Back to Journals » Biologics: Targets and Therapy » Volume 14
Glycopeptides as Potential Interventions for COVID-19
Authors Demsie DG , Gebre AK , Yimer EM , Alema NM , Araya EM , Bantie AT, Allene MD , Gebremedhin H, Yehualaw A , Tafere C , Tadese HT , Amare B , Weldekidan E , Gebrie D
Received 13 May 2020
Accepted for publication 11 August 2020
Published 13 October 2020 Volume 2020:14 Pages 107—114
DOI https://doi.org/10.2147/BTT.S262705
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Doris Benbrook
Desalegn Getnet Demsie, 1 Abadi Kahsu Gebre, 2 Ebrahim M Yimer, 3 Niguse Meles Alema, 1 Ephrem Mebrahtu Araya, 1 Abere Tilahun Bantie, 4 Mengesha Dessie Allene, 5 Hagazi Gebremedhin, 2 Adane Yehualaw, 6 Chernet Tafere, 6 Haileslassie Tesfay Tadese, 7 Bekalu Amare, 2 Etsay Weldekidan, 1 Desye Gebrie 8, 9
1Adigrat University, College of Medicine and Health Sciences, Department of Pharmacy, Adigrat, Ethiopia; 2Mekelle University, College of Health Sciences, Department of Pharmacology and Toxicology, Mekelle, Ethiopia; 3Wollo University, College of Medicine and Health Sciences, Department of Pharmacy, Dessie, Ethiopia; 4Adigrat University, College of Medicine and Health Sciences, Department of Anesthesia, Adigrat, Ethiopia; 5Debre Berhan University, School of Medicine, College of Medicine, Department of Anesthesia, Debre Berhan, Ethiopia; 6Bahir Dar University, College of Health Sciences, Department of Pharmacy, Bahir Dar, Ethiopia; 7Adigrat University, College of Medicine and Health Sciences, Department of Nursing, Adigrat, Ethiopia; 8Mekelle University, College of Health Sciences, Department of Social Pharmacy and Pharmacoepidemiology, Mekelle, Ethiopia; 9Addis Ababa University, College of Health Sciences, Center for Innovative Drug Development and Therapeutic Trials for Africa, Addis Ababa, Ethiopia
Correspondence: Desalegn Getnet Demsie Email [email protected]
Abstract: Coronavirus disease 2019 (COVID-19), an infectious disease that primarily attacks the human pulmonary system, is caused by a viral strain called severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). The outbreak emerged from Wuhan, China, and later spread throughout the world. Until the first week of May 2020, over 3.7 million cases had been reported worldwide and more than 258,000 had died due to the disease. So far, off label use of various drugs has been tried in many clinical settings, however, at present, there is no vaccine or antiviral treatment for human and animal coronaviruses. Therefore, repurposing of the available drugs may be promising to control emerging infections of SARS-COV2; however, new interventions are likely to require months to years to develop. Glycopeptides, which are active against gram-positive bacteria, have demonstrated significant activity against viral infections including SARS-COV and MERS-COV and have a high resemblance of sequence homology with SARS-COV2. Recent in vitro studies have also shown promising activities of aglycon derivative of glycopeptides and teicoplanin against SARS-COV2. Hydrophobic aglycon derivatives and teicoplanin, with minimal toxicity to human cell lines, inhibit entry and replication of SARS-COV2. These drugs block proteolysis of polyprotein a/b with replicase and transcription domains. Teicoplanin use was associated with complete viral clearance in a cohort of patients with severe COVID-19 symptoms. This review attempts to describe the activity, elucidate the possible mechanisms and potential clinical applications of existing glycopeptides against corona viruses, specifically SARS-COV2.
