Back to Journals » Cancer Management and Research » Volume 13
GSG2 Promotes Development and Predicts Poor Prognosis of Ovarian Cancer
Authors Huang Y, Liu Y, Zhu K, Ma X, Lu R, Zhang M
Received 30 July 2020
Accepted for publication 17 December 2020
Published 19 January 2021 Volume 2021:13 Pages 499—508
DOI https://doi.org/10.2147/CMAR.S274807
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 4
Editor who approved publication: Dr Xueqiong Zhu
Yan Huang,1,2,* Yixuan Liu,2,3,* Keyu Zhu,2,3,* Xiaolu Ma,2,3 Renquan Lu,2,3 Meiqin Zhang1,2
1Department of Gynecologic Oncology, Fudan University Shanghai Cancer Center, Fudan University, Shanghai, People’s Republic of China; 2Department of Oncology, Shanghai Medical College, Fudan University, Shanghai, People’s Republic of China; 3Department of Clinical Laboratory, Fudan University Shanghai Cancer Center, Fudan University, Shanghai, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Renquan Lu
Department of Clinical Laboratory, Fudan University Shanghai Cancer Center, 270 Dong’An Road, Shanghai 200032, People’s Republic of China
Tel +86 18017312789
Email [email protected]
Meiqin Zhang
Department of Gynecologic Oncology, Fudan University Shanghai Cancer Center, 270 Dong’An Road, Shanghai 200032, People’s Republic of China
Tel +86 18017312218
Fax +86 21-64175590
Email [email protected]
Purpose: Ovarian cancer is one of the most common malignant tumors in gynecology, whose treatment was seriously limited by the unclear understanding of molecular mechanism in disease development. GSG2, also known as Haspin, is a novel molecule found to be involved in human cancers.
Materials and Methods: In this study, immunohistochemical analysis was used to detect GSG2 expression in ovarian cancer tissues and corresponding normal tissues. Statistical analysis was performed to construct relationship between GSG2 and tumor characteristics as well as prognosis. Ovarian cell model with GSG2 knockdown was constructed through lentivirus-mediated transfection of shRNA, which was used in MTT assay, colony formation assay and flow cytometry for investigating the role of GSG2 in ovarian cancer. A human apoptosis antibody array was used to identify potential downstream apoptosis-related proteins of GSG2.
Results: The results demonstrated the upregulation of GSG2 in ovarian cancer, whose expression was positively related to tumor grade and AJCC stage, and negatively correlated with patients’ prognosis. Moreover, knockdown of GSG2 inhibited ovarian cancer development through suppressing cell growth and inducing cell apoptosis. Further exploration revealed that a variety of apoptosis-related and PI3K signaling pathway-related proteins may be implicated in the GSG2 induced regulation of ovarian cancer.
Conclusion: In summary, it was illustrated that GSG2 was involved in the development of ovarian cancer, which has the potential to become therapeutic target and prognostic indicator in ovarian cancer treatment.
