Back to Journals » Infection and Drug Resistance » Volume 13
Genetic and Virulence Characteristics of Linezolid and Pretomanid Dual Drug-Resistant Strains Induced from Mycobacterium tuberculosis in vitro
Authors Hu M , Fu L, Wang B, Xu J, Guo S, Zhao J, Li Y, Chen X, Lu Y
Received 6 April 2020
Accepted for publication 16 May 2020
Published 12 June 2020 Volume 2020:13 Pages 1751—1761
DOI https://doi.org/10.2147/IDR.S257145
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Suresh Antony
Minghao Hu,1 Lei Fu,1 Bin Wang,1 Jian Xu,1 Shaochen Guo,1 Jiaojie Zhao,1 Yuanyuan Li,1 Xiaoyou Chen,2 Yu Lu1
1Department of Pharmacology, Beijing Key Laboratory of Drug Resistance Tuberculosis Research, Beijing Tuberculosis and Thoracic Tumor Research Institute, Beijing Chest Hospital, Capital Medical University, Beijing 101149, People’s Republic of China; 2Department of Tuberculosis, Beijing Chest Hospital, Capital Medical University, Beijing Tuberculosis and Thoracic Tumor Research Institute, Beijing 101149, People’s Republic of China
Correspondence: Yu Lu; Xiaoyou Chen Email [email protected]; [email protected]
Purpose: Linezolid (LZD) and pretomanid (PA-824) are promising candidates in regimens for the treatment of drug-resistant tuberculosis. However, research on LZD and PA-824 dual drug-resistant (LPDR) strains is rarely reported. This study aimed to investigate the genotypic and virulence characteristics of LPDR strains.
Methods: To obtain the LPDR strains (marked as LP or PL strains), we used a two-way induction method, namely, we first induced LZD- or PA-824-resistant mutants from the parental Mycobacterium tuberculosis (MTB) strain H37Rv in vitro, then we obtained the LPDR strains from induction of LZD- or PA-824-resistant mutants. Mutations in rplC, rrl, or ddn and fgd1 were identified in all mutants. To investigate the virulence of these strains, six strains were selected as representative strains, including LZD-resistant strains, PA-824-resistant strains and LPDR strains. We performed the animal survival study as virulence of MTB can be measured as survival time of an animal after being infected.
Results: We induced 38 mutant strains of LZD and PA-824 mono or dual drug resistance from H37Rv in vitro. The mutation frequency of rplC (C154R) gene in LPDR strains was 100% and 86%, respectively. In the animal survival study, animals infected with different drug-resistant strains survived significantly longer than those infected with H37Rv; animals infected with LPDR strains and PA-824-resistant strains survived similarly and both of which survived significantly shorter than those infected with LZD-resistant strains.
Conclusion: Our study showed that rplC gene had a high mutation frequency in LPDR strains. The virulence of LPDR strains was similar to PA-824-resistant strains, and the virulence of the LZD-resistant strains was weaker than PA-824-resistant strains.
Keywords: Mycobacterium tuberculosis, linezolid, PA-824, dual drug resistance, genotype, virulence
Introduction
Tuberculosis (TB) is a chronic infectious disease caused by Mycobacterium tuberculosis (MTB) and is the leading cause of death from a single infectious agent (above HIV/AIDS). The global burden of TB remains high, with an estimated 10 million incident cases and 1.5 million deaths in 2018.1 Drug-resistant TB continues to be a public health crisis; globally, approximately 500,000 people developed TB that was resistant to rifampicin (RR-TB), of which 78% had multidrug-resistant TB (MDR-TB).1 The prevention and treatment of drug-resistant tuberculosis continues to be a serious challenge facing medical workers. In 2018, global treatment success rates of MDR/RR-TB was relatively low at 56%.1 However, the existing drug combinations for MDR-TB often have adverse reactions with the use of second-line drugs and usually require long treatment durations. Moreover, only three new anti-TB drugs have been approved in the past 40 years, including bedaquiline, delamanid, and pretomanid (PA-824, Pa) (recently approved by the US Food and Drug Administration under the Limited Population for Antibacterial and Antifungal Drugs pathway, as part of the BPaL combination).2 The lack of new drugs targeting MDR-TB makes the disease more difficult to treat.
