Back to Journals » Infection and Drug Resistance » Volume 14
In vitro and in vivo Effect of Antimicrobial Agent Combinations Against Carbapenem-Resistant Klebsiella pneumoniae with Different Resistance Mechanisms in China
Authors Liu E, Jia P, Li X, Zhou M , Kudinha T , Wu C, Xu Y, Yang Q
Received 16 November 2020
Accepted for publication 7 January 2021
Published 5 March 2021 Volume 2021:14 Pages 917—928
DOI https://doi.org/10.2147/IDR.S292431
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Prof. Dr. Héctor Mora-Montes
Enbo Liu, 1,* Peiyao Jia, 1, 2,* Xue Li, 1, 3,* Menglan Zhou, 1, 2 Timothy Kudinha, 4, 5 Chuncai Wu, 1 Yingchun Xu, 1 Qiwen Yang 1
1Department of Clinical Laboratory, State Key Laboratory of Complex Severe and Rare Diseases, Peking Union Medical College Hospital, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing, 100730, People’s Republic of China; 2Graduate School, Peking Union Medical College, Chinese Academy of Medical Sciences, Beijing, People’s Republic of China; 3Department of Clinical Laboratory, Beijing Anzhen Hospital, Capital Medical University, Beijing, People’s Republic of China; 4School of Biomedical Sciences, Charles Sturt University, Orange, 2800, Australia; 5Pathology West, NSW Health Pathology, Orange, 2800, Australia
*These authors contributed equally to this work
Correspondence: Qiwen Yang Tel +86-10-6915-9763
Fax +86-10-6915-9766
Email [email protected]
Objective: This study aimed to evaluate the in vitro and in vivo effects of different combinations of antimicrobial agents against carbapenemase-producing and non-producing Klebsiella pneumoniae from China.
Methods: A checkerboard assay of meropenem (MEM), amikacin (AK), tigecycline (TGC), colistin (COL) and their combinations was carried out against 58 clinical carbapenem-resistant K. pneumoniae (CRKp) isolates, including 11 carbapenemase-non-producing K. pneumoniae isolates and 21 isolates producing KPC-2 enzyme, 11 NDM-1, 13 IMP, one VIM-1 and one OXA-48. The checkerboard assay was analyzed by the fractional inhibitory concentration index (FICI). A time–kill assay and Galleria mellonella infection model were conducted to evaluate the in vitro and in vivo effects of the four drugs alone and in combination.
Results: In the checkerboard assay, TGC+AK and MEM+AK combinations showed the highest synergistic effect against KPC-2 and NDM-1 carbapenemase-producing isolates, with synergy+partial synergy (defined as FICI < 1) rates of 76.2% and 71.4% against KPC-2 producers, and 54.5% and 81.8% against NDM-1 producers. TGC+AK and MEM+COL combinations showed the highest rate of synergistic effect against IMP-producing isolates. Against carbapenemase-non-producing isolates, TGC+COL and TGC+AK combinations showed the highest rate of synergy effect (63.6% and 54.5%). MEM+AK showed a synergistic effect against one VIM-1 producer (FICI=0.31) and an additivite effect (FICI=1) against one OXA-48 producer. In the time–kill assay, COL+AK, COL+TGC, COL+MEM and AK+TGC showed good synergistic effects against the KPC-2-producing isolate D16. COL+MEM and COL+TGC combinations showed good effects against the NDM-1-producing isolate L13 and IMP-4-producing isolate L34. Against the carbapenemase-non-producing isolate Y105, MEM+TGC and COL+AK showed high synergistic effects, with log 10CFU/mL decreases of 6.2 and 5.5 compared to the most active single drug. In the G. mellonella survival assay, MEM-based combinations had relatively high survival rates, especially when combined with colistin, against KPC-2 producers (90% survival rate) and with amikacin against metallo-beta-lactamase producers (95– 100% survival rate).
Conclusion: Our study suggests that different antimicrobial agent combinations should be considered against CRKp infections with different resistance mechanisms.
Keywords: resistance mechanisms, time–kill curve assay, carbapenem-resistant Klebsiella pneumoniae, CRKP, antimicrobial agent combinations, Galleria mellonella infection model
Introduction
Carbapenem-resistant Enterobacteriaceae (CRE), including carbapenem-resistant Klebsiella pneumoniae (CRKp), are classified by the US Centers for Disease Control and Prevention (CDC) as urgent antimicrobial-resistant bacteria.1 CRKp has been increasingly reported in the past decade.2 The production of carbapenemases, especially KPC-2, NDM-1 and IMP-type carbapenemases, is the most important resistance mechanism of CRKp in China.3 Some studies have also reported that the loss of outer membrane proteins, such as OmpK35 and OmpK36, could contribute to carbapenem resistance in CRKp.
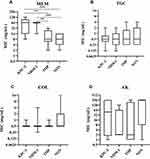 |
Figure 1 The difference in MIC values between different types of carbapenemase. (A) Meropenem (MEM); (B) tigecycline (TGC); (C) colistin (COL); (D) amikacin (AK). Notes: ***p<0.001; **p<0.01. |
Infections with CRKp bacteria, especially coexisting virulence factors, are associated with high mortality rates and the treatments are extremely limited.4,5 Antimicrobial combination therapy is reported as an effective method against CRE infection.6–9 Although some studies have evaluated the effects of combination against KPC-producing isolates,10–12 it is still not clear whether the different types of carbapenemase affect the efficacy of combination therapy using in vitro and in vivo assays.
