Back to Journals » OncoTargets and Therapy » Volume 7
High expression of Sonic Hedgehog signaling pathway genes indicates a risk of recurrence of breast carcinoma
Authors Jeng K , Sheen I, Jeng W, Yu M, Hsiau H, Chang F
Received 18 September 2013
Accepted for publication 2 November 2013
Published 27 December 2013 Volume 2014:7 Pages 79—86
DOI https://doi.org/10.2147/OTT.S54702
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 4
Kuo-Shyang Jeng,1 I-Shyan Sheen,2 Wen-Juei Jeng,2 Ming-Che Yu,3 Hsin-I Hsiau,3 Fang-Yu Chang3
1Department of Surgery, Far Eastern Memorial Hospital, Taipei, 2Department of Internal Medicine, Chang-Gung Memorial Hospital, Linkou Medical Center, Chang-Gung University, Tao-Yuan, 3Department of Medical Research, Far Eastern Memorial Hospital, Taipei, Taiwan
Background: Abnormal activation of the Sonic Hedgehog (SHH) signaling pathway contributing to carcinogenesis of some organs has been reported in the literature. We hypothesize that activation of the SHH pathway contributes to the recurrence of breast carcinoma.
Methods: Fifty consecutive patients with invasive breast carcinoma following curative resection were enrolled in this prospective study. The ratios of messenger RNA (mRNA) expression for Sonic Hedgehog (SHH), patched homolog-1 (PTCH-1), glioma-associated oncogene-1 (GLI-1), and smoothened (SMOH) were measured between breast carcinoma tissue and paired noncancerous breast tissue. These ratios were compared with their clinicopathologic characteristics. These factors and the mRNA ratios were compared between patients with recurrence and those without recurrence.
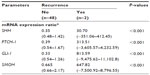
Results: The size of the invasive cancer correlated significantly with the ratio of SHH mRNA (P=0.001), that of PTCH-1 mRNA (P=0.005), and that of SMOH mRNA (P=0.021). Lymph node involvement correlated significantly with the ratio of SMOH mRNA (P=0.041). The correlation between Her-2 neu and the ratio of GLI-1 mRNA was statistically significant (P=0.012). Each ratio of mRNA of SHH, PTCH-1, GLI-1, and SMOH correlated significantly with cancer recurrence (P<0.001 for each).
Conclusion: We suggest that high expression of SHH mRNA, PTCH-1 mRNA, GLI-1 mRNA, and SMOH mRNA in breast cancer tissue correlates with invasiveness and is a potential biomarker to predict postoperative recurrence.
Keywords: SHH pathway, breast carcinoma, prediction, recurrence
Introduction
Rates of breast cancer, a leading cause of death in Taiwan and many other countries, have increased rapidly in recent times.1 In spite of advances in diagnostic tools and surgical techniques, and chemotherapy, radiotherapy, and targeted therapy, the potential for cancer recurrence remains.2 Early prediction of those who are likely to experience postoperative recurrence is a challenge. For those with a high potential of recurrence, postoperative adjuvant therapies have to be introduced early. This makes it important to identify new biomarkers in predicting patient prognosis.
The Sonic Hedgehog (SHH) signaling pathway plays a critical role in organizing cell growth and differentiation during embryonic tissue patterning,3–5 and is important in mouse mammary gland development.6 Disruption of the Patched homolog-1 (PTCH-1) or glioma-associated oncogene-2 (GLI-2) gene results in severe defects in ductal morphogenesis, such as ductal dysplasia, similar to human breasts.6,7 In vitro research shows that disruptions of these genes also occur in breast carcinoma.7 These implicate the potential role of the SHH pathway in breast oncogenesis. However, whether the SHH pathway contributes to cancer recurrence remains unknown.
We hypothesize that SHH activation contributes to the recurrence of breast carcinoma. This prospective study was designed to assess the correlation between mRNA expression of SHH, PTCH-1, glioma-associated oncogene-1 (GLI-1), and smoothened (SMOH) and postoperative outcomes.