Keywords: ovarian cancer, GSG2, prognosis, cell proliferation, cell apoptosis
Introduction
Ovarian cancer is one of the most common malignant tumors in gynecology. The incidence of ovarian cancer ranks third, while the mortality rate is the highest among female malignant tumors of the reproductive system.1 Ovarian cancer has the characteristics of concealed onset, high malignancy, and easy metastasis, which causes its poor prognosis.2 Although early-stage ovarian cancer is relatively easy to cure, patients are often in the middle and late stages of the disease at the time of diagnosis because the ovary is located in the deep pelvic cavity and has no specific clinical symptoms in the early stage of the disease.3 Although the treatment of ovarian cancer such as surgery, radiotherapy and chemotherapy has been greatly developed and advanced in recent years, the prognosis of advanced ovarian cancer has not been significantly improved. In recent years, the emergence of molecular-targeted therapy has brought a glimmer of light to the treatment of ovarian cancer, especially advanced ovarian cancer.4 Although researchers have developed a variety of targeted drugs for ovarian cancer and applied them in clinical treatment, the improvement of prognosis is still far from satisfactory.5,6 Therefore, in-depth analysis of the molecular mechanism of ovarian cancer development and the search for regulatory factors that play an important role as drug targets are of great significance for improving the accuracy and efficiency of ovarian cancer targeted therapy.5
GSG2 was first discovered in mouse sperm cells and was later defined as the protein kinase Haspin. The C-terminus of GSG2 has an atypical kinase domain.7 The crystal structure of GSG2 shows that the domain is always active and open, and there is no activation sequence of the conventional kinase domain.8 During mitosis, GSG2 can specifically phosphorylate the third threonine of histone (H3T3) to localize Aurora B to the centromere.9 Survivin recognizes H3T3ph to bring the CPC (chromosomal passenger complex) to the centromere region, thereby promoting the metaphase arrangement of chromosomes and the cell cycle.10–12 Although the function of GSG2 has not been clearly illustrated so far, some studies have shown that it plays a role as a cancer-promoting factor in various human tumors13 such as prostate cancer,14 gallbladder cancer15 and pancreatic cancer.16 Several studies have shown that its small molecule inhibitor CHR-6494 shows anti-proliferative activity in various tumor cell lines.17 Therefore, this work aims to clarify its regulatory effect on ovarian cancer.
In this study, it was found that GSG2 was significantly upregulated in ovarian cancer tissues compared with normal tissues. The depletion of GSG2 in ovarian cancer cells could inhibit ovarian cancer cell growth and promote cell apoptosis. In other words, the results herein identified GSG2 as a potential therapeutic target in the treatment of ovarian cancer.
Materials and Methods
Immunohistochemical Staining
Tissue microarray containing 115 tumor tissues of ovarian cancer patients (including 6 tissues with AJCC stage I, 24 tissues with AJCC stage II, 60 tissues with AJCC stage III and 23 tissues with AJCC stage IV) and 58 normal ovarian tissues were obtained from Shanghai Outdo Biotech Company (Cat. #HOvaC160Su01). All specimens were dewaxed and rehydrated, and treated with citric acid buffer for antigen retrieval at 100°C for 10 min. Subsequently, GSG2 antibody (1:200, Cat. #bs-15413R, Bioss) was added at 4°C overnight, followed by washing by PBS and the addition of secondary antibody (1:400, Cat. #Ab111909, Abcam) was added. Slices were stained with DAB and hematoxylin and the images were taken under the photomicroscope. Staining percentage scores were classified as: 1 (1%-24%), 2 (25%-49%), 3 (50%-74%) and 4 (75%-100%) and staining intensity were scored 0 (Signalless color) to 3 (light yellow, brown and dark brown). The IHC scores were determined by percentage and intensity scores.
Lentiviral Vector Construction
GSG2 was used as the template to design RNA interference target sequences (10055: 5ʹ-CCACAGGACAATGCTGAACTT-3ʹ, 00176: 5ʹ-AAGGAAACTGGTGGTGGGAAA-3ʹ, 00177: 5ʹ-AGGGATTGACTTAGAGCAAAT-3ʹ). The sequences were ligated to the linearized BR-V108 vector (Shanghai Yibeirui Biomedical science and Technology Co., Ltd.), which was linearized by restriction endonuclease Age I and EcoR I (Cat. # R3552L and R3101L, NEB). The restriction site overhang was added to the ends of the oligos. The ligation products were transformed into E.coli and positive clones were selected and verified by PCR (primer-F: 5ʹ-CCTATTTCCCATGATTCCTTCATA-3ʹ; primer-R: 5ʹ-GTAATACGGTTATCCACGCG-3ʹ). High purity plasmids were extracted by EndoFree midi Plasmid Kit (Cat. #DP118-2, TIANGEN). Afterwards, 293T cells were co-transfected with BR-V108, Helper 1.0 and Helper 2.0 plasmids to obtain lentivirus.
Cell Culture and Cell Transfection
The human ovarian epithelial cell line IOSE08 and human ovarian cancer cell lines 3AO, HO-8910, OVCAR-3 were obtained from ATCC. 3AO and HO-8910 cells were cultured in 90% RPMI-1640 containing 10% FBS in 6-well plates and OVCAR-3 were cultured in 90% DMEM containing 10% FBS. The culture environment was moist air containing 5% CO2 at 37°C.
