Back to Journals » OncoTargets and Therapy » Volume 14
Increased HSF1 Promotes Infiltration and Metastasis in Cervical Cancer via Enhancing MTDH-VEGF-C Expression
Authors Shi X, Deng Z, Wang S, Zhao S, Xiao L, Zou J, Li T, Tan S, Tan S, Xiao X
Received 29 November 2020
Accepted for publication 6 February 2021
Published 26 February 2021 Volume 2021:14 Pages 1305—1315
DOI https://doi.org/10.2147/OTT.S291812
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Prof. Dr. Nicola Silvestris
Xueyan Shi,1 Zhenghao Deng,2 Shouman Wang,3 Shuai Zhao,1,4 Lan Xiao,5 Jiang Zou,1 Tao Li,1 Sichuang Tan,6 SipAin Tan,1 Xianzhong Xiao1
1Sepsis Translational Medicine, Key Laboratory of Hunan Province, Xiangya School of Medicine, Central South University, Changsha, Hunan, 410008, People’s Republic of China; 2Department of Pathology, Xiangya School of Medicine, Central South University, Changsha, Hunan, 410008, People’s Republic of China; 3Department of Breast Surgery, Xiangya Hospital, Central South University, Changsha, Hunan, 410008, People’s Republic of China; 4Department of Pathology, Tianjin Medical University Cancer Institute and Hospital, Tianjin, 300060, People’s Republic of China; 5Department of Traditional Chinese Medicine, The Third Xiangya Hospital, Central South University, Changsha, Hunan, 410008, People’s Republic of China; 6Department of Thoracic Surgery, Second Xiangya Hospital, Central South University, Changsha, Hunan, 410008, People’s Republic of China
Correspondence: SipAin Tan Tel +86 – 13975805079
Email [email protected]
Sichuang Tan Tel +86 – 13975806723
Email [email protected]
Purpose: To explore the molecular mechanism of promoting cervical cancer by HSF1 in vivo and in vitro.
Methods: The expression of HSF1 in 110 paraffin-embedded cervical cancer sections of different grades was examined via immunohistochemistry analyses. Expression of HSF1 downstream targets Metadherin (MTDH), VEGF-C and CD31 were studied using immunohistochemistry analyses. HSF1 transcriptional activity in the MTDH promoter region was detected by EMSA, CHIP and luciferase. Cell proliferation and clonality were detected by MTT and clonal formation assay. Cell migration and invasion ability were investigated by scratch analysis and transwell assay. HSF1-mediated tumorigenesis in vivo was examined in xenograft models.
Results: HSF1 expression of cervical cancer cell line was increased compared to normal human cervical tissues. HSF1 enhanced the expression of MTDH, VEGF-C and CD31. HSF1 can combine with MTDH promoter to promote the expression of MTDH. HSF1 enhanced HeLa cell proliferation and clone formation. Furthermore, HSF1 increased HeLa cells migration and invasion in vitro. In the transplanted tumor model, HSF1 inhibited tumor growth in vivo after interference, and reduced the expression of MTDH, VEGF-C and CD31.
Discussion: HSF1 can promote the proliferation, metastasis and invasion of cervical cancer.
Keywords: HSF1, cervical cancer tissue, metastasis, MTDH, transcriptional regulation
Introduction
An epidemic study identified the uterine cervix carcinoma as one of the causes of the highest cancer mortality among women worldwide.1,2 There is an urgent need to identify more biological mark in cervical cancer. Heat shock transcription factor 1 (HSF1) is an important transcription factor for regulating heat shock response or stress response in the body. It can play other biological functions by combining with the heat shock element (HSE) in the promoter regions of other genes to affect their expression. Inducted heat shock proteins (e.g. HSP70) has been proved to be one of the main biological functions of HSF1.3–5 Current research of HSF1 has indicated its overexpression in various human tumors,6,7 however, there is no HSF1 immunohistochemistry expression report in uterine cervix carcinoma. Therefore, a systematic HSF1 expression in clinical samples was carried out. MTDH and VEGF-C expression were further characterized to explore the oncogenic traits of HSF1 in cervical cancer. Acting as a cell surface protein, MTDH is implicated in the oncogenesis and angiogenesis of various human cancers.8 Overexpression of AEG-1 was found correlated with the cervical cancer clinical staging and tumor differentiation. The main cause of MTDH overexpression is up-regulation of transcription.9 Our further clarification proved MTDH overexpression is correlated with HSF1 expression enhancement. Luciferase report assay, chromatin immunoprecipitation assay(CHIP) and electrophoretic mobility shift assay(EMSA) proved that HSF1 can regulate the expression of MTDH at the transcriptional level. Thus, it is concluded that increased HSF1 regulated the metastasis of cervical cancer via increasing MTDH-VEGF-C expression.