In order to identify the best combination regimen that could provide alternative treatments for carbapenemase-producing K. pneumoniae, we investigated the effectiveness of single drug and combinations of meropenem, colistin, amikacin and tigecycline against CRKp with different resistance mechanisms (carbapenemase-producing K. pneumoniae and carbapenemase-non-producing K. pneumoniae) using an in vitro checkerboard test and time–kill assays, and an in vivo Galleria mellonella infection model.13,14
Materials and Methods
Bacterial Isolates
A total of 58 non-repetitive carbapenem-resistant clinical K. pneumoniae isolates were collected from 11 teaching hospitals in China from 2015 to 2018. Carbapenemase genes were detected by polymerase chain reaction and sequenced as previously described.15–17 All strains were identified by the Vitek 2 system (bioMérieux, Marcy-l’Étoile, France), and MALDI-TOF MS apparatus (Bruker Biotyper; Bruker Daltonik, Bremen, Germany) (Supplementary Table 1). All the K. pneumoniae isolates were further characterized by multilocus sequence typing (MLST) (http://bigsdb.pasteur.fr/klebsiella/klebsiella.html). We prefer to collect isolates with different ST types to avoid outbreak strains.
Antimicrobial Susceptibility Testing
Meropenem (MEM), amikacin (AK) and colistin (COL) were obtained from the National Institute for the Control of Pharmaceutical and Biological Products (Beijing, China). Tigecycline (TGC) was purchased from Sigma-Aldrich (Saint Louis, MO, USA). Drugs were diluted in sterile cation-adjusted Mueller–Hinton broth (BD BBL, Sparks, MD, USA) and the concentration ranges of each drug were as follows: meropenem (0.25–32 mg/L), amikacin (0.5–64 mg/L), colistin (0.06–8 mg/L), tigecycline (0.03–4 mg/L). The bacterial inoculum was 5×105 CFU/mL. Escherichia coli strain ATCC 25922 and P. aeruginosa ATCC 27853 were used as the quality control strains in each batch.
The minimum inhibitory concentrations (MICs) of the four antimicrobials were determined by the broth microdilution method according to CLSI recommendations.18 CLSI criteria were used to interpret the antimicrobial susceptibility of meropenem, amikacin and colistin, whereas the FDA breakpoint was used for tigecycline.
Checkerboard Test
Six different combinations of antimicrobials were selected and tested by the checkerboard test: MEM plus AK, MEM plus COL, MEM plus TGC, AK plus COL, AK plus TGC and COL plus TGC. The information on the six isolates is shown in Table 1. The MICs of drug combinations were determined after incubation at 35°C for 18–24 h in ambient air.
 |
Table 1 Isolates Used in the Time–Kill Assay and G. mellonella Survival Assay |
The interaction between antimicrobial agents was determined based on the calculated fractional inhibitory concentration index (FICI). The FIC of each drug was calculated as a ratio of the MIC of drug A (or B) when used in the combination (combo) and the MIC of drug A (or B) when tested alone, according to the following formula: FICI=FICA+FICB=MIC(A–combo)/MIC(A–alone)+MIC(B–combo)/MIC(B–alone).19 The results were interpreted as follows: synergism, FICI ≤0.5; partial synergism, 0.5<FICI<1; additivity, FICI=1; indifference, 1<FICI<4; and antagonism, FICI ≥4.20
Time–Kill Assay
The time–kill activities of antimicrobial drugs against six clinical isolates were assessed. The selection criteria were based on different resistance mechanisms (Table 1). Drug concentrations were selected based on clinically achievable serum levels of each drug as defined by a literature review.21–25 All drugs were tested alone and in combination at fixed concentrations of meropenem 8 mg/L, colistin 2 mg/L, tigecycline 0.25 mg/L and amikacin 16 mg/L. Experiments were carried out with a starting inoculum of 5×105 CFU/mL. Tubes were incubated at 35°C with shaking. Samples were taken out at 0, 2, 4, 8, 16 and 24 h, serially diluted, plated and counted. The lower limit of accurately quantifiable CFU using was 1 log10 CFU/mL of viable bacteria per mL.25
Synergy was defined as ≥2 log10 CFU/mL decrease at 24 h for the antimicrobial combination compared with the most active single agent. No interaction was defined as <2 log10 CFU/mL increase or decrease at 24 h for the drug combination in comparison with the most active antibiotic alone. Antagonism was defined as ≥2 log10 CFU/mL increase between the combination and the most active single drug alone. Bactericidal activities of single drug or combinations were defined as a decrease of ≥3 log10CFU/mL compared with the untreated control at the start of each assay from the original inocula, whereas bacteriostatic activity was defined as <3 log10 CFU/mL decrease.26
In vivo G. mellonella Survival Assay
The in vivo G. mellonella survival assay was conducted as previously described.22,27 Six clinical K. pneumoniae isolates with different resistance mechanisms were selected (Table 1). Standardized larvae were purchased from KaideRuixin Co. (Tianjin, China). The larvae are 5–6 instars, about 2–3 cm long, weighing about 250 mg, and have good activity and a creamy color. Caterpillars were inoculated with 10 μL of K. pneumoniae at a concentration of 1×107–1×108 CFU/mL (optimal infection dose of each strain causing approximately 80% lethality within 3 days) into the last left proleg using a 50 μL Hamilton syringe. Antibiotics were given as 10 μL injections either alone or in combination, into another proleg within half an hour after infection. The following dosages were based on human doses: meropenem 15 mg/kg, colistin 2.5 mg/kg, tigecycline 1 mg/kg, amikacin 15 mg/kg.28,29 The larvae were observed for survival every 12 h for 3 days. Each treatment group had 20 caterpillars. Mock-inoculated (sterile saline) larvae were used as controls. Experiments were performed in triplicate and the results of any experiment in which two or more larvae died in any control group were discarded. The time–kill assay and G. mellonella survival assay were performed in triplicate.