Prepared lentivirus was transfected to the HO-8910 cells using Lipofectamine 2000 transfection reagent (Thermo Fisher Scientific, USA). After 72 h, the expression of green fluorescent protein was observed under a fluorescence microscope and the transfection efficiency was evaluated.
Quantitative Real-Time PCR
Total RNA was extracted from shGSG2 and shCtrl HO-8910 cell samples using Trizol, then the concentration and quality of RNA was detected by Nanodrop 2000/2000C. cDNA for qRT-PCR was obtained by reverse transcription using Promega M-MLV kit. PCR was performed by using AceQ qPCR SYBR Green Master Mix (Vazyme) and Real-time PCR instrument VII7 (ABI). The PCR cycling conditions were as follows: Initial denaturation at 95°C for 30 sec, followed by 45 cycles at 95°C for 5 sec, 60°C for 30 sec, 95°C for 15 sec, 55°C for 30 sec and 95°C for 15 sec. 2−∆∆CT method were applied for analyzing with GAPDH as the reference control. Primers used in PCR was as follows:
GSG2, upstream sequence (5ʹ-3ʹ): GGAAGGGGTGTTTGGCGAAGT, downstream sequence (5ʹ-3ʹ): TGAGGAGCAAGGGAGGGTAAG;
GAPDH, upstream sequence (5ʹ-3ʹ): TGACTTCAACAGCGACACCCA, downstream sequence (5ʹ-3ʹ): CACCCTGTTGCTGTAGCCAAA.
Western Blot and Antibody Array Assay
Western blot assay was applied to determine the GSG2 protein expression levels in shGSG2 HO-8910 cells. Cells were collected and lysed with 1 × cold lysis buffer. Protein was extracted with BCA protein detection kit (Cat. #23225, HyClone-Pierce). 10% SDS-PAGE was used for Western analysis. Blots were transferred to a PVDF membrane and incubated with blocking liquid for 1 h. GSG2 antibody (1:1000, Cat. #ab21686, Abcam), CCND1 antibody (1:3000, Cat. #ab134175, Abcam), CDK6 antibody (1:1000, Cat. #ab151247, Abcam), MAPK9 antibody (1:3000, Cat. #ab76125, Abcam), PIK3CA antibody (1:1000, Cat. #ab40776, Abcam), and internal standard GAPDH antibody (1:3000, Cat. #AP0063, Bioworld) were added overnight at 4°C. After incubated with secondary antibody HRP goat anti-rabbit IgG polyclonal (1:3000, Cat. # A0208, Beyotime) for 1 h at room temperature, the membrane was color developed using ECL-PLUS/Kit (Cat. #RPN2232, Amersham).
Human Apoptosis Antibody Array was purchased from Abcam (Cat. Ab134001) for detecting the expression of apoptosis-related proteins according to the manufacturer’s instructions.
Apoptotic Assay
The proportion of shGSG2 HO-8910 cell apoptosis was detected by flow cytometry. HO-8910 cells were seeded and cultured for 5 days in a 6 cm-well dish. Afterwards, cells were digested with trypsin and re-suspended with 1 × binding buffer. Before detected by FACScan, cells were stained by Annexin V-APC for 15 min in the dark. Cell phase percentage was determined to assess the apoptotic rate.
MTT Assay
Proliferation of shGSG2 HO-8910 cells was detected via MTT assay with five 96-well plates. 2500 cells/well were seeded to the 96-well plates for culturing. In the next 5 days (one time one day), 20 μL 5 mg/mL MTT solution was added into per well, and 4 h later, the culture medium was discarded and 100 μL DMSO was added for shaking 5 min. The OD490 was measured with a Microplate Reader and the cell viability was calculated.