Materials and methods
Eligibility and exclusion criteria
Sixty patients with invasive ductal carcinoma of the breast undergoing surgery were screened prospectively for entry into this institutional review board (Far Eastern Memorial Hospital Research Ethics Review Committee) approved study between September 2008 and December 2009. Exclusion criteria included preoperative neoadjuvant therapies, previous mastectomy (recurrent category), a nearest resection margin of less than 5 mm, carcinoma in situ, refusal to participate, and lack of attendance for regular postoperative follow-up. After exclusion, 50 consecutive female patients of mean age 54.61±10.25 years were enrolled.
Operative procedures and methods
Surgical procedures included 20 modified radical mastectomies and 30 partial mastectomies plus axillary lymph node dissection. At the end of each operation, we did a 0.5 × 0.5 cm sized tissue biopsy from both cancerous and noncancerous portions of the resected specimens. The noncancerous biopsy site was at least 2 cm away from the cancer margin. The instruments were changed between biopsy procedures to avoid transfer of cancer cells into noncancerous tissues.
Formalin-fixed, paraffin-embedded sections of obtained tissues were stained with hematoxylin for histology according to World Health Organization breast carcinoma histologic classification criteria.8 The clinical stage was categorized as the American Joint Committee on Cancer TNM classification (7th edition).8 Clinicopathologic characteristics regarding age, invasive tumor size, lymph node metastasis, lymphovascular invasion, and estrogen receptor, progesterone receptor, and human epidermal growth factor receptor-2 (Her-2neu) status were obtained from clinical and pathology reports (Table 1). All patients agreed to participate in this study and signed an informed consent before surgery.
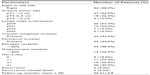  | Table 1 Clinicopathologic characteristics of patients (n=50) |
Detection of human mRNA for SHH, PTCH-1, GLI-1, and SMOH
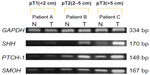
The examination included extraction of RNA and reverse transcription and amplification of cDNA for SHH, PTCH-1, GLI-1, SMOH, and glyceraldehyde-3-phosphate dehydrogenase (GAPDH) by real-time polymerase chain reaction (PCR).
Extraction of RNA and reverse transcription PCR
RNA was extracted from tumor tissue and noncancerous tissue using the innuPREP RNA mini kit (Analytik, Jena AG, Jena, Germany). All the tissues were homogenized in lysis buffer at room temperature for 5 minutes, and then centrifuged at 12,000 rpm for 2 minutes. Each sample was mixed with 70% alcohol and then centrifuged. We added a first wash buffer and centrifuged for one minute, then a second wash buffer and repeated the procedure. After removing the ethanol completely, the samples were centrifuged at 13,000 rpm for 3 minutes. To elute the RNA, we added RNase-free water, incubated the samples for 3 minutes, and then centrifuged them again.
cDNA was synthesized from 1 μg of mRNA using a high capacity cDNA reverse transcription kit (Applied Biosystems, Foster City, CA, USA). The reverse transcription reaction solution consisted of 2.0 μL 10 × reverse transcription buffer, 0.8 μL 100 mM dNTP mix, 2.0 μL 10 × reverse transcription random primers, and 1.0 μL of MultiScribe reverse transcriptase (Applied Biosystems). The RNA solution was mixed with reverse transcription solution and incubated at 25°C for 10 minutes, 37°C for 120 minutes, and 85°C for 5 seconds.
