Back to Journals » International Journal of Nanomedicine » Volume 15
Inhibition of Glioma Cells’ Proliferation by Doxorubicin-Loaded Exosomes via Microfluidics
Authors Thakur A , Sidu RK , Zou H, Alam MK , Yang M, Lee Y
Received 2 June 2020
Accepted for publication 31 August 2020
Published 28 October 2020 Volume 2020:15 Pages 8331—8343
DOI https://doi.org/10.2147/IJN.S263956
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Prof. Dr. Anderson Oliveira Lobo
Supplementary video of "Glioma cells’ proliferation inhibited by DOX-loaded exosomes" [ID 263956].
Views: 195
Abhimanyu Thakur,1 Rakesh Kumar Sidu,1 Heng Zou,2 Md Kowsar Alam,2 Mengsu Yang,2 Youngjin Lee1
1Department of Neuroscience, City University of Hong Kong, Kowloon Tong, Hong Kong SAR; 2Department of Biomedical Sciences, City University of Hong Kong, Kowloon Tong, Hong Kong SAR
Correspondence: Youngjin Lee
Department of Neuroscience, City University of Hong Kong, Yuen Building, City University of Hong Kong 83, Tat Chee Avenue, Kowloon Tong, Hong Kong SAR Tel +852-3442-4313
Fax +852-3442-0549
Email [email protected]
Background: Malignant glioma is a fatal brain cancer. Accumulated evidence has demonstrated that exosomes can cross the blood–brain barrier (BBB), suggesting their potential use as drug delivery vehicles to glioma. Therefore, various loading methods of anticancer agents into exosomes have been developed. However, the loading efficiency of anticancer drugs, such as doxorubicin (DOX) and paclitaxel (PTX), into exosomes is relatively low, thus challenging to improve the drug delivery efficiency to glioma cells (GMs) via exosomes.
Methods: To improve the loading efficiency of doxorubicin into exosomes, a microfluidic device (Exo-Load) was developed. Next, to increase the exosomal delivery of doxorubicin to GMs, autologous exosomes were used for its loading via Exo-Load. Briefly, exosomes from SF7761 stem cells-like- and U251-GMs were isolated and characterized by nano-tracking analysis (NTA), transmission electron microscopy (TEM), and immunogold EM. Finally, doxorubicin was successfully loaded into exosomes with saponin by Exo-Load, and the uptake and functionality of doxorubicin-loaded exosomes for parent GMs were evaluated.
Results: The loading efficiency of DOX into SF7761 stem cells-like- and U251-GMs-derived-exosomes were 19.7% and 7.86% via Exo-Load at the injection flow rate of 50 μL/min, respectively. Interestingly, the loading efficiency of DOX into U251 GMs-derived exosomes was significantly improved to 31.98% by a sigmoid type of Exo-Load at the injection flow rate of 12.5 μL/min. Importantly, DOX-loaded GMs-derived exosomes via Exo-Load inhibited parent GMs’ proliferation more than heterologous GMs, supporting exosomes’ homing effect.
Conclusion: This study revealed that DOX and PTX could be loaded in exosomes via Exo-Load, demonstrating that Exo-Load could be a potential drug-loading device into exosomes with further optimization. This study also demonstrated that the delivery of DOX to SF7761 GMs via their daughter exosomes was much more efficient rather than U251 GMs-derived exosomes, supporting that the use of autologous exosomes could be better for glioma drug targeting.
Keywords: glioma cells, exosomes, drug loading, microfluidics, doxorubicin, paclitaxel
Introduction
Glioma, a major brain tumor, originates from glial stem cells in the central nervous system (CNS).1 Despite rapid progress in its research, the treatment of glioma with potent anticancer drugs, such as doxorubicin (DOX) and paclitaxel (PTX), has a major obstacle in the presence of the blood–brain barrier (BBB), which blocks their penetration into the CNS.2 Therefore, synthetic nanoparticles and liposomes capable of crossing the BBB have been tested for the delivery of these anticancer drugs to glioma, showing their potential application in glioma therapy. However, the synthetic exogenous vehicles generate a broad range of neurotoxicity. For example, they form aggregates and agglomerates in the brain, inducing unexpected neurotoxicity.3 In addition, unmodified liposome-mediated drug delivery is limited owing to the shorter circulation time and instability.4–6 Interestingly, recent reports have demonstrated that exosomes, naturally released small vesicles from every cell type, have great potential as drug delivery vehicles to the CNS without producing significant side-effects as seen in the synthetic vehicles.7 Moreover, the homing capability of exosomes has gained attention as efficient drug delivery vehicles, supporting that autologous exosomes can be better vehicles for drug targeting.8
Exosomes, nano-sized extracellular vesicles with a size range of 30–200 nm, transfer various peptides, fatty acids, and nucleic acids, including microRNA (miRNA), to neighboring cells as the cargo shuttles responsible for intercellular communication.9–11 Particularly, exosomes have the capacity to cross the BBB, and they can be internalized into specific neural cells by endocytosis, pinocytosis, and phagocytosis. Therefore, exosomes have been investigated as vehicles to deliver a BBB-impermeable drug, or RNA interference (RNAi), to targeted brain cells, including glioma cells (GMs),12,13 Importantly, exosomes-mediated delivery of DOX to GMs effectively prevented tumor growth in a zebrafish model,14,15 However, much remains to be elucidated for the development of optimal exosomes-based loading and delivery of anticancer drugs to the CNS. Therefore, exosomes isolated from SF7761 human pediatric diffuse intrinsic pontine GMs, stem-like cells,16 and U251 GMs were investigated for the delivery of DOX and PTX to GMs autologously or heterologously.
For the better delivery of an anticancer drug to glioma as well as the maintenance of effective drug-level in glioma tissues, enhancing the loading efficiency of an anticancer agent into exosomes is crucial. Therefore, to improve the loading efficiency of therapeutic drugs into exosomes, various methods, including electroporation, incubation, and chemical reagents, have been investigated.17–23 (Supplementary Table S1). Particularly, with the progress of micro- and nano-fabrication technologies, the application of microfluidics to drug loading and delivery to cells24,25 has been investigated. However, microfluidics-based drug loading into exosomes has not been much studied and progressed. Microfluidics offers several advantages, including simple, efficient setup, controlled parameters such as flow rate and pressure, to obtain optimal conditions for drug loading.