Materials and Methods
Human Tissue Specimens
110 paraffin-embedded cervical cancer samples were collected at the Department of Gynecology and Obstetrics of Xiangya Hospital from January 2017 to August, 2017. All specimens had not adopted radiotherapy and chemotherapy before surgery. Clinical and pathological data of the 110 patients with cervical cancer, including age, infiltration, lymph node metastasis, clinical stage and cancer differentiation grade, were collected. The clinical stage and cancer differentiation grade were classified according to the 2009 International Federation of Gynecology and Obstetrics(FIGO) staging system. Before the clinical material was used for research purposes, the Research Ethics Committee of Xiangya Hospital approved this, and that it was conducted in accordance with the Declaration of Helsinki. The ethical code is 2021-KT01.
Plasmids
The siRNA sequence (5′ - CCAAGTACTTCAAGCACAA −3′) matches the 331 to 350 nucleotide of the human HSF1 gene (NM_ 005526), was selected for the construction of HSF1 shRNA plasmid. The HSF1 shRNA and NC plasmids were constructed as described respectively.10 Generation of the pGL3- MTDH promoter constructs was as follows:
5′-AGGTACCGAGCTCTTACGCGTGCCCACGGGCT′TTCCAACTTTTG-3′ (sense), 5′-GCTTACTTAGATCGCAGATCTGCGATCCACGCCGCGCGC-3′ (antisense), the PCR fragment was cloned into the pGL3. The accuracy of the pGL3-MTDH (- 670 to +1) construct was verified by sequencing (data not shown).
Immunohistochemical Analyses
Paraffin sections were dewaxed with 100% xylene and rehydrated with a percentage of ethanol series. Then, remove the wax and rinse again, add 3%H2O2, soak for 10min, then pour out H2O2, add citric acid buffer, cook in microwave oven for 3min (medium heat), cool to room temperature, pour out the citric acid buffer, add serum to block, dilute with HSF1 antibody (1: 250 dilution; Cell Signaling Technology) and place at 4°C shaker for one night. After washing, add the secondary antibody and incubated at room temperature for 2h.The sections were observed with DAB (3, 30-Diaminobenzidine), hematoxylin reverse staining, and the staining reaction was observed under microscope.
CHIP Assays
CHIP analysis was performed according to the Beyotime Biotech (Shanghai, China). After the DNA was precipitated by HSF1 and control antibodies, PCR was performed to detect the HSF1 binding site at position - 0.5 kb within the MTDH promoter (GenBank accession number: NC_ 000008. 11), the PCR primers as follows: 5′-gccaggctggcaactggtaggcacg-3′ (sense) and 5′-ccgttactgtagaccagaagctg-3′ (antisense), and a 424 bp PCR product was produced.
EMSA Analysis
Nuclear protein was extracted from three different HeLa cells according to the manufacturer’s instructions (Light Shift Chemiluminescent EMSA Kit, PIERCE). DNA probes were generated according to the - 450 to - 438 bp of the human MTDH promoter, HRP-labeled oligonucleotides sequences (5′-gagctcaTTCTGGAAGGATCcaactgg-3′), and mutant sequence (5′- tattaacCTCTGCACCAGCTgttagtt-3′). The nucleoprotein extraction and binding reactions were completed as described earlier,9 EMSA was performed. The nucleoprotein extraction and oligonucleotide mixture were separated on a non-denaturing PAGE gel. The strips were examined by enhanced chemiluminescent (ECL) assay kit (Pierce).
