Back to Journals » OncoTargets and Therapy » Volume 13
High Expression of miR-196b Predicts Poor Prognosis in Patients with Ovarian Cancer
Authors Li Y, Li J, Liu Z, Zhang Y
Received 27 March 2020
Accepted for publication 20 August 2020
Published 1 October 2020 Volume 2020:13 Pages 9797—9806
DOI https://doi.org/10.2147/OTT.S254942
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr William C. Cho
Yang Li,1 Jing Li,2 Zirong Liu,1 Yamin Zhang1
1Department of Hepatobiliary Surgery, Tianjin First Central Hospital, Tianjin 300192, People’s Republic of China; 2Department of Gynaecology, Tianjin First Central Hospital, Tianjin 300192, People’s Republic of China
Correspondence: Yamin Zhang Department of Hepatobiliary Surgery
Tianjin First Central Hospital, Tianjin 300192, People’s Republic of China
Email [email protected]
Background/Aims: To analyze the clinical significance of miR-196b expression in ovarian cancer and predict the function and possible mechanism of miR-196b.
Methods: Both Kaplan–Meier (K-M) and Cox proportional hazards regression model were used to analyze the prognostic factors of patients with ovarian cancer. MiR196-b was modulated in ovarian cancer cells, and the cell viability, cell cycle, and cell cycle-related gene expression were analyzed. The target genes of miR-196b were then predicted and checked the relationship between the target genes.
Results: MiR-196b was an independent risk factor, while high expression of miR-196b was associated with poor prognosis of ovarian cancer. MiR-196b overexpression increased cancer cell proliferation. Cdkn1b, as one of the targets of miR-196b, was related to cell viability and mitosis.
Conclusion: High expression of miR-196b was significantly associated with poor prognosis of the patients with ovarian cancer. MiR-196b could increase the cell proliferation of ovarian cancer by modulating Cdkn1b expression.
Keywords: microRNAs, MiR-196b, ovarian cancer, Cdkn1b, proliferation
Introduction
Ovarian cancer is a common gynecological malignant tumor in the gynecological. In developed and developing countries, ovarian cancer is expected to rank fifth and eighth, respectively, in new cancer cases, with the sixth and eighth mortality rates each year.1 Because most patients with ovarian cancer are diagnosed at an advanced stage, the prognosis is poor, and the 5-year survival rate is only 45%.2 Although only 30% of patients with distant metastatic ovarian cancer can survive for 5 years or longer, this proportion is higher than advanced cervical cancer, colon cancer, kidney cancer, and pancreatic cancer.3 It is, therefore, necessary to look for a prognosis Indicator of ovarian cancer. MicroRNAs (miRNAs) are non-coding short-chain (19–22 nt) RNAs that regulate gene expression at transcriptional and/or post-transcriptional levels.4 Approximately 60% of mRNA is currently believed to be controlled by microRNAs, which play a major role in regulating tumorigenesis, growth, proliferation, migration, invasion, apoptosis, etc.4,5 miR-196b has a different function in various tumors. For example, high expression in gastric cancer can promote the metastasis and proliferation of gastric cancer.6 However, it can prevent tumor proliferation, invasion, and migration in cervical cancer.7 In ovarian cancer, miR-196b has been found to be highly expressed in recurrent ovarian cancer than in primary ovarian cancer. MiR-196b may be used as an indicator for predicting recurrent ovarian cancer, but research related to the prognosis of ovarian cancer is very rare.8 In this study, we analyzed the ovarian cancer dataset based on the TCGA database, examined the clinical significance of miR-196b expression in ovarian cancer, and looked for potential new target for prognosis prediction in patients with ovarian cancer.
Methods and Information
Patients and Samples
Level 3 data for miRNAseq, mRNAseq, and clinical data with ovarian cancer have been downloaded from the TCGA database. Patients participating in this study had data regarding overall survival (OS), miRNAseq, and mRNA seq. The sample data included the age of diagnosis, the largest diameter of the tumor, the stage and grade of the tumor, and the presence or absence of residual tumors in the study of the clinical characteristics. After sorting the data, we obtained 492 cases of miR-196b expression, and clinical feature matching data, 414 cases of gene expression and clinical characteristics matching data, 414 cases of gene expression, and microRNAs expression matching data.
