Scientific, Inc.) at 4°C for 4 h with rotation. Subsequently, the beads were washed using lysis buffer containing proteinase K and 10% SDS, then the supernatants were obtained and the RNA was isolated and coprecipitated RNA was detected using qRT-PCR assays.
Tumor Xenograft Experiment
Ten female BALB/c nude mice (6–8 weeks old, 18–22 g) were bought from the Animal Center of Southern Medical University. OCI-LY7 cells (2x106) that were stably transfected with lv-sh-LINC00908 or lv-sh-NC were injected subcutaneously into the left flank of the mice (n=5 mice per group). Tumor sizes were measured every week. After 4 weeks, the mice were anesthetized and the tumor tissues were collected and weighed, and RT-qPCR was conducted to examine the expression of LINC00908 and miR-671-5p. This study was approved by the Ethic and Research Committees of Nanfang Hospital, Southern Medical University. The guidelines for the welfare of experimental animals are GB/T 35892-2018 standard issued by the General Administration of Quality Supervision, Inspection and Quarantine of the People’s Republic of China.
Statistical Analysis
The experiments were repeated at least three times. All results are presented as the mean ± SD. All statistical analyses were conducted using GraphPad Prism software. One-way ANOVA with Bonferroni post hoc test or two-tailed Student’s t-test were used for the comparisons among groups. In addition, Pearson’s coefficient correlation analysis was performed for expression correlation analysis. P < 0.05 was recognized to indicate a statistically significant difference.
Results
LINC00908 Expression is Elevated in DLBCL
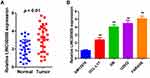
qRT-qPCR was conducted to assess the relative expression levels of DLBCL in 28 pairs of DLBCL and adjacent normal tissues. LINC00908 expression was markedly increased in DLBCL tissues compared with in paired normal tissues (Figure 1A). Moreover, the expression levels of LINC00908 in four DLBCL cell lines and in human lymphoblastoid B cells were examined. The results indicated that the DLBCL cell lines exhibited markedly higher expression levels of LINC00908 compared with the human lymphoblastoid B cell line (Figure 1B). Later on, 2 DLBCL cell lines, FARAGE and U2932, were applied in the following experiments because they were verified to express the highest LINC00908 level among 4 DLBCL cell lines.
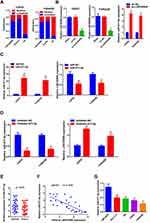
Reciprocal Modulation Between LINC00908 and miR-671-5p
To investigate the potential mechanism of LINC00908 in the development of DLBCL, we determined the subcellular localization of LINC00908. The results showed that LINC00908 was mostly distributed in the cytoplasm (Figure 2A), which suggested that LINC00908 might exert its biological function by sponging miRNA. miRcode (http://www.mircode.org) was used to conduct bioinformatics analysis, and the results indicated that miR-671-5p was a potential target of LINC00908. To confirm this hypothesis, the correlation between LINC00908 and miR-671-5p was explored. The present results demonstrated that miR-671-5p expression was markedly increased following knockdown of LINC00908 (Figure 2B). Furthermore, LINC00908 expression was markedly decreased when the cells were transfected with miR-671-5p mimic (Figure 2C), while the expression levels of LINC00908 were markedly increased when the cells were transfected with miR-671-5p inhibitor (Figure 2D). In addition, miR-671-5p expression was determined in DLBCL tissues, and the results indicated that miR-671-5p expression was down-regulated in DLBCL tissues (Figure 2E). Notably, miR-671-5p expression was negatively correlated with LINC00908 expression (Figure 2F). Additionally, DLBCL cell lines exhibited markedly lower expression levels of miR-671-5p compared with human lymphoblastoid B cell line (Figure 2G).
