Back to Journals » Psoriasis: Targets and Therapy » Volume 8
Limited clinical utility of HLA-Cw6 genotyping for outcome prediction in psoriasis patients under ustekinumab therapy: a monocentric, retrospective analysis
Authors Anzengruber F, Ghosh A , Maul JT, Drach M, Navarini AA
Received 3 January 2018
Accepted for publication 30 January 2018
Published 23 March 2018 Volume 2018:8 Pages 7—11
DOI https://doi.org/10.2147/PTT.S161437
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 4
Editor who approved publication: Prof. Dr. Uwe Wollina
Florian Anzengruber,1,2 Adhideb Ghosh,1–3 Julia-Tatjana Maul,1,2 Mathias Drach,1,2 Alexander A Navarini1–3
1Department of Dermatology, University Hospital Zurich, Zurich, Switzerland; 2Faculty of Medicine, University of Zurich, Zurich, Switzerland; 3Competence Center Personalized Medicine University of Zurich/Swiss Federal Institute of Technology (ETH) Zurich, Switzerland
Purpose: Several studies have suggested that an HLA-Cw6+ allele can predict an improved outcome of treatment in psoriasis patients. The aim of the study was to assess whether the published association between HLA-Cw6 allele carriers and response to ustekinumab has the potential to impact treatment decisions.
Patients and methods: Differences in Psoriasis Activity and Severity Index 50, 75, and 90; Nail Psoriasis Severity Index; and Dermatology Life Quality Index at 16 weeks were evaluated between HLA-Cw6 allele carriers vs. non-carriers. Thirty patients with moderate-to-severe psoriasis under treatment with ustekinumab were included in our study.
Results: There was no difference between the two groups with respect to Psoriasis Activity and Severity Index 50, 75, and 90 or in terms of change in Nail Psoriasis Severity Index or Dermatology Life Quality Index.
Conclusion: In our retrospectively analyzed cohort, we could not detect the previously reported better response in HLA-Cw6+ vs. HLA-Cw6− patients.
Keywords: human leukocyte antigen, Cw6, genetic variations, HLA-Cw6, ustekinumab
Introduction
More than 36 genetic loci are associated with psoriasis,1 and of these, HLA-C*06, also known as HLA-Cw6 or PSORS1 (6p21.33), has the strongest disease association.2–4 HLA-Cw6+ patients tend to have an earlier onset of disease and more severe manifestations.5–8 Previous studies have suggested that HLA-Cw6 genotyping could possibly play a role in predicting therapy outcome to the interleukin-12/23 antagonist, ustekinumab, making pharmacogenomics a promising field of research.9–14
In 2013, Talamonti et al published a cohort study with 51 Caucasian patients, in which the treatment response of psoriasis patients under ustekinumab was analyzed with respect to three genetic variations: TNFAIP3 rs610604, late cornified envelope 3B/3C, and HLA-Cw6. While the TNFAIP3 rs610604 polymorphism and the late cornified envelope 3B/3C gene deletions were not found to be associated with statistically significant differences in outcome, patients with a polymorphism on the HLA-Cw6 allele showed increased responses.12
Similar results were observed by Chiu et al in a study of 66 Chinese patients. Other clinical variables were analyzed in a multivariate model. However, age, age of onset, body weight, Psoriasis Activity and Severity Index (PASI) at baseline, psoriatic arthritis, previous biologics used, and previous traditional systemic treatment were not significantly associated with the treatment outcome of the interleukin-12/23 blocker, whereas the Cw*0602 status was.11
Additionally, the presence of a GG genotype on the IL12B rs6887695 single nucleotide polymorphism (SNP) and the absence of an AA genotype on the IL12B rs3212227 single-nucleotide polymorphism have been shown to be associated with a superior treatment outcome. This study also confirmed previous data on HLA-Cw6, as mentioned above.10
The association was also retrospectively analyzed in 601 patients, which showed that there is a superior outcome in HLA-Cw6+ patients; however, the difference observed in this study between the groups was only modest.9
The aim of our study was to determine if there is an association between HLA-Cw6 allele carriers and response to ustekinumab, retrospectively, in our cohort of patients. Our primary endpoint was PASI 75 (a 75% change in PASI) at week 16. Secondary outcomes included PASI 50, 90, as well as changes in Nail Psoriasis Severity Index (NAPSI) or Dermatology Life Quality Index (DLQI) at week 16.
Patients and methods
Patients
Thirty patients with moderate-to-severe psoriasis treated with ustekinumab between January 2015 and March 2017 were included. Prior authorization for the study was obtained from the cantonal ethic commission, Kantonale Ethikkommision Zürich, (BASEC-Nr. 2017-00595), and all participants needed to sign an informed consent “Einrichtung einer dermatologischen Biobank (version 6.0 from 20 February 2015)”. The patients’ medical data were encrypted, and confidentiality was thus ensured. For all patients, ustekinumab was a second-line treatment. First-line treatments included UVA/UVB light therapy, methotrexate (MTX), acitretin, and/or cyclosporine A. Fumaric acid esters are not available in Switzerland. Ustekinumab was administered in the standard dosage (<100 kg: 45 mg, >100 kg: 90 mg; month 0, 1, 3).