Cell Culture and Transfection
The human ovarian cancer cell line CAOV3 (ATCC, USA) was cultured with Dulbecco’s Modified Eagle’s Medium (Gibco, Thermo Fisher Scientific Inc., USA) added with 10% fetal bovine serum (FBS, Gibco, Thermo Fisher Scientific Inc., USA) and A2780 (Sigma, Mo. the USA) were grown in RPMI-1640 medium (Sigma, Mo. the USA) supplemented by 10% FBS and 2mM Glutamax (Thermo Fisher Scientific Inc., USA). Both cell lines maintained less than ten passage times.
MiR-196b mimic (miR-196b+), miRNA mimic negative control #1 (miR-C), miR-196b inhibitor (miR-196b-), miRNA inhibitor negative control (miR-C-), Small interfering RNA (siRNA) against Cdkn1b (si-Cdkn1b), siRNA negative control (si-Ctrl) were all obtained from Guangzhou RIBOBIO Company (Guangzhou, China)(The final concentration was 50nM). Cell transfection was performed by using Lipofectamine 2000 (Invitrogen, Thermo Fisher Scientific Inc., USA) as advised by the manufacturer.
Target Site Cloning and Luciferase Reporter Assays
In order to investigate whether miR-196b modulates Cdkn1b expression by binding to its 3ʹ untranslated region (3ʹUTR), luciferase assays were performed using a psiCHECK2 plasmid (Promega, Madison, WI). The Cdkn1b 3ʹUTR was cloned into the luciferase vector psiCHECK2. Based on the starBase (v2.0) database, miR196b has a binding site for 3ʹUTR of Cdkn1b. Mutagenesis of Cdkn1b binding sites was performed using the KOD Plus Mutagenesis kit (Toyobo, Osaka, Japan) based on the instructions. The sequence of the clone has been verified by DNA sequencing. CAOV3 and A2780 cells were seeded in 6-well plates and then transfected with psiCHECK2-Cdkn1b containing the 3ʹUTR of Cdkn1b and psiCHECK2-Cdkn1b-Mut containing mutated miR-196b binding sites in the Cdkn1b 3ʹUTR. Transfection was performed in the presence of miR-196b mimic or miRNA mimic negative control by using Lipofectamine 2000. The cells were harvested forty-eight hours after the transfection. Renilla luciferase activity was assessed using a Lumat LB9501 instrument (Berthold, Bad Wildbad, Germany), and the results were normalized using firefly luciferase activity.
Cell Viability
Cell counting kit-8 (CCK8, Dojindo Molecular Technologies, Inc. Tokyo, Japan) was used to check cell viability. Cells were incubated with a 10% CCK8-culture medium for two hours in an incubator. Absorbance was measured at 450 nm using a plate reader (SpectraMax i3, Molecular Devices, Tokyo, Japan).
Cell-Cycle Analysis by Flow Cytometry
Cells were harvested and washed twice with ice-cold PBS and capped with 70% ethanol. 1x106 cells were suspended in 500 ul of RNase A/PI solution (20ug/mL RNase A and 50 ug/mL PI in PBS) and incubated at 4°C for thirty minutes in the dark. These cells have been analyzed by flow cytometry.
RT-PCR Analysis
The RNA, including miRNA of each sample, was harvested by using the RNeasy Mini Kit (Qiagen, Hilden, Germany) as recommended by the manufacturer. cDNA was then synthesized by using a reverse transcription kit (Applied Biosystems, Foster City, CA, USA) or TaqMan microRNA Reverse Transcription Kit (Applied Biosystems, Foster City, CA, USA). The following TaqMan assays (assay ID) were used: cyclin-dependent kinase (CDK) 4 (Hs00123089_CE), cyclin D1 (Hs00108480_CE), miR-196b (002215). GAPDH (Hs00123089_CE) or U6 (001973) was selected as internal controls. StepOnePlus Real-Time polymerase chain reaction (PCR) System (Applied Biosystems) has been used to perform qRT-PCR.