Microfluidics-assisted drug loading into liposomes for various cancer therapy has been studied for several decades.26 However, drug loading into exosomes via microfluidics has not been investigated.27 Therefore, a microfluidic device, Exo-Load, was developed in this study to test whether DOX and PTX could be loaded into exosomes as well as the high loading efficiency of a drug into exosomes could be achieved by it. In addition, saponin has been extensively used as a permeabilizing agent, changing the properties of the cellular membrane.28 Therefore, we utilized saponin along with DOX to enhance its loading efficiency into exosomes by the combined effect of increased permeabilization- and shear stress-induced stimulation in the microchannel of the Exo-Load microfluidic device (Figure 1). And, finally, the uptake and functionality of DOX-loaded exosomes by Exo-Load was comprehensively investigated.
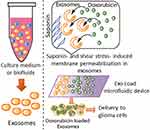 |
Figure 1 Drug loading into exosomes by Exo-Load. Schematic representation of the doxorubicin loading into exosomes via the Exo-Load microfluidic device in the presence of saponin. |
Materials and Methods
Cell Culture
Human pediatric diffuse intrinsic pontine SF7761 GMs (gift from Dr. Kui Ming Chan, BMS, City University of Hong Kong) were cultured in StemPro NSC SFM (Cat# A10509-01, Invitrogen), as described previously.29,30 U251 GMs (Guangzhou Cellcook Biotech Co., Ltd., China) were cultured in Dulbecco’s modified eagle medium (DMEM) with high glucose (Cat# 10569–010, Life Technologies) supplemented with 10% fetal bovine serum (FBS), 100 units/mL penicillin, and 100 μg/mL streptomycin. Both cell lines were cultured and maintained at 37°C and 5% CO2. Both the cells were authenticated by the short tandem repeats (STR) profile.
Isolation of Exosomes
Exosomes from SF7761 GMs were isolated by a modified differential ultracentrifugation method similar to the previous report.10 In brief, the conditioned medium from their 24 hours culture of SF7761 GMs was first centrifuged at 300×g for 10 minutes. The subsequent supernatant was further centrifuged at 16,500×g for 20 minutes, followed by filtration of succeeding supernatant through a 0.2 µm filter to remove microvesicles (MVs) and apoptotic bodies. Finally, the supernatant was subjected to ultra-centrifugation at 120,000×g for 70 minutes to isolate exosomes.
Total Exosome Isolation (TEI) method was utilized in the isolation of U251 GMs-derived exosomes from the conditioned medium of their 24 hours culture (Cat# 4478359, Thermo Scientific).31
Size-Distribution Analysis of Exosomes
Size and concentration of exosomes were analyzed via using a Malvern Nanosight NS300 nano tracking analyzer (NTA) with a syringe pump as described previously.10,31
Transmission Electron Microscopy (TEM) and Immunogold-EM
The morphology and size of exosomes were characterized by the analysis of TEM images (FEI/Philips Tecnai 12 BioTWIN) and exosome identity was confirmed by the analysis of immunogold-EM as described previously,10,32 In brief, for the detection of CD63 of GMs-derived exosomes by immunogold-EM, the blocked grids, containing fixed exosomes with 2% paraformaldehyde (PFA), were transferred to a drop of the anti‐CD63 antibody (dilution=1:100) in PBS/0.5% bovine serum albumin (BSA), and were further incubated for 1 hour. The exosomes on the grids were then washed with PBS/0.5% BSA five times for 3 minutes, followed by their incubation with 10 nm gold‐labeled secondary antibody in PBS/0.5% BSA for 30 minutes, and then washed five times for 3 minutes with 100 µL drops of PBS/0.5% BSA. The exosomes on the grids were stained with 2% uranyl acetate and then observed under a transmission electron microscope (FEI/Philips Tecnai 12 BioTWIN).
Designing and Simulation of the Exo-Load Microfluidic Device
The design-patterns of Exo-Load were drawn by CoralDraw software version X7. The modeling of fluid flow inside the microfluidic channels of Exo-Load was simulated by COMSOL Multiphysics (version 4.4, COMSOL, Inc., Burlington, MA, USA). The 3D-simulation model of the microfluidic channels was constructed for numerical analysis according to the dimension of Exo-Load listed in Supplementary Figure S1. The standard Navier−Stokes equation was employed to simulate the streamlines. The symmetry of boundary condition was set to the no-slip on the walls of microfluidic channels. Similarly, fluid-velocity distribution in the microchannel was simulated to test the mixture of DOX and saponin with exosomes, using the parameters of inlet velocity and channel height. The volume of fluid was estimated based on the dimension of the microchannel.
Fabrication of the Exo-Load Microfluidic Device
Polydimethylsiloxane (PDMS) was used for fabricating the microfluidic device, Exo-Load, by the soft UV lithography technique as described previously.33 In brief, the photomask was first created, and the prototyping process was based on the standard process of silicon soft lithography with slight modifications,34,35 such as adding several tweaks to the process. The photomask created over the printed circuit board (PCB) by Kinston was exposed to UV irradiation for 3 minutes. After photo etching, the PCB board was developed for further chemical etching using Developer 50. Developer 50 treatment was continued till the copper layer of the PCB board was exposed. Thereafter, the PCB board was immersed in 2% FeCl3 (W/V) solution in distilled water for 45 minutes to achieve a microchannel feature with a height of approximately 20 µm. After the etching process, the photomask was washed with acetone and dried under nitrogen gas. Soft lithography was followed by curing polydimethylsiloxane (PDMS) (1:10 w/w) on the etched PCB board with desired features. The inlet and outlet holes (Supplementary Figure S1A) were made using a 0.5 mm diameter biopsy punch. The Exo-load device is sealed with a clean glass slide (Sail brand Microscope slides, Cat# 7101) with the Plasma Cleaner at 800 mTorr for 3 minutes and was fitted with Polyethylene (PE) tubing of 0.58 cm diameter (BD INTRAMEDICTM Polyethylene Tubing, 0.023”x0.038”, Inner diameter x Outer diameter).