Colony Formation Assay
Lentivirus transfected HO-8910 cells in the logarithmic growth phase were trypsinized, resuspended and seeded into 6-well plates with 800 cells per well and further cultured in the RPMI-1640 medium with 10% FBS for 9 days. The culture medium was exchanged every 72 h. Cell clones were photographed under a fluorescence microscope (Olympus, Tokyo). Cells were fixed in 4% paraformaldehyde and stained in 500 μL Giemsa. Colony forming rate was calculated as colony number/inoculated cell number × 100%.
Statistical Analysis
All our cell experiments were performed in triplicate and the data were expressed as mean ± SD (n ≥ 3) and statistical analyzed using SPSS 20.0 (SPSS) and GraphPad Prism software 7.0. The χ2 test, Mann–Whitney U-test, Spearman rank correlation analysis, Kaplan–Meier survival analysis were applied to estimate the association between the expression level of GSG2 and clinical characteristics. t-test were used to compare the difference. P values < 0.05 were defined as significant.
Results
GSG2 Was Upregulated in Ovarian Cancer and Predicted Poor Prognosis
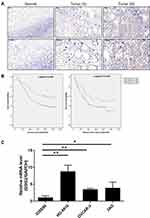
In order to explore the role of GSG2 in the development and progression of ovarian cancer, clinical ovarian cancer and normal tissues were collected for immunohistochemical staining analysis. As shown by representative images in Figure 1A, the expression of GSG2 was obviously higher in ovarian cancer tissues with advanced stage compared with that with early stage or normal tissues. After dividing all specimens into high expression group and low expression group based on the median of IHC score, the relationship between GSG2 and tumor characteristics was assessed. The results not only demonstrated the upregulation of GSG2 in ovarian cancer but also revealed the significant relationship between GSG2 expression and pathological grade and AJCC stage (Table 1–3). Moreover, the negative association between GSG2 and overall survival as well as disease-free survival was further elucidated by Kaplan–Meier survival analysis (Figure 1B). On the other hand, we detected the endogenous expression of GSG2 in ovarian epithelial cell IOSE80 and ovarian cancer cell HO-8910, 3AO and OVCAR-3, which showed that GSG2 expression was higher in ovarian cancer cells than normal cells, and higher in HO-8910 cells than 3AO and OVCAR-3 cells (Figure 1C). Collectively, these results showed that GSG2 may be involved in the development of ovarian cancer and could potentially act as a prognostic indicator.
 |
Table 1 Expression Patterns of GSG2 in Ovarian Cancer Tissues and Normal Tissues Revealed in Immunohistochemistry Analysis |
 |
Table 2 Relationship Between GSG2 Expression and Tumor Characteristics in Patients with Ovarian Cancer |
 |
Table 3 Relationship Between GSG2 Expression and Tumor Characteristics in Patients with Ovarian Cancer Analyzed by Spearman Rank Correlation Analysis |
The Construction of GSG2 Knockdown Cell Line
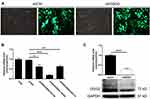
For launching loss-of-function experiments to illustrate the functions of GSG2 in ovarian cancer, GFP (green fluorescent protein) labelled lentivirus vectors expressing GSG2-silencing shRNAs were prepared and transfected into HO-8910 cells. The observation of GFP signal in >80% cell population 24 h after transfection was set as the threshold of successful transfection (Figure 2A). Among 3 shRNAs designed for knocking down GSG2, RNAi-10055 with the highest knockdown efficiency, as evaluated by qPCR, was used in following experiments without further illustration (Figure 2B). Then, qPCR and Western blotting were simultaneously performed to verify the downregulation of GSG2 in shGSG2 group compared with shCtrl group (Figure 2C). All these results indicated that HO-8910 cell model with GSG2 knockdown was successfully constructed and could be used for subsequent investigations.