Western Blotting
Cells were lysed using the RIPA buffer (Thermo, MA, USA) containing both the protease inhibitor cocktail (Sigma, MO, USA) and the PhosSTOP phosphatase inhibitor cocktail (Roche, Tokyo, Japan). Protein concentrations were then measured using the BCA kit (Thermo, MA, USA), and equal amounts of extracted proteins were separated by 10% SDS-PAGE gels and transferred to PVDF membranes (Bio-Rad, USA). The membranes were incubated either with the Cdkn1b (1:2500, 3698, Cell Signaling Technology, Inc. Tokyo, Japan) or β-actin (1:3000, 3700, Cell Signaling Technology, Inc. Tokyo, Japan) primary antibody and appropriate HRP-conjugated secondary antibody (1:3000). Chemiluminescence (GE, UK) was used to detect the proteins.
Data Analysis
Statistical analysis of patients with ovarian cancer data was performed using SPSS (21.0, IBM, NY, USA). Survival difference analysis was performed using the Kaplan–Meier method and Log rank test. Multivariate Cox proportional hazard models have been used to analyze risk factors for prognosis. Student’s t-test or One-way ANOVA was used to compare the difference between groups. Bilateral P values < 0.05 have been statistically significant.
Results
Clinical Features of TCGA Dataset in Patients with Ovarian Cancer
The basic clinical data for these patients are summarized in Table 1. The initial tissue diagnosis of patients with ovarian cancer had an average of 59.7 years (30–87 years), as listed in Table 2. Among them, bilateral patients accounted for 69.5%, stage III, and IV patients accounted for 93.1%, G3 and G4 accounted for 84.8%, and no residual tumors accounted for 19.5% after diagnosis. The data suggest that it is difficult to diagnose the disease at an early stage, and most of the patients with ovarian cancer are detected at the later stage.
 |
Table 1 Data Definition Rules |
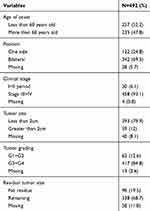 |
Table 2 Clinical Data of 492 Patients with Ovarian Cancer |
The Expression of miR-196b is an Independent Risk Factor for Predicting the Prognosis of Patients with Ovarian Cancer
After analyzing the relationship between expression levels of microRNA and OS of ovarian cancer patients, we found that patients with high expression of miR-196b had a poor prognosis. K-M and Cox proportional hazard models have been used to analyze whether miR-196b is a reliable indicator for determining the prognosis of ovarian cancer patients. In the single-factor K-M survival analysis results (Table 3), the age of patients (P = 0.04), clinical-stage (P = 0.011), residual tumors (P < 0.001), and miR-196b (P = 0.004) affected the survival of patient. The multivariate Cox proportional hazard analysis model, as shown in Table 4, suggested that miR-196b, clinical stage, and residual tumors are independent variables that influence the prognosis of patients with ovarian cancer. Figure 1 shows the K-M curve of miR-196b. Results suggest that patients with low expression of miR-196b have a longer OS (log-rank, P < 0.05).
 |
Table 3 Single-Factor Survival Analysis of 492 Patients with Ovarian Cancer |
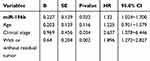 |
Table 4 Multivariate Cox Proportional Risk Regression Analysis in 492 Patients with Ovarian Cancer |
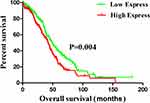 |
Figure 1 The overall survival of patient with ovarian cancer cut-off by miR-196b expression. |
MiR-196b Increased Cancer Cell Proliferation
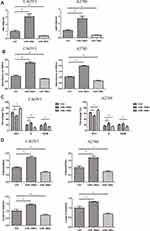
To detect the effect of miR-196b on ovarian cancer, we have overexpressed or suppressed miR196b in ovarian cancer cells CAOV3 and A2780. The overexpression and suppression of the miR-196b were confirmed by PCR analysis, as presented in Figure 2A. After the overexpression of miR-196b, cell viability was significantly increased compared to the control group. In contrast, the suppression of miR-196b remarkably decreased cell viability, as shown in Figure 2B. Further, the distribution of cell cycle revealed that after miR-196b overexpression, more cells were in the S phase, and many more cells remained in the G0/1 phase when miR-196b was suppressed, as compared to the control (Figure 2C). When miR-196b was modulated, the expression of CDK4 and cyclin D1 showed changes, as shown in Figure 2D.