Drug Loading into Exosomes by Exo-Load
Fluid flow was controlled via using two syringe pumps (RWD Life Science Co., Ltd) and 1 mL syringe. Three syringes were used to dispense 1 mL of drug, exosomes, and saponin into the three inlets, respectively, and the mixture was collected at the three outlets which could be represented from Supplementary Figure 1A.
As described previously,17 the loading efficiency of DOX into exosomes with incubation, electroporation, and sonication was relatively low. To facilitate its loading efficiency, SF7761 GMs-derived or U251 GMs-derived exosomes (1 mL with 4.0×106 particles/mL) were injected into the central inlet channel of Exo-Load device via using a peristaltic pump through valve V3 at the flow rate of 2.4 ms−1 at room temperature. Concurrently, DOX (1 mL with 22 µM) (Abcam, Cat# ab120629) and saponin (1 mL with 18 µM) (Sigma Aldrich, Cat# 47036) were injected via two side channels, respectively, with the same flow rate, for the increased permeabilization of exosome membrane during drug loading. The mixtures from the outlets were collected in 1.5 mL centrifuge tubes (for three outlets). The mixtures were centrifuged at 10,000×g, 4°C for 1 hour, and the supernatant was analyzed for the measurement of free drug concentration using a UV-Visible spectrophotometer and HPLC. Similarly, PTX was loaded into SF7761 GMs-derived exosomes via a linear Exo-Load device.
Determination of the Loading Efficiency of DOX in Exosomes
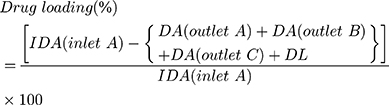
The uploaded concentration of DOX and PTX in exosomes via Exo-Load were indirectly determined by an Ultra-Violet Visible Scanning Spectrophotometer (Shimadzu 1700) at 490 nm or high-performance liquid chromatography (HPLC) (Waters HPLC-PDA-FLR fraction collector system).36 In brief, standard curves of absorbance vs concentration for the quantification of DOX and PTX were created via plotting absorbance in response to the range of 0–50 µM DOX and PTX in 1% DMSO. The concentration of free DOX and PTX in the supernatant of exosome solution was calculated according to the absorbance value obtained at λmax corresponding to the reference DOX and PTX concentration in the standard curve or HPLC. The loading efficiency of DOX and PTX into exosomes was calculated using the following formula:
where IDA is the initial drug amount, DA is the drug amount in supernatant (nmole), and DL is the amount of drug loss during loading through Exo-Load (nmole).
Study of Drug Release Profile
The release of Dox and PTX were evaluated as described previously.22 In short, 4 mL of Exo-Dox solution was transferred into a dialysis tube (M.W. cut-off: 14 kDa; Shanghai. Green Bird Technology Development Co., Ltd., China). The tube was first placed into 10 mL of PBS buffer (pH 7.4) or acetate buffer (pH 5.0). The release of DOX was performed at 37°C. At selected time intervals (3, 6, 12, 24, and 36 hours), the dialysate was removed for the analysis of absorbance and replaced with a fresh buffer solution. The concentrations of DOX were obtained based on the standard curves.
Determination of Exosomes Uptake into Recipient Cells
DOX-loaded SF7761 GMs-derived exosomes were labelled via using Exo-green (Cat# EXOG200A-1, System Biosciences) as per the manufacturer’s instructions. Equal amounts of labelled SF7761 exosomes were incubated with SF7761- or U251- GMs for 4–24 hours, and they were counterstained with phalloidin-rhodamine, a red fluorescent marker for labeling the membrane of cells, and DAPI, a blue fluorescent marker for labeling nuclei of cells. The uptake of DOX-loaded SF7761 GMs-exosomes (Exo-Dox) was imaged with SF7761- and U251- GMs using a confocal microscope (Zeiss Laser Scanning Microscope LSM 880 NLO with Airyscan) at 60x objective setup, as described previously.37,38
Immunocytochemistry
Immunocytochemistry was done as described previously.38 Briefly, SF7761- and U251- GMs were grown on poly-D-lysine coated-glass coverslips until they reached 70% confluence, followed by treatment with Exo-Green labeled SF7761 GMs-derived EXO-DOXs. Subsequently, GMs were washed with PBS, fixed with 4% paraformaldehyde, and permeabilized with 0.1% Triton X-100 in PBS solution for 10 minutes. Blocking was done with 1% BSA in PBS for 1 hour and primary antibody was incubated at RT for 2 hours. Rhodamine phalloidin-conjugated primary antibody (Cat# R415, Invitrogen) solution was dissolved in blocking solution (1:200 dilution) and washes were performed with PBS. Coverslips were mounted in a Vectashield mounting medium with DAPI (Cat# H-1200, Vector Laboratories), and examined on a Zeiss Laser Scanning Microscope LSM 880 NLO with Airyscan.
Flow Cytometry
To analyze the uptake of DOX into SF7761 GMs-derived exosomes, the exosomes (labeled with BODIPY® TR ceramide for membrane staining: red) alone and exosomes loaded with DOX (red) were examined with a Cytoflex flow cytometer, after capturing on 4 µm latex beads (Thermo Scientific). The gate was set on microbeads, and exosomes captured on microbeads based on forward vs side scatter (FSC vs SSC) within the gated population were collected per sample. Uptake of DOX into exosomes was measured by the shift of peak red fluorescence intensity, calculated by the geometric mean of the population.
Cell Viability Assay
MTT cell viability assay was performed as described previously.39 Briefly, GMs (10,000 cells per well) were seeded in a 96-well microtiter tissue culture plate. After 48 hours of culture, the medium was removed, and GMs were washed with PBS. Subsequently, GMs were incubated with 20 μL of exosomes, DOX (concentration=4.3 µM), and EXO-DOXs (concentration of DOX=4.3 µM) for 24 hours; control was treated with vehicles (DMSO in medium). After incubation, medium in GMs’ culture was removed, and MTT (5 mg/mL in DMSO) solution was added to each well. Cells were then incubated at 37°C for 4 hours. Next, medium in each well was removed, and 150 µL of DMSO was added to each well and mixed thoroughly. Finally, the plates were incubated at 37°C for 10 minutes, and absorbance was measured on a microtiter plate reader at 570 nm.