Among the repurposed and new anti-TB drugs, linezolid (LZD) and PA-824 have demonstrated antimycobacterial efficacy.3–6 LZD is an oxazolidinone class of anti-TB drug, which inhibits protein synthesis. LZD was classified as group 5 (last group) for the treatment of MDR-TB by WHO in 2014, whereas it is now classified as group A—the preferred drug group for MDR-TB and RR-TB treatment.7 Therefore, LZD plays a critical role in the treatment of drug-resistant tuberculosis. PA-824 belongs to the nitroimidazole class of antibiotics, which is presently undergoing Phase 3 trials for the treatment of drug-resistant TB. Fortunately, a few currently completed or ongoing clinical trials may bring hope for the treatment of MDR-TB based on new and repurposed anti-TB drugs. Such as the inexpensive and short (9-month) Bangladesh regimen which contains clofazimine yielded an 84% successful treatment outcome,8 a shorter (6-month) bedaquiline-containing regimen (levofloxacin, LZD, and clofazimine as an optimized background regimen) reached a 73% treatment success rate in South Africa.9 Further, the first clinical trial named Nix-TB trial (NCT02333799), which aims to test a novel regimen BPaL, consists of three drugs (bedaquiline, PA-824, and LZD) and holds the potential to be a shorter, all-oral, and affordable treatment for all types of TB, especially for MDR–TB and extensively drug-resistant tuberculosis (XDR-TB). Encouragingly, the Nix-TB clinical study reached a durable high cure rate (90%) at 6 months after completing treatment from 98 patients in the intention-to-treat analysis.2 Clearly, the combined regimen of LZD and PA-824 in the treatment of drug-resistant TB has great prospect. However, because the Nix-TB trial is a simplified three-drug combination, more attention to the occurrence of drug resistance in this regimen is required. It has been reported that the frequency of spontaneous mutant MTB against PA-824 is relatively high (9.0 × 10–7), which is only slightly lower than that of isoniazid (1.3 × 10−6).10 Moreover, induction in vitro is also prone to producing PA-824-resistant MTB strains,11,12 which indicates that MTB resistant to PA-824 is likely to occur in clinical practice in the future. Indeed, with the gradual application of LZD in the clinical treatment of anti-TB, LZD-resistant MTB clinical isolates have also been reported.4,13-16 Recently, isolates with acquired resistance to both bedaquiline and LZD or bedaquiline and delamanid (analogue of PA-824) were obtained from patients in clinical trials.17,18 All of these studies indicate that long-term use of the Nix-TB regimen may lead to the emergence of double-resistant mutant strains against both LZD and PA-824. Thus, we should pay more attention to the occurrence of LZD and PA-824 dual-resistant (LPDR) TB strains in this simplified three-drug regimen.
Future research on the Nix-TB trial is expected to be applied to clinical practice. However, research on LPDR-TB strains is rarely reported. Usually, drug resistance mutations in the chromosome of MTB may have multiple effects on fitness cost, such as a reduction in the in vitro growth rate or reduced virulence or so-called compensatory mutations (ameliorate the fitness cost).19 Moreover, different strains of MTB may have variable degrees of virulence. Studies have shown that increasing degrees of drug resistance in clinic isolates (ie MDR or XDR-TB) is associated with reduced virulence.20 Attenuation of virulence in laboratory-manipulated MTB was also reported.21 But there is little information regarding the degree of virulence of the LPDR strains. Whether LPDR strains have similar biological characteristics to MDR-TB strains remains to be investigated. Generally, the standard terms “mortality” can be used for a description of virulence of MTB and can be defined as below: mortality is the percentage of infected animals that die and is also measured as survival time of an animal after being infected, another important parameter is bacterial load or burden.22 In this study, we first obtained the LPDR-TB strains through an induction approach in vitro and assessed their genetic characteristics.23 We then conducted growth analysis and compared the virulence with that of a mouse infection model for representative strains, which may have important implications for early detection of LPDR strains in clinical settings and help to predict their replication and virulence in TB patients.
Materials and Methods
Bacterial Strains and Growth Conditions
The M. tuberculosis strains used was the laboratory strain H37Rv (ATCC 27,294; American Type Culture Collection, Rockville, MD), which was frozen in aliquots and subcultured in Middlebrook 7H9 broth (Difco, USA) supplemented with 10% oleic acid-albumin-dextrose-catalase (OADC) (Becton-Dickinson, USA) and 0.05% Tween 80 at 37°C without disturbance for 15 days. Colony-forming unit (CFU) counting was carried out by plating serial 10-fold dilutions of the cultures on 7H10 agar medium supplemented with OADC. Colonies were counted after incubation at 37°C for 3 to 4 weeks and CFU counts were expressed as mean±standard deviation (SD). M. tuberculosis strains were diluted in phosphate buffered saline (PBS) before infection.