Cdkn1b as the Target Modulated by miR-196b
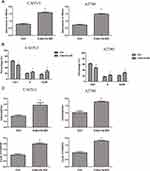
Based on the previous report, Cdkn1b (p27Kip1), known as the tumor-suppressive protein, could be the target of miR-196b.9 After the miR-196b mimic., we first checked the Cdkn1b expression by PCR and found that Cdkn1b expression was significantly decreased. Besides, the protein level of Cdkn1b was found to be reduced after miR-196b overexpression, as presented in Figure 3A and B.
A luciferase reporter system was used to confirm the relationship between miR-196b and Cdkn1b. Figure 3C depicts the pairing between the targets miR-196b and Cdkn1b. The predicted miR-196b recognition sites in the Cdkn1b 3′ UTRs were cloned into a psiCHECK2 plasmid (psiCHECK2-Cdkn1b WT). Furthermore, site-directed mutagenesis (pmirCHECK2-Cdkn1b MT) generated the mutated (MT) miR-196b binding sites of Cdkn1b. Then, the vectors were co-transfected into CAOV3 and A2780, which were already transfected with pre-miR-196b or pre-miR-control. MiR-196b inhibited luciferase activity in cells transfected with psiCHECK2-Cdkn1b WT. In contrast, transfection of miR-196b did not reduce luciferase activity in cells transfected with psiCHECK2-Cdkn1b MT vector, while the transfection of pre-miR-control did not reduce luciferase activity in cells transfected with WT or MT (Figure 3D).
MiR-196b Modulated Cancer Cell Proliferation Through Cdkn1b
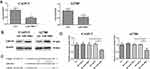
We found whether the miR196b modulated cancer cell proliferation through Cdkn1b. It was observed that Cdkn1b was destroyed by CAOV3 and A2780 cells through the siRNA transfection of Cdkn1b. Thus, cell proliferation remarkably increased after the suppression of Cdkn1b, as depicted in Figure 4A. The cell-cycle analysis also showed that the percentage of S and G2/M phase increased after suppressing Cdkn1b (Figure 4B). The PCR analysis of cell cycle-related genes further confirmed that Cdkn1b could inhibit cancer cell proliferation (Figure 4C).
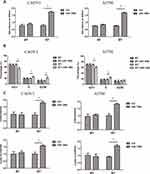
To clarify the relationship between miR-196b and Cdkn1b on cancer cell proliferation, transfection of miR-196b in Cdkn1b 3ʹ-UTR wild or mutated CAOV3 and A2780 cells was performed. The cell viability showed that the overexpression of miR-196b did not affect the cell proliferation of Cdkn1b 3ʹ-UTR mutated cancer cells in the same way, as did in Cdkn1b 3ʹ-UTR wild types (Figure 5A). The cell cycle distribution and cell cycle-related genes expression also showed the same tendency (Figure 5B and C). These demonstrate that the miR-196b triggers cancer cell proliferation by inhibiting Cdkn1b expression.
Discussion
In the human genome, there are only about 2% of gene-expressing proteins, and in recent years, more and more attention has been paid to gene functions in the remaining 98% non-protein-coding region.10 MicroRNAs are non-coding small RNAs that play an essential role in the development of tumors. Research on microRNAs is expected to set new targets for the diagnosis and treatment of tumors.
