Back to Journals » International Journal of General Medicine » Volume 14
Impact of PCSK9, WDR12, CDKN2A, and CXCL12 Polymorphisms in Jordanian Cardiovascular Patients on Warfarin Responsiveness and Sensitivity
Authors Ibdah RK , AL-Eitan LN , Alrabadi NN, Almasri AY, Alnaamneh AH, Khasawneh RH , Alghamdi MA
Received 16 October 2020
Accepted for publication 17 December 2020
Published 14 January 2021 Volume 2021:14 Pages 103—118
DOI https://doi.org/10.2147/IJGM.S287238
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Scott Fraser
Rasheed K Ibdah,1 Laith N AL-Eitan,2 Nasr N Alrabadi,3 Ayah Y Almasri,2 Adan H Alnaamneh,2 Rame H Khasawneh,4 Mansour A Alghamdi5,6
1Division of Cardiology, Department of Internal Medicine, Faculty of Medicine, Jordan University of Science and Technology, Irbid, Jordan; 2Department of Biotechnology and Genetic Engineering, Jordan University of Science and Technology, Irbid, Jordan; 3Department of Pharmacology, Faculty of Medicine, Jordan University of Science and Technology, Irbid, Jordan; 4Department of Hematopathology, King Hussein Medical Center (KHMC), Jordan Royal Medical Services (RMS), Amman, Jordan; 5Department of Anatomy, College of Medicine, King Khalid University, Abha, Saudi Arabia; 6Genomics and Personalized Medicine Unit,College of Medicine, King Khalid University, Abha, Saudi Arabia
Correspondence: Laith N AL-Eitan
Department of Biotechnology and Genetic Engineering, Jordan University of Science and Technology. P.O.Box 3030, Irbid 22110, Jordan
Tel +962-2-7201000 Ext. 23464
Fax +962-2-7201071
Email [email protected]
Background: The main objective of this study is sought to determine the impacts of PCSK9, WDR12, CDKN2A, and CXCL12 polymorphisms on warfarin sensitivity and responsiveness in Jordanian cardiovascular patients during the initiation and stabilization phases of therapy.
Methods: This study took place at the anticoagulation clinic at Queen Alia Heart Institute (QAHI) in Jordan. DNA samples were collected from 212 cardiovascular patients and 213 healthy controls. Genomic SNPs genotyping was conducted using the MassARRAY System at the Australian Genome Research Facility.
Results: This study assessed 10 polymorphisms (rs11206510 within the PCSK9 gene, rs6725887 and rs7582720 within the WDR12 gene, rs4977574, rs10757278, and rs1333049 within the CDKN2A gene, rs2862116, rs7906426, rs1746048, and rs268322 within the CXCL12 gene) in 212 Jordanian cardiovascular patients. Carriers of CDKN2A rs1333049, rs10757278, and PCSK9 rs11206510 polymorphisms had an increased risk of resistance during the initiation phase of warfarin therapy compared to those who do not carry it, or those who are carrying one polymorphism only (P < 0.05), while carriers of CXCL12 rs7906426 polymorphism had similar increased risk but during the stabilization phase of warfarin therapy (P < 0.05).
Conclusion: Carriers of CXCL12 rs2862116 polymorphism had an increased risk to be warfarin extensive responders compared to those with no or only one polymorphism (P = 0.01). However, the presence of PCSK9 rs11206510 polymorphism affects the warfarin maintenance doses (P ˃ 0.0001).