Statistical Analysis
Data were analyzed using GraphPad Prism, version 7.01 software. Results were expressed as the mean±SE of three replicates. The difference between control and experimental group(s) was analyzed by using either Student's t-test or one way-ANOVA test. P<0.05 and P<0.01 were considered statistically significant.
Results
Characterization of SF7761 Stem-Like- and U251-GMs-Derived Exosomes
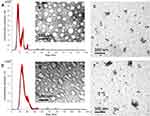
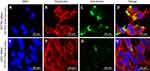
Exosomes released from SF7761- and U251- GMs were isolated by a differential ultracentrifugation and a TEI method. The size, morphology, and the presence of CD63, a representative exosome marker, of isolated exosomes were further analyzed by NTA, TEM, and immunogold EM, respectively. SF7761 GMs-derived exosomes were mainly 100 nm in size (Figure 2A), mostly round in shape (Figure 2B), and CD63-positive in immunogold EM (Figure 2C). In contrast, U251 GMs-derived exosomes with CD63-positive dots (Figure 2E) were mainly 150 nm in size (Figure 2D) with heterogeneous shape (Figure 2E). Particularly, SF7761 GMs appeared to release more exosomes than that of U251 GMs (Figure 2A, D), although statistical analysis with multiple cell culture conditions for each cell type were not further pursued.
COMSOL Simulation of the Exo-Load Microfluidic Device
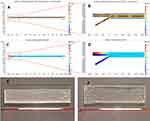
To predict whether DOX, saponin, and exosomes could be mixed in the microchannel of the Exo-Load microfluidic device, COMSOL Multiphysics software was used. COMSOL simulation was based on a laminar flow model due to the small dimensions of the microchannel (length; l=5.4 mm, and height; h=20 µm). Exo-Load was designed to have three inlets (marked as inlet a, b, and c) for uploading DOX (or PTX), saponin, and exosomes which could be exchanged with each other through the pillars between three microchannels, and three outlets (marked as outlet a, b, and c) for taking DOX-loaded exosomes. The velocity field of Exo-Load showed higher magnitude of the fluid-velocity in the middle channel at approximately 3 m/s than that in the outer walls at 0.5 m/s (Figure 3A and B), indicating that exosomes in the middle channel had higher shear stress (Figure 3E) and the drug flowed at inlet-a (Supplementary Figure S1A) could be distributed and mixed gradually throughout the channels. Additionally, the fluid-concentration profile of Exo-Load (Figure 3C and D) indicated that the initial concentration of drug, 4.5 mol/m3, was distributed and reached a final concentration of drug, 2 mol/m3. Moreover, the study demonstrated that the mixing of each component within the microchannel was a gradual and steady process after the perfusion of each component with a constant velocity at 50 µL/min. Interestingly, COMSOL simulation showed that Exo-Load with two inlet-channels also had higher velocity in the middle channel as well (Supplementary Figure S1B). Based on COMSOL simulation, the Exo-Load microfluidic device was fabricated via using the soft lithography method (Supplementary Figure S1C) with the design (Figure 2E, Supplementary Figure S1D-E). To enhance drug–exosomes interactions in the microchannel, the sigmoid type of Exo-Load was also fabricated (Figure 3F). And, actual fluid flow inside the microchannel and the mixing effect between the channels were investigated by the introduction of trypan blue dye in inlet A. The region of interest (ROI) was adjusted at the middle of the microchannel such that the three channels could be viewed at the same time. 0.4% Trypan blue dissolved in 1 mL PBS was perfused through each Inlet- A, B, and C, respectively, via using syringe pumps (RWD Life Science Co., Ltd) (Supplementary Figure S1D-E). Supplementary video S1 was recorded simultaneously with the start of perfusion into the microfluidics. Supplementary video S1 shows the laminar flow and mixing phenomena along the pillars of the microchannel. This observation supports the hypothesis of the design aimed for loading drugs into exosomes by their mixture in the microchannel (Supplementary video S1).
Loading of DOX and PTX in GMs-Derived Exosomes
Through the process of optimization, injection volume and concentration of DOX (1 mL, 22 µM, 22 nmole), saponin (1 mL, 18 µM, 18 nmole), and exosomes (1 mL, 4.0×106 particles/mL) at the flow rate 50 µL/min were used in the Exo-Load-based DOX loading into GMs-derived exosomes. The basic functionality of Exo-Load was tested in the initial optimized condition; nonetheless it could be better with further optimization in the drug loading into exosomes.
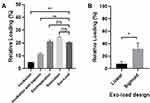
Based on the equation described in the method, the loading efficiency of a drug into GMs- derived exosomes by Exo-Load was calculated. The loading efficiency of DOX in SF7761 GMs-derived exosomes was 19.7% at the injection flow rate, 50 µL/min, which was compared with the drug loading efficiency obtained with conventional techniques, such as simple incubation, incubation with saponin, electroporation, and sonication (Figure 4A). In-detail data for the calculation of the loading efficiency of drug into exosomes was indicated in Supplementary Table S2. In addition, HPLC analysis in Supp Table S3 indicates that the loading efficiency of DOX into exosomes is around 16% (Supplementary Figure S2). Particularly, the loading efficiency of PTX, a hydrophilic drug, into SF7761 GMs-derived exosomes was 17.7% (Supplementary Figure S3). Conclusively, both DOX and PTX could be significantly loaded into SF7761 GMs-derived exosomes by Exo-Load. Furthermore, confocal microscopy analysis of DOX-loaded exosomes visualized that Exo-Load made the loading of DOX in SF7761 GMs-derived exosomes (Supplementary Figure S4A-C). At the same time, flow cytometry analysis showed the shift of exosome fluorescence intensity from 104 to 107 after the addition of DOX to SF7761 GMs-derived exosomes (Supplementary Figure S4D). Altogether, it was evident that DOX was successfully loaded into SF7761 GMs-derived exosomes in the presence of saponin via Exo-Load. However, the loading efficiency of DOX in U251 GMs-derived exosomes was relatively low, such as 7.85% and 7.86% at the injection flow rate, 12.5 µL/min and 50 µL/min, respectively (Figure 4A–B). To improve the loading efficiency of drug into exosomes, the sigmoid type of Exo-Load was developed and tested to load DOX into U251 GMs-derived exosomes. The loading efficiency of DOX in U251 GMs-derived exosomes was enhanced, reaching 31.98% at the injection flow rate of 12.5 µL/min (Figure 4B).