 |
Table 2 Susceptibility of 58 Clinical CRKps to Four Antimicrobial Agents |
Statistics
The differences in MIC values between different type of carbapenemase were estimated by Kruskal-Wallis test for one-way analysis of variance (ANOVA) followed by Dunn's multiple comparisons tests. Statistical significance was defined for an overall error at the 0.05 level (95% confidence interval). Survival rates were calculated and represented using GraphPad Prism 7 (GraphPad, La Jolla, USA). The log-rank (Mantel–Cox) test was used to compare survival rates between different treatment groups. P values of <0.05 were considered statistically significant.
Results
Bacterial Isolates
From 2015 to 2018, a total of 58 non-repetitive carbapenem-resistant clinical K. pneumoniae isolates were collected from 11 teaching hospitals in China. The CRKP isolates were recovered from various clinical specimens, including sputum (n=17), blood (n=11), urine (n=10), abscesses (n=6), drainage (n=6), bile (n=4), bronchoalveolar lavage fluid (n=3) and cerebrospinal fluid (n=1).
Antimicrobial Susceptibility of Tested Strains
All strains were non-susceptible to meropenem (MIC ranged from 2 to >256 mg/L). Twenty-six isolates (44.8%) were resistant to amikacin (MIC ranged from 1 to >256 mg/L), whereas only four isolates (6.9%) were resistant to colistin and five isolates were non-susceptible to tigecycline. The 50% MIC (MIC50) values of MEM, TGC, COL and AK were 16, 0.5, 0.5 and 32 mg/L, respectively. The 90% MIC (MIC90) values of MEM, TGC, COL and AK were 64, 2, 1 and >256 mg/L, respectively (Table 2).
KPC-2-producing isolates and NDM-producing isolates were distributed with higher MIC values for meropenem than IMP-producing isolates and carbapenemase-non-producing isolates, although there was no difference in susceptibility (Figure 1).
Resistance Mechanisms of Tested Isolates
Fifty-eight clinical isolates were screened for their resistance mechanisms and it was confirmed that 21 (36.2%) isolates carried blaKPC-2, 11 (19.0%) isolates carried blaNDM-1, 13 (22.4%) isolates carried blaIMP (eight isolates carried blaIMP-4, four isolates carried blaIMP-8 and one isolate carried blaIMP-26), one (1.7%) isolate carried blaVIM-1, one (1.7%) isolate carried blaOXA-48 and 11 (19.0%) isolates carried no carbapenemase genes but had porin (ompK35 and/or ompK36) loss.
MLST
Among the 58 clinical CRKP isolates, the ST types of enrolled isolates were diverse (35 distinct STs were observed). ST11 (n=9, 16%) was the most predominant clone, followed by ST17 (n=4, 7%), ST48 (n=3, 5%), ST4928 (n=3), ST4930 (n=3) and another 30 STs (n=36).
Twenty-one KPC-2-producing isolates belonged to 17 ST types, while 11 NDM producers belonged to six ST types, 13 IMP producers belonged to eight ST types and 11 carbapenemase non-producers belonged to seven ST types. One VIM-1 producer was ST54 type and one OXA-48-producer was ST353 type.
Checkuerboard Test
The results of the checkerboard synergy are shown in Table 3. Detailed FICI values are shown in Supplementary Table 2. Different combinations of antimicrobial agents had different effects on various carbapenemase types.
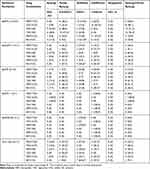 |
Table 3 In vitro Combination Effect of Different Regimens Against CRKps with Different Resistance Mechanisms Using the Checkerboard Assay |
Against KPC-2 or NDM-1 carbapenemase-producing isolates, TGC+AK and MEM+AK combinations showed the highest synergistic effect. TGC+AK combination showed a synergy+partial synergy rate of 76.2% against KPC-2-producing isolates and 54.5% against NDM-1-producing isolates. The MEM+AK regimen showed a synergy+partial synergy rate of 71.4% against KPC-2-producing isolates and 81.8% against NDM-1-producing isolates. The other four combination regimens mostly exhibited additive or indifferent effects. No combination exhibited an antagonistic effect.