Minimum Inhibitory Concentration (MIC90) Determination
LZD (Ark Pharm, US) and PA-824 (Biochempartner, China) MICs were determined by the microplate Alamar Blue assay,24,25 using 2-fold dilutions ranging from 5 to 0.039 μg/mL and 2.5 to 0.019 μg/mL, respectively. M. tuberculosis (100 μL containing 2 × 105 CFU) was added to wells, yielding a final testing volume of 200 μL. The plates were incubated for 7 days at 37°C after which 12.5 μL of 20% Tween 80 and 20 μL of Alamar Blue were added to all wells. After incubation at 37°C for another 24 h, fluorescence was measured at an excitation wavelength of 530 nm and an emission wavelength of 590 nm. The MIC was defined as the lowest concentration eliciting a reduction in fluorescence of ≥90% relative to the mean fluorescence of replicate drug-free controls. M. tuberculosis H37Rv was used as a drug-susceptible control. The same methods were used to determine the MICs of LZD and PA-824 against mono or dual drug-resistant MTB strains.
Generation of Induced LPDR Mutants
The parental M. tuberculosis strain H37Rv was grown in Middlebrook 7H9 broth after thawing to late-log-phase. Cultures of 1 × 108 CFU/mL were obtained when the optical density (OD) at 570 nm, measured by a microplate reader, was 0.1. Cultures were diluted to a concentration of 104–105 CFU/mL with 1 × PBS and 200 μL spread on 7H10 agar plates (containing 10% OADC enrichment and 50 μg of cycloheximide, 200 U of polymyxin B, 50 μg of carbenicillin, and 20 μg of trimethoprim/mL) with 0.5 × MIC concentrations of LZD (0.15 μg/mL) or PA-824 (0.07 μg/mL) and drug-free control plates. All plates were incubated at 37°C for 3 to 4 weeks after which 2–3 well-grown single colonies were picked from the plates with LZD or PA-824 containing. Then, each single colony was homogenized, diluted, and repeated subcultures were carried out on three 7H10 agar plates; a plate with 2- fold higher drug (LZD or PA-824), a plate with the growth permitting drug concentration (same as which growth was picked from) and a drug-free control.23 This process was repeated until concentrations of 8 times MIC were reached for LZD or PA-824. Finally, LZD/PA-824-resistant mutants that grew well on 7H10 agar plates containing a concentration of 8 times MIC LZD or PA-824 were isolated.
The LZD-resistant mutants were respectively cultured to logarithmic growth phase and diluted to a concentration of 104–105 CFU/mL. 200 μL of the bacterial suspension was applied to 7H10 agar plates containing PA-824 at a concentration of 0.5 × MIC and LZD at a concentration of 8 times MIC, and drug-free control plates. All plates were incubated at 37°C for 3 to 4 weeks after which 2–3 well-grown single colonies were picked from the plates containing LZD and PA-824. Then, each single colony was homogenized, diluted, and repeated subcultures were respectively inoculated on three 7H10 agar plates; a plate with 2-fold increasing concentrations of PA-824 while the concentration of LZD remained unchanged, a plate with the growth permitting drug concentration (same as which growth was picked from) and a drug-free control.23 This process was repeated until concentrations of 8 times MIC were reached for LZD and PA-824. Finally, LZD and PA-824 dual drug-resistant MTB strains (LPDR mutants, marked as LP strains) were isolated and grown on 7H10 agar plates containing 8 times MIC of LZD and PA-824. A similar method was used to induce LPDR mutants, marked as PL strains from PA-824-resistant mutants. Experimental overview for generation of induced LPDR mutants is shown in Figure 1.