Higher Uptake of DOX-Loaded Autologous Exosomes by Parent GMs
Several studies demonstrated that autologous exosomes could be used as ideal delivery vehicles of therapeutic agents to parent cells with a homing effect.40 Therefore, to determine whether the uptake of DOX-loaded SF7761 GMs-derived exosomes (SF7761 GMs-derived EXO-DOXs) into SF7761- and U251- GMs could be different from each other; Exo-green-labeled SF7761 GMs-derived EXO-DOXs were incubated with SF7761- or U251- GMs for 4–24 hours. Exosomes were labeled with Exo-green (green) to clearly determine the distribution of SF7761 GMs-derived EXO-DOXs inside cells. SF7761- and U251- GMs were visualized with the counterstaining with phalloidin-rhodamine, a red fluorescent marker for labeling the membrane of cells, and DAPI, a blue fluorescent marker for labeling the nuclei of cells, although DOX produced intrinsic fluorescence (weak red) (Figure 5). The result at multiple time points after the co-incubation of exosomes and recipient cells revealed that the uptake of SF7761 GMs-derived EXO-DOXs by SF7761 GMs were much higher than that of U251 GMs (Figure 5A–H, data not shown), supporting the previous observation, higher uptake of autologous exosomes in other cancer cells,19,41 Most SF7761 GMs with SF7761 GMs-derived EXO-DOXs were dead during co-incubation and even survived SF7761 GMs with SF7761 GMs-derived EXO-DOXs appeared to be unhealthy in terms of morphology.
Exosomes-Mediated Delivery of DOX Inhibits the Proliferation of GMs
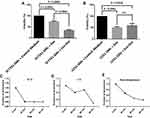
To determine the effect of SF7761 GMs-derived EXO-DOXs on the viability of cancer cells, SF7761- and U251- GMs were incubated with cell culture medium (control), DOX, or SF7761 GMs-derived EXO-DOXs in a 96-well plate, and the viability was measured after 4 hours-incubation (Figure 6A–B). The result indicated that SF7761 GMs-derived EXO-DOXs treatment was much more effective to inhibit GMs’ proliferation rather than only DOX treatment. Furthermore, it was found that of the number of survived SF7761 GMs was much more significantly reduced with treatment with SF7761 GMs-derived EXO-DOXs than that of U251 GMs (Figure 6A–B), which was also correlated with the higher uptake of SF7761 GMs-derived EXO-DOXs by autologous SF7761 GMs.
Finally, to examine the stability of SF7761 GMs-derived EXO-DOXs, the amount of DOX released from SF7761 GMs-derived EXO-DOXs into the supernatant with respect to time was measured.36 Interestingly, SF7761 GMs-derived EXO-DOXs were stable in every condition; in particular, release of DOX from EXO-DOXs at 37°C, body temperature, was below 15% during 90 minutes-incubation. At 4°C, stability of EXO-DOXs were well maintained during 60 minute-incubation (Figure 6C–E). The release profile of DOX and PTX from SF7761-derived exosomes was determined at two different conditions, pH 7.4 and pH 5, by the quantification of free drug in the supernatant using a dialysis tube method (Molecular weight cut-off 14 kDa) as described previously.21,22 The release pattern was investigated at pH 5.0 because it was a common pH in the hypoxic tumor microenvironment. Interestingly, the release of DOX and PTX from exosomes was enhanced at pH 5.0 as compared to at pH 7.4 (Supplementary Figure S5A and B). In general, the release profile of DOX and PTX from exosomes was slightly different from each other. In the early phase (below 6 hours) at pH 7.4, DOX and PTX had similar release patterns; however, PTX release from exosomes was saturated (lag phase) after 12 hours, in contrast to the continuous release of DOX from exosomes, indicating that DOX might be more encapsulated in the inner space of exosomes. In addition, the TEM examination of size and morphology of EXO-DOXs suggested that the overall size of exosomes was slightly increased upon loading with DOX with good stability (Supplementary Figure 5C-F).