Western Blotting
Cell proteins were lysed with RIPA lysates (P0013, Beyotime, China), Ultrasonic was performed for 15s and centrifuged for 10min in a 4°C centrifuge at 12000rpm, protein concentrations were determined by bicinchoninic acid protein assay kit (Pierce, USA), configure the protein sample quantity, heating for 10 min at 95°C. Configuration 10% SDS-PAGE and sample run electrophoresis, sealed in TBST containing 5% skim milk powder for 1.5h, and incubated overnight with primary antibody. After washes with TBST, horseradish peroxidase-conjugated anti-rabbit or anti-mouse IgG was used as the secondary antibody. Filters were probed with primary antibodies after the electrophoresis and transfection: HSF1 (Cell signaling), CD31 (Abcam), MTDH and VEGFC (Bioworld Technology, Inc). Anti-actin antibody was used as internal reference.
In vitro Wound Healing Assay
We seeded the HeLa, HeLa–shHSF1 and HeLa–NC cells in 24-well plates (1 x105 cells/well) for wound healing assay. Edge distance measurement and wound line scratch were performed as described previously.11 Representative photos of cells migrated into the wounds were captured and the length of the scratch was calculated 24hr, 48hr later. Wound closure rate was calculated as: (0hr scratch width − 48hr scratch width)/ 0hr scratch width × 100%.
Τranswell Invasion Assay
A 24-Well microchamber (Corning, New York, USA) with 5-um-pore polycarbonate membrane was used for the experiment. Apply glue to the bottom of the chamber and place in an incubator at 37°C. 6 × 104 cells were placed in serum-free medium, and the lower chamber was filled with 500 mL of media containing 10% FBS. Then, incubation at 37°C for 24h, the cells on the lower surface of the membrane were fixed and stained with 0.2% crystal violet. The cell counts were observed under a microscope, and 5–10 fields were taken at high magnification.
Subcutaneous Tumor Model
BALB/c nude mice four to five-week-old (Shanghai SLAC Laboratory Animal Co. Ltd., China) were used in the nude mouse tumorigenesis assay. Stably transfected cells (3 ×106 cells/mouse) were injected subcutaneously into axilla. Then, 42 days of tumor implantation, the nude mice were sacrificed, then dissected the xenografts, the tumor volume and weight were measured as follows: volume = 0.5 ×a×b2, where a is the longest diameter and b is the transverse diameter. This research project has been approved by the Laboratory Animal Welfare and Ethical Committee of Central South University, in line with the welfare and ethical principles of laboratory animal proposed by the World Organization for Animal Health. The ethical code is 2018sydw077.
Statistical Analyses
All experiments were repeated at least three times. SPSS 17.0 software (SPSS, Inc., Chicago, IL, USA) and GraphPad Prism 6 (GraphPad Software Inc., CA, USA) were used for statistical analyses. Differences between groups were analyzed using the χ2 test, Fishers exact test or one-way analysis of variance, according to which P< 0.05 was considered statistically significant.
Results
Enhancive HSF1 Was Related to Tumor Metastasis, Infiltration, Clinical Stage and Differentiation in Cervical Cancer
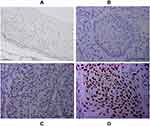
Immunohistochemical analysis of 110 cases of different grades of cervical cancer paraffin section. As indicated in Figure 1, the normal cervical tissue displayed a weak expression while the poorly differentiated ones displayed higher expression of HSF1. HSF1 positive expression was mainly concentrated in the nucleus. 56.4% percent of the sections showed high HSF1 expression, while the remaining 48 cases (43.6%) displayed low HSF1 expression. However, the results showed that HSF1 expression was not obviously associated with patient age. The high HSF1 expression was obviously correlated with clinical stage (p<0.001), cancer differentiation (p<0.001), cancer infiltration (P< 0.01), and lymph node metastasis (p<0.001) (Table 1). Our immunohistochemistry work on the cervical cancer displayed that HSF1 is a potent oncogene, whose high expression is related to cancer infiltration, differentiation and lymph node metastasis of cervical cancer.