Keywords: corona virus, SARS-COV, SARS-COV2, COVID19, glycopeptides
Introduction
Coronaviruses (CoVs), spherical and enveloped viruses with a diameter of 100–160 nm, cause respiratory and gastro-intestinal infections both in humans and animals.1 The family of corona viruses is classified into α, β, γ and δ genus. Both α and β genus infect mammals, whereas γ and δ have been thought pathogenic only to birds.2
Previously, seven CoVs have been identified as human-susceptible viruses, of these α-CoVs, HCoV-229E and HCoV-NL63, and β–CoVs, HCoV-HKU1 and HCoV-OC43 are less pathogenic to humans and usually cause common cold or mild respiratory infections, particularly in individuals with strong immune status.3–5 However, certain vulnerable groups such as patients with co-morbidities and elderly people may experience severe infections.5 The other three, SARS-CoV, Middle East Respiratory Syndrome Corona Virus (MERS-CoV), and SARS-CoV2, emerged in 2002, 2012, and 2019, respectively, and can cause severe respiratory illness and fatalities.6
SARS-CoV2 epidemic has diffused to many counties in the world from the epicenter Wuhan city, China; it was initially named as 2019-Novel Corona Virus (2019-nCoV) on January 12, 2020 by the World Health Organization (WHO).2,7 Later on, WHO designated the name of the disease as Corona Virus Disease-2019 (COVID-19)9 on February 11, 20202,7 that recently led to huge impacts on humans' health globally,8 indicating that coronaviruses pose a continuous pandemic threat to humans. On January 30, 2020, the WHO declared COVID-19 as a public health emergency of international concern.9
COVID-19 initially resulted in a cluster of viral pneumonia cases10 and later the number of cases escalated in China and globally.11 In less than 3 months, the virus advanced to the whole world and has led to global shutdown.12 Until the first week of May 2020, over 3.7 million cases had been reported worldwide. Currently, the United States of America (USA) (>1.2 million) has the largest number of cases followed by nearly 260,000 cases in Spain and 213,000 cases in Italy. The death toll from the disease surpassed 258,000 and nearly 1,240,000 had recovered from the disease.13 The case fatality rate of COVID-19 is 4% (Surveillances, 2020) compared to a rate of 9.65% reported due to SARS.14
The ongoing outbreak of COVID-19 has challenged the health care systems and communities in many countries15 for secondary transmissions have occurred through, for example, travellers and the speed of transmission is accelerating.16 The outbreak has negatively affected service-oriented economies17,18 by interrupting global trade and supply chains, depressing asset prices, and forcing multinational businesses to make hard decisions with limited information.19,20 Due to the ongoing progress of the disease, many nations have declared lockdown strategies and ordered people to stay at home; they have also allocated budgets for pandemic preparedness.20
Initially, most patients infected with the virus were reportedly exposed to Huanan seafood market and wet animal market in Wuhan City, Hubei Province suggesting a potential zoonotic origin.6,10 Genome sequencing and evolutionary analysis demonstrated 96.2% of genetic similarity between SAR S-CoV-2 and bat CoV RaTG13. This hinted that SAR S-CoV-2 might be transmitted from the natural reservoir, bat, to human via unknown intermediate hosts.21 Then person-to-person transmission has been reported both in hospital, family settings and communities,22 where symptomatic, presymptomatic and asymptomatic individuals transmit the virus via droplets from coughing or sneezing or direct contact with another person.6
Most patients with COVID19 experience fever, cough, difficulty of breathing and fatigue or myalgia. Less frequent symptoms including increased sputum production, headache, hemoptysis and diarrhea have also been reported. This viral pneumonia may complicate to acute respiratory distress syndrome (ARDS), acute cardiac injury, and secondary infection.6,10 These symptoms and complications are highly pronounced in individuals with comorbidities such as cardiovascular and metabolic comorbidities.7,23 Among patients diagnosed with COVID-19, 17.1%, 16.4%, and 9.7% of the cases had hypertension, other cardiovascular disease and diabetes, respectively, and acute cardiac injury was reported in at least 8.0% of the individuals.12
Several measures have already been implemented to prevent and control the disease; however, the management and control of local transmission after importation heavily rely on a country’s health capacity on the application and execution of strict measures of detection, prevention, and control.24,25 As of nations’ awareness of the epidemic, drugs such as chloroquine phosphate,26 lopinavir/ritonavir,18,24 broad-spectrum antibiotics, corticosteroids, interferons, ribavirin, and mycophenolate mofetil have shown some activity against COVs infection.27 Nevertheless, effective vaccines or antiviral therapeutic agents have not been approved to treat COVID-19 till now28 and the treatment approaches were not selected based on well-organized clinical trials.27
Glycopeptides had demonstrated anti-SARS-COV, anti-MERS-COV activities, and are effective against other superimposed viral and bacterial infections. Given that SARS-COV and MERS-COV have high genetic resemblance with SARS-COV2, drugs which are effective against these viruses could be considered for 2019-nCoV treatment.12 More recent evidence also showed that teicoplanin potently inhibited SARS-COV2 infection in in vitro studies.29 In addition, teicoplanin exhibited promising clinical outcomes in a cohort of patients who received a dose of 6 mg/kg every 24 hours (loading dose every 12 hours for three doses) for 7–12 days.30
Assuming the urgency of treatment for 2019-nCoV outbreak, the potential to repurpose existing glycopeptides or in development could be considered as potential drugs for 2019-nCoV infections. Thus, the aim of this review was to describe the activity, elucidate the possible mechanism and potential to repurpose existing glycopeptides against coronaviruses, especially SARS-COV2.