Quantification of gene expression by real-time PCR
Real-time PCR was carried out on a LightCycler 480 (Roche, Mannheim, Germany) using SYBR Green PCR mix (Clontech, Palo Alto, CA, USA). The specific primer sequences were: SHH (forward) 5′-GAA AGC AGA GAA CTC GGT GG-3′ and (reverse) 5′-GGA AAG TGA GGA AGT CGC TG-3′; PTCH-1 (forward) 5′-CTC CCA AGC AAA TGT ACG AGC A-3′ and (reverse) 5′-TGA GTG GAG TTC TGT GCG ACA C-3′; GLI-1 (forward) 5′-CTC CCG AAG GAC AGG TAT GTA AC-3′ and (reverse) 5′-CCC TAC TCT TTA GGC ACT AGA GTT G-3′; SMOH (forward) 5′-GGG AGG CTA CTT CCT CAT CC-3′ and (reverse) 5′-GGC AGC TGA AGG TAA TGA GC-3′; and GAPDH (forward) 5′-CAC CAC CAA CTG CTT AG-3′ and (reverse) 5′-CTT CAC CAC CTT CTT GAT G-3′. The housekeeping gene GAPDH was used as a loading control. PCR conditions were as follows: one cycle at 95°C for one minute followed by 40 cycles at 95°C for 10 seconds, 58°C for 5 seconds, and 72°C for 20 seconds. The specificity of the PCR products was tested by dissociation curves. The crossing points of primer probes were normalized to GAPDH. Relative values of transcripts were calculated using the equation 2 – ΔΔCt, where ΔCt is equal to the difference in crossing point for target and reference and relative quantization according to the following equation:
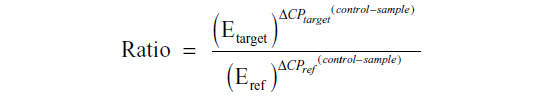
Western blot analysis
All tissues were homogenized. After evaporation of liquid nitrogen, lysis buffer containing protease inhibitor was added. The samples were placed on ice for 10 minutes before centrifugation. The supernatants containing proteins were harvested, and protein concentration was determined using the bicinchoninic acid assay. Protein samples were diluted in buffer, boiled for 10 minutes, loaded onto gels, and electrophoresed. Separated proteins were transferred to polyvinylidene fluoride membranes (Millipore, Bedford, MA, USA). These membranes were then blocked with 5% milk and 0.1% Tween-20 (Sigma-Aldrich, St Louis, MO, USA) in Tris-buffered saline at pH 7.4 for one hour prior to overnight incubation with the primary antibody. The antibodies included a rabbit polyclonal antibody against human SHH (1843-1, 1:1,000, Epitomics, Burlingame, CA, USA), human PTCH-1 (NBP1-71662, 1:500, Novus Biologicals, Littleton, CO, USA), human GLI-1 (AB3444, 1:1000), and human SMOH (NBP1-01011, 1:500, Novus Biologicals), respectively, and a mouse monoclonal antibody against human actin (AC-15, 1:1,000). Blots were washed and incubated with the appropriate peroxidase-conjugated secondary antibody. Immunorecognition was done using enhanced chemiluminescence. Images were captured and analyzed densitometrically using MultiGauge version 3.0 software (Fuji Photo Film Co. Ltd., Tokyo, Japan). The grayscale values of the bands for SHH, PTCH-1, GLI-1, and SMOH were normalized to determine the protein level. The experiments were repeated three times independently.
Ratio measurement
After examination, the ratios of mRNA expression in cancer tissue and in noncancerous tissue were measured for SHH, PTCH-1, GLI-1, and SMOH.
Follow-up
After discharge, the patients were followed up regularly at the outpatient clinic (mean 30.2±14.8 months, range 36–48 months) for periodic assessment, including breast ultrasonography, abdominal ultrasonography, serum CA153 and carcinoembryonic antigen (every 3 months during the first 3 years, and every 6 months thereafter), chest X-ray, and a whole body bone scan (every 6 months for one year, then annually). Magnetic resonance imaging or positron emission tomography-computed tomography (PET-CT) scan was done selectively if there was suspicion of recurrence.
Confirmation of recurrence depends upon two points. One is from the imaging studies, including ultrasonography, chest X-ray, whole body bone scan, magnetic resonance imaging, or PET-CT scan. The other is the tissue proof. Biopsy is a key procedure to establish the pathologic diagnosis, if it is feasible. The pathologist usually performs the immunohistochemical stain, estrogen receptor, progesterone receptor and the Her-2 neu for the detected lesion between recurrence and a second primary breast tumor.
Outcome measures
Irrespective of location, detection of a tumor on any imaging was defined as recurrence.
Statistical analysis
The correlation between recurrence, clinicopathologic variables, and the ratio of each mRNA was analyzed. Comparisons between groups were performed using the chi-square test (or Fisher’s exact test) for continuous variables. Statistical analysis was performed by analysis of variance using Statistical Package for the Social Sciences version 10 software (SPSS Inc, Chicago, IL, USA). Statistical significance was considered at P<0.05.