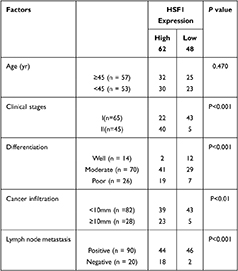 |
Table 1 Correlation Between the Expression of HSF1 and Clinic Pathological Parameters in Cervical Cancer Patients |
VEGF-C and MTDH Displayed Enhanced Expression in Cervical Cancer
Angiogenesis is one of the key factors responsible for cancer infiltration and metastasis. To clarify the biological function of HSF1 played in cervical cancer, CD31, the vessel intensity marker, was found to be positively co-related with the HSF1 expression (Figure 2A and B). VEGF-C has been confirmed to promote angiogenesis, lymph node metastasis, and infiltration, which was also found to be increased in cervical cancer (Figure 2E and F).12 However, there is no putative HSF1 binding site (the DNA binding sequence is GAANNTCC) in – 3000nt 5′-flanking region of VEGF-C and CD31. MTDH, the upstream transcriptional regulatory protein of VEGF-C, was strongly related to lymph node metastasis and infiltration in cervical cancer. Enhanced MTDH was also proved to be positive co-related with HSF1 expression (Figure 2C and D). Further analyses were conducted to verify the relationship between HSF1 and MTDH.
HSF1 Transcriptionally Regulate the MTDH Expression
Bioinformatics analyses identified a highly putative HSE core sequence in the - 450bp ~ - 438bp sequence of the 5′ promoter region of MTDH (Figure 3C). Luciferase report assay was performed to prove the transcriptional control of MTDH by HSF1. As can be seen by Figure 3D, fold increase of luciferase can be detected after the HSF1 expression plasmid and MTDH-pGL3 plasmid were transfected into HeLa cells. EMSA was used to detect whether MTDH could bind to HSF1 in vitro. As shown in Figure 3A, the labeled probe could bind to HSF1, but the cold probe could not bind to HSF1, and the mutant probe could still bind to HSF1, proving the specificity of this binding site. To demonstrate the regulation of MTDH by HSF1 in vivo, we performed the CHIP assay (Figure 3B). DNA fragments were almost undetectable with normal IgG antibodies, whereas DNA fragments were detectable when anti-HSF1 antibodies were used, and transfection of constitutively active HSF1 version obviously increased the number of PCR fragments of the MTDH promoter (Figure 3B). These results suggested that HSF1 could bind with MTDH in vivo and in vitro to participate in its transcriptional regulation.
Knocking Down HSF1 Reduced the Migration and Invasion Capabilities of HeLa
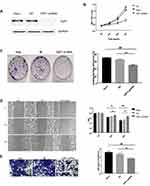
HSF1-shRNA was transfected into HeLa cells, Western blot displayed that the expression of HSF1 in HSF1-shRNA cells was reduced, indicating that HSF1 interference was successful (Figure 4A). MTT assay showed that cell activity of HSF1-shRNA cells decreased after interference compared with cells in HeLa and NC groups (Figure 4B). Cloning experiments showed that the proliferation of HSF1-shRNA cells decreased after interference compared with cells in HeLa and NC (Figure 4C). Transwell and wound closure assays showed that knocking down HSF1 can obviously suppress the invasion and migration of HeLa cells (Figure 4D and E). These results suggested that HSF1 knockout in HeLa cells could significantly decrease the degree of malignancy in vitro.
Knocking Down HSF1 Reduced the Malignancy of HeLa in vivo
The effect of reduced HSF1 expression on HeLa cells in vivo was verified by tumor transplantation experiments. After HSF1 was knocked out, the growth of HeLa cells in vivo was obviously suppressed, and the growth curves of both the untreated group and the NC group at day 48 showed a rapid increase in tumor volume (Figure 5 and B). In addition, HSF1 knockout tumors were obviously smaller in diameter (188.18 ± 20.15 mm3) and weight (0.23 ± 0.091 g) than those of HeLa (889.16 ± 40.94mm3, 0.64 ± 0.12g) and empty vector-transformed tumors (988.18 ± 55.19mm3, 0.73 ± 0.1g) (Figure 5C and D). These data suggest that downregulation of HSF1 can significantly inhibit tumor growth in vivo.