Keywords: warfarin, cardiovascular disease, PCSK9, WDR12, CDKN2A, CXCL1
Introduction
Warfarin is the most oral anticoagulant used to control blood coagulation in patients with cardiovascular diseases (CVDs) such as venous thrombosis, atrial fibrillation, patients with a thrombosis history, and cardiac valve replacements.1,2 Warfarin effect as an anticoagulant is monitored using the international normalized ratio (INR), which is the measurement of the patient’s thrombotic status, and is targeted to be kept within a strict therapeutic range. This is necessary especially with the close relationship between the risk of hemorrhage and the high INR measurements and, on the other hand, between the risk of stroke or thromboembolism and the low INR measurements.3,4 Warfarin controls the thrombotic status and maintains the INR within the therapeutic range by inhibiting the vitamin K epoxide reductase (VKOR) enzyme, which is integral for the production of active clotting factors such as II, VII, IX, and X.5,6 Although Warfarin is the most commonly used anticoagulant since 1954,7 the use of this type of anticoagulant is hindered by the high variability in response between different patients and the required dosage adjustment. This may be attributed to the differences in age, diet, liver and renal functions, concomitant diseases, and the use of concomitant medications.8–11
Genetic variability was suggested to be able to affect and change the trend of response and patients’ sensitivity to warfarin therapy.12,13 Tens of genes may play integral roles in warfarin metabolisms such as VKORC1, CYP2C9, FVII, and APOE.14 Among these genes, the polymorphism of CYP2C9 and VKORC1 genes are considered the main sources of those genetic variations that influencing warfarin metabolism. Concomitantly, many studies have been extensively conducted on both of them and in different populations or ethnic groups.15 Studying more genes that may influence CVD or warfarin metabolism can help in finding more accurate therapeutic strategies with minimum adverse side-effects and pave the way for a more proper application of personalized medicine in therapeutics, which is advantageous over the traditional approach of the trial-and-error.16,17 Therefore, four genes (PCSK9, WDR12, CDKN2A, and CXCL12) were analyzed to determine if they can have any effect on the sensitivity and responsiveness of warfarin. Pro-protein-convertase subtilisin-Kexin type 9 (PCSK9) gene was found to be associated with Coronary heart disease (CHD) in some populations, due to its ability to modulate cholesterol metabolism by controlling LDL receptor (LDLR).18,19 D Repeat Protein 12 (WDR12) gene encodes a protein that has integral roles in a wide range of cellular processes, including proliferation and cell division, cell cycle progression, ribosome biogenesis, and regulate lipid level.20–22 Several studies found that the polymorphisms within the WDR12 gene were involved in some CVD in the white population.23,24 Moreover, the cyclin-dependent kinase inhibitor 2A gene (CDKN2A), a tumor suppressor gene that is involved in the regulation of many cellular processes including proliferation, aging, and apoptosis, was found to be related to different diseases including CVD.25,26 Also, the Chemokine (C-X-C motif) ligand 12 (CXCL12) gene, a family member of the CXC-chemokines, was found to be associated with CVD through arterial remodeling and thickening.22,27 We hypothesized that there is a correlation between the genomic polymorphisms found in the PCSK9, WDR12, CDKN2A, and CXCL12 genes with warfarin sensitivity and responsiveness. To test this hypothesis, genotyping of the genomics variations was analyzed and determined in the PCSK9, WDR12, CDKN2A, and CXCL12 genes, and thus, understand the impacts of PCSK9, WDR12, CDKN2A, and CXCL12 polymorphisms on the CVD’s risks in Jordanian patients, and to analyze their effects on warfarin response and sensitivity in those patients.
Patients and Methods
Study Design and Population
A population of 425 subjects including 212 cardiovascular patients and 213 healthy controls was recruited for the aim of this study. The healthy control subjects were older than 18 years of age and/or were free of any CVD. After being informed about the aims of the study and signing the proper informed consent, the cardiovascular patients from the anticoagulation clinics at the Queen Alia Heart Institute (QAHI) with matched ages were recruited. Those selected patients started the warfarin therapy at any time between January 2014 and November 2015 and continued on warfarin for 3 or more months. Those patients who did not visit the anticoagulation clinic regularly have lost their clinical data, receiving concomitant mediations that are interacting with warfarin, pregnant women and alcoholic abusers were all excluded from the study. Both the institutional review board (IRB) committees at Jordan University of Science and Technology and the Royal Medical Services approved this study with ethical code number 13/78/2014. This study was also conducted in accordance with the Declaration of Helsinki.
Out of the 350 cardiovascular patients who were initially examined, 50 patients were excluded for not fulfilling the study criteria, and 300 patients were referred for participation in this study (Figure 1). Then after, during the study processes, 80 patients were excluded due to their refusal, insufficient follow-up, or inability to continue on the scheduled therapeutic program for the targeted period. Therefore, 220 patients eventually participated in the study. At the genotyping step, eight more patients were subsequently excluded due to the failure of genotyping. Finally, only 139 out of the 212 remaining participants were able to reach the therapeutic stabilization phase. Patients’ demographics and clinical characteristics for each group were listed in a previous study by AL-Eitan et al (2019).13
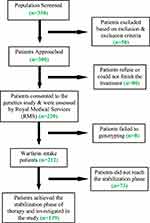 |
Figure 1 A flow chart demonstrating the study flow and design. |
Data Collection and Follow-Up Time
The blood samples were collected from patients during their visits to the anticoagulant clinic and from all healthy control subjects to the determination of the venous INR and SNP genotypes within PCSK9, WDR12, CDKN2A, and CXCL12 genes. Data including relevant demographic and clinical characteristics were recorded. The dose for warfarin and the INR values were recorded in both the initiation (when first used) and the stabilization (the INR is stabilized within the therapeutic range for 3 months) phases.