Results
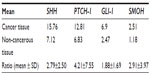
All patients survived, but morbidity was noted in two patients (seroma, subsided after conservative treatment). Two patients (4%) had recurrence during follow-up. Table 2 shows the mean values for SHH mRNA, PTCH-1 mRNA, SMOH mRNA, and GLI-1 mRNA detected in cancer tissue and noncancerous tissue. The mean ratios of SHH mRNA, PTCH-1 mRNA, GLI-1 mRNA, and SMOH mRNA between cancer tissue and noncancerous tissue were 2.79±2.50, 4.21±7.55, 1.88±1.69, and 2.91±3.97, respectively.
Table 3 indicates the correlation between clinicopathologic characteristics and the ratio of each mRNA. The size of invasive cancer correlated significantly with mRNA ratios for SHH (P=0.001), PTCH-1 (P=0.005), and SMOH (P=0.021), respectively (Figure 1). Lymph node involvement correlated significantly with the ratio of SMOH mRNA (P=0.041) while Her-2 neu correlated significantly with the ratio of GLI-1 mRNA (P=0.012).
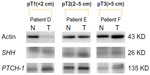
Compared with the mRNA values, the amounts of protein detected by Western blot were relatively small. The mean values for the tumors were 5.26 for SHH, 1.80 for PTCH-1, and 3.16 for GLI-1, whereas those for noncancerous tissue were 3.80 for SHH, 1.55 for PTCH-1, and 1.97 for GLI-1. The amounts of protein detected for SMOH were too small to measure, but were higher in tumors, albeit not significantly so (Figure 2), while the ratio of cancer/non-cancerous tissue of protein for GLI-1 correlated significantly with expression of Her-2 neu (P=0.012). Table 4 shows a comparison of the ratios of mRNA between those with recurrence and those without recurrence. Each ratio is statistically significant (both P<0.001). Table 5 shows the significant factors affecting recurrence, including invasive tumor size (P=0.024), and mRNA ratios for SHH, PTCH-1, GLI-1, and SMOH (P<0.001 for each).
Discussion
Our study shows that, compared with paired noncancerous tissue, a higher expression of SHH mRNA, PTCH-1 mRNA, GLI-1 mRNA, and SMOH mRNA in breast cancer tissue is associated with an increased risk of recurrence (Tables 4 and 5). Excluding GLI-1, all correlated significantly with the size of the invasive cancer (Table 2). In addition, SMOH mRNA correlated with lymph node involvement (Table 2). Both tumor size and lymph node involvement are invasive characteristics and significant prognostic determinants.9 As Souzaki et al have mentioned, the SHH pathway mediates progression from noninvasive cancer to invasive cancer.10
The mRNA for each gene was expressed in over 98% of cancer tissue. This high expression is similar to that reported by Kubo et al11 and Cui et al,12 but different from that reported by Mukherjee et al.13 We attribute this discrepancy to two possible factors. One is the different examination methods used. Real-time PCR is more sensitive than conventional PCR or Western blotting.14 The other reason is the different clinicopathologic stages of the patients. All of our patients had invasive carcinoma of variable size (T1, 54%, Table 1), similar to those in the studies of Kubo et al and Cui et al,11,12 whereas the stage of patients in the study by Mukherjee et al was more advanced.13
GLI-1 is a well known target gene in the SHH pathway.15,16 The expression of nuclear GLI-1 is positively associated with SHH in breast tissues.12 Aberrant activation of the SHH pathway leads GLI-1 into the nucleus, promoting gene transcription and maintaining the biological behavior of cancer.15,16 GLI-1 contributes to the proliferation, survival, and migration of inflammatory breast cancer.17
According to ten Haaf et al, GLI-1 expression is associated with an unfavorable outcome of invasive cancer.18 Similarly, our recurrent cases had a higher expression of GLI-1 mRNA (Tables 4 and 5). Overexpression of GLI-1 in other cancers also adversely affects the risk of recurrence.19,20
Kubo et al reported that nuclear GLI-1 correlates with estrogen receptor status.11 Koga et al mentioned a link between the SHH pathway and estrogen receptor alpha.21 Kameda et al suggested this pathway as a therapeutic target for those with estrogen receptor-negative carcinoma.22 In our cases, GLI-1 did not correlate with estrogen receptor or progesterone receptor status, but correlated with Her-2 neu.