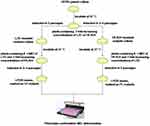 |
Figure 1 Experimental overview for generation of induced LZD and PA-824 dual drug-resistant MTB strains (LPDR mutants, marked as LP or PL strains) in vitro from M. tuberculosis H37Rv. |
DNA Sequencing
Extraction of genomic DNA from MTB strains was performed with freshly cultured bacteria, as reported previously.15 Crude DNA served as the template for PCR amplification to generate gene fragments from LZD and Pa mutant isolates. Sequencing of PCR products was performed using the Sanger method, with primers designed to be specific for rplC, rrl, or ddn and fgd1. The primers used for PCR were rplC-F (5′-TCCGCTCACCGCATAAGTACA-3′) and rplC-R (5′-CGATGTTGGCCGGGACGT-3′); rrl-F1 (5′-CCTGAGGCAACACTCGGACTT-3′) and rrl-R1 (5′-ACGGATTTGCCTATCGCTCT-3′); and rrl-F2 (5′-CCTAAGGCGAGGCCGACA G-3′) and rrl-R2 (5′-GGCCGCCGT AACTCTATGC-3′).26 The ddn gene was PCR amplified using primers ddn-F (5′-CGAGCGCACCGACCA GAGC-3′) and ddn-R (5′-GCATGGCCCGCAGGTGGACAA-3′); and fgd1-F (5′-CGTGGCCGC GAGCGAGGTGAA-3′) and fgd1-R (5′-CGCCCGAACCGTCAACAACACTGG-3′).27 PCR products were sent to Rui Biotech Company for sequencing. The sequencing results were analyzed by alignment against corresponding sequences of the reference M. tuberculosis strain H37Rv (ATCC 27294).
In keeping with the MIC values and typical mutation types of all the drug-resistant strains, we chose six mutant strains (including two LZD-resistant strains, two PA-824-resistant strains, and two LPDR strains; H37Rv strain as control) as the representatives for the following experiments.
Growth Kinetics of Drug-Resistant TB Strains
Bacterial suspensions of the six representative strains were diluted to a final concentration of 1 × 106 CFU/mL in a total volume of 20 mL and inoculated in 7H9 liquid medium for 35 days. Cultures were sampled at 0, 7, 14, 21, 28, and 35 days, diluted in PBS, and inoculated onto 7H10 solid medium for colony enumeration. Experiments were performed in triplicate and repeated twice.
Mice and Bacterial Infection
Specific pathogen free (SPF) BALB/c female mice (Beijing Vital River Laboratory Animal Technology Co., Ltd. China), weighing 16 to 19 g, were used in the present study. A total of 168 BALB/c mice were infected intravenously via the tail vein with 1 × 107 CFU of MTB strains per mouse. Mice were divided into seven groups and each group contained twenty-four mice. Eight mice each group were sacrificed at 4 weeks post-infection, of which body weight and weights of each lung and spleen were measured, and an organ coefficient was obtained by organ weight/body weight, as reported previously.28 Among them, lungs and spleens of 5 mice each group were removed, homogenized, and serially diluted in 10-fold steps in PBS. Aliquots (100 μL) were spread onto 7H10 plates in duplicate. The plates were incubated for organ CFU counts. The remaining 16 mice each group were allocated to the animal survival studies for 85 days post-infection. All animal studies were approved by the Ethics Committee of Beijing Chest Hospital, Capital Medical University. All procedures involving animal treatment were guided by the US Government Principles for the Utilization and Care of Vertebrate Animals Used in Testing, Research, and Training and were in accordance with the Public Health Service Policy on Humane Care and Use of Laboratory Animals.
Statistical Analysis
Organ CFU counts were log transformed before analysis and mean CFU counts were compared by one-way analysis of variance with the least significant difference (LSD) post hoc test to control for multiple comparisons. Survival analyses were performed using the Kaplan–Meier method and the log rank test was used to compare the observed differences in survival.29 All data were analyzed using SPSS software version 22 (IBM Corp., Armonk, NY, USA) and GraphPad Prism 7 (GraphPad Software, Inc.) with P values of <0.05 being considered significant.