SNP Selection and Genotyping
In this study, 10 SNPs within PCSK9, WDR12, CDKN2A, and CXCL12 genes were obtained from public databases, including the SNP database of the National Center for Biotechnology Information (NCBI) (http://www.ncbi.nlm.nih.gov/SNP/) and the Applied Biosystems SNP database (http://www.appliedbiosystems). Information about the aforementioned SNPs, including the genes, their SNP IDs, and their loci are shown in (Table S1). The Wizard Genomic DNA Purification Kit (Promega) was used for genomic DNA extraction. The genotype profile of the samples that fulfill the quantitative requirements was identified on a MassARRAY_ System (iPLEX GOLD) (Sequenom, San Diego, CA, USA) at the Australian Genome Research Facility (AGRF). Information is available upon request about the MassARRAYTM system protocols and the used genes primers. The Sequenom MassARRAYTM system protocol consists of five steps: Template Amplification, Dephosphorylation, SBE reactions. Sample conditioning and transfer and Genotype calling and bioinformatics.
Outcome Measures
This study focused on the evaluation of the warfarin responsiveness and sensitivity during both the initiation and stabilization phases of therapy. Therefore, patients were classified into two categories. The first category was distinguished based on warfarin sensitivity according to Gordon's study (2009).28 In this category, patients were classified into three groups:
- Extensive metabolizer or warfarin resistance group, requiring a higher dose (>49 mg/week) of warfarin to achieve the therapeutic INR.
- Warfarin response group or moderate metabolizers, needing an intermediate dose of 21–49 mg/week.
- Warfarin-sensitive or poor metabolizers, requiring a low dose of warfarin (<21 mg/week).
The second category was based on warfarin responsiveness, patients were divided into three groups according to Higashi et al (2002):29
- Good responders who have an INR value within the target therapeutic range.
- Poor responders (The value of the INR beneath the targeted therapeutic range).
- Ultra-responders (INR over the therapeutic range).
Finally, the maintenance dose during the stabilization phase is defined as the mean average of the administered doses to the patient during that time. All unchanged weekly doses based on the measurements of the INR for no less than two consecutive visits were used to calculate the stable maintenance dose.
Statistical Analysis
The Statistical Package for the Social Sciences (SPSS) version 21.0 was used to conduct all statistical analyses. Microsoft Excel was used to calculate simple call rates and discrepancies. The Pearson X2 test was used to calculate the deviation from Hardy Weinberg Equilibrium (HWE). Moreover, the Court lab-HW calculator was used to calculate the Minor allele frequencies (MAF) and HWE p-values for genotypic distribution. To test for the association between the studied SNPs and the warfarin response, different analyses tests were conducted, including the chi-square (Kruskal–Wallis and Tukey Pairwise comparison) tests. The haplotype genetic analysis test was performed using SNPStats software (https://www.snpstats.net/start.htm).
Results
Study Group
A total of 212 cardiovascular patients with warfarin intake finally succeeded in participation in the current study. The samples from those patients were valid to be used in the investigation for the association of PCSK9, WDR12, CDKN2A, and CXCL12 polymorphisms with the risks of CVD. Besides, the effect and association between those SNPs and both the sensitivity and responsiveness profile of warfarin were studied at both phases of therapy; the initiation and the stabilization phases. However, only 139 of the participants reached the late stage of therapy stabilization and, therefore, were included in the evaluation of the effect of these polymorphisms on the responsiveness and sensitivity to warfarin. Overall, 18.7% of the participants were poor metabolizers, 18.0% extensive metabolizers, and 63.3% good metabolizers, with an average age of 54.2, 47.6, and 53.2 years, respectively. The demographic characteristics, indications for anticoagulation therapy, and warfarin required dose for each group were summarized in a previous study by Al-Eitan et al.30
All 10 candidate SNPs that were investigated in this study were following the HWE except WDR12 rs7582720 SNP. All SNPs were polymorphic and surpassed the tests for quality control with both a low discrepancy rate and a high accuracy. The minor alleles and their frequencies for the studied SNPs are shown in (Table S2).