PTCH-1 promotes tumor formation by inhibiting expression of Fas, which mediates apoptosis.15,16,23 PTCH-1 regulates cell cycle progression and predisposes cells to proliferative and expansive behavior.15,16,24 Our patients with recurrence showed overexpression of PTCH-1. High PTCH-1 also affects metastasis of other cancers.19,25 SMOH encoding of a transmembrane protein acting in a receptor complex was affected by PTCH-1.15,16,18,23 Tao et al studied patients having triple-negative breast cancer by immunohistochemical analysis of factors in the SHH pathway.26 They observed that expression of PTCH-1 was significantly decreased in breast cancer compared with mammary hyperplasia. They hypothesized that continuous activation of mutated Sonic Hedgehog signaling may transform breast stem cells to oncogenesis and malignant progression.26
The SHH pathway is activated mainly in cancer stem cells and not in each cancer cell.27 Cancer stem cells capable of initiating and sustaining the cancer are usually highly proliferative and invasive.28–33 Their presence has been reported in various malignancies, including those of the breast.28–35 However, from the perspective of amount, cancer stem cells are only a small population among all cancer cells.27–33 Whether activation of the SHH pathway occurs early or late in carcinogenesis remains a matter of debate.12,36,37 Some research has demonstrated that increased SHH is associated with early pTNM stage, indicating that its upregulation may occur early.33,36,37 Whereas SHH also activates in late or recurrent stages in other malignancies.19,20,37–39 We found that overexpression of these mRNA genes correlates with larger tumor size and recurrence (Tables 3 and 4). We believe that this pathway activates not only early on in cancer stem cells to enhance growth, but also later on to accelerate progression and recurrence. MDA-MB-231 cells expressing SHH enhance chemotactic migration and the degradation/invasion of extracellular matrix proteins.33
In both in vivo and in vitro studies, blockade of SHH pathway may inhibit tumor growth.40 Cancer recurrence is complicated and contributed to by circulating cancer cells detached from the primary tumor, the molecular signature of the cancer, and the susceptibility of the microenvironment of the remnant tissues.41,42 Cui et al showed that the SHH gene is important in vascular formation, thus contributing to recurrence.12 Upregulation of the SHH pathway in a tumor may impact the microenvironment.
For further clinical application, we suggest an mRNA study of PTCH-1 and GLI-1 from core needle biopsy tissue (if feasible) as a reference for prognostication. This would provide useful information for both surgeons and patients before selecting a treatment option. In addition, using specific SHH pathway inhibitors to mitigate cancer recurrence is potentially a therapeutic strategy.40,43
Only two patients had cancer recurrence. We attribute to this to two reasons. One is that the follow-up period is not long, and the other is that the number of study patients is not large, and these are the limitations of this study. However, it does give us information on activation of SHH in the recurrence pathway. From this study, we suggest that overexpression of SHH mRNA, PTCH-1 mRNA, GLI-1 mRNA, and SMOH mRNA in breast cancer tissues is a potential biomarker for prediction of postoperative recurrence.
Acknowledgment
This work was supported by a grant from the Far Eastern Memorial Hospital (FEMH-2011-C-017).
Disclosure
The authors declare that they have no conflicts of interest to report in this work.