Against IMP-carbapenemase-producing isolates, TGC+AK and MEM+COL combinations showed the highest rate of synergy effect, with both synergy+partial synergy rate of 61.5%, followed by MEM+AK combination (46.2%), AK+COL (46.2%), TGC+COL (30.8%) and MEM+TGC (15.4%). No combination exhibited an antagonistic effect. Against VIM-1-producing isolates, TGC+COL (FICI=0.5) and MEM+AK (FICI=0.31) combinations showed synergistic effects, while the other four combinations showed indifference. Against OXA-48-producing isolated, two combinations, MEM+TGC and MEM+AK, showed additivity (FICI=1), while the others showed indifference. Against carbapenemase-non-producing isolates, TGC+COL and TGC+AK combinations showed the highest rate of synergy effect, with synergy+partial synergy rates of 63.6% and 54.5%, followed by MEM+TGC combination (45.5%), MEM+AK (45.5%), MEM+COL (36.4%) and AK+COL (9.1%). No combination exhibited an antagonistic effect.
Time–Kill Assay
The time–kill assays for six clinical isolates treated with antimicrobials at clinically achievable serum levels are shown in Figure 2.
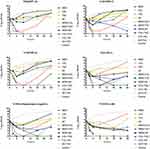 |
Figure 2 In vitro time–kill assay using meropenem (MEM), colistin (COL), tigecycline (TGC) and amikacin (AK), either alone or in combination, against six CRKps with different resistance mechanisms |
Against KPC-2-producing isolate D16, colistin-based antimicrobial combinations, including COL+AK, COL+TGC and COL+MEM, showed a highly synergistic effect, with a log10 CFU/mL decrease of 7.4 at 24 h compared to the corresponding most active single drug. AK+TGC showed a 1.8 log10 CFU/mL decrease compared to single amikacin.
Against NDM-1-producing isolate L13, MEM+AK, COL+MEM and COL+TGC showed the highest synergistic effect, with log10 CFU/mL decreases of 6.5, 6.3 and 6.0 separately compared to the most active single drug.
Against IMP-4-producing isolate L34, MEM+COL and COL+TGC showed synergistic effects, with log10 CFU/mL decreases of 6.3 and 2.1 compared to the most active single drug.
Against VIM-1-producing isolate P5, COL+AK, MEM+TGC and AK+TGC showed synergistic effects, with log10 CFU/mL decreases of 6.4, 4.56 and 3.1 compared to the most active single drug.
Against OXA-48-producing isolate P13, COL+TGC and MEM+AK showed synergistic effects, with log10CFU/mL decreases of 4.2 and 2.1 compared to the most active single drug.
Against carbapenemase-non-producing isolate Y105, MEM+TGC and COL+AK showed highly synergistic effects, with log10 CFU/mL decreases of 6.2 and 5.5 compared to the most active single drug. COL+TGC, MEM+AK and MEM+COL also showed slight synergistic effects.
In vivo Survival Tests in Larvae
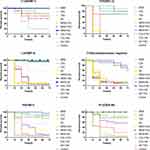
Galleria mellonella larvae infected with six clinical strains were used to evaluate the in vivo interactions of drug combinations. Survival rates were used as the primary index to assess the in vivo interactions (Figure 3).
Against all the five isolates with different types of carbapenemases (KPC-2, NDM-1, IMP-4, VIM-1, OXA-48), MEM+AK, MEM+TGC and MEM+COL showed high survival rates (ranging from 70% to 100%) after 72 h of inoculation, compared to the control. And AK+TGC showed a high survival rate against IMP-4 (60%), VIM-1 (60%) and OXA-48 (85%). Against carbapenemase-non-producing isolate Y105, MEM+AK showed the highest survival rate (85%), followed by the MEM+TGC group (60%).
Against KPC-2-producing isolate D16, the control group without antimicrobial agents indicated that this strain a hypervirulent one. The MEM+COL group showed the highest survival rate (90%) after 72 h of inoculation, followed by the MEM+AK group (80%) and MEM+TGC group (70%). COL+TGC, COL+AK and AK+TGC groups showed inferior survival rates.
Against NDM-1-producing isolate L13, the MEM+TGC group and MEM+AK group showed the highest survival rate (100%) after 72 h of inoculation, followed by the MEM+COL group (80%).
Against IMP-4-producing isolate L34, the MEM+AK and MEM+COL group showed the highest survival rate (100%) after 72 h of inoculation, followed by the MEM+TGC group (95%) and AK+TGC group (60%).
Against VIM-1-producing isolate P5, the MEM+AK and MEM+TGC groups showed the highest survival rates (95% and 90% separately) after 72 h of inoculation, followed by the MEM+COL group (85%) and AK+TGC group (60%).
Against OXA-48-producing isolate P13, the MEM+AK and MEM+TGC groups showed the highest survival rate (100%) after 72 h of inoculation, followed by the AK+TGC group (85%) and MEM+COL group (80%).