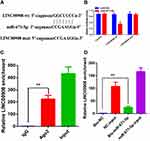
miR-671-5p is a Direct Target of LINC00908
A dual-luciferase reporter assay was conducted to further assess whether miR-671-5p was a direct target of LINC00908. The data of the luciferase assay revealed that miR-671-5p mimic markedly decreased luciferase reporter expression in the cells transfected with LINC00908-wt but not in the cells transfected with LINC00908-mut or NC (Figure 3A and B). It is widely acknowledged that miRNAs function by regulating RNA-induced silencing complex (RISC).8 Ago2, a key component of RISC, exerts crucial roles in RNA cleavage. Therefore, a RIP assay was conducted to determine whether miR-671-5p regulated miRNAs function by regulating RNA-induced silencing complex (RISC) via RISC formation. As the results demonstrated, compared with NC (IgG), LINC00908 and miR-671-5p were preferentially enriched in anti-Ago2 antibody-incubated beads (Figure 3C). To further investigate whether LINC00908 and miR-671-5p bind to each other, RNA pull-down assay was also performed. The present results demonstrated that compared with NC, the biotin-labeled miR-671-5p mimic pulled down more LINC00908 (Figure 3D). Collectively, these data revealed that LINC00908 bound to miR-671-5p directly.
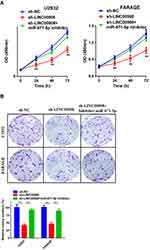
LINC00908 and miR-671-5p Effects on DLBCL Cell Proliferation and Invasion
CCK-8 assays and colony formation assays were conducted to explore the effects of LINC00908 and miR-671-5p on DLBCL cell proliferation. The results suggested that the proliferation of DLBCL cells in the sh- LINC00908 group was markedly impaired compared with that of cells in the NC group, but the inhibitory effect was reversed when the cells were co-transfected with si-LINC00908 and the miR-671-5p inhibitor (Figure 4). Transwell invasion assays were performed to explore the effects of LINC00908 and miR-671-5p on DLBCL cell invasion. The results suggested that the invasion of DLBCL cells in the sh- LINC00908 group was markedly impaired compared with that of cells in the NC group, but the inhibitory effect was reversed when the cells were co-transfected with si-LINC00908 and the miR-671-5p inhibitor (Figure 5). Collectively, all these results demonstrated that the oncogenic function of LINC00908 in DLBCL involved negative regulation of miR-671-5p.
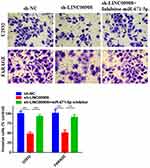 |
Figure 5 The role of LINC00908 and miR-671-5p in DLBCL cell invasion. Transwell assay was performed to determine cell invasion of different groups. **p < 0.01. |
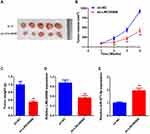
LINC00908 Silencing Limits DLBCL Tumor Growth in vivo
To further verify the in vitro results, a subcutaneous xenograft tumor model was established by injecting stable sh-NC or sh- LINC00908 cells into nude mice. Consistent with the in vitro findings, the in vivo experiments indicated that the volume and weight of tumors in sh-LINC00908 group were markedly reduced compared with those in the sh-NC group (Figures 6A–C). Furthermore, the expression levels of LINC00908 were decreased in the sh-LINC00908 group tumors, while miR-671-5p expression was increased in the sh-LINC00908 group tumors compared with in the sh-NC group tumors (Figure 6D and E). Therefore, it was concluded that LINC00908 silencing limited DLBCL tumor growth in vivo.
Discussion
In recent years, accumulating evidence has revealed fundamental roles of lncRNAs in the regulation of numerous physiological and pathological processes, such as cell differentiation, proliferation and apoptosis, and the initiation and progression of human cancer.9–11 At present, it is widely acknowledged that numerous lncRNAs are involved the progression of DLBCL. For example, lncRNA SMAD5-AS1 inhibited proliferation of DLBCL as a ceRNA of to increase APC expression and limiting Wnt/β-catenin pathway.3 LncRNA MALAT1 promoted tumorigenesis and immune escape of DLBCL by sponging miR-195.12 LncRNA TUG1 promoted tumor growth of DLBCL by promoting ubiquitination of MET.13 However, the exact biological function and mechanism of the majority of lncRNAs in DLBCL remain unclear.