References
Parkin DM, Bray F, Ferlay J, Pisani P. Estimating the world cancer burden: Globocan 2000. Int J Cancer. 2001;15(94):153–156. | |
Nechuta S, Lu W, Zheng Y, et al. Comorbidities and breast cancer survival: a report from the Shanghai Breast Cancer Survival Study. Breast Cancer Res Treat. 2013;139(1):227–235. | |
Walterhouse DO, Yoono JW, Iannaccone PM. Developmental pathways: sonic hedgehog-patched-GLI. Environ Health Perspect. 1999;107(3):167–171. | |
Agathocleous M, Locker M, Harris WA, Perron M. A general role of hedgehog in the regulation of proliferation. Cell Cycle. 2007;6(2):156–159. | |
Ingham PW, McMahon AP. Hedgehog signaling in animal development: paradigms and principles. Gene Dev. 2001;15(23):3059–3087. | |
García-Zaragoza E, Pérez-Tavarez R, Ballester A, et al. Intraepithelial paracrine hedgehog signaling induces the expansion of ciliated cells that express diverse progenitor cell markers in the basal epithelium of the mouse mammary gland. Dev Biol. 2012;372(1):28–44. | |
Fiaschi M, Rozell B, Bergström A, Toftgård R, Kleman MI. Targeted expression of GLI1 in the mammary gland disrupts pregnancy-induced maturation and causes lactation failure. J Biol Chem. 2007;282(49):36090–36101. | |
Edge S, Byrd DR, Compton CC, Fritz AG, Greene FL, Trotti A, editors. AJCC Cancer Staging Manual. 7th Ed. New York: Springer; 2010. | |
Georgescu R, Coros MF, Stolnicu S, et al. Prognostic factors in breast cancer. Rev Med Chir Soc Med Nat Iasi. 2012;116(1):262–267. | |
Souzaki M, Kubo M, Kai M, et al. Hedgehog signaling pathway mediates the progression of non-invasive breast cancer to invasive breast cancer. Cancer Sci. 2011;102(2):373–381. | |
Kubo M, Nakamura M, Tasaki A, et al. Hedgehog signaling pathway is a new therapeutic target for patients with breast cancer. Cancer Res. 2004;64(17):6071–6074. | |
Cui W, Wang LH, Wen YY, et al. Expression and regulation mechanisms of sonic hedgehog in breast cancer. Cancer Sci. 2010;101(4):927–933. | |
Mukherjee S, Frolova N, Sadlonova A, et al. Hedgehog signaling and response to cyclopamine differ in epithelial and stromal cells in benign breast and breast cancer. Cancer Biol Ther. 2006;5(6):674–683. | |
Pfaffl MW. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001;29(9):e45. | |
Bale AE, Yu KP. The hedgehog pathway and basal cell carcinomas. Hum Mol Genet. 2001;10(7):757–762. | |
Johnson RL, Rothman AL, Xie J, et al. Human homolog of parched, a candidate gene for the basal cell nevus syndrome. Science. 1996;272(5268):1668–1671. | |
Thomas ZI, Gibson W, Sexton JZ, et al. Targeting GLI1 expression in human inflammatory breast cancer cells enhances apoptosis and attenuates migration. Br J Cancer. 2011;104(10):1575–1586. | |
ten Haaf A, Bektas N, von Serenyi S, et al. Expression of the glioma-associated oncogene homolog (GLI) 1 in human breast cancer is associated with unfavourable overall survival. BMC Cancer. 2009;9:298. | |
Jeng KS, Sheen IS, Jeng WJ, et al. High expression of patched homolog-1 messenger RNA and glioma-associated oncogene-1 messenger RNA of sonic hedgehog signaling pathway indicates a risk of postresection recurrence of hepatocellular carcinoma. Ann Surg Oncol. 2013;20(2):464–473. | |
Mori Y, Okumura T, Tsunoda S, Sakai Y, Shimada Y. GLI-1 expression is associated with lymph node metastasis and tumor progression in esophageal squamous cell carcinoma. Oncology. 2006;70(5):378–389. | |
Koga K, Nakamura M, Nakashima H, et al. Novel link between estrogen receptor alpha and hedgehog pathway in breast cancer. Anticancer Res. 2008;28(2A):731–740. | |
Kameda C, Tanaka H, Yamasaki A, et al. The Hedgehog pathway is a possible therapeutic target for patients with estrogen receptor-negative breast cancer. Anticancer Res. 2009;29(3):871–879. | |