Results
Phenotypic and Genotypic Resistance Characteristics of Drug-Resistant TB Strains
The H37Rv in vitro induction approach yielded 7 LZD-resistant strains within 5 passages (L1–L7); the MIC values of these mutants against LZD ranged from 5.99 μg/mL to 18.28 μg/mL. We also obtained 5 PA-824-resistant strains within 3–4 passages (P1–P5); the MIC values against PA-824 for all strains were more than 20 μg/mL. Moreover, 26 LPDR mutants within 8–9 passages were obtained (LP1–LP19 and PL1–PL7); the MIC values against LZD ranged from 4.88 μg/mL to 17.04 μg/mL and >20 μg/mL for PA-824 (except LP7, which was 18.60 μg/mL). As shown in Table 1, the MIC of the induced drug-resistant strains against both LZD and PA-824 were more than 8 times higher than that of wild type strain before induction. Therefore, the LPDR strains with phenotypic drug resistance were successfully obtained in the present study. The sequencing results of the rplC, rrl, ddn, and fgd1 genes were compared with the standard sequences of H37Rv (NCBI Reference Sequence: NC_000962.3). We found that the in vitro induced LZD-resistant strains all harbored rplC gene mutations (C154R) but no rrl gene mutations, the PA-824-resistant strains all harbored ddn gene mutations but no fgd1 gene mutations, and the LPDR mutations all occurred in the rplC gene (except PL2), with some harboring mutations in the ddn or fgd1 genes. Overall, the mutation frequency of rplC, ddn and fgd1 gene in different LPDR strains (marked as LP or PL strains) were 100%, 37%, 11% and 86%, 100%, 14%, respectively.
 |
Table 1 Linezolid (LZD) and Pretomanid (PA-824) Mono- or Dual-Resistant Mutant Strains of M. tuberculosis |
The Growth Kinetics of Drug-Resistant TB Strains
Referring to the MIC values of obtained mutants against LZD and PA-824, as well as the mutations of related genes, we selected L1 and L3 as LZD-resistant representative mutants, P1 and P3 as PA-824-resistant representative mutants, LP11 and PL7 as LDPR representative mutants, and H37Rv as the control strain. For the six representative strains, growth curves were plotted as shown in Figure 2. There were no significant differences in CFUs between different resistant strains (including LZD-resistant strains, PA-824-resistant strains, LPDR strains) and H37Rv.
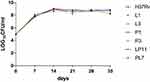 |
Figure 2 Growth of mutant strains through colony forming units (CFU) counts in an M. tuberculosis H37Rv background. |
Survival of BALB/c Mice Infected with Drug-Resistant TB Strains
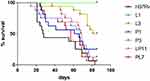
To compare the virulence of the six representative strains, BALB/c mice were infected intravenously via the tail vein and the survival of BALB/c mice post-infection were observed and recorded as shown in Figure 3. The median survival times of BALB/c mice were 27.5, undefined, 83, 63.5, 67, 68.5, and 57 days in the H37Rv, L1, L3, P1, P3, LP11 and PL7 groups, respectively. The median survival time of the mice infected with MTB mutant strains (L, P, LP, and PL groups) were significantly longer than that of H37Rv group (P < 0.0001). Additionally, the L group had a significantly longer median survival time than the P (L3 versus P3, P < 0.01), LP (L3 versus LP11, P < 0.01), or PL groups (L3 versus PL7, P < 0.001). The median survival time of LP group was similar to that of the P group (LP11 versus P1, P >0.05) and longer than that of the PL group but no difference statistically (LP11 versus PL7, P >0.05).
Organ Coefficient and CFU Counts of BALB/c Mice
To evaluate the effect of infection of the six representative strains on organs, we measured the weights of the lungs and spleens. When BALB/c mice were infected after 4 weeks, mice from the H37Rv group had the lowest body weight (16.95±2.83 g), while the L3 group (19.52 ± 1.66 g) mice were heavier and the L1 (19.98±0.58 g) group mice were the heaviest, both of which were significantly different to the H37Rv group (both P < 0.01). The body weights of the P1 group (17.34±2.31 g), P3 group (17.39±1.36 g), LP11 group (17.68±1.38 g), and PL7 group (17.77±1.55 g) mice were between the H37Rv group mice and the L group (L1 and L3) mice; the difference between the P group (P1 & P3) and L group was statistically significant (P < 0.05) as shown in Figure 4A. There were some differences in organ coefficients of the lung weights as indicated in Figure 4B, with the H37Rv group having the highest lung coefficient (3.39±1.39%), while the L1 group (2.06±0.21%; P < 0.01) and L3 group (2.35±0.63%; P < 0.05) had the lowest lung coefficients, the difference between the P1 group (2.88±1.22%) and L1 group was statistically significant (P < 0.05).The lung coefficients between the H37Rv group and the L groups in the P1 group, P3 group, LP11 group, and PL7 group were 2.88±1.22%, 2.72±0.42%, 2.82±0.38%, and 2.68±0.47%, respectively. For the organ coefficients of the spleen, the L1 group (2.66±0.34%) and L3 group (2.77±0.84%) had the lowest spleen coefficients; the P1 group (3.55±0.85%), P3 group (4.15±0.36%), LP11 group (3.89±0.31%), and PL7 group (3.60±0.41%) were slightly higher than the H37Rv group (3.32±0.69%). There was a statistically significant difference between the L1 group versus the H37Rv group (P < 0.05), and the L3 group versus the P1 or PL7 groups (P < 0.05, P < 0.01 respectively) as shown in Figure 4C. For the total lung viable CFU counts (lg CFU), as shown in Figure 4D, there was no significant difference between the groups except for the L1 group, which was lower. For the total spleen viable CFU—L3 group (8.56±0.31), P1 group (8.67±0.19), P3 group (8.63±0.31), and LP11 group (8.71±0.22)—counts were slightly higher than the H37Rv group (8.45±0.36), the L1 group (8.22±0.28), and the PL7 group (8.50±0.20), but there was no significant difference between the groups as shown in Figure 4E.