Association of PCSK9, WDR12, CDKN2A, and CXCL12 SNPs with CVD
Among 10 SNPs of PCSK9, WDR12, CDKN2A, and CXCL12 genes, only rs4977574 SNP of CDKN2A gene showed a significant association with CVD (P = 0.04). Genotypic and allelic frequencies at PCSK9, WDR12, CDKN2A, and CXCL12 gene loci in patients and control subjects are shown in (Table 1). Moreover, significant differences were found between CXCL12 haplotypes (CTAA and CTAG) and CVD among patients and controls with (P = 0.04) and (P ˂ 0.0001), respectively (Table S3).
 |
Table 1 The Allelic and Genotypic Distributions of PCSK9, WDR12, CDKN2A, and CXCL12 SNPs in 212 Cardiovascular Patients and 213 Healthy Controls |
Association of PCSK9, WDR12, CDKN2A, and CXCL12 Polymorphisms with Warfarin Sensitivity
No significant association between all 10 SNPs and warfarin sensitivity among the three warfarin-sensitive groups during the initiation phase of therapy were found (P ˃ 0.05) (Table 2). Carriers of the wild-type (CC) of rs11206510 SNP in the PCSK9 gene had a significantly higher risk (60.0%) of being resistant to warfarin (P = 0.03) compared to the carriers of the other polymorphisms (CT) and (CT) (Table 2). For the CDKN2A polymorphisms, carriers of the wild-type (CC) of rs1333049 SNP had a significantly higher risk (28.3%) of being resistant to warfarin (P = 0.04) compared with carriers of (GC) (11.2%) and (GG) (16.0%) polymorphisms (Table 2). Moreover, carriers of the CDKN2A rs10757278 SNP (GG) had a significantly higher risk (30.4%) of being resistant to warfarin (P = 0.01) compared to wild-type (AA) subjects (12.2%) or carriers of the polymorphism (GA) (12.1%) (Table 2). However, a significant association was found between CDKN2A haplotype (CGA) (P = 0.0001) and warfarin sensitivity (Table S4). Also, two haplotypes of the CXCL12 gene (TCAA and CTAG) showed significant associations with warfarin sensitivity (P = 0.03) and (P = 0.04), respectively (Table S4). Moreover, no significant association was found between CXCL12 polymorphism and sensitivity to warfarin (P ˃ 0.05) (Table 3) during the stabilization phase of therapy. But carrier of (GG) genotype of CXCL12 rs7906426 SNP had a significantly higher risk of being resistant to warfarin compared to the carriers of the other genotypes (P = 0.04) (Table 3).
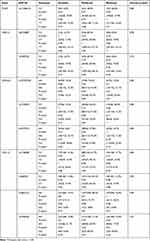 |
Table 2 Association of PCSK9, WDR12, CDKN2A, and CXCL12 SNPs with Warfarin Sensitivity During the Initiation Phase of Therapy |
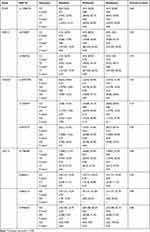 |
Table 3 Association of PCSK9, WDR12, CDKN2A, and CXCL12 SNPs with Warfarin Sensitivity During the Stabilization Phase of Therapy |
Association of PCSK9, WDR12, CDKN2A and CXCL12 SNPs with Variability on Warfarin Required Doses
No significant differences were found between PCSK9, WDR12, CDKN2A, and CXCL12 polymorphisms and warfarin Initiation doses (P ˃ 0.05) (Table 4). In contrast, a significant association was observed between rs11206510 SNP of the PCSK9 gene and warfarin Maintenance doses (P ˃ 0.0001) (Table 4). The post-hock multiple comparisons test was used to the Association of PCSK9, WDR12, CDKN2A and CXCL12 SNPs with variability on warfarin required doses, the test reveals that there was a significant association between rs11206510 SNP of the PCSK9 gene and warfarin Initiation and Maintenance doses (P ˃ 0.05) (Table S5).