Athar M, Li C, Tang X, et al. Inhibition of smoothened signaling prevents ultraviolet B-induced basal cell carcinomas through regulation of Fas expression and apoptosis. Cancer Res. 2004;64(20):7545–7552. | |
Adolphe C, Hetherington R, Ellis T, Wainwright B. Patched1 functions as a gatekeeper by promoting cell cycle progression. Cancer Res. 2006;66(4):2081–2088. | |
You S, Zhou J, Chen S, et al. PTCH1, a receptor of hedgehog signaling pathway, is correlated with metastatic potential of colorectal cancer. Ups J Med Sci. 2010;115(3):169–175. | |
Tao Y, Mao J, Zhang Q, Li L. Overexpression of Hedgehog signaling molecules and its involvement in triple-negative breast cancer. Oncol Lett. 2011;2(5):995–1001. | |
Tian F, Mysliwietz J, Ellwart J, Gamarra F, Huber RM, Bergner A. Effects of the Hedgehog pathway inhibitor GDC-0449 on lung cancer cell lines are mediated by side populations. Clin Exp Med. 2012;12(1):25–30. | |
Mishra L, Banker T, Murray J, et al. Liver stem cells and hepatocellular carcinoma. Hepatology. 2009;49(1):318–329. | |
Chiba T, Kita K, Zheng YW, et al. Side population purified from hepatocellular carcinoma cells harbors cancer stem cell-like properties. Hepatology. 2006;44(1):240–251. | |
Suetsugu A, Nagaki M, Aoki H, Motohashi T, Kunisada T, Moriwaki H. Characterization of CD133+ hepatocellular carcinoma cells as cancer stem/progenitor cells. Biochem Biophys Res Commun. 2006;351(4):820–824. | |
Ma S, Chan KW, Hu L, et al. Identification and characterization of tumorigenic liver cancer stem/progenitor cells. Gastroenterology. 2007;132(7):2542–2556. | |
Yang ZF, Ngai P, Ho DW, et al. Identification of local and circulating cancer stem cells in human liver. Hepatology. 2008;47(3):919–928. | |
Lu ZH, Jia J, Ren J, Ma B, DI LJ, Song GH. [Detection of breast cancer stem cells and the expression of key molecules in hedgehog signaling pathway]. Beijing Da Xue Xue Bao. 2008;40(5):480–485. Chinese. | |
Li Z, Liu Y, Tuve S, et al. Toward a stem cell gene therapy for breast cancer. Blood. 2009;113(2):5423–5433. | |
Clayton S, Mousa SA. Therapeutics formulated to target cancer stem cells: is it in our future? Cancer Cell Int. 2011;11:7. | |
Huang S, He J, Zhang X, et al. Activation of the hedgehog pathway in human hepatocellular carcinomas. Carcinogenesis. 2006;27(7):1334–1340. | |
Thayer SP, di Magliano MP, Heiser PW, et al. Hedgehog is an early and late mediator of pancreatic cancer tumorigenesis. Nature. 2003;425(6960):851–856. | |
Harris LG, Pannell LK, Singh S, Samant RS, Shevde LA. Increased vascularity and spontaneous metastasis of breast cancer by hedgehog signaling mediated upregulation of cyr61. Oncogene. 2012;31(28):3370–3380. | |
Kim JY, Ko GH, Lee YJ, et al. Prognostic value of sonic hedgehog protein expression in gastric cancer. Jpn J Clin Oncol. 2012;42(11):1054–1059. | |
Jeng KS, Sheen IS, Jeng WJ, et al. Blockade of sonic hedgehog pathway effectively inhibits the growth of hepatoma in mice – an “in vivo” study. Oncol Lett. 2012;4(6):1158–1162. | |
Meng S, Tripathy D, Frenkel EP, et al. Circulating tumor cells in patients with breast cancer dormancy. Clin Cancer Res. 2004;10(24):8152–8162. | |
Méhes G, Witt A, Kubista E, Ambros PF. Circulating breast cancer cells are frequently apoptotic. Am J Pathol. 2001;159(1):17–20. | |
Zhang X, Harrington N, Moraes RC, Wu MF, Hilsenbeck SG, Lewis MT. Cyclopamine inhibition of human breast cancer cell growth independent of Smoothened (Smo). Breast Cancer Res Treat. 2009;15(3):505–521. |
 © 2013 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2013 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.