LINC00908 was first found that it was associated with prognosis of glioma patients and may serve as a potential biomarkers of glioma.14 Moreover, it was reported that LINC00908 was highly expressed in HCC tissues and high LINC00908 expression was associated with progression and metastasis of HCC patients. The in vitro and in vivo studies further demonstrated that LINC00908 promoted the development of HCC through increasing SOX4 stability via inhibiting proteasomal degradation.6 Besides, Shan et al reported that LINC00908 enhanced CRC cell proliferation and limited CRC cell apoptosis via regulating KLF5 expression by sponging miR-143-3p.7 Nevertheless, Fan et al found that LINC00908 inhibited prostate cancer progression by negatively regulating microRNA-483-5p and then elevate the expression of TSPYL5.5 Obviously, the biological roles and mechanism of LINC00908 in human diseases including DLBCL are largely unknown. The current study aimed to investigate the exact roles and potential mechanism of LINC00908 in DLBCL. First, it was observed that the LINC00908 expression was up-regulated in clinical DLBCL tissues and cell lines. Second, our study also evaluated the biological effects of LINC00908 silencing on DLBCL cell proliferation and invasion in vitro, and tumor growth in vivo. The results of a series of experiments revealed that LINC00908 silencing inhibited cell proliferation and invasion in vitro and tumor growth in vivo in DLBCL.
The competing endogenous RNA (ceRNA) theory, wherein miRNA is sequestered by lncRNA which serves as a molecular sponge, has attracted increasing attention.15,16 For instance, lncRNA MYLK-AS1 facilitated HCC progression and angiogenesis by targeting miR-424-5p/E2F7 axis and activating VEGFR-2 signaling pathway.17 lncRNA CASC9 promoted bladder cancer proliferation and epithelial-mesenchymal transition by sponging miR-758-3p and upregulating TGF-β2.18 Therefore, it was predicted that miR-671-5p may be a direct target of LINC00908. In this study, LINC00908 was revealed to be a ceRNA by directly targeting miR-671-5p. Previous studies have demonstrated that miR-671-5p involved the development of several types of human cancer. For instance, miR-671-5p promoted colon cancer cells proliferation, migration, and invasion of by targeting TRIM67.19 DLEU1 aggravated osteosarcoma carcinogenesis through modulating miR-671-5p/DDX5 axis.20 MiRNA-671-5p facilitated prostate cancer progression metastasis via regulating NFIA/CRYAB axis.21 The results of the qRT-qPCR indicated the reciprocal inhibitory effect of LINC00908 and miR-671-5p. Furthermore, the effect of LINC00908 silencing on DLBCL cell proliferation and invasion was reversed by miR-671-5p inhibitor. The results of the luciferase assay showed that miR-671-5p directly bound to LINC00908. Additionally, the data of RIP and RNA pull-down assays revealed that miR-671-5p modulated LINC00908 in a RISC-dependent manner. Collectively, these data demonstrated that LINC00908 acts as an endogenous sponge of miR-671-5p, and that LINC00908 and miR-671-5p negatively regulate each other in DLBCL.
To conclude, the present study provides new insights into the role of LINC00908 in DLBCL development through a mechanism through sponging miR-671-5p. These findings also suggested that LINC00908 may serve as a new potential therapeutic target of DLBCL.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Galaznik A, Huelin R, Stokes M, et al. Systematic review of therapy used in relapsed or refractory diffuse large B-cell lymphoma and follicular lymphoma. Future Sci OA. 2018;4(7):FSO322. doi:10.4155/fsoa-2018-0049
2. Tang X, Long Y, Xu L, Yan X. LncRNA MORT inhibits cancer cell proliferation and promotes apoptosis in mantle cell lymphoma by upregulating miRNA-16. Cancer Manag Res. 2020;12:2119–2125. doi:10.2147/CMAR.S233859