Glycopeptides (GPs) as Potential Drugs for Coronavirus Treatment
Insight into the potential use of GPs against SARS-COV2 emerged after in vitro and animal studies demonstrated the effect of vancomycin, teicoplanin, ristomycin, eremimycin-type aglycons, and their semisynthetic derivatives to treat or prevent viral infections caused by Coronaviridae, especially, SARS-CoV and MERS-COV.31 Studies also indicated that GPs, teicoplanin and its derivatives, were effective against enveloped viruses such as ebola envelope pseudotyped viruses (pEBOV), human immunodeficiency virus (HIV),32,33 human hepatitis C virus (HCV),34,35 influenza viruses,36 dengue virus and other flu viruses,37 SARS-CoV and feline (FIPV).38 GPs, including DA-40926 showed antiviral activity with different degrees of inhibition in cell culture.32 High genetic similarity (79.5%) between SARS-COV and 2019-nCOV and their ability to recognize the same enzyme, ACE2, strengthen the speculation that GPs may exert anti-SARS-COV2 activity.39
Particularly, the aglycons (Figure 1), onto which various functional groups attach, preferably hydrophobic residue, have marked activity against human immunodeficiency virus type1 (HIV1) and human immunodeficiency virus type 2 (HIV2) at low micromolar concentrations (1–5 uM). Nevertheless, some of the compounds were toxic for human cells even at low drug concentrations.32 Cell toxicity was avoided (IC50> 500M), by removing all the carbohydrate residues, but anti-HIV-1 activity (EC50~50–65 M) was achieved at relatively high concentrations.40 In later studies, these GPs devoid of carbohydrate residues exhibited no inhibitory action to SARS-CoV and FIPV. Interestingly, modifications made to the basic structure lead to the synthesis of aglycon derivatives, and glycopeptide antibiotics with a modified or partially destroyed peptide core, with potent activity against SARS-CoV and FIPV in cell culture.40
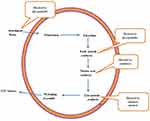 |
Figure 1 Major targets for anti-SARS-COV2 agents. |
Teicoplanin and its derivatives (dalbavancin, oritavancin, and telavancin) mainly target SARS-COV2 and other enveloped virus entry (Figure 2), an ideal drug target for antiviral activity,28 in to the cytoplasm of host cells.41 Another study on COVs also confirmed that the main molecular event associated with the action of GPs and aglycons is inhibition of virus entry into the host cells.32 In fact, virus entry into the host cells requires several biochemical and molecular changes.42,43
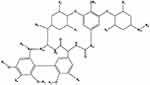 |
Figure 2 Chemical structure of a teicoplanin derivative aglycon. |
GPs inhibit cathepsin L which is involved in the proteolysis of S protein, a spike protein that is selected and targeted for recognition of host cells by nCoV-2019.44,45 S protein priming by cathepsin L and other cellular proteases, trans-membrane serine protease 2 (TMPRSS2), furin and trypsin is fundamental to virus attachment and fusion28,46 (Figure 2) of viral and host cell membranes. Enzymatic modification of S protein by TMPRSS2 accompanies cleavage-mediated conformational changes to both S protein and ACE2, which allows the host cell membrane to invaginate for initiation of viral endocytosis into the endosomes and the lysosomes.42
Subsequently, S protein undergoes a series of enzymatic cleavages and modifications by endo-lysosomal cysteine proteases, cathepsin L, and to a lesser degree by cathepsin B in the endosomes.41 All the aforementioned process could not be assumed without proteolysis modification of S protein and ACEI with accompanied attachment and fusion. Thus, cathepsin L inhibition could interfere with entry of SARS-CoV-2 into the host cells.43
According to in vitro studies, teicoplanin potently inhibited attachment of SARS-CoV2 virus to ACE2 receptors of the host, indicating that this glycopeptide blocks entrance of 2019-nCoV-Spike-pseudoviruses into the cytoplasm (IC50, 1.66 μM).44,45 Previously teicoplanin, has routinely been used in clinics to treat infection of human cells by MERS-COV, pEBOV, SARS-COV44 by specifically inhibiting the activity of cathepsin L, opening a novel avenue for the development of additional glycopeptides as potential inhibitors of cathepsin L-dependent viruses.21,41