Statistical analysis
HLA-Cw6 allele positive patients were compared to wild-type patients for PASI 50, 75, 90, NAPSI, and DLQI at 16 weeks after the onset of therapy. Statistical analysis was conducted with Student’s t-test using GraphPad Prism.
DNA extraction and genotyping
Genomic DNA was isolated from EDTA tubes with venous blood. A primer-specific polymerase chain reaction (Olerup SSP®, Vienna, Austria) was used for HLA-C*06 genotyping.
Results
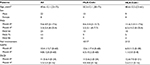
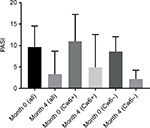
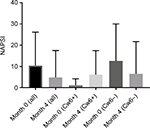
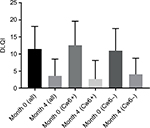
Thirty patients were included in this retrospective study (Table 1), with almost three times as many men as women. At the onset, the average PASI was higher among HLA-Cw6+ patients, which was expected (11 vs. 8.6). The reduction in PASI 75 after 4 months was larger in the HLA-Cw6− participants (34.4%) as compared to HLA-Cw6+ patients (25.6%); however, this did not reach statistical significance (p=0.173). PASI 50, 75, and 90 was achieved by 52.9%, 76.5%, and 29.4% of patients in the HLA-Cw6− group, compared to 76.9%, 38.5%, and 23.1% of patients in the HLA-Cw6+ group, respectively (Figure 1; Table 1). More patients with the HLA-Cw6+ allele had nail psoriasis (10/17), while in the other group, only two out of 13 showed nail involvement. The Nail Psoriasis Severity was higher among Cw6− patients; however, this could largely be due to the small sample size. The effect of ustekinumab between the groups showed no significant difference (p=0.374) regarding nail improvement (Figure 2). Life quality was similarly improved in both the HLA-Cw6− and Cw6+ patient groups (11.5–4.0 vs. 12.6–2.6, p=0.534; Figure 3).
Discussion
In this relatively small retrospective study, we observed a similar level of therapeutic response in psoriasis patients treated with ustekinumab regardless of their HLA-Cw6 status. This is in contrast to the current literature that contains several publications indicating a superior treatment outcome in HLA-Cw6+ patients. A power analysis showed that the small cohort provided only little power (0.55 for PASI, 0.17 for NAPSI, and 0.11 for DLQI) to detect any differences. In addition, the PASI at the start of ustekinumab therapy was on average 8.6 and 11.0, respectively, therefore rather low in comparison to other studies.
However, as we set out to investigate whether on a day-to-day basis, the determination of the Cw6 allele may be of clinical utility or not, our data are relevant for the situation of making a treatment decision in a single patient. Even though in larger cohorts, a small signal for an additional therapeutic benefit of ustekinumab in Cw6+ patients was demonstrated, our findings suggest that Cw6 genotyping is probably of too limited value for routine integration into the clinical laboratory workup of moderate-to-severe patients.
It has to be emphasized that in the most recent study published by Li et al,9 the reported difference between Cw6-positive and -negative groups was only moderate. This difference does not seem sufficient to guide clinical decisions or predictions based on the patient’s Cw6 genotype.
While several studies have previously shown a superior treatment response to ustekinumab in HLA-Cw6+ patients,9–13 the data concerning tumor necrosis factor-alpha antagonists and HLA-Cw6 were conflicting.15,16 In a study by Gonzalez-Lara et al, Cw6+ patients responded better to tumor necrosis factor-alpha antagonists as compared to Cw6− patients.16 A higher response rate of HLA-Cw6+ patients to MTX was seen in one study. Interestingly, HLA-Cw6 status (odds ratio: 2.75) was a more important predictor than even the dose of MTX (odds ratio: 1.08).17 To date, no published studies have assessed the influence of HLA-Cw6 on secukinumab, ixekizumab, or apremilast therapy responses.
As personalized medicine is moving forward at a rapid pace, we are hard-pressed to find biomarkers to predict both therapeutic success and side effects of old and new drugs. HLA-Cw6 certainly is a candidate allele for such a need, but may on its own not mitigate the whole risk. In a previously published case report of homozygous twins, both positive for a HLA-Cw6 mutation, both of them responded well to ustekinumab therapy. Surprisingly, one of the twins was thereafter stable over time (PASI 5), while the other’s symptoms worsened (from PASI 9 to 33).18
Our study has the weakness of including only 30 patients. As it is a retrospective analysis and the genotyping was only done post hoc, observer bias is largely ruled out. It will be interesting to see further studies, especially on secukinumab, ixekizumab, or apremilast; but the quest for a gene aiding us in our choice of therapy and flawlessly predicting therapy outcome is not over yet.