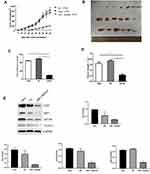 |
Figure 5 Continued. |
HSF1 Depleted Xenograft Tissues Displayed Decreased MTDH and VEGF-C Expression
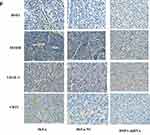
Western blot detected the expression of HSF1, CD31, MTDH and VEGF-C expression in the in vivo xenograft tissue. As indicated in Figure 5E, significant reduction of MTDH, CD31 and VEGF-C were observed in the HSF1 depleted xenograft tissues. MTDH displayed a significant expression reduction as HSF1, which was consistent with previous in vitro experiments. The expressions of MTDH, CD31 and VEGF-C in the transplanted tumor were detected by immunohistochemistry (Figure 5F). Positive staining was observed in both HeLa and HeLa NC groups, while the positive staining was obviously reduced in HSF1-shRNA group. Similarly, the expression of MTDH, CD31 and VEGF-C were downregulated in the HSF1-shRNA group.
Discussion
Cervical cancer is the fourth most common malignancy in women worldwide and is a major global health challenge. Each year, more than half a million women are diagnosed with cervical cancer, and more than 300,000 people worldwide die from the disease.13
The treatment of cervical cancer has developed from using cytotoxic therapy to targeting specific molecular subtypes, but the mortality rate of cervical cancer is still very high, which is attributed to the lack of sensitive early biomarkers on the one hand, and the vulnerability of cervical cancer to invasion and metastasis on the other hand.14 Current research of cervical cancer has identified many diagnostic and disease follow-up markers due to its high mortality,15,16 however, there are still more screening markers of cervical carcinoma need to be clearly established. As an important transcription factor, HSF1 is an endogenous anti-injury mechanism formed in the long-term evolutionary process of the body, which can induce the expression of downstream genes and regulate the stress response by binding with the HSE in the promoter of target genes.3 However, the study of HSF1 in cervical cancer has not been reported. For the first time, HSF1 displayed a significant enhanced expression in cervical cancer via immunohistochemistry. Increased HSF1 was associated with cervical cancer infiltration, lymph node metastasis, clinical stage and cancer differentiation. Previous research proved increased VEGF-C expression accounts for the increased angiogenesis and metastasis in cervical cancer.17,18 The microvessel intensity was found related to HSF1 expression. Increased VEGF-C was detected in cervical cancer, however, there is no transcription binding site in the promoter region of VEGF-C, which abolish the possibility of direct transcriptional control of VEGF-C by HSF1. Acting as a transcriptional protein implicated in the human oncogenesis and angiogenesis of various cancers, MTDH can upregulate the expression of VEGF by activating NF-kB pathway. Metadherin is highly expressed in cervical cancer through activation of ERK /NF-κB pathway to give chemical resistance to cervical cancer cells.19 Statistical analysis identified positive relation between HSF1, MTDH, and VEGF-C protein in cervical cancer sections. MTDH, and VEGF-C also reduced its expression in the in vivo HSF1 depleted HeLa xenograft tissue. Bioinformatics analysis identified putative HSE core sequence in the - 450bp ~ - 438bp sequence of the 5′ promoter region of MTDH. Luciferase report assay, CHIP and EMSA proved the direct binding of HSF1 of the core HSE sequence of MTDH. VEGF can inhibit T lymphocyte infiltration in tumor microenvironment by inhibiting endothelial cell activation induced by NF-kB.20 Gang et al found that MTDH can regulate the expression of VEGF in head and neck squamous cell carcinoma.21 Considering the relationship between MTDH, NF-kB and VEGF-C,22,23 it is proposed that HSF1 regulate the angiogenesis and metastasis of cervical cancer via MTDH-NF-kB-VEGF-C pathway. Current research identified that HSF1 can regulate HSPs, various proteins regulating cellular metabolism and signaling pathways, miRNAs and lncRNAs. HSF1 transcriptome is profoundly different in malignant cells compared with heat shock treated cells.24–26 In the tumorigenesis and progression, HSF1 plays multifaceted roles in cancer via regulating various proteins concerning critical cellular processes, for example, maintain homeostasis, inhibit apoptosis, and promote proliferation.27
For the first time, clinical investigation identified increased HSF1 in cervical cancer, increased HSF1 expression was co-connected with cervical cancer infiltration, lymph node metastasis, clinical stage and cancer differentiation. Knocking down HSF1 successfully reduces the malignancy of HeLa in vivo and in vitro. MTDH was proved to be a target molecule regulated by HSF1 transcription. It is proved that HSF1 acts as an oncogene in cervical cancer via transcriptionally regulating MTDH. HSF1 can be used as a therapeutic molecule via suppressing the expression of MTDH.