The other targets for GPs are SARS-CoV encoded proteases. The genetic information of CoVs encoded in a large polyprotein with a replicase domain that shares two-thirds of the genome length and the structural regions. The nonstructural protein 5 (nsp5), the main protease (Mpro), picornavirus3C-like cysteine protease (3CLpro), and the papain-like cysteine protease (PLpro) involve in the extensive proteolytic processing of nonstructural and structural polyproteins, providing the functional proteins for SARS-CoV viral replication and propagation.47–50 SARS-CoV 3CLpro, potently inhibited a glycopeptide derivative (Figure 1),31 and was considered as a promising target for anti-SARS therapy through its interference with virus propagation process and replication (Figure 2). Following large protein proteolysis, replication of the virus will proceed by the action of viral RNA-dependent RNA polymerase.51 Remdesivir, monophosphoramidate prodrug of an adenosine analog, blocked CoVs replication through targeting the key replication enzyme, RNA polymerase in vitro and animal studies.52
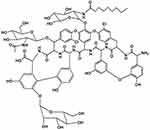 |
Figure 3 Chemical structure of teicoplanin (a glycopeptide antibiotic). |
Moreover, GPs inhibit transcription and replication-competent virus-like particles, with an IC 50 as low as 330 nM.53 With this, teicoplanin (Figure 3) efficiently inhibited enveloped human respiratory syncytial virus, but not non-enveloped viruses (IC50 as low as 330 Nm),54 indicating that the agent probably targets common components of enveloped viruses,53 including SARS-CoV2.28
Viral infection activates multiple pathways, like virus-encoded or virus-induced protein kinases which are considered as targets for antimicrobials and antiviral agents.55–58 Protein kinases (PKs) of mature virions are thought to facilitate virus entry and replication of human pathogenic viruses. According to more recent studies, glycopeptide antibiotics, teicoplanin and eremomycin inhibit protein kinases, a key regulator of cell signaling.59 Replacing the sugar moiety with hydrophobic residue in to aglycons and pseudo aglycons of glycopeptides is a means to synthesize glycopeptide aglycon derivatives with efficient activity against PKs of the viruses.37 Another in vitro study also confirmed that various hydrophobic residues of teicoplanin aglycon carboxamides which were effective against PKs of SARS-COV, successfully inhibited all tested human PKs at low micromolar IC50 values.59
Clinical Application of Glycopeptides
Based on the aforementioned collective evidences, glycopeptides have been utilized as a potential antiviral agent or as intervention for likely superimposed bacterial pneumonia. Indeed, a number of reports have suggested that bacterial copathogens such as Staphylococcus aureus, Streptococcus pneumoniae, Staphylococcus aureus, and Klebsiella pneumoniae, among others,60,61 and viruses such as influenza, rhinovirus/enterovirus, influenza B virus, and human immunodeficiency virus62 are commonly identified in SARS-COV2 respiratory infections. These bacterial pathogens cause infections in up to 7.1% of COVID-19 cases, and have led to the empiric use of broad spectrum antibiotics in the majority of patients (71.3%).61,63
Besides bacterial co-infection, the empiric use of broad spectrum antibiotics may be attributed to indistinguishable symptoms of hospital-acquired and ventilator-associated pneumonia, and viral infections.7,64 In this regard, teicoplanin has been recommended as promising potential antiviral and antibacterial agent in some studies. A recent cohort study was carried out on 21 severe COVID-19 patients who received teicoplanin 6 mg/kg every 24 hours (loading dose every 12 hours for three doses) for 7–12 days, suggested that the drug might be considered as a potential intervention for COVID-19 following achievement of a complete viral clearance.30 Importantly, the anti SARS-COV2 activity of aglycons and teicoplanin is associated with reduced risk of nephrotoxicity and less frequent cutaneous rash, red man syndrome, and total adverse events.65