 |
Table 4 Association of PCSK9, WDR12, CDKN2A, and CXCL12 SNPs with Variability on Warfarin Required Doses |
Association of PCSK9, WDR12, CDKN2A, and CXCL12 Polymorphisms and Warfarin Responsiveness
There were no significant differences between the frequencies for the polymorphisms among the three groups: poor, good, and ultra-responders during the Initiation Phase of Therapy (P ˃ 0.05) (Table 5). In contrast, a significant association was observed between CXCL12 rs2862116 SNP and warfarin responsiveness during the stabilization phase of therapy (P = 0.01) (Table 6). In fact, carriers of this SNP in the wild-type (AA) and the polymorphic genotype (AG) were considered as extensive responders (Table 6). Moreover, a significant association was observed between the CAG genetic haplotype of the CXCL12 gene and warfarin responsiveness (P = 0.029) (Table S6).
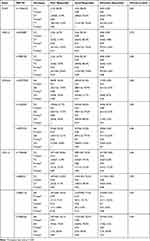 |
Table 5 Association of PCSK9, WDR12, CDKN2A, and CXCL12 SNPs with Warfarin Responsiveness During the Initiation Phase of Therapy |
 |
Table 6 Association of PCSK9, WDR12, CDKN2A, and CXCL12 SNPs with Warfarin Responsiveness During the Stabilization Phase of Therapy |
Association of PCSK9, WDR12, CDKN2A, and CXCL12 Polymorphisms and INR Values
A significant difference was observed between INR values measured at the initiation phase of therapy and CDKN2A rs4977574 SNP (P = 0.04) (Table 7), and this SNP has remained significant even after the post-Hock multiple comparisons test with p<0.05 (Table S7). Moreover, A significant difference was found between Maintenance INR values and rs2862116 SNP of the CXCL12 gene (P = 0.04; Table 7).
 |
Table 7 Association of PCSK9, WDR12, CDKN2A, and CXCL12 SNPs with INR Treatment Outcome |
Discussion
In the current study, researchers examined the role of the PCSK9, WDR12, CDKN2A, and CXCL12 polymorphisms in an increased risk of CVD, in addition to warfarin sensitivity and response in patients from Jordan with CVD during both phases of warfarin therapy; initiation and stabilization phases. Several studies identified the association of these genes with an increased risk of CVD in a certain population. For example, according to Schunkert (2011), rs11206510 SNP of the PCSK9 gene was associated with increased risk of CHD in the European population,24 as such, the rs11206510-C allele was accompanied with the early-onset CHD, but not with overall CHD in Han Chinese according to Xu (2010).31 On the other hand, another study by Lv et al demonstrated that the rs11206510 SNP was significantly accompanied by CHD in Han Chinese.32 Similar to these conflicting results, some studies found that the WDR12 polymorphisms were involved in CVD in some populations, while other studies failed to find the association between these polymorphisms and the disease. PCSK9 polymorphisms were not associated with coronary artery disease risk in Southern Chinese Han population.33 Moreover, the association of the WDR12 polymorphisms with CVD was proved by Schunkertet al (2010).24 While López -Mejías et al were not able to confirm this association of the WDR12 gene with CVD in Southern European patients.34 Moreover, CDKN2A and CXCL12 polymorphisms were turned up to be accompanied by CAD in different populations.25,27 Therefore, the association between these genes and CVD in the Jordanian population was assessed, it was found that only CDKN2A rs4977574 SNP was associated with CVD, possibly due to the role of the CDKN2A gene in controlling the cell cycle, and controlling the G1 phase progression pathway, that enhances the functions of the genes which is integral for cell proliferation and cell cycle progression.35 Increasing this pathway activity in cardiovascular tissues leads to a high proliferation rate, and expedites remodeling and hypertrophy of both the vascular and cardiac tissues, thus, increasing the patients' susceptibility to CAD.36
In addition to the CTAA and CTAG genetic haplotype blocks of the CXCL12 gene that showed significant association with CVD, this gene plays an integral role in the proliferation and accumulation of progenitor cells of the smooth muscles, cell arrest, angiogenesis, and cell survival, that may explain the involvement of this gene in CVD.37 The allelic frequencies of the polymorphisms in our Jordanian population were approximately in agreement with those found in other ethnicities, for instance, the PCSK9 rs11206510 T allele, that can be found in about 81% of Caucasians, 94% of Chinese Han population, and 83% in our population, and the CXCL12 rs1746048 C allele, that can be found in about 84% of Caucasians, 64% of Chinese Han population, and 77% in our population. These contradictory results reflect the heterogeneity of CVD, as such, many interactions between a variety of genetic and the environmental factors are involved in the pathogenesis of CVD. Restricted to our knowledge, we could not find valid studies that can show any association between PCSK9, WDR12, CDKN2A, and CXCL12 polymorphisms and the warfarin metabolism in Jordanian patients with CVD. The results out of the current pharmacogenetic study indicate that no significant association was found of PCSK9, WDR12, CDKN2A, and CXCL12 polymorphisms with warfarin sensitivity among all categorical groups (resistant, moderate, and sensitive) during both phases of the therapy; the initiation and the stabilization phases (P ˃ 0.05). However, significant associations were found between PCSK9 rs11206510 and CDKN2A rs1333049 SNPs and resistance to warfarin, indicated that patients with these polymorphisms required higher warfarin doses to achieve its therapeutic effect. Moreover, carrying a CDKN2A CGA and CXCL12 (TCAA and CTAG) genetic haplotype blocks were associated significantly with the sensitivity of warfarin.