Against carbapenemase-non-producing isolate Y105, MEM+AK showed the highest survival rate (85%) after 72 h of inoculation, followed by the MEM+TGC group (60%).
Discussion
Infections caused by CRKp are severe and associated with limited treatment options; therefore, some researchers are looking for new future therapeutic weapons against strains of multidrug-resistant K. pneumoniae, such as essential oils from parts of or whole plants, poma inhibitor drugs and combinations of different agents.30–33 Among them, combined antimicrobial therapy is increasingly used as the first-line treatment for CRKp to reduce the dosage of single antimicrobial agents,34 and is associated with higher survival.35 Although many studies have reported antimicrobial agent combinations against CRKP, there has been little research into therapy against CRKPs with different resistance mechanisms. In this study, we evaluated the synergistic effect of different combinations of clinically important antimicrobials used for CRKp infections by the checkerboard combination test, time–kill test and in vivo G. mellonella larvae infection model.
KPC-type carbapenemase is the most common transmissible class A carbapenemase in Enterobacteriaceae worldwide, while KPC-2 is common in China. In a report from the China CRE Network, KPC-2 enzyme was the most common carbapenemase and was identified in 78/155 isolates.36 KPC carbapenemases are capable of hydrolyzing all β-lactams and always lead to high-level resistance to several antimicrobials. In our study, TGC+AK and MEM+AK combinations showed the highest synergistic effect against KPC-2-producing isolates (synergy+partial synergy rate of 76.2% and 71.4%), which is in agreement with previous studies.37,38 At the same time, against selected isolate D16, we also found a synergistic effect of the COL+AK and COL+MEM combinations in the checkerboard test, which is also consistent with the time–kill assay and G. mellonella model. In our study, colistin and tigecycline showed high antimicrobial susceptibility against all CRKP isolates (93.1% and 91.3%, respectively), which was consistent with the high susceptibility rates of TGC (88.6%) and colistin (73.9%) to CRE in another survey.39 However, inferior clinical outcomes and colistin resistance with colistin monotherapy were frequently observed in patients infected with KPC-producing K. pneumoniae.40,41 Qureshi et al’s study reported that combination therapy with colistin-polymyxin B or tigecycline and a carbapenem improved survival in bacteremia due to KPC-producing K. pneumoniae.40 A clinical report published in 2019 showed that a TGC and MEM combination therapy failed in an infection caused by KPC-2-producing K. pneumoniae.42 In this study, apart from the colistin and meropenem combination, AK combined with TGC or MEM showed in vitro synergistic effects as well, which may give physicians more therapeutic options.
The class B metallo-beta-lactamases are a complex group of enzymes that can hydrolyze all β-lactams except for the monobactams, including carbapenems, and are not inhibited by commercially available β-lactamase inhibitors. Notable transmissible MBL genes in Enterobacteriaceae include IMP-type, VIM-type and NDM-type.43
In our study, NDM-1-producing strains generally showed moderate to high levels of meropenem resistance (MICs of 16–64 mg/L). We found that amikacin showed a low resistance rate to NDM-1 producers, which was different from reports from India and the UK.36,44 Regarding the best combination evaluation, we considered that the MEM+AK combination showed good in vitro and in vivo effects.
Klebsiella pneumoniae isolates producing IMP-type and VIM-type enzymes are rare in China. The 11 IMP-producing strains in this study showed low-level resistance to meropenem, in agreement with a previous study.45 Against IMP-carbapenem-producing isolates, TGC+AK and MEM+COL combinations in the checkerboard test showed the highest rate of synergy effect, with a synergy+partial synergy rate of 61.5%. MEM+COL also showed good effects in the time–kill model and G. mellonella model.
Only one VIM-1-producing strain was enrolled in this study because this enzyme is very rare in K. pneumoniae in China. TGC+COL and MEM+AK combinations showed synergistic effects, while MEM+AK combination also had a good effect in the time–kill model and G. mellonella model.
OXA-48 enzymes hydrolyze penicillins at a high level and carbapenems at a low level, while sparing extended-spectrum cephalosporins. In China, OXA-48-producing K. pneumoniae strains are very rare. In our study, only one OXA-48-producing isolate was enrolled. MEM+TGC and MEM+AK showed additivite effects in the checkerboard test. MEM+AK showed good synergistic effects in the time–kill test and the G. mellonella model.
In addition to carbapenemase production, porin loss also plays an important role in carbapenem resistance. In K. pneumoniae isolates, the carbapenemase-non-producing CRKp always lack their OmpK35 and OmpK36 porins. In our study, TGC+COL and TGC+AK combinations showed the highest rate of synergy effect, with synergy+partial synergy rates of 63.6% and 54.5%, which indicates that a tigecycline-based regimen may be effective against this type of CRKp.
In this study, we found that MEM+AK or MEM+COL combination always showed synergistic effects against CRKps, even though these strains had a low or high level of resistance to meropenem. Previous studies have indicated the mechanisms by which meropenem disrupts the formation of cell walls and helps amikacin to act on the bacteria. Carbapenem is a β-lactam antibiotic which binds to penicillin-binding proteins and prevents peptidoglycan synthesis, leading to lytic cell death.46 Amikacin is an aminoglycoside antibiotic due to its polycationic nature, whereby aminoglycosides first bind to the anionic compounds found on the bacterial surface, then aberrant cytoplasmic membrane proteins cause damage to the integrity of the cytoplasmic membrane, facilitating the entry of aminoglycoside molecules in abundant quantities.47 In our study, we also found a good effect of tigecycline plus amikacin, which may be useful in clinical guidance in the selection of combination drugs.