Conclusion and Future Perspectives
Since no vaccine or drugs are currently available for treatment and prevention of coronavirusinfections, antibacterial drugs that have previous reports of activities against viruses of similar species of coronavirus could be effective against COVID-19 and other coronaviruses strengthening the arsenal of existing antiviral options. For instance, glycopeptide antibiotics and their derivatives such as teicoplanin, oritavancin and dalbavancin could be repurposed for COVID-19 treatment because these antibiotics have been shown to inhibit several coronaviruses like SARS-COV, MERS-COV, SARS-COV2 and other enveloped viruses in the laboratory. Glycopeptides and their derivatives have demonstrated inhibitory action on transcription-replication competent of virus-like particles. This may give a hint towards the impeding action of glycopeptides on replication of SARS-COV2. Recent clinical evidence has also shown that complete viral clearance was achieved following administration of teicoplanin for a cohort of patients. We recommend further in vitro studies to rule out their effectiveness against COVID-19.
Author Contributions
All authors made a significant contribution to the work reported, whether that is in the conception, study design, execution, acquisition of data, analysis and interpretation, or in all these areas; took part in drafting, revising or critically reviewing the article; gave final approval of the version to be published; have agreed on the journal to which the article has been submitted; and agree to be accountable for all aspects of the work.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Salata C, Calistri A, Parolin C, Palu G. Coronaviruses: a paradigm of new emerging zoonotic diseases. Pathog Dis. 2019;77(9):ftaa006. doi:10.1093/femspd/ftaa006
2. Cui J, Li F, Shi ZL. Origin and evolution of pathogenic coronaviruses. Nat Rev Microbiol. 2019;17(3):181–192. doi:10.1038/s41579-018-0118-9
3. Ksiazek TG, Erdman D, Goldsmith CS, et al. A novel coronavirus associated with severe acute respiratory syndrome. N Engl J Med. 2003;348(20):1953–1966. doi:10.1056/NEJMoa030781
4. Zhong NS, Zheng BJ, Li YM, et al. Epidemiology and cause of severe acute respiratory syndrome (SARS) in Guangdong, People’s Republic of China, in February, 2003. Lancet. 2003;362(9393):1353–1358. doi:10.1016/S0140-6736(03)14630-2
5. Carlos WG, Dela Cruz CS, Cao B, Pasnick S, Jamil S. Novel wuhan (2019-nCoV) coronavirus. Am J Respir Crit Care Med. 2020;201(4):P7–P8. doi:10.1164/rccm.2014P7
6. Yin Y, Wunderink RG. MERS, SARS and other coronaviruses as causes of pneumonia. Respirology. 2018;23(2):130–137. doi:10.1111/resp.13196
7. Zhu N, Zhang D, Wang W, et al. A novel coronavirus from patients with pneumonia in China, 2019. N Engl J Med. 2020;382(8):727–733. doi:10.1056/NEJMoa2001017
8. Channappanavar R, Perlman S. Pathogenic human coronavirus infections: causes and consequences of cytokine storm and immunopathology. Semin Immunopathol. 2017;39(5):529–539. doi:10.1007/s00281-017-0629-x
9. Lai CC, Shih TP, Ko WC, Tang HJ, Hsueh PR. Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and corona virus disease-2019 (COVID-19): the epidemic and the challenges. Int J Antimicrob Agents. 2020;55(3):105924. doi:10.1016/j.ijantimicag.2020.105924
10. Huang C, Wang Y, Li X, et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet. 2020;395(10223):497–506. doi:10.1016/S0140-6736(20)30183-5
11. United States Center for Disease Control and Prevention (US-CDC). Confirmed 2019-nCoV cases globally. Availabe from: https://www.cdc.gov/coronavirus/2019-ncov/locations-confirmed-cases.html#map.