Conclusion
In our opinion, routine HLA-Cw6 genotyping does not impact our clinical routine.
Disclosure
AAN is funded by the Promedica and Bruno - Bloch Foundation. He is on the advisory board of Janssen, AbbVie, Novartis, Celgene, MSD, Pfizer. FA is funded by Hochspezialisierte Medizin 2 from the canton of Zurich; he is also funded by Forschungskredit of the University of Zurich. FA received honoraria from AbbVie, Novartis, Celgene and Eli Lilly. The authors report no other conflicts of interest in this work.
References
O’Rielly DD, Rahman P. Genetics of psoriatic arthritis. Best Pract Res Clin Rheumatol. 2014;28(5):673–685. | ||
Nair RP, Stuart P, Henseler T, et al. Localization of psoriasis-susceptibility locus PSORS1 to a 60-kb interval telomeric to HLA-C. Am J Hum Genet. 2000;66(6):1833–1844. | ||
Duffin KC, Woodcock J, Krueger GG. Genetic variations associated with psoriasis and psoriatic arthritis found by genome-wide association. Dermatol Ther. 2010;23(2):101–113. | ||
Mabuchi T, Ota T, Manabe Y, et al. HLA-C*12:02 is a susceptibility factor in late-onset type of psoriasis in Japanese. J Dermatol. 2014;41(8):697–704. | ||
Ruiz-Romeu E, Ferran M, Sagrista M, et al. Streptococcus pyogenes-induced cutaneous lymphocyte antigen-positive T cell-dependent epidermal cell activation triggers TH17 responses in patients with guttate psoriasis. J Allergy Clin Immunol. 2016;138(2):491–499.e496. | ||
Sokolik R, Gebura K, Iwaszko M, et al. Significance of association of HLA-C and HLA-E with psoriatic arthritis. Hum Immunol. 2014;75(12):1188–1191. | ||
Sathishkumar D, George R, Daniel D, Peter JV. Clinical profile of childhood-onset psoriasis and prevalence of HLA-Cw6: a hospital-based study from India. Postgrad Med J. 2015;91(1076):309–314. | ||
Stuart PE, Nair RP, Hiremagalore R, et al. Comparison of MHC class I risk haplotypes in Thai and Caucasian psoriatics shows locus heterogeneity at PSORS1. Tissue Antigens. 2010;76(5):387–397. | ||
Li K, Huang CC, Randazzo B, et al. HLA-C*06:02 Allele and response to IL-12/23 inhibition: results from the ustekinumab phase 3 psoriasis program. J Invest Dermatol. 2016;136(12):2364–2371. | ||
Galluzzo M, Boca AN, Botti E, et al. IL12B (p40) gene polymorphisms contribute to ustekinumab response prediction in psoriasis. Dermatology. 2016;232(2):230–236. | ||
Chiu HY, Wang TS, Chan CC, Cheng YP, Lin SJ, Tsai TF. Human leucocyte antigen-Cw6 as a predictor for clinical response to ustekinumab, an interleukin-12/23 blocker, in Chinese patients with psoriasis: a retrospective analysis. Br J Dermatol. 2014;171(5):1181–1188. | ||
Talamonti M, Botti E, Galluzzo M, et al. Pharmacogenetics of psoriasis: HLA-Cw6 but not LCE3B/3C deletion nor TNFAIP3 polymorphism predisposes to clinical response to interleukin 12/23 blocker ustekinumab. Br J Dermatol. 2013;169(2):458–463. | ||
Talamonti M, Galluzzo M, Chimenti S, Costanzo A. HLA-C*06 and response to ustekinumab in Caucasian patients with psoriasis: outcome and long-term follow-up. J Am Acad Dermatol. 2016;74(2):374–375. | ||
Gupta R, Debbaneh MG, Liao W. Genetic epidemiology of psoriasis. Curr Dermatol Rep. 2014;3(1):61–78. | ||
Caldarola G, Sgambato A, Fanali C, et al. HLA-Cw6 allele, NFkB1 and NFkBIA polymorphisms play no role in predicting response to etanercept in psoriatic patients. Pharmacogenet Genomics. 2016;26(9):423–427. | ||
Gonzalez-Lara L, Batalla A, Coto E, et al. The TNFRSF1B rs1061622 polymorphism (p.M196R) is associated with biological drug outcome in Psoriasis patients. Arch Dermatol Res. 2015;307(5):405–412. | ||
Indhumathi S, Rajappa M, Chandrashekar L, Ananthanarayanan PH, Thappa DM, Negi VS. Pharmacogenetic markers to predict the clinical response to methotrexate in south Indian Tamil patients with psoriasis. Eur J Clin Pharmacol. 2017;73(8):965–971. | ||
Burlando M, Cozzani E, Campisi C, Di Costanzo A, Parodi A. Different responses to ustekinumab of two HLA-Cw6-positive homozygous twins with psoriasis. Acta Derm Venereol. 2016;96(6):858–859. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.