In addition to the results mentioned above, patients were divided into three groups of responders (extensive, good, and poor) based on the average measurements of their INR during both phases of therapy; the initiation and the stabilization phases. The study findings indicated that there are no significant differences between all groups and with all polymorphisms during the imitation phase, and only CXCL12 rs2862116 SNP showed a significant difference during the stabilization phase. Moreover, only CDKN2A CAG genetic haplotype block affects warfarin responsiveness in Jordanian patients. According to current results, there was not any significant influence of any of those polymorphisms on the dose of warfarin when considering the initiation phase of therapy. However, during the stabilization phase, only the PCSK9 rs11206510 SNP was able to demonstrate a significant association with warfarin doses. Moreover, only CDKN2A rs4977574 SNP showed a significant difference during the initiation phase concerning the INR measurements, in contrast, CXCL12 rs2862116 SNP showed a significant difference with regards to the maintenance measurements of the INR. It is also possible that the Jordanian patients carry further gene mutations that can be involved in the pathway of coagulation. One of the limitations of this study is the number of individuals involved in the study. Therefore, further investigations with a larger sample size in different Arab populations are required to confirm the findings of this study.
Conclusions
In the present study, the frequencies of PCSK9, WDR12, CDKN2A, and CXCL12 polymorphisms were examined in the Jordanian population and, then after, we examined their effects on the sensitivity and responsiveness to warfarin during both phases of therapy; the initiation and the stabilization phases. In conclusion, this study demonstrated a significant association between the CDKN2A rs4977574 SNP and the risk of CVD in the Jordanian population. Moreover, the presence of certain genotypes of the CDKN2A rs1333049 and rs10757278, and the PCSK9 rs11206510 polymorphisms significantly affected the resistance profile of warfarin during the initiation phase of therapy, while the CXCL12 rs2862116 SNP significantly affected responsiveness to warfarin during the stabilization phase of therapy. Moreover, PCSK9 rs11206510, CDKN2A rs4977574, and CXCL12 rs2862116 SNPs showed significant differences in warfarin maintenance doses, initiation INR, and maintenance INR, respectively. Further studies with a larger sample size and possibly different populations will be required to confirm our results.
Acknowledgments
We grant our thanks to the Anti-coagulant clinic in Queen Alia Heart Institute for their generous help during this study.
Funding
This project was funded under grant number (203/2014) and supported by the Deanship of Research at Jordan University of Science and Technology.
Disclosure
The authors declared no conflict of interest.