Abbreviations
HSF1, heat shock transcription factor 1; MTDH, metadherin; FIGO, Federation of Gynecology and Obstetrics; ChIP, Chromatin immunoprecipitation; EMSA, electrophoretic mobility shift assay; ECL, enhanced chemiluminescent.
Acknowledgments
This work was supported by the grants from National Natural Science Foundation of China (No. 81671895; 81871610; 81471897; 81870071); Natural Science Foundation of Hunan Province, China, (No. 2019JJ40393); Fundamental Research Funds of Central South University for Postgraduate Students (No. 1053320191465).
Author Contributions
All authors made substantial contributions to conception and design, acquisition of data, or analysis and interpretation of data; took part in drafting the article or revising it critically for important intellectual content; agreed to submit to the current journal; gave final approval of the version to be published; and agree to be accountable for all aspects of the work.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Eoh KJ, Nam EJ, Kim SW, et al. Nationwide comparison of surgical and oncologic outcomes in endometrial cancer patients undergoing robotic, laparoscopic, and open surgery: a Population-Based Cohort Study. Cancer Res Treat. 2020. doi:10.4143/crt.2020.802
2. Jiang X, Tang H, Chen T. Epidemiology of gynecologic cancers in China. J Gynecol Oncol. 2018;29(1):e7. doi:10.3802/jgo.2018.29.e7
3. Gomez-Pastor R, Burchfiel ET, Thiele DJ. Regulation of heat shock transcription factors and their roles in physiology and disease. Nat Rev Mol Cell Biol. 2018;19(1):4–19. doi:10.1038/nrm.2017.73
4. Li J, Labbadia J, Morimoto RI. Rethinking HSF1 in stress, development, and organismal health. Trends Cell Biol. 2017;27(12):895–905. doi:10.1016/j.tcb.2017.08.002
5. Paszek A, Kardynska M, Bagnall J, et al. Heat shock response regulates stimulus-specificity and sensitivity of the pro-inflammatory NF-kappaB signalling. Cell Commun Signal. 2020;18(1):77. doi:10.1186/s12964-020-00583-0
6. Jin X, Moskophidis D, Mivechi NF. Heat shock transcription factor 1 is a key determinant of HCC development by regulating hepatic steatosis and metabolic syndrome. Cell Metab. 2011;14(1):91–103. doi:10.1016/j.cmet.2011.03.025
7. Tsukao Y, Yamasaki M, Miyazaki Y, et al. Overexpression of heat-shock factor 1 is associated with a poor prognosis in esophageal squamous cell carcinoma. Oncol Lett. 2017;13(3):1819–1825. doi:10.3892/ol.2017.5637
8. Liang Y, Fu D, Hu G. Metadherin: an emerging key regulator of the malignant progression of multiple cancers. Thorac Cancer. 2011;2(4):143–148. doi:10.1111/j.1759-7714.2011.00064.x
9. Jayamohan S, Kannan M, Moorthy RK, et al. Dysregulation of miR-375/AEG-1 axis by human papillomavirus 16/18-E6/E7 promotes cellular proliferation, migration, and invasion in cervical cancer. Front Oncol. 2019;9:847. doi:10.3389/fonc.2019.00847
10. Tan J, Tan S, Zheng H, et al. HSF1 functions as a transcription regulator for Dp71 expression. Cell Stress Chaperones. 2015;20(2):371–379. doi:10.1007/s12192-014-0558-8
11. Tan S, Tan J, Tan S, et al. Decreased Dp71 expression is associated with gastric adenocarcinoma prognosis. Oncotarget. 2016;7(33):53702–53711. doi:10.18632/oncotarget.10724
12. Tan S, Tan S, Chen Z, et al. Knocking down Dp71 expression in A549 cells reduces its malignancy in vivo and in vitro. Cancer Invest. 2016;34(1):16–25. doi:10.3109/07357907.2015.1084002