12. Li C, Yang Y, Ren L. Genetic evolution analysis of 2019 novel coronavirus and coronavirus from other species. Infect Genet Evol. 2020;82:104285. doi:10.1016/j.meegid.2020.104285
13. Coronavirus Updates. Available from: https://www.worldometers.info/coronavirus/.
14. Wong G, Liu W, Liu Y, et al. MERS, SARS, and Ebola: the role of super-spreaders in infectious disease. Cell Host Microbe. 2015;18(4):398–401. doi:10.1016/j.chom.2015.09.013
15. Kim JY, Choe PG, Oh Y, et al. The first case of 2019 novel coronavirus pneumonia imported into Korea from Wuhan, China: implication for infection prevention and control measures. J Korean Med Sci. 2020;35(5).
16. Corman VM, Landt O, Kaiser M, et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Eurosurveillance. 2020;25(3):2000045. doi:10.2807/1560-7917.ES.2020.25.3.2000045
17. Gormsen NJ, Koijen RS. Coronavirus: impact on stock prices and growth expectations. University of Chicago, Becker Friedman Institute for Economics Working Paper; 2020 Mar 17(2020–22).
18. Fernandes N. Economic effects of coronavirus outbreak (COVID-19) on the world economy. SSRN. 2020;3557504.
19. Ayittey FK, Ayittey MK, Chiwero NB, Kamasah JS, Dzuvor C. Economic impacts of Wuhan 2019‐nCoV on China and the world. J Med Virol. 2020;92(5):473–475. doi:10.1002/jmv.25706
20. Koonin LM. Novel coronavirus disease (COVID-19) outbreak: now is the time to refresh pandemic plans. J Bus Contin Emer Plan. 2020;13(4):1–5.
21. Zhou P, Yang XL, Wang XG, et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020;579(7798):270–273. doi:10.1038/s41586-020-2012-7
22. Chan JF, Yuan S, Kok KH, et al. A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: a study of a family cluster. Lancet. 2020;395(10223):514–523. doi:10.1016/S0140-6736(20)30154-9
23. Chen Y, Liu Q, Guo D. Emerging coronaviruses: genome structure, replication, and pathogenesis. J Med Virol. 2020;92(4):418–423. doi:10.1002/jmv.25681
24. Gilbert M, Pullano G, Pinotti F, et al. Preparedness and vulnerability of African countries against importations of COVID-19: a modelling study. Lancet. 2020;395(10227):871–877. doi:10.1016/S0140-6736(20)30411-6
25. WHO Regional Office for Africa (WHO-AFRO) WHO ramps up preparedness for novel coronavirus in the African region. Available from: https://www.afro.who.int/news/who-ramps-preparedness-novel-coronavirus-african-region.
26. Gao J, Tian Z, Yang X. Breakthrough: chloroquine phosphate has shown apparent efficacy in treatment of COVID-19 associated pneumonia in clinical studies. Biosci Trends. 2020;14(1):72–73. doi:10.5582/bst.2020.01047
27. Rabaan AA, Alahmed SH, Bazzi AM, Alhani HM. A review of candidate therapies for Middle East respiratory syndrome from a molecular perspective. J Med Microbiol. 2017;66(9):1261–1274. doi:10.1099/jmm.0.000565
28. Heymann DL, Shindo N. COVID-19: what is next for public health? Lancet. 2020;395(10224):542–545. doi:10.1016/S0140-6736(20)30374-3
29. Hoffmann M, Kleine-Weber H, Krüger N, et al. The novel coronavirus 2019 (2019-nCoV) uses the SARS-coronavirus receptor ACE2 and the cellular protease TMPRSS2 for entry into target cells. BioRxiv. 2020.
30. Ceccarelli G, Alessandri F, d’Ettorre G, et al. Is Teicoplanin A complementary treatment option for Covid-19? The question remains. Int J Antimicrob Agents. 2020;56(2):106029. doi:10.1016/j.ijantimicag.2020.106029
31. Kuo CJ, Liang PH. Characterization and inhibition of the main protease of severe acute respiratory syndrome coronavirus. Chem Bio Eng Rev. 2015;2(2):118–132.