The miR-196 family includes miR-196a and miR-196b, and the mature miR-196a (3ʹ-GGGUUGUUGUACUUUGAUGGAU-5ʹ) has a base, which is different from miR-196b (3ʹ-GGGUUGUUGUCCUU-UGAUGGAU-5ʹ).11 Although MiR-196b is abnormally expressed in various tumors, its role in different types of tumors is still controversial. Studies have reported that high expression of miR-196b in osteosarcoma,12 gastric cancer,6 malignant glioma,13 pancreatic cancer,14 breast cancer,15 MLL-reconstituted leukemia,16 esophageal cancer,17 colon cancer,18 and lung cancer19 is associated with poor prognosis, indicating that miR-196b may have a cancer-promoting effect. The low expression of miR-196b in melanoma,20,21 cervical cancer,7 and liver cancer,22 is associated with poor prognosis, suggesting that it may have a tumor suppressor effect. In a recent study of ovarian cancer, the expression of miR-196b in recurrent ovarian cancer is higher than that in primary ovarian cancer,8 which could be a marker for recurrence of ovarian cancer. One study reported that overexpression of miR-196b may contribute to invasion activities of cancer cells in recurrent epithelial ovarian cancer by regulating the Homeobox A9 gene.23 The methylation status of the miR-196b promoter region affects the level of miR-196b expression in tumors. In the study of gastric cancer, oral cancer, and melanoma, hypomethylation of the miR-196b promoter region increased the expression of miR-196b.6,24 Another interesting study showed that 17 beta-estradiol (E(2)) alters microRNA (miRNA) expression profiles in the adult zebrafish (Danio rerio) and miR-196b was one of the most up-regulated miRNAs.25 Hypermethylation of miR-196b promoter region in chronic leukemia, malignant glioma, prostate cancer, hepatocellular carcinoma, cervical cancer reduces miR-196b expression.7,26–29
A variety of miR-196b target genes have recently been identified, including HOXA9, c-Myc, RUNX2, HOXB7, SOCS2.19,30–32 MiR-196b plays a role in different tumors by regulating these target genes. In this study, we identified that Cdnk1b is negatively correlated with miR-196b. Cdkn1b, cyclin-dependent kinase inhibitor 1B (p27kip1), is a cyclin/cyclin-dependent kinase complex inhibitor.33,34 Reduced levels of Cdkn1b have been observed in cancers, including premalignant ovarian tumors.35,36 As shown in our present research, miR-196b, which is highly expressed in ovarian tumors, is negatively correlated with Cdkn1b expression. Cdkn1b is involved in cell differentiation, proliferation, apoptosis, cell-cell adhesion, and growth inhibition.37 In our research, suppressed Cdkn1b reduced the cancer cell proliferation by inhibiting cell mitosis, which has been shown to be a cell-cycle stop, including down-regulation of cell-cycle related genes.
Conclusion
miR-196b was significantly associated with overall survival in patients with ovarian cancer, and ovarian cancer patients with high expression of miR-196b had a poor prognosis. The results of this study suggested that miR-196b might be a new marker for prognosis prediction. At the same time, Besides, the miR-196b facilitated the proliferation of cancer cells by targeting Cdkn1b.
Disclosure
The authors declare that they have no competing interests in this work.
References
1. Bray F, Ferlay J, Soerjomataram I, et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424. doi:10.3322/caac.21492
2. Jemal A, Siegel R, Ward E, et al. Cancer statistics, 2008. CA Cancer J Clin. 2008;58:71–96. doi:10.3322/CA.2007.0010
3. Keegan TH, Ries LA, Barr RD, et al. Comparison of cancer survival trends in the United States of adolescents and young adults with those in children and older adults. Cancer. 2016;122:1009–1016. doi:10.1002/cncr.29869
4. Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–297. doi:10.1016/S0092-8674(04)00045-5
5. Bracken CP, Scott HS, Goodall GJ. A network-biology perspective of microRNA function and dysfunction in cancer. Nat Rev Genet. 2016;17:719–732.
6. Lee SW, Park KC, Kim JG, et al. Dysregulation of MicroRNA-196b-5p and MicroRNA-375 in gastric cancer. J Gastric Cancer. 2016;16:221–229. doi:10.5230/jgc.2016.16.4.221
7. How C, Hui AB, Alajez NM, et al. MicroRNA-196b regulates the homeobox B7-vascular endothelial growth factor axis in cervical cancer. PLoS One. 2013;8:e67846. doi:10.1371/journal.pone.0067846
8. Chong GO, Jeon HS, Han HS, et al. Differential MicroRNA expression profiles in primary and recurrent epithelial ovarian cancer. Anticancer Res. 2015;35:2611–2617.