13. DeMei M, XiangXin L, YongPing X, YongXia Y, YunHai Y, Lin Z. Vascular endothelial growth factor C expression is closely correlated with lymph node recurrence and poor prognosis in patients with early stage cervical cancer. J Int Med Res. 2013;41(5):1541–1549. doi:10.1177/0300060513493038
14. Cohen PA, Jhingran A, Oaknin A, Denny L. Cervical cancer. Lancet. 2019;393(10167):169–182. doi:10.1016/S0140-6736(18)32470-X
15. Zacharakis N, Chinnasamy H, Black M, et al. Immune recognition of somatic mutations leading to complete durable regression in metastatic breast cancer. Nat Med. 2018;24(6):724–730. doi:10.1038/s41591-018-0040-8
16. Valenti G, Vitale SG, Tropea A, Biondi A, Lagana AS. Tumor markers of uterine cervical cancer: a new scenario to guide surgical practice? Updates Surg. 2017;69(4):441–449. doi:10.1007/s13304-017-0491-3
17. Abera GB, Abebe SM, Werku AG. Demand for cervical cancer screening in Tigray Region of Ethiopia in 2018: a Community-Based Cross-Sectional Study. Int J Womens Health. 2020;12:795–804. doi:10.2147/IJWH.S255548
18. Tang J, Yang Z, Wang Z, et al. Foxp3 is correlated with VEGF-C expression and lymphangiogenesis in cervical cancer. World J Surg Oncol. 2017;15(1):173. doi:10.1186/s12957-017-1221-5
19. He M, Cheng Y, Li W, et al. Vascular endothelial growth factor C promotes cervical cancer metastasis via up-regulation and activation of RhoA/ROCK-2/moesin cascade. BMC Cancer. 2010;10:170. doi:10.1186/1471-2407-10-170
20. Zhang J, Zhang Y, Liu S, et al. Metadherin confers chemoresistance of cervical cancer cells by inducing autophagy and activating ERK/NF-κB pathway. Tumor Biol. 2013;34(4):2433–2440. doi:10.1007/s13277-013-0794-z
21. Huang H, Langenkamp E, Georganaki M, et al. VEGF suppresses T-lymphocyte infiltration in the tumor microenvironment through inhibition of NF-kappaB-induced endothelial activation. FASEB J. 2015;29(1):227–238. doi:10.1096/fj.14-250985
22. Zhu GC, Yu CY, She L, et al. Metadherin regulation of vascular endothelial growth factor expression is dependent upon the PI3K/Akt pathway in squamous cell carcinoma of the head and neck. Medicine (Baltimore). 2015;94(6):e502. doi:10.1097/MD.0000000000000502
23. Yu JQ, Zhou Q, Zhu H, Zheng FY, Chen ZW. Overexpression of astrocyte elevated gene-1 (AEG-1) in cervical cancer and its correlation with angiogenesis. Asian Pac J Cancer Prev. 2015;16(6):2277–2281. doi:10.7314/APJCP.2015.16.6.2277
24. Li WF, Dai H, Ou Q, Zuo GQ, Liu CA. Overexpression of microRNA-30a-5p inhibits liver cancer cell proliferation and induces apoptosis by targeting MTDH/PTEN/AKT pathway. Tumour Biol. 2016;37(5):5885–5895. doi:10.1007/s13277-015-4456-1
25. Li Y, Xu D, Bao C, et al. MicroRNA-135b, a HSF1 target, promotes tumor invasion and metastasis by regulating RECK and EVI5 in hepatocellular carcinoma. Oncotarget. 2015;6(4).
26. Puustinen MC, Sistonen L. Molecular mechanisms of heat shock factors in cancer. Cells. 2020;9(5). doi:10.3390/cells9051202
27. Mendillo ML, Santagata S, Koeva M, et al. HSF1 drives a transcriptional program distinct from heat shock to support highly malignant human cancers. Cell. 2012;150(3):549–562. doi:10.1016/j.cell.2012.06.031
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.