Discussion
One of the major hurdles in glioma treatment is the delivery of effective anticancer drugs, such as DOX and PTX, to the GMs through the BBB.42 Interestingly, GMs release a tremendous amount of exosomes,43,44 which have strong potential as drug delivery vehicles because of their nano size (30–200 nm) and lipophilic biophysical property, allowing them to cross the BBB, enabling them to reach targeted cells in the brain, including GMs.45 Furthermore, anticancer drugs, microRNA, and RNAi, can be loaded into autologous or engineered exosomes which can be specifically transported to targeted neural cell types without producing many side-effects,46,47 Therefore, in this study, exosomes were first isolated and characterized from two different human glioma cell lines, pediatric diffuse intrinsic pontine SF7761 stem cell-like GMs and U251 GMs, to determine whether GMs-derived exosomes could be used as good drug delivery vehicles. The result indicated that SF7761 GMs-derived exosomes were relatively homogenous, round in shape, and were higher in their release from SF7761 GMs, compared with exosome release from U251 GMs, demonstrating the different size, morphology, and release capacity of GMs-derived exosomes depending on their parent GMs (Figure 2A-E), which could be important information in determining the phenotype of cancer as well as developing new delivery vehicles for anticancer drugs. Exosomes have been investigated as drug delivery vehicles for a decade,27,48 and, therefore, various conventional techniques have been developed for the loading of a drug into exosomes, which includes incubation with saponin, electroporation, and sonication,17–20 (Supplementary Table S1). However, improving both drug loading efficiency in exosomes and stability of drug-loaded exosomes has been much challenged to enable the better drug delivery to the specific targeted cells. Therefore, the microfluidics-based loading of a drug in exosomes was tested in this study. Some reports demonstrated that microfluidics could provide a better performance in loading a drug in cells better than conventional techniques due to its capability of precise and controlled setup that could be easily customized to achieve enhanced drug loading.49,50 In the present work, the linear and sigmoid types of Exo-Load were developed to increase the loading efficiency of DOX and PTX into GMs-derived exosomes (Figure 3A–F). Particularly in this study, saponin was utilized as a membrane permeabilizing agent.20,27 In addition, the number of exosomes (1 mL of 4.0×106 particles) injected into Exo-Load was optimized. With the experimental conditions as noted (at the flow rate 50 µL/min), DOX loading efficiency for SF7761 GMs-derived exosomes (19.7%) by the linear type of Exo-Load was better than that of U251 GMs-derived exosomes (7.86%) (Figure 4A–B), demonstrating that SF7761 GMs-derived exosomes had different membrane properties and could be better in the loading of a drug into exosomes, as similar to that of hematopoietic stem cells- and mesenchymal stem cells (MSCs)- derived exosomes in the loading and delivery of a drug into exosomes.51–53 Not only DOX but also PTX, a relatively more hydrophilic drug, was loaded well into SF7761 GMs-derived exosomes, revealing that its loading efficiency was around 17.7% (Supplementary Figure S3). Importantly, the sigmoid type of Exo-Load improved the loading efficiency of DOX into U251 GMs-derived exosomes at the biggest value, 31.98%, at the injection flow rate, 12.5 µL/min (Figure 4B), presumably due to the enhanced mixing effect and high retention time, indicating that Exo-Load-based drug loading into exosomes could be further improved by the additional optimization of loading conditions in the future. To test their stability as drug delivery vehicles, SF7761 GMs-derived EXO-DOXs were incubated in different temperatures. They were stable at 37°C for at least 90 minutes (Figure 6C–E), demonstrating its potential use in vivo. Finally, the uptake and effect of SF7761 GMs-derived EXO-DOXs in the autologous SF7761- and heterologous U251- GMs were investigated. Autologous uptake of SF7761 GMs-derived EXO-DOXs in SF7761 GMs was more significant than heterologous uptake of SF7761 GMs-derived EXO-DOXs in U251 GMs (Figure 5A–H), indicating the presence of a specific mechanism for their autologous interaction, which provided great insights into the development of the specific delivery vehicles of anticancer drugs to parent GMs (Figure 5A–E). More importantly, SF7761 GMs-derived EXO-DOXs inhibited the proliferation of SF7761 GMs more than U251 GMs, which was correlated with the result of uptake assay. Distinctively, SF7761 GMs-derived EXO-DOXs was also much more potent in the inhibition of SF7761 GMs than only DOXs (Figure 6A–B), indicating that they were functional as efficient delivery vehicles for anticancer drugs. It may be very interesting in the future to investigate whether SF7761 GMs-derived EXO-DOXs can be effective in reducing in vivo tumor progression of a mouse model of glioma.
Conclusion
In the current study, BBB-impermeable two anticancer agents, DOX and PTX, were successfully loaded into SF7761 stem cell-like GMs-derived exosomes by the Exo-Load microfluidic device in the presence of saponin, a permeabilization agent, and shear stress in microfluidic channels. Loading performance of DOX in U251 GMs-derived exosomes by the sigmoid type of Exo-Load was better than the linear type of Exo-Load, suggesting that Exo-Load-based drug loading into exosomes could be promising through further improvement and optimization processes. Importantly, the uptake of SF7761 GMs-derived EXO-DOXs by autologous SF7761 GMs was much more than heterologous U251 GMs, leading to a potent anti-proliferation effect for SF7761 GMs. In conclusion, the present study for the first time proposed an Exo-Load (microfluidic) device-based loading of anticancer agents into exosomes, and it further demonstrated that the autologous uptake of EXO-DOXs could efficiently inhibit parent GMs’ proliferation. Therefore, it would be interesting to test whether EXO-DOXs could inhibit tumor growth in a mouse model of glioma by its delivery through the olfactory route with a nasal spray formulation or tail-vein injection in the future.
Acknowledgments
The project was supported by the City University of Hong Kong (Grant No. 9610340 and 7200472), Early Career Scheme (ECS)-UGC (Grant No. 21102517) and General Research Fund (GRF)-UGC (Grant No. 11103918) of Hong Kong Research Grant Council awarded to Dr. Youngjin Lee at City University of Hong Kong. This work was also supported by grants from the General Research Fund (CityU_11319516) and the Research Impact Fund (R1020-18F) of Hong Kong Research Grant Council, and the Basic Research Projects of Shenzhen Knowledge Innovation Program (JCYJ20180307123759162) awarded to Dr. Mengsu Yang at City University of Hong Kong.
Author Contributions
AT and YL designed the experiments and wrote the manuscript. AT, RKS, HZ, and MKA performed microfluidic-related experiments, COMSOL simulation was done by HZ, and the remaining experiments were done by AT. All authors contributed to the data analysis, drafting, or revising of the article, have agreed on the journal to which the article will be submitted, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Disclosure
The authors report no conflicts of interest in publishing this work. A conference abstract related to this research work has been presented at the American Association for Cancer Research (AACR) Annual meeting-2018, Chicago, IL, USA. In: Proceedings of the AACR Annual Meeting 2018; 2018 Apr 14-18; Chicago, IL. Philadelphia (PA): AACR; Cancer Res 2018;78(13 Suppl): Abstract nr 3720 (DOI: 10.1158/1538-7445.AM2018-3720).