9. Meyer SE, Muench DE, Rogers AM, et al. miR-196b target screen reveals mechanisms maintaining leukemia stemness with therapeutic potential. J Exp Med. 2018;215:2115–2136. doi:10.1084/jem.20171312
10. Alexander RP, Fang G, Rozowsky J, et al. Annotating non-coding regions of the genome. Nat Rev Genet. 2010;11:559–571. doi:10.1038/nrg2814
11. Chen C, Zhang Y, Zhang L, et al. MicroRNA-196: critical roles and clinical applications in development and cancer. J Cell Mol Med. 2011;15:14–23. doi:10.1111/j.1582-4934.2010.01219.x
12. Zhang C, Yao C, Li H, et al. Combined elevation of microRNA-196a and microRNA-196b in sera predicts unfavorable prognosis in patients with osteosarcomas. Int J Mol Sci. 2014;15:6544–6555. doi:10.3390/ijms15046544
13. Awuah PK, Rhieu BH, Singh S, et al. beta-Catenin loss in hepatocytes promotes hepatocellular cancer after diethylnitrosamine and phenobarbital administration to mice. PLoS One. 2012;7:e39771. doi:10.1371/journal.pone.0039771
14. Zhang Y, Li M, Wang H, et al. Profiling of 95 microRNAs in pancreatic cancer cell lines and surgical specimens by real-time PCR analysis. World J Surg. 2009;33:698–709. doi:10.1007/s00268-008-9833-0
15. Li Y, Zhang M, Chen H, et al. Ratio of miR-196s to HOXC8 messenger RNA correlates with breast cancer cell migration and metastasis. Cancer Res. 2010;70:7894–7904. doi:10.1158/0008-5472.CAN-10-1675
16. Popovic R, Riesbeck LE, Velu CS, et al. Regulation of mir-196b by MLL and its overexpression by MLL fusions contributes to immortalization. Blood. 2009;113:3314–3322. doi:10.1182/blood-2008-04-154310
17. Warnecke-Eberz U, Chon SH, Holscher AH, et al. Exosomal onco-miRs from serum of patients with adenocarcinoma of the esophagus: comparison of miRNA profiles of exosomes and matching tumor. Tumour Biol. 2015;36:4643–4653. doi:10.1007/s13277-015-3112-0
18. Ge J, Chen Z, Li R, et al. Upregulation of microRNA-196a and microRNA-196b cooperatively correlate with aggressive progression and unfavorable prognosis in patients with colorectal cancer. Cancer Cell Int. 2014;14:128. doi:10.1186/s12935-014-0128-2
19. Yu SL, Lee DC, Sohn HA, et al. Homeobox A9 directly targeted by miR-196b regulates aggressiveness through nuclear Factor-kappa B activity in non-small cell lung cancer cells. Mol Carcinog. 2016;55:1915–1926. doi:10.1002/mc.22439
20. Braig S, Mueller DW, Rothhammer T, et al. MicroRNA miR-196a is a central regulator of HOX-B7 and BMP4 expression in malignant melanoma. Cell Mol Life Sci. 2010;67:3535–3548. doi:10.1007/s00018-010-0394-7
21. Micevic G, Muthusamy V, Damsky W, et al. DNMT3b modulates melanoma growth by controlling levels of mTORC2 component RICTOR. Cell Rep. 2016;14:2180–2192. doi:10.1016/j.celrep.2016.02.010
22. Rebucci M, Sermeus A, Leonard E, et al. miRNA-196b inhibits cell proliferation and induces apoptosis in HepG2 cells by targeting IGF2BP1. Mol Cancer. 2015;14:79. doi:10.1186/s12943-015-0349-6
23. Chong GO, Jeon HS, Han HS, et al. Overexpression of microRNA-196b accelerates invasiveness of cancer cells in recurrent epithelial ovarian cancer through regulation of homeobox A9. Cancer Genomics Proteomics. 2017;14:137–141.