Knockdown of GSG2 Inhibited Cell Proliferation and Colony Formation
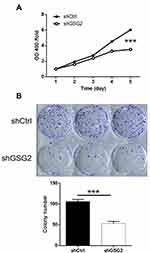
MTT assay was utilized to study the effects of GSG2 knockdown on cell proliferation. Through detecting cell viability of shGSG2 and shCtrl cells in 5 days of culturing, it was demonstrated that the proliferation rate of cells with downregulated GSG2 was significantly restrained (Figure 3A). Moreover, colony formation assay was applied to assess the anchorage-independent growth of HO-8910 cells with or without GSG2 knockdown. As shown in Figure 3B, cells in shGSG2 group formed much less colonies than that in shCtrl group, indicating the suppression of colony formation upon GSG2. Altogether, we demonstrated the inhibition of ovarian cancer cell growth by GSG2 knockdown, which was in consistent with our deduction.
Knockdown of GSG2 Promoted Cell Apoptosis of Ovarian Cancer Through Regulating Apoptosis-Related Proteins
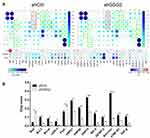
Finally, we evaluated the role of GSG2 in ovarian cancer cell apoptosis through flow cytometry. The results indicated that GSG2 knockdown significantly promoted cell apoptosis, which was also in consistent with the above results (Figure 4, table 2, table 1, table 3). Moreover, a human apoptosis antibody array was used to identify the differential expression of 43 apoptosis-related proteins in shCtrl and shGSG2 groups of HO-8910 cells. As shown in Figure 5, it was manifested that GSG2 knockdown may promote cell apoptosis through upregulating pro-apoptosis proteins including Bad and FasL, and downregulating anti-apoptosis proteins including Bcl-2, Bcl-w, cIAP-2, HSP27, HSP60, HSP70, IGF-II, IGFBP-2, Survivin, sTNF-R1 and TNF-β. Moreover, several well-known tumor regulator proteins such as CCND1, CDK6, MAPK9 and PIK3CA were also subjected to detection. CCND1, CDK6 and PIK3CA, which were generally PI3K signaling-related proteins, were downregulated in shGSG2 cells, while MAPK9 was upregulated (Figure 6).
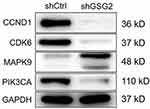 |
Figure 6 GSG2 regulated ovarian cancer through regulating cancer-related proteins. The expression of CCND1, CDK6, MAPK9 and PIK3CA was detected in ovarian cancer cells with or without GSG2 knockdown. |
Discussion
Ovarian cancer is one of the gynecological malignancies with the highest morbidity and mortality. However, because of the lack of specific diagnostic markers, rapid progress and easy metastasis, the treatment of ovarian cancer is still a major problem in the research field of ovarian cancer. Recently, the emergence and development of molecular-targeted therapy has brought a new approach to the treatment of ovarian cancer, which is based on the understanding of the molecular mechanism of ovarian cancer. Therefore, more and more attention has been paid in the exploration of novel molecules involved in the development and progression of ovarian cancer. For example, Zhang’s group found that nucleolar spindle-associated protein 1 (NuSAP1) was significantly upregulated in ovarian cancer, knockdown of which could induce inhibition of cell proliferation and migration, arrest of cell cycle, and promotion of cell apoptosis.18 Gao et al suggested that TCF12 may be involved in the growth and metastasis of ovarian cancer, which could be used as a potential prognostic indicator for ovarian cancer.19 Recently, SNCG was identified as a key regulator in the progression and metastasis of high-grade serous ovarian cancer, of which PI3K/Akt signaling pathway acted as the potential downstream mechanism.20
The biological functions of GSG2, also named as Haspin, in mitosis have been comprehensively investigated. Recently, due to the discovery of anti-tumor activity of GSG2 inhibitor such as CHR-6494, more and more attention has been paid to the role of GSG2 in development of malignant tumors.17 Yu et al reported, in prostate cancer cells, the inhibition of proliferation and migration and promotion of apoptosis, suggesting GSG2 as a tumor promotor in prostate cancer.14 Similar functions could also be observed in pancreatic cancer16 and gallbladder carcinoma.16 In this study, our group discerned GSG2 as a novel regulator of ovarian cancer, thus a potential therapeutic target for treatment and prognostic indicator. The detection and analysis of GSG2 expression in clinical specimens revealed that it was highly expressed in ovarian cancer, positively correlated with pathological grade and AJCC stage, and negatively associated with prognosis of ovarian cancer patients. The role of GSG2 in ovarian cancer was further confirmed by in vitro verifications, which showed that GSG2 knockdown inhibited ovarian cancer cell proliferation and promoted cell apoptosis. Moreover, several pro-apoptosis proteins such as Bad and FasL, and a variety of anti-apoptosis proteins including Bcl-2, Bcl-w, cIAP-2, HSP27, HSP60, HSP70, IGF-II, IGFBP-2, Survivin, sTNF-R1 and TNF-β were found to be involved in the regulation of cell apoptosis by GSG2 knockdown.