References
1. Ansell J, Hirsh J, Hylek E, et al. Pharmacology and management of the vitamin K antagonists: American college of chest physicians evidence-based clinical practice guidelines. Chest. 2008;133(6):160S–198S. doi:10.1378/chest.08-0670
2. Kim Y, Smith A, Wu AH. C3435T polymorphism of MDR1 gene with warfarin resistance. Clin Chim Acta. 2013;425:34–36. doi:10.1016/j.cca.2013.07.010
3. Hirsh J, Dalen JE, Anderson DR, et al. Oral anticoagulants: mechanism of action, clinical effectiveness, and optimal therapeutic range. Chest. 2001;119(1):8S–21S. doi:10.1378/chest.119.1_suppl.8S
4. Eriksson N, Wallentin L, Berglund L, et al. Genetic determinants of warfarin maintenance dose and time in therapeutic treatment range: a RE-LY genomics substudy. Pharmacogenomics. 2016;17(13):1425–1439. doi:10.2217/pgs-2016-0061
5. Rost S, Fregin A, Ivaskevicius V, et al. Mutations in VKORC1 cause warfarin resistance and multiple coagulation factor deficiency type 2. Nature. 2004;427(6974):537–541. doi:10.1038/nature02214
6. Yin T, Miyata T. Warfarin dose and the pharmacogenomics of CYP2C9 and VKORC1—rationale and perspectives. Thromb Res. 2007;120(1):1–10. doi:10.1016/j.thromres.2006.10.021
7. Ravina E. The Evolution of Drug Discovery: From Traditional Medicines to Modern Drugs. John Wiley & Sons; 2011.
8. Hirsh J. American Heart Association/American College of Cardiology foundation guide to warfarin therapy. J Am Coll Cardiol. 2003(41):1633–1652.
9. Visser LE, Bleumink GS, Trienekens PH, et al. The risk of overanticoagulation in patients with heart failure on coumarin anticoagulants. Br J Haematol. 2004;127(1):85–89. doi:10.1111/j.1365-2141.2004.05162.x
10. Ansell J, Hirsh J, Poller L, et al. The pharmacology and management of the vitamin K antagonists: the seventh ACCP conference on antithrombotic and thrombolytic therapy. Chest. 2004;126(3):204S–233S. doi:10.1378/chest.126.3_suppl.204S
11. Lip GY, Frison L, Halperin JL, Lane DA. Comparative validation of a novel risk score for predicting bleeding risk in anticoagulated patients with atrial fibrillation: the HAS-BLED (Hypertension, abnormal renal/liver function, stroke, bleeding history or predisposition, labile INR, elderly, drugs/alcohol concomitantly) score. J Am Coll Cardiol. 2011;57(2):173–180. doi:10.1016/j.jacc.2010.09.024
12. AL-Eitan L, Almasri A, Khasawneh R. Impact of CYP2C9 and VKORC1 polymorphisms on warfarin sensitivity and responsiveness in Jordanian cardiovascular patients during the initiation therapy. Genes. 2018;9(12):578. doi:10.3390/genes9120578
13. Al-Eitan LN, Almasri AY, Al-Habahbeh SO. Impact of a variable number tandem repeat in the CYP2C9 promoter on warfarin sensitivity and responsiveness in Jordanians with cardiovascular disease. Pharmgenomics Pers Med. 2019;12:15–22. doi:10.2147/PGPM.S189838
14. Hughes DA, Pirmohamed M. Warfarin Pharmacogenetics. Springer; 2007:899–902.
15. Li C, Schwarz UI, Ritchie MD, et al. Relative contribution of CYP2C9 and VKORC1 genotypes and early INR response to the prediction of warfarin sensitivity during initiation of therapy. Blood. 2009;113(17):3925–3930. doi:10.1182/blood-2008-09-176859
16. Xie H-G, Frueh FW. Pharmacogenomics Steps Toward Personalized Medicine. Per Med. 2005;2(4):325–337. doi:10.2217/17410541.2.4.325.