24. Hou YY, You JJ, Yang CM, et al. Aberrant DNA hypomethylation of miR-196b contributes to migration and invasion of oral cancer. Oncol Lett. 2016;11:4013–4021. doi:10.3892/ol.2016.4491
25. Cohen A, Shmoish M, Levi L, et al. Alterations in micro-ribonucleic acid expression profiles reveal a novel pathway for estrogen regulation. Endocrinology. 2008;149:1687–1696. doi:10.1210/en.2007-0969
26. Liu Y, Zheng W, Song Y, et al. Low expression of miR-196b enhances the expression of BCR-ABL1 and HOXA9 oncogenes in chronic myeloid leukemogenesis. PLoS One. 2013;8:e68442. doi:10.1371/journal.pone.0068442
27. Shen J, Wang S, Zhang YJ, et al. Genome-wide aberrant DNA methylation of microRNA host genes in hepatocellular carcinoma. Epigenetics. 2012;7:1230–1237. doi:10.4161/epi.22140
28. Lakomy R, Sana J, Hankeova S, et al. MiR-195, miR-196b, miR-181c, miR-21 expression levels and O-6-methylguanine-DNA methyltransferase methylation status are associated with clinical outcome in glioblastoma patients. Cancer Sci. 2011;102:2186–2190. doi:10.1111/j.1349-7006.2011.02092.x
29. Hulf T, Sibbritt T, Wiklund ED, et al. Discovery pipeline for epigenetically deregulated miRNAs in cancer: integration of primary miRNA transcription. BMC Genomics. 2011;12:54. doi:10.1186/1471-2164-12-54
30. Ren W, Wu S, Wu Y, et al. MicroRNA-196a/-196b regulate the progression of hepatocellular carcinoma through modulating the JAK/STAT pathway via targeting SOCS2. Cell Death Dis. 2019;10:333. doi:10.1038/s41419-019-1530-4
31. Bai X, Meng L, Sun H, et al. MicroRNA-196b inhibits cell growth and metastasis of lung cancer cells by targeting Runx2. Cell Physiol Biochem. 2017;43:757–767. doi:10.1159/000481559
32. Abe W, Nasu K, Nakada C, et al. miR-196b targets c-myc and Bcl-2 expression, inhibits proliferation and induces apoptosis in endometriotic stromal cells. Hum Reprod. 2013;28:750–761. doi:10.1093/humrep/des446
33. Guan X, Chen L, Wang J, et al. Mutations of phosphorylation sites Ser10 and Thr187 of p27Kip1 abolish cytoplasmic redistribution but do not abrogate G0/1 phase arrest in the HepG2 cell line. Biochem Biophys Res Commun. 2006;347:601–607. doi:10.1016/j.bbrc.2006.06.114
34. Slingerland J, Pagano M. Regulation of the cdk inhibitor p27 and its deregulation in cancer. J Cell Physiol. 2000;183:10–17. doi:10.1002/(SICI)1097-4652(200004)183:1<10::AID-JCP2>3.0.CO;2-I
35. Sui L, Dong Y, Ohno M, et al. Implication of malignancy and prognosis of p27(kip1), Cyclin E, and Cdk2 expression in epithelial ovarian tumors. Gynecol Oncol. 2001;83:56–63. doi:10.1006/gyno.2001.6308
36. Fredersdorf S, Burns J, Milne AM, et al. High level expression of p27(kip1) and cyclin D1 in some human breast cancer cells: inverse correlation between the expression of p27(kip1) and degree of malignancy in human breast and colorectal cancers. Proc Natl Acad Sci U S A. 1997;94:6380–6385. doi:10.1073/pnas.94.12.6380
37. He W, Wang X, Chen L, et al. A crosstalk imbalance between p27(Kip1) and its interacting molecules enhances breast carcinogenesis. Cancer Biother Radiopharm. 2012;27:399–402. doi:10.1089/cbr.2010.0802
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.