As a well-known proto-oncogene, CCND1, also known as cyclin D1, is widely amplified in human cancers including ovarian cancer.21 It also mediated the regulation of ovarian cancer by many different molecules.22 In our study, we also deduced that CCND1 may function as a potential downstream of GSG2 in ovarian cancer. CDK6 is another protein that is believed to possess important functions in tumor progression and is promising therapeutic target for cancer treatment.23 Previous study and our results both indicated the implication of CDK6 in the regulation of ovarian cancer.24 We found that MAPK9 (JNK2) was upregulated in ovarian cancer cells with relatively low GSG2 expression, which was in consistent with previous report.25 Finally, as a critical member in PI3K/Akt signaling pathway, oncogene PIK3CA was also found to be suppressed in shGSG2 cells.26
Conclusions
In conclusion, GSG2 was identified as a potential therapeutic target and prognostic indicator of ovarian cancer. We demonstrated the upregulation of GSG2 in ovarian cancer, whose expression was positively related to tumor grade and AJCC stage, and negatively correlated with patients’ prognosis. The loss-of-function study also proved the involvement of GSG2 in ovarian cancer development through regulating apoptosis-related proteins and downstream pathways such as PI3K signaling.
Data Sharing Statement
Not applicable.
Ethics Approval and Informed Consent
This study was approved by Ethics committee of Fudan university. Informed consents were collected from all the patients in this study.
Consent for Publication
Not applicable.
Funding
There is no funding to report.
Disclosure
The authors report no conflicts of interest for this work.
References
1. Webb PM, Jordan SJ. Epidemiology of epithelial ovarian cancer. Best Pract Res Clin Obstet. 2017;41:3–14. doi:10.1016/j.bpobgyn.2016.08.006
2. Stewart C, Ralyea C, Lockwood S. Ovarian cancer: an integrated review. Semin Oncol Nurs. 2019;35(2):151–156. doi:10.1016/j.soncn.2019.02.001
3. Narod S. Can advanced-stage ovarian cancer be cured? Nat Rev Clin Oncol. 2016;13(4):255–261. doi:10.1038/nrclinonc.2015.224
4. Grunewald T, Ledermann JA. Targeted therapies for ovarian cancer. Best Pract Res Clin Obstet. 2017;41:139–152. doi:10.1016/j.bpobgyn.2016.12.001