References
1. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2016. CA Cancer J Clin. 2016;66:7–30. doi:10.3322/caac.21332
2. Gan HK, Rosenthal MA, Cher L, et al. Management of glioblastoma in Victoria, Australia (2006–2008). J Clin Neurosci. 2015;22(9):1462–1466. doi:10.1016/j.jocn.2015.03.029
3. Lai RC, Yeo RWY, Tan KH, Lim SK. Exosomes for drug delivery — a novel application for the mesenchymal stem cell. Biotechnol Adv. 2013;31:543–551. doi:10.1016/j.biotechadv.2012.08.008
4. Wang X, Wang Y, Chen ZG, Shin DM. Advances of cancer therapy by nanotechnology. Cancer Res Treat. 2009;41:1. doi:10.4143/crt.2009.41.1.1
5. Torchilin VP. Recent advances with liposomes as pharmaceutical carriers. Nat Rev Drug Discov. 2005;4:145–160. doi:10.1038/nrd1632
6. Çağdaş M, Sezer AD, Bucak S. Liposomes as potential drug carrier systems for drug delivery. Appl Nanotechnol. 2014. doi:doi.10.5772/58459
7. Tan S, Wu T, Zhang D, Zhang Z. Cell or cell membrane-based drug delivery systems. Theranostics. 2015;5:863–881. doi:10.7150/thno.11852
8. Bunggulawa EJ, Wang W, Yin T, et al. Recent advancements in the use of exosomes as drug delivery systems. J Nanobiotechnology. 2018;16:81. doi:10.1186/s12951-018-0403-9
9. Raposo G, Stoorvogel W. Extracellular vesicles: exosomes, microvesicles, and friends. J Cell Biol. 2013;200:373–383. doi:10.1083/jcb.201211138
10. Thakur A, Qiu G, NG S-p G, et al. Direct detection of two different tumor-derived extracellular vesicles by SAM-AuNIs LSPR biosensor. Biosens Bioelectron. 2017;94:400–407. doi:10.1016/j.bios.2017.03.036
11. Thakur A, Qiu G, NG S-P, Wu C-ML, Lee Y. Detection of membrane antigens of extracellular vesicles by surface plasmon resonance. J Lab Precis Med. 2017;2:98. doi:10.21037/jlpm.2017.12.08
12. Jiang L, Vader P, Schiffelers RM. Extracellular vesicles for nucleic acid delivery: progress and prospects for safe RNA-based gene therapy. Gene Ther. 2017;24:157–166. doi:10.1038/gt.2017.8
13. Alvarez-Erviti L, Seow Y, Yin H, Betts C, Lakhal S, Wood MJA. Delivery of siRNA to the mouse brain by systemic injection of targeted exosomes. Nat Biotechnol. 2011;29:341–345. doi:10.1038/nbt.1807
14. Yang T, Martin P, Fogarty B, et al. Exosome delivered anticancer drugs across the blood-brain barrier for brain cancer therapy in Danio Rerio. Pharm Res. 2015;32:2003–2014. doi:10.1007/s11095-014-1593-y
15. Syn NL, Wang L, Chow EK-H, Lim CT, Goh B-C. Exosomes in cancer nanomedicine and immunotherapy: prospects and challenges. Trends Biotechnol. 2017;35:665–676. doi:10.1016/j.tibtech.2017.03.004
16. Taylor IC, Hütt-Cabezas M, Brandt WD, et al. Disrupting NOTCH slows diffuse intrinsic pontine glioma growth, enhances radiation sensitivity, and shows combinatorial efficacy with bromodomain inhibition. J Neuropathol Exp Neurol. 2015;74:778–790. doi:10.1097/NEN.0000000000000216
17. Haney MJ, Klyachko NL, Zhao Y, et al. Exosomes as drug delivery vehicles for Parkinson’s disease therapy. J Control Release. 2015;207:18–30. doi:10.1016/j.jconrel.2015.03.033
18. Kim MS, Haney MJ, Zhao Y, et al. Development of exosome-encapsulated paclitaxel to overcome MDR in cancer cells. Nanomed Nanotechnol Biol Med. 2016;12:655–664. doi:10.1016/j.nano.2015.10.012
19. Saari H, Lázaro-Ibáñez E, Viitala T, Vuorimaa-Laukkanen E, Siljander P, Yliperttula M. Microvesicle- and exosome-mediated drug delivery enhances the cytotoxicity of Paclitaxel in autologous prostate cancer cells. J Control Release. 2015;220:727–737. doi:10.1016/j.jconrel.2015.09.031
20. Fuhrmann G, Serio A, Mazo M, Nair R, Stevens MM. Active loading into extracellular vesicles significantly improves the cellular uptake and photodynamic effect of porphyrins. J Control Release. 2015;205:35–44. doi:10.1016/j.jconrel.2014.11.029
21. Wei H, Chen J, Wang S, et al. A nanodrug consisting of doxorubicin and exosome derived from mesenchymal stem cells for osteosarcoma treatment in vitro. Int J Nanomedicine. 2019;14:8603–8610. doi:10.2147/IJN.S218988
22. Yang L, Han D, Zhan Q, et al. Blood TfR+ exosomes separated by a pH-responsive method deliver chemotherapeutics for tumor therapy. Theranostics. 2019;9:7680–7696. doi:10.7150/thno.37220
23. Salarpour S, Forootanfar H, Pournamdari M, Ahmadi-Zeidabadi M, Esmaeeli M, Pardakhty A. Paclitaxel incorporated exosomes derived from glioblastoma cells: comparative study of two loading techniques. DARU J Pharm Sci. 2019;27:533–539. doi:10.1007/s40199-019-00280-5
24. Maeki M. Microfluidics for pharmaceutical applications. Microfluid Pharm Appl, Elsevier. 2019;101–119. doi:10.1016/B978-0-12-812659-2.00004-1
25. Damiati S, Kompella U, Damiati S, Kodzius R. Microfluidic devices for drug delivery systems and drug screening. Genes (Basel). 2018;9:103. doi:10.3390/genes9020103
26. Casadó A, Sagristá ML, Mora Giménez M. A novel microfluidic liposomal formulation for the delivery of the SN-38 camptothecin: characterization and in vitro assessment of its cytotoxic effect on two tumor cell lines. Int J Nanomedicine. 2018;13:5301–5320. doi:10.2147/IJN.S166219