Conclusion
In summary, we have determined superior antimicrobial agent combinations against CRKp isolates by different resistance mechanisms. The combinations of MEM+AK, MEM+COL and TGC+AK showed synergistic effects and could provide alternative treatments for carbapenemase-producing K. pneumoniae, while TGC+COL performed better against carbapenemase-non-producing K. pneumoniae, which will be helpful in clinical settings where different CRKps are encountered. However, additional clinical studies are required to confirm the efficacy of these combinations.
Ethics Approval and Consent to Participate
The protocol has been reviewed by the human research ethics committee of the Institutional Review Board (IRB) of the Peking Union Medical College Hospital (Ethics Approval Number: S-K1167), and since all bacterial strains were from residual samples used in clinical diagnosis, it was determined that they met the criteria for exemption. This project did not involve any patient information, nor did it affect the normal diagnosis and treatment of patients. After consultation with the IRB, formal ethical approval was reviewed and waived, and written patient consent was not required.
Author Contributions
All authors made substantial contributions to conception and design, acquisition of data, or analysis and interpretation of data; took part in drafting the article or revising it critically for important intellectual content; agreed to submit to the current journal; gave final approval of the version to be published; and agree to be accountable for all aspects of the work.
Funding
This study was supported by Pfizer Inc. Further support was provided by the National Natural Science Foundation of China (82072318), National Key Research and Development Program of China (2018YFE0101800, 2018YFC1200100, 2018YFC1200105), Chinese Academy of Medical Sciences (CAMS) Initiative for Innovative Medicine (Grant No. 2016-I2M-3-014). This work is supported by Beijing Key Clinical Specialty for Laboratory Medicine - Excellent Project (No. ZK201000).
Disclosure
The authors report no conflicts of interest in this work. The funders had no role in the study design, collection, and analysis of data, interpretation of results, or preparation of the manuscript.
References
1. Centres for Disease Control and Prevention (US). Antibiotic Resistance Threats in the United States. Centers for Disease Control and Prevention; 2013.
2. Nordmann P, Naas T, Poirel L. Global spread of Carbapenemase-producing Enterobacteriaceae. Emerg Infect Dis. 2011;17(10):1791–1798. doi:10.3201/eid1710.110655
3. Zhang R, Liu L, Zhou H, et al. Nationwide surveillance of clinical Carbapenem-resistant Enterobacteriaceae (CRE) strains in China. EBioMedicine. 2017;19:98–106. doi:10.1016/j.ebiom.2017.04.032
4. Munoz-Price LS, Poirel L, Bonomo RA, et al. Clinical epidemiology of the global expansion of Klebsiella pneumoniae carbapenemases. Lancet Infect Dis. 2013;13(9):785–796. doi:10.1016/S1473-3099(13)70190-7
5. Fasciana T, Gentile B, Aquilina M, et al. Co-existence of virulence factors and antibiotic resistance in new Klebsiella pneumoniae clones emerging in south of Italy. BMC Infect Dis. 2019;19(1):928. doi:10.1186/s12879-019-4565-3
6. Wang X, Wang Q, Cao B, et al. Retrospective observational study from a chinese network of the impact of combination therapy versus monotherapy on mortality from Carbapenem-resistant Enterobacteriaceae Bacteremia. Antimicrob Agents Chemother. 2019;63(1).