5. Guan L, Lu Y. New developments in molecular targeted therapy of ovarian cancer. Discov Med. 2018;26(144):219–229.
6. Lim HJ, Ledger W. Targeted therapy in ovarian cancer. Womens Health. 2016;12(3):363–378. doi:10.2217/whe.16.4
7. Eswaran J, Patnaik D, Filippakopoulos P, et al. Structure and functional characterization of the atypical human kinase haspin. Proc Natl Acad Sci. 2009;106(48):20198. doi:10.1073/pnas.0901989106
8. Ghenoiu C, Wheelock MS, Funabiki H. Autoinhibition and polo-dependent multisite phosphorylation restrict activity of the histone H3 kinase haspin to mitosis. Mol Cell. 2013;52(5):734–745. doi:10.1016/j.molcel.2013.10.002
9. Zhou L, Tian X, Zhu C, Wang F, Higgins JM. Polo-like kinase-1 triggers histone phosphorylation by haspin in mitosis. EMBO Rep. 2014;15(3):273–281. doi:10.1002/embr.201338080
10. Wang F, Dai J, Daum JR, et al. Histone H3 thr-3 phosphorylation by haspin positions aurora B at centromeres in mitosis. Science. 2010;330(6001):231–235. doi:10.1126/science.1189435
11. Dai J, Higgins JMG. Haspin: a mitotic histone kinase required for metaphase chromosome alignment. Cell Cycle. 2005;4(5):665–668. doi:10.4161/cc.4.5.1683
12. Yu F, Jiang Y, Lu L, et al. Aurora-A promotes the establishment of spindle assembly checkpoint by priming the Haspin-Aurora-B feedback loop in late G2 phase. Cell Discov. 2017;3(1):16049. doi:10.1038/celldisc.2016.49
13. Wang P, Hua X, Bryner YH, Liu S, Gitter CB, Dai J. Haspin inhibition delays cell cycle progression through interphase in cancer cells. J Cell Physiol. 2020;235(5):4508–4519. doi:10.1002/jcp.29328
14. Yu F, Lin Y, Xu X, et al. Knockdown of GSG2 inhibits prostate cancer progression in vitro and in vivo. Int J Oncol. 2020;57(1):139–150. doi:10.3892/ijo.2020.5043
15. Zhu D, Gu X, Lin Z, Yu D, Wang J, Li L. HASPIN is involved in the progression of gallbladder carcinoma. Exp Cell Res. 2020;390(2):111863. doi:10.1016/j.yexcr.2020.111863
16. Han X, Kuang T, Ren Y, Lu Z, Liao Q, Chen W. Haspin knockdown can inhibit progression and development of pancreatic cancer in vitro and vivo. Exp Cell Res. 2019;385(1):111605. doi:10.1016/j.yexcr.2019.111605
17. Amoussou NG, Bigot A, Roussakis C, Robert JH. Haspin: a promising target for the design of inhibitors as potent anticancer drugs. Drug Discov Today. 2018;23(2):409–415. doi:10.1016/j.drudis.2017.10.005
18. Zhang Y, Huang K, Cai H, Chen S, Sun D, Jiang P. The role of nucleolar spindle-associated protein 1 in human ovarian cancer. J Cell Biochem. 2020. doi:10.1002/jcb.29661
19. Gao S, Bian T, Zhang Y, Su M, Liu Y. TCF12 overexpression as a poor prognostic factor in ovarian cancer. Pathol Res Pract. 2019;215(9):152527. doi:10.1016/j.prp.2019.152527
20. Zhang J, Liu X, Li C, et al. SNCG promotes the progression and metastasis of high-grade serous ovarian cancer via targeting the PI3K/AKT signaling pathway. J Exp Clin Canc Res. 2020;39(1):79. doi:10.1186/s13046-020-01589-9
21. Qie S, Diehl JA. Cyclin D1, cancer progression, and opportunities in cancer treatment. J Mol Med. 2016;94(12):1313–1326. doi:10.1007/s00109-016-1475-3
22. Jia Y, Shi H, Cao Y, Feng W, Li M, Li X. PDZ and LIM domain protein 4 suppresses the growth and invasion of ovarian cancer cells via inactivation of STAT3 signaling. Life Sci. 2019;233:116715. doi:10.1016/j.lfs.2019.116715
23. Sherr CJ, Beach D, Shapiro GI. Targeting CDK4 and CDK6: from discovery to therapy. Cancer Discov. 2016;6(4):353. doi:10.1158/2159-8290.CD-15-0894
24. Zhu X, Yan S, Xiao S, Xue M. Knockdown of ALPK2 inhibits the development and progression of ovarian cancer. Cancer Cell Int. 2020;20(1):267. doi:10.1186/s12935-020-01347-z
25. Chauhan SC, Vannatta K, Ebeling MC, et al. Expression and functions of transmembrane mucin MUC13 in ovarian cancer. Cancer Res. 2009;69(3):765. doi:10.1158/0008-5472.CAN-08-0587
26. Noorolyai S, Shajari N, Baghbani E, Sadreddini S, Baradaran B. The relation between PI3K/AKT signalling pathway and cancer. Gene. 2019;698:120–128. doi:10.1016/j.gene.2019.02.076
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.