3. Zhao CC, Jiao Y, Zhang YY, et al. Lnc SMAD5-AS1 as ceRNA inhibit proliferation of diffuse large B cell lymphoma via Wnt/beta-catenin pathway by sponging miR-135b-5p to elevate expression of APC. Cell Death Dis. 2019;10(4):252. doi:10.1038/s41419-019-1479-3
4. Zhu Q, Li Y, Guo Y, et al. Long non-coding RNA SNHG16 promotes proliferation and inhibits apoptosis of diffuse large B-cell lymphoma cells by targeting miR-497-5p/PIM1 axis. J Cell Mol Med. 2019;23(11):7395–7405. doi:10.1111/jcmm.14601
5. Fan L, Li H, Zhang Y. LINC00908 negatively regulates microRNA-483-5p to increase TSPYL5 expression and inhibit the development of prostate cancer. Cancer Cell Int. 2020;20(1):10. doi:10.1186/s12935-019-1073-x
6. Hu X, Li Q, Zhang J. The long noncoding RNA LINC00908 facilitates hepatocellular carcinoma progression via interaction with Sox-4. Cancer Manag Res. 2019;11:8789–8797. doi:10.2147/CMAR.S216774
7. Shan TD, Tian ZB, Li Q, et al. Long intergenic noncoding RNA 00908 promotes proliferation and inhibits apoptosis of colorectal cancer cells by regulating KLF5 expression. J Cell Physiol. 2021;236(2):889–899. doi:10.1002/jcp.29899
8. Filipowicz W, Bhattacharyya SN, Sonenberg N. Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nat Rev Genet. 2008;9(2):102–114. doi:10.1038/nrg2290
9. Zhang Y, Liu Q, Liao Q. Long noncoding RNA: a dazzling dancer in tumor immune microenvironment. J Exp Clin Cancer Res. 2020;39(1):231. doi:10.1186/s13046-020-01727-3
10. Feng Y, Wu M, Hu S, Peng X, Chen F. LncRNA DDX11-AS1: a novel oncogene in human cancer. Hum Cell. 2020;33(4):946–953. doi:10.1007/s13577-020-00409-8
11. Wang Y, Zhang M, Zhou F. Biological functions and clinical applications of exosomal long non-coding RNAs in cancer. J Cell Mol Med. 2020;24(20):11656–11666. doi:10.1111/jcmm.15873
12. Wang QM, Lian GY, Song Y, Huang YF, Gong Y. LncRNA MALAT1 promotes tumorigenesis and immune escape of diffuse large B cell lymphoma by sponging miR-195. Life Sci. 2019;231:116335. doi:10.1016/j.lfs.2019.03.040
13. Cheng H, Yan Z, Wang X, et al. Downregulation of long non-coding RNA TUG1 suppresses tumor growth by promoting ubiquitination of MET in diffuse large B-cell lymphoma. Mol Cell Biochem. 2019;461(1–2):47–56. doi:10.1007/s11010-019-03588-7
14. Song L, Zhang S, Duan C, et al. Genome-wide identification of lncRNAs as novel prognosis biomarkers of glioma. J Cell Biochem. 2019;120(12):19518–19528. doi:10.1002/jcb.29259
15. Thomson DW, Dinger ME. Endogenous microRNA sponges: evidence and controversy. Nat Rev Genet. 2016;17(5):272–283. doi:10.1038/nrg.2016.20
16. Tay Y, Rinn J, Pandolfi PP. The multilayered complexity of ceRNA crosstalk and competition. Nature. 2014;505(7483):344–352. doi:10.1038/nature12986
17. Teng F, Zhang JX, Chang QM, et al. LncRNA MYLK-AS1 facilitates tumor progression and angiogenesis by targeting miR-424-5p/E2F7 axis and activating VEGFR-2 signaling pathway in hepatocellular carcinoma. J Exp Clin Cancer Res. 2020;39(1):235. doi:10.1186/s13046-020-01739-z
18. Zhang Z, Chen F, Zhan H, et al. lncRNA CASC9 sponges miR7583p to promote proliferation and EMT in bladder cancer by upregulating TGFbeta2. Oncol Rep. 2021;45(1):265–277. doi:10.3892/or.2020.7852
19. Jin W, Shi J, Liu M. Overexpression of miR-671-5p indicates a poor prognosis in colon cancer and accelerates proliferation, migration, and invasion of colon cancer cells. Onco Targets Ther. 2019;12:6865–6873. doi:10.2147/OTT.S219421
20. Chen X, Zhang C, Wang X. Long noncoding RNA DLEU1 aggravates osteosarcoma carcinogenesis via regulating the miR-671-5p/DDX5 axis. Artif Cells Nanomed Biotechnol. 2019;47(1):3322–3328. doi:10.1080/21691401.2019.1648285
21. Zhu Z, Luo L, Xiang Q, et al. MiRNA-671-5p promotes prostate cancer development and metastasis by targeting NFIA/CRYAB axis. Cell Death Dis. 2020;11(11):949. doi:10.1038/s41419-020-03138-w
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.