7. Tumbarello M, Viale P, Viscoli C, et al. Predictors of mortality in bloodstream infections caused by Klebsiella pneumoniae carbapenemase-producing K. pneumoniae: importance of combination therapy. Clin Infect Dis. 2012;55(7):943–950. doi:10.1093/cid/cis588
8. Daikos GL, Tsaousi S, Tzouvelekis LS, et al. Carbapenemase-producing Klebsiella pneumoniae bloodstream infections: lowering mortality by antibiotic combination schemes and the role of carbapenems. Antimicrob Agents Chemother. 2014;58(4):2322–2328. doi:10.1128/AAC.02166-13
9. Falagas ME, Lourida P, Poulikakos P, Rafailidis PI, Tansarli GS. Antibiotic treatment of infections due to carbapenem-resistant Enterobacteriaceae: systematic evaluation of the available evidence. Antimicrob Agents Chemother. 2014;58(2):654–663. doi:10.1128/AAC.01222-13
10. Nath S, Moussavi F, Abraham D, Landman D, Quale J. In vitro and in vivo activity of single and dual antimicrobial agents against KPC-producing Klebsiella pneumoniae. J Antimicrob Chemother. 2017;73(2):431–436. doi:10.1093/jac/dkx419
11. Lawandi A, Leite G, Cheng MP, Lefebvre B, Longtin J, Lee TC. In vitro synergy of β-lactam combinations against KPC-producing Klebsiella pneumoniae strains. J Antimicrob Chemother. 2019;74(12):3515–3520. doi:10.1093/jac/dkz389
12. Hagiya H, Aoki K, Akeda Y, et al. In vitro effectiveness of meropenem and cefmetazole combination treatment against KPC-2-producing enterobacteriaceae. Microbial Drug Resist (Larchmont, NY). 2019;25(6):839–845. doi:10.1089/mdr.2018.0397
13. Krezdorn J, Adams S, Coote PJ. A Galleria mellonella infection model reveals double and triple antibiotic combination therapies with enhanced efficacy versus a multidrug-resistant strain of Pseudomonas aeruginosa. J Med Microbiol. 2014;63(7):945–955. doi:10.1099/jmm.0.074245-0
14. Wand ME, McCowen JWI, Nugent PG, Sutton JM. Complex interactions of Klebsiella pneumoniae with the host immune system in a Galleria mellonella infection model. J Med Microbiol. 2013;62(12):1790–1798. doi:10.1099/jmm.0.063032-0
15. Poirel L, Walsh TR, Cuvillier V, Nordmann P. Multiplex PCR for detection of acquired carbapenemase genes. Diagn Microbiol Infect Dis. 2011;70(1):119–123. doi:10.1016/j.diagmicrobio.2010.12.002
16. Landman D, Bratu S, Quale J. Contribution of OmpK36 to carbapenem susceptibility in KPC-producing Klebsiella pneumoniae. J Med Microbiol. 2009;58(10):1303–1308. doi:10.1099/jmm.0.012575-0
17. Zhou M, Wang D, Kudinha T, Yang Q, Yu S, Xu YC. Comparative evaluation of four phenotypic methods for detection of class A and B carbapenemase-producing enterobacteriaceae in China. J Clin Microbiol. 2018;56(8). doi:10.1128/JCM.00395-18
18. CLSI. Performance Standards for Antimicrobial Susceptibility Testing Twenty-Second Informational Supplement M100-S21. Wayne, PA: Clinical and Laboratory Standards Institute; 2012.
19. Bae S, Kim MC, Park SJ, et al. In vitro synergistic activity of antimicrobial agents in combination against clinical isolates of colistin-resistant Acinetobacter baumannii. Antimicrob Agents Chemother. 2016;60(11):6774–6779. doi:10.1128/AAC.00839-16
20. Bai Y, Liu B, Wang T, et al. In Vitro activities of combinations of rifampin with other antimicrobials against multidrug-resistant Acinetobacter baumannii. Antimicrob Agents Chemother. 2015;59(3):1466–1471. doi:10.1128/AAC.04089-14
21. Albur M, Noel A, Bowker K, MacGowan A. Bactericidal activity of multiple combinations of tigecycline and colistin against NDM-1-producing Enterobacteriaceae. Antimicrob Agents Chemother. 2012;56(6):3441–3443. doi:10.1128/AAC.05682-11
22. Alhadab AA, Ahmed MA, Brundage RC. Amikacin pharmacokinetic-pharmacodynamic analysis in pediatric cancer patients. Antimicrob Agents Chemother. 2018;62(4). doi:10.1128/AAC.01781-17
23. Markou N, Markantonis SL, Dimitrakis E, et al. Colistin serum concentrations after intravenous administration in critically ill patients with serious multidrug-resistant, gram-negative bacilli infections: a prospective, open-label, uncontrolled study. Clin Ther. 2008;30(1):143–151. doi:10.1016/j.clinthera.2008.01.015
24. Rodvold KA, Gotfried MH, Cwik M, Korth-Bradley JM, Dukart G, Ellis-Grosse EJ. Serum, tissue and body fluid concentrations of tigecycline after a single 100 mg dose. J Antimicrob Chemother. 2006;58(6):1221–1229. doi:10.1093/jac/dkl403
25. Gaibani P, Lombardo D, Lewis RE, et al. In vitro activity and post-antibiotic effects of colistin in combination with other antimicrobials against colistin-resistant KPC-producing Klebsiella pneumoniae bloodstream isolates. J Antimicrob Chemother. 2014;69(7):1856–1865. doi:10.1093/jac/dku065
26. Cebrero-Cangueiro T, Álvarez-Marín R, Labrador-Herrera G, et al. In vitro activity of pentamidine alone and in combination with aminoglycosides, tigecycline, rifampicin, and doripenem against clinical strains of carbapenemase-producing and/or Colistin-resistant Enterobacteriaceae. Front Cell Infect Microbiol. 2018;8:363. doi:10.3389/fcimb.2018.00363
27. Champion OL, Titball RW, Bates S. Standardization of G. mellonella larvae to provide reliable and reproducible results in the study of fungal pathogens. J Fungi (Basel, Switzerland). 2018;4(3). doi:10.3390/jof4030108
28. Benthall G, Touzel RE, Hind CK, et al. Evaluation of antibiotic efficacy against infections caused by planktonic or biofilm cultures of Pseudomonas aeruginosa and Klebsiella pneumoniae in Galleria mellonella. Int J Antimicrob Agents. 2015;46(5):538–545. doi:10.1016/j.ijantimicag.2015.07.014
29. Betts JW, Phee LM, Hornsey M, Woodford N, Wareham DW. In vitro and in vivo activities of tigecycline-colistin combination therapies against carbapenem-resistant Enterobacteriaceae. Antimicrob Agents Chemother. 2014;58(6):3541–3546. doi:10.1128/AAC.02449-14
30. Donadu MG, Trong LN, Viet HD, et al. Phytochemical compositions and biological activities of essential oils from the leaves, rhizomes and whole plant of hornstedtia bella škorničk. Antibiotics (Basel, Switzerland). 2020;9(6).