32. Balzarini J, Pannecouque C, De Clercq E, et al. Antiretroviral activity of semisynthetic derivatives of glycopeptide antibiotics. J Med Chem. 2003;46(13):2755–2764. doi:10.1021/jm0300882
33. Preobrazhenskaya MN, Olsufyeva EN. Polycyclic peptide and glycopeptide antibiotics and their derivatives as inhibitors of HIV entry. Antiviral Res. 2006;71(2–3):227–236. doi:10.1016/j.antiviral.2006.04.008
34. Obeid S, Printsevskaya SS, Olsufyeva EN, et al. Inhibition of hepatitis C virus replication by semi-synthetic derivatives of glycopeptide antibiotics. J Antimicrob Chemother. 2011;66(6):1287–1294. doi:10.1093/jac/dkr104
35. Maieron A, Kerschner H. Teicoplanin therapy leading to a significant decrease in viral load in a patient with chronic hepatitis C. J Antimicrob Chemother. 2012;67(10):2537–2538. doi:10.1093/jac/dks217
36. Bereczki I, Kicsák M, Dobray L, et al. Semisynthetic teicoplanin derivatives as new influenza virus binding inhibitors: synthesis and antiviral studies. Bioorg Med Chem Lett. 2014;24(15):3251–3254. doi:10.1016/j.bmcl.2014.06.018
37. De Burghgraeve T, Kaptein SJ, Ayala-Nunez NV, et al. An analogue of the antibiotic teicoplanin prevents flavivirus entry in vitro. PLoS One. 2012;7(5):e37244. doi:10.1371/journal.pone.0037244
38. Balzarini J, Keyaerts E, Vijgen L, et al. Inhibition of feline (FIPV) and human (SARS) coronavirus by semisynthetic derivatives of glycopeptide antibiotics. Antiviral Res. 2006;72(1):20–33. doi:10.1016/j.antiviral.2006.03.005
39. Batlle D, Wysocki J, Satchell K. Soluble angiotensin-converting enzyme 2: a potential approach for coronavirus infection therapy? Clin Sci. 2020;134(5):543–545. doi:10.1042/CS20200163
40. Printsevskaya SS, Solovieva SE, Olsufyeva EN, et al. Structure-activity relationship studies of a series of antiviral and antibacterial aglycon derivatives of the glycopeptide antibiotics vancomycin, eremomycin, and dechloroeremomycin. J Med Chem. 2005;48(11):3885–3890. doi:10.1021/jm0500774
41. Nal B, Chan C, Kien F, et al. Differential maturation and subcellular localization of severe acute respiratory syndrome coronavirus surface proteins S, M and E. J Gen Virol. 2005;86(5):1423–1434. doi:10.1099/vir.0.80671-0
42. Yang N, Shen HM. Targeting the endocytic pathway and autophagy process as a novel therapeutic strategy in COVID-19. Int J Biol Sci. 2020;16(10):1724. doi:10.7150/ijbs.45498
43. He X, Lau EH, Wu P, et al. Temporal dynamics in viral shedding and transmissibility of COVID-19. Nat Med. 2020;1–4.
44. Zhang J, Ma X, Yu F, et al. Teicoplanin potently blocks the cell entry of 2019-nCoV. BioRxiv. 2020.
45. Baron SA, Devaux C, Colson P, Raoult D, Rolain JM. Teicoplanin: an alternative drug for the treatment of coronavirus COVID-19. Int J Antimicrob Agents. 2020;105944.