27. Antimisiaris S, Mourtas S, Marazioti A. Exosomes and exosome-inspired vesicles for targeted drug delivery. Pharmaceutics. 2018;10:218. doi:10.3390/pharmaceutics10040218
28. Jamur MC, Oliver C. Permeabilization of Cell Membranes. 2010:63–66. doi:10.1007/978-1-59745-324-0_9
29. Thakur A, Qiu G, Xu C, et al. Label-free sensing of exosomal MCT1 and CD147 for tracking metabolic reprogramming and malignant progression in glioma. Sci Adv. 2020;(6):eaaz6119. doi:10.1126/sciadv.aaz6119
30. Chan KM, Han J, Fang D, Gan H, Zhang Z. A lesson learned from the H3.3K27M mutation found in pediatric glioma. Cell Cycle. 2013;12:2546–2552. doi:10.4161/cc.25625
31. Qiu G, Thakur A, Xu C, Ng S-P, Lee Y, Wu C-ML. Detection of glioma-derived exosomes with the biotinylated antibody-functionalized titanium nitride plasmonic biosensor. Adv Funct Mater. 2019;29:1806761. doi:10.1002/adfm.201806761
32. Théry C, Amigorena S, Raposo G, Clayton A. Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Curr Protoc Cell Biol. 2006;30:
33. Ketpun P, Tongmanee B, Piyaviriyakul S, et al. A potential application of triangular microwells to entrap single cancer cells: a canine cutaneous mast cell tumor model. Micromachines. 2019;10:841. doi:10.3390/mi10120841
34. Zou H, Yue W, Yu W-K, et al. Microfluidic platform for studying chemotaxis of adhesive cells revealed a gradient-dependent migration and acceleration of cancer stem cells. Anal Chem. 2015;87:7098–7108. doi:10.1021/acs.analchem.5b00873
35. Li C-W, Yang J, Yang M. Dose-dependent cell-based assays in V-shaped microfluidic channels. Lab Chip. 2006;6:921. doi:10.1039/b600058d
36. Sun D, Zhuang X, Xiang X, et al. A novel nanoparticle drug delivery system: the anti-inflammatory activity of curcumin is enhanced When encapsulated in exosomes. Mol Ther. 2010;18:1606–1614. doi:10.1038/mt.2010.105
37. Zhang X, Shastry S, Bradforth SE, Nadeau JL. Nuclear uptake of ultrasmall gold-doxorubicin conjugates imaged by fluorescence lifetime imaging microscopy (FLIM) and electron microscopy. Nanoscale. 2015;7:240–251. doi:10.1039/C4NR04707A
38. Escrevente C, Keller S, Altevogt P, Costa J. Interaction and uptake of exosomes by ovarian cancer cells. BMC Cancer. 2011;11:108. doi:10.1186/1471-2407-11-108
39. Rawat A, Majumder QH, Ahsan F. Inhalable large porous microspheres of low molecular weight heparin: in vitro and in vivo evaluation. J Control Release. 2008;128:224–232. doi:10.1016/j.jconrel.2008.03.013
40. Campanella C, Caruso Bavisotto C, Logozzi M, et al. On the choice of the extracellular vesicles for therapeutic purposes. Int J Mol Sci. 2019;20:236. doi:10.3390/ijms20020236
41. Emam SE, Abu Lila AS, Elsadek NE, et al. Cancer cell-type tropism is one of crucial determinants for the efficient systemic delivery of cancer cell-derived exosomes to tumor tissues. Eur J Pharm Biopharm. 2019;145:27–34. doi:10.1016/j.ejpb.2019.10.005
42. Wang D, Wang C, Wang L, Chen Y. A comprehensive review in improving delivery of small-molecule chemotherapeutic agents overcoming the blood-brain/brain tumor barriers for glioblastoma treatment. Drug Deliv. 2019;26:551–565. doi:10.1080/10717544.2019.1616235
43. Kucharzewska P, Christianson HC, Welch JE, et al. Exosomes reflect the hypoxic status of glioma cells and mediate hypoxia-dependent activation of vascular cells during tumor development. Proc Natl Acad Sci. 2013;110(18):7312–7317. doi:10.1073/pnas.1220998110
44. Skog J, Würdinger T, van Rijn S, et al. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat Cell Biol. 2008;10:1470–1476. doi:10.1038/ncb1800
45. Villa F, Quarto R, Tasso R. Extracellular vesicles as natural, safe and efficient drug delivery systems. Pharmaceutics. 2019;11:557. doi:10.3390/pharmaceutics11110557
46. You B, Xu W, Zhang B. Engineering exosomes: a new direction for anticancer treatment. Am J Cancer Res. 2018;8:1332–1342.
47. Ohno S, Kuroda M. Exosome-Mediated Targeted Delivery of miRNAs, 2016 261–270. doi:10.1007/978-1-4939-3753-0_19.
48. Ha D, Yang N, Nadithe V. Exosomes as therapeutic drug carriers and delivery vehicles across biological membranes: current perspectives and future challenges. Acta Pharm Sin B. 2016;6:287–296. doi:10.1016/j.apsb.2016.02.001
49. Calistri NL, Kimmerling RJ, Malinowski SW, et al. Microfluidic active loading of single cells enables analysis of complex clinical specimens. Nat Commun. 2018;9:4784. doi:10.1038/s41467-018-07283-x
50. Duncombe TA, Tentori AM, Herr AE. Microfluidics: reframing biological enquiry. Nat Rev Mol Cell Biol. 2015;16:554–567. doi:10.1038/nrm4041
51. Yeo RWY, Lai RC, Zhang B, et al. Mesenchymal stem cell: an efficient mass producer of exosomes for drug delivery. Adv Drug Deliv Rev. 2013;65:336–341. doi:10.1016/j.addr.2012.07.001
52. Liu C, Su C. Design strategies and application progress of therapeutic exosomes. Theranostics. 2019;9:1015–1028. doi:10.7150/thno.30853
53. Munoz JL, Bliss SA, Greco SJ, Ramkissoon SH, Ligon KL, Rameshwar P. Delivery of functional anti-miR-9 by mesenchymal stem cell–derived exosomes to glioblastoma multiforme cells conferred chemosensitivity. Mol Ther Nucl Acids. 2013;2:e126. doi:10.1038/mtna.2013.60
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.