31. Usai D, Donadu M, Bua A, et al. Enhancement of antimicrobial activity of pump inhibitors associating drugs. J Infect Dev Ctries. 2019;13(2):162–164. doi:10.3855/jidc.11102
32. Mohamed SH, Mohamed MSM, Khalil MS, Azmy M, Mabrouk MI. Combination of essential oil and ciprofloxacin to inhibit/eradicate biofilms in multidrug-resistant Klebsiella pneumoniae. J Appl Microbiol. 2018;125(1):84–95. doi:10.1111/jam.13755
33. Trong LN, Viet HD, Quoc Doan T, et al. Biological activities of essential oils from leaves of Paramignya trimera (Oliv.) Guillaum and Limnocitrus littoralis (Miq.) swingle. Antibiotics (Basel, Switzerland). 2020;9(4).
34. Schultsz C, Geerlings S. Plasmid-mediated resistance in enterobacteriaceae: changing landscape and implications for therapy. Drugs. 2012;72(1):1–16. doi:10.2165/11597960-000000000-00000
35. Trecarichi EM, Tumbarello M. Therapeutic options for carbapenem-resistant Enterobacteriaceae infections. Virulence. 2017;8(4):470–484. doi:10.1080/21505594.2017.1292196
36. Zhang Y, Wang Q, Yin Y, et al. Epidemiology of Carbapenem-resistant Enterobacteriaceae infections: report from the China CRE network. Antimicrob Agents Chemother. 2018;62(2).
37. Petersen PJ, Labthavikul P, Jones CH, Bradford PA. In vitro antibacterial activities of tigecycline in combination with other antimicrobial agents determined by chequerboard and time-kill kinetic analysis. J Antimicrob Chemother. 2006;57(3):573–576. doi:10.1093/jac/dki477
38. Zhang J, Yu L, Fu Y, et al. Tigecycline in combination with other antibiotics against clinical isolates of carbapenem-resistant Klebsiella pneumoniae in vitro. Ann Palliat Med. 2019;8(5):622–631. doi:10.21037/apm.2019.09.11
39. Sader HS, Castanheira M, Flamm RK, Mendes RE, Farrell DJ, Jones RN. Tigecycline activity tested against carbapenem-resistant Enterobacteriaceae from 18 European nations: results from the SENTRY surveillance program (2010–2013). Diagn Microbiol Infect Dis. 2015;83(2):183–186. doi:10.1016/j.diagmicrobio.2015.06.011
40. Qureshi ZA, Paterson DL, Potoski BA, et al. Treatment outcome of bacteremia due to KPC-producing Klebsiella pneumoniae: superiority of combination antimicrobial regimens. Antimicrob Agents Chemother. 2012;56(4):2108–2113. doi:10.1128/AAC.06268-11
41. Zarkotou O, Pournaras S, Voulgari E, et al. Risk factors and outcomes associated with acquisition of colistin-resistant KPC-producing Klebsiella pneumoniae: a matched case-control study. J Clin Microbiol. 2010;48(6):2271–2274. doi:10.1128/JCM.02301-09
42. Bi S, Yao X, Huang C, et al. Antagonistic effect between tigecycline and meropenem: successful management of KPC-producing Klebsiella pneumoniae infection. Infection. 2019;47(3):497–500. doi:10.1007/s15010-019-01274-w
43. Logan LK, Weinstein RA. The epidemiology of Carbapenem-resistant Enterobacteriaceae: the impact and evolution of a global menace. J Infect Dis. 2017;215(suppl_1):S28–s36. doi:10.1093/infdis/jiw282
44. Kumarasamy KK, Toleman MA, Walsh TR, et al. Emergence of a new antibiotic resistance mechanism in India, Pakistan, and the UK: a molecular, biological, and epidemiological study. Lancet Infect Dis. 2010;10(9):597–602. doi:10.1016/S1473-3099(10)70143-2
45. Franceschini N, Caravelli B, Docquier JD, et al. Purification and biochemical characterization of the VIM-1 metallo-beta-lactamase. Antimicrob Agents Chemother. 2000;44(11):3003–3007. doi:10.1128/AAC.44.11.3003-3007.2000
46. Kohanski MA, Dwyer DJ, Collins JJ. How antibiotics kill bacteria: from targets to networks. Nat Rev Microbiol. 2010;8(6):423–435. doi:10.1038/nrmicro2333
47. Ramirez MS, Tolmasky ME. Amikacin: uses, resistance, and prospects for inhibition. Molecules (Basel, Switzerland). 2017;22(12):2267. doi:10.3390/molecules22122267
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.