46. Beniac DR, Andonov A, Grudeski E, Booth TF. Architecture of the SARS coronavirus prefusion spike. Nat Struct Mol Biol. 2006;13(8):751–752. doi:10.1038/nsmb1123
47. Chou KC, Wei DQ, Zhong WZ. Binding mechanism of coronavirus main proteinase with ligands and its implication to drug design against SARS. Biochem Biophys Res Commun. 2003;308(1):148–151. doi:10.1016/S0006-291X(03)01342-1
48. Yang H, Yang M, Ding Y, et al. The crystal structures of severe acute respiratory syndrome virus main protease and its complex with an inhibitor. Proc Natl Acad Sci. 2003;100(23):13190–13195. doi:10.1073/pnas.1835675100
49. Tsaytler P, Harding HP, Ron D, Bertolotti A. Selective inhibition of a regulatory subunit of protein phosphatase 1 restores proteostasis. Science. 2011;332(6025):91–94. doi:10.1126/science.1201396
50. De Clercq E. Potential antivirals and antiviral strategies against SARS coronavirus infections. Expert Rev Anti Infect Ther. 2006;4(2):291–302. doi:10.1586/14787210.4.2.291
51. Brown AJ, Won JJ, Graham RL, et al. Broad spectrum antiviral remdesivir inhibits human endemic and zoonotic deltacoronaviruses with a highly divergent RNA dependent RNA polymerase. Antiviral Res. 2019;169:104541. doi:10.1016/j.antiviral.2019.104541
52. Reina J. Remdesivir, the antiviral hope against SARS-CoV-2. Rev Esp Quimioter. 2020;33:176–179.
53. Wang Y, Cui R, Li G, et al. Teicoplanin inhibits Ebola pseudovirus infection in cell culture. Antiviral Res. 2016;125:1–7. doi:10.1016/j.antiviral.2015.11.003
54. Zhou N, Pan T, Zhang J, et al. Glycopeptide antibiotics potently inhibit cathepsin L in the late endosome/lysosome and block the entry of Ebola virus, Middle East respiratory syndrome coronavirus (MERS-CoV), and severe acute respiratory syndrome coronavirus (SARS-CoV). J Biol Chem. 2016;291(17):9218–9232. doi:10.1074/jbc.M116.716100
55. Leader DP. Viral protein kinases and protein phosphatases. Pharmacol Ther. 1993;59(3):343–389. doi:10.1016/0163-7258(93)90075-O
56. Chao SH, Price DH. Flavopiridol inactivates P-TEFb and blocks most RNA polymerase II transcription in vivo. J Biol Chem. 2001;276(34):31793–31799. doi:10.1074/jbc.M102306200
57. Schang LM. Advances on cyclin-dependent kinases (CDKs) as novel targets for antiviral drugs. Curr Drug Targets Infect Disord. 2005;5(1):29–37. doi:10.2174/1568005053174609
58. Mohr EL, McMullan LK, Lo MK, et al. Inhibitors of cellular kinases with broad-spectrum antiviral activity for hemorrhagic fever viruses. Antiviral Res. 2015;120:40–47. doi:10.1016/j.antiviral.2015.05.003
59. Cozza G, Fortuna M, Meggio F, et al. Hydrophobic derivatives of glycopeptide antibiotics as inhibitors of protein kinases. Biochemistry (Moscow). 2018;83(10):1222–1230. doi:10.1134/S0006297918100073
60. Kim SH, Kang CI, Huh K, et al. Evaluating the optimal dose of teicoplanin with therapeutic drug monitoring: not too high for adverse event, not too low for treatment efficacy. Eur J Clin Microbiol Infect Dis. 2019;38:2113–2120. doi:10.1007/s10096-019-03652-6
61. Langford BJ, So M, Raybardhan S, et al. Bacterial co-infection and secondary infection in patients with COVID-19: a living rapid review and meta-analysis. Clin Microbiol Infect. 2020. doi:10.1016/j.cmi.2020.07.016
62. Lai CC, Wang CY, Hsueh PR. Co-infections among patients with COVID-19: the need for combination therapy with non-anti-SARS-CoV-2 agents? J Microbiol Immunol Infect. 2020. doi:10.1016/j.jmii.2020.05.013
63. Rawson TM, Moore LS, Zhu N, et al. Bacterial and fungal co-infection in individuals with coronavirus: A rapid review to support COVID-19 antimicrobial prescribing. Clini Infect Dis. 2020. doi:10.1093/cid/ciaa530
64. Zhou F, Yu T, Du R, et al. Articles Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: a retrospective cohort study. Lancet. 2020;6736:1–9.
65. Cavalcanti AB, Goncalves AR, Almeida CS, Bugano DD, Silva E Teicoplanin versus vancomycin for proven or suspected infection. Cochrane Database Syst Rev. 2010;(6):CD007022. doi:10.1002/14651858.CD007022.pub2.
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