Discussion
To obtain the LPDR TB strains, we used a two-way induction method. In vitro, we first induced LZD or PA-824-resistant strains and then induced LPDR strains (LP or PL). As previously reported,30 the frequency of spontaneous mutation for LZD was 2×10−8–5×10−9 in vitro, which is lower than PA-824-resistant mutants (9.0×10−7). It is not surprising that the induction of LZD-resistant mutants was relatively difficult (5 passages). However, with a relatively low frequency of spontaneous mutation for LZD, we were still able to obtain LPDR strains within 8–9 passages. This implies that LPDR strains may emerge from the usage of LZD and PA-824 for a long time in clinical application.
For the LZD-resistant and LPDR strains, all mutants harbored the rplC (C154R) mutation but not the rrl mutation. This is consistent with previous studies that rplC (C154R) is the dominant mutation in LZD laboratory or clinical isolates.4,13,18,31 As reported, ddn, fgd1, or other genes (fbiA, fbiB, fbiC; not detected previously) are relative genes conferring PA-824 resistance in MTB;11,12 most mutant strains in the present study harbored ddn or fgd1 mutations in the PA-824-resistant and LPDR strains. Of note, two different mutations (V61G, Y89H) in 5 PA-824-resistant mutants were identified in ddn, which seems inconsistent with previous studies that diversity of mutations in ddn were discovered in 53 spontaneous in vitro-selected PA-824-resistant mutants.11 This may be due to a relatively few strains of mutants we finally obtained (5 versus 53), since four other different mutations (88S→STOP, G81S, I102T, T56A) in the LPDR strains (marked as LP strains) were identified in ddn. It should be noted that a few LPDR strains, such as LP11 and LP12, had no mutations in the ddn or fgd1 genes, which may indicate that these strains may have other known genes associated with PA-824 resistance or they have other resistance mechanisms. These LPDR strains will provide the possibility for the exploration of new mechanisms of LZD and PA-824 dual resistance, and which will be the focus of our subsequent research.
Based on the different frequencies of LZD and PA-824 mutants (the induction of LZD-resistant mutants was relatively difficult), we speculate that the development of PA-824 resistance may be the first step in the evolution of LPDR strain, just as the evolution of MDR develops from isoniazid resistance as a common first step.32,33 This suggests that the occurrence of PA-824 resistance should be monitored dynamically in the treatment of MDR-TB. In addition, the mutation frequency of rplC gene in different LPDR strains (marked as LP or PL strains) was 100% and 86%, respectively. This suggests that rplC (C154R) may be one of the dominant mutations in LPDR strains. Therefore, it will be of interest to determine whether rplC mutation can be used to predict the occurrence of LPDR strains in the Nix-TB regimen in the future. Notably, unlike new anti-TB drug PA-824, LZD can also be used for infections caused by Gram-positive bacteria, such as community-acquired pneumonia. Thus, patients may have started taking LZD due to related diseases before anti-TB treatment. In such cases, determination of rplC mutation may have important implications for early detection of LPDR strains.
In terms of growth characteristics of the six representative strains, our results is consistent with previous studies, which showed that mutations in rrl are usually associated