17. AL-Eitan LN, Tarkhan AH. Practical challenges and translational issues in pharmacogenomics and personalized medicine from 2010 onwards. Curr Pharmacogenomics Person Med. 2017;14(1):7–17. doi:10.2174/1875692115666161215103842
18. Horton JD, Cohen JC, Hobbs HH. Molecular biology of PCSK9: its role in LDL metabolism. Trends Biochem Sci. 2007;32(2):71–77. doi:10.1016/j.tibs.2006.12.008
19. Huijgen R, Boekholdt SM, Arsenault BJ, et al. RETRACTED: plasma PCSK9 levels and clinical outcomes in the TNT (treating to new targets) trial: a nested case-control study. J Am Coll Cardiol. 2012;59(20):1778–1784. doi:10.1016/j.jacc.2011.12.043
20. Hölzel M, Rohrmoser M, Schlee M, et al. Mammalian WDR12 is a novel member of the Pes1–Bop1 complex and is required for ribosome biogenesis and cell proliferation. J Cell Biol. 2005;170(3):367–378. doi:10.1083/jcb.200501141
21. Blattmann P, Schuberth C, Pepperkok R, et al. RNAi–based functional profiling of loci from blood lipid genome-wide association studies identifies genes with cholesterol-regulatory function. PLoS Genet. 2013;9(2):e1003338. doi:10.1371/journal.pgen.1003338
22. Zabalza M, Subirana I, Lluis-Ganella C, et al. Association between coronary artery disease genetic variants and subclinical atherosclerosis: an association study and meta-analysis. Rev Esp Cardiol. 2015;68(10):869–877. doi:10.1016/j.rec.2014.10.023
23. Kathiresan S, Voight BF, Purcell S, et al. Genome-wide association of early-onset myocardial infarction with single nucleotide polymorphisms and copy number variants. Nat Genet. 2009;41(3):334–341.
24. Schunkert H, König IR, Kathiresan S, et al. Large-scale association analysis identifies 13 new susceptibility loci for coronary artery disease. Nat Genet. 2011;43(4):333–338. doi:10.1038/ng.784
25. Samani NJ, Erdmann J, Hall AS, et al. Genomewide association analysis of coronary artery disease. N Eng J Med. 2007;357(5):443–453. doi:10.1056/NEJMoa072366
26. Lettre G, Palmer CD, Young T, et al. Genome-wide association study of coronary heart disease and its risk factors in 8090 African Americans: the NHLBI CARe Project. PLoS Genet. 2011;7(2):e1001300. doi:10.1371/journal.pgen.1001300
27. Farouk SS, Rader DJ, Reilly MP, et al. CXCL12: a new player in coronary disease identified through human genetics. Trends Cardiovasc Med. 2010;20(6):204–209. doi:10.1016/j.tcm.2011.08.002
28. Consortium IWP. Estimation of the warfarin dose with clinical and pharmacogenetic data. N Eng J Med. 2009;360(8):753–764.
29. Higashi MK, Veenstra DL, Kondo LM, et al. Association between CYP2C9 genetic variants and anticoagulation-related outcomes during warfarin therapy. JAMA. 2002;287(13):1690–1698. doi:10.1001/jama.287.13.1690
30. Al-Eitan LN, Almasri AY, Khasawneh RH. Effects of CYP2C9 and VKORC1 polymorphisms on warfarin sensitivity and responsiveness during the stabilization phase of therapy. Saudi Pharm J. 2019;27(4):484–490. doi:10.1016/j.jsps.2019.01.011
31. Xu C, Wang F, Wang B, et al. Minor allele C of chromosome 1p32 single nucleotide polymorphism rs11206510 confers risk of ischemic stroke in the Chinese Han population. Stroke. 2010;41(8):1587–1592. doi:10.1161/STROKEAHA.110.583096
32. Lv X, Zhang Y, Rao S, et al. Joint effects of genetic variants in multiple loci on the risk of coronary artery disease in Chinese Han subjects. Circ J. 2012; 76(8):1987–1992. doi:10.1253/circj.cj-12--0156.
33. Cai G, Yu L, Huang Z, et al. Serum PCSK9 levels, but not PCSK9 polymorphisms, are associated with CAD risk and lipid profiles in southern Chinese Han population. Lipids Health Dis. 2018;17(1):213. doi:10.1186/s12944-018-0859-5
34. Schunkert H, Erdmann J, Samani NJ. Genetics of myocardial infarction: a progress report. Eur Heart J. 2010;31(8):918–925. doi:10.1093/eurheartj/ehq038
35. Dimova DK, Dyson NJ. The E2F transcriptional network: old acquaintances with new faces. Oncogene. 2005;24(17):2810–2826. doi:10.1038/sj.onc.1208612
36. Pilbrow AP, Folkersen L, Pearson JF, et al. The chromosome 9p21. 3 coronary heart disease risk allele is associated with altered gene expression in normal heart and vascular tissues. PLoS One. 2012;7(6):e39574. doi:10.1371/journal.pone.0039574
37. Döring Y, Pawig L, Weber C, et al. The CXCL12/CXCR4 chemokine ligand/receptor axis in cardiovascular disease. Front Physiol. 2014;5:212. doi:10.3389/fphys.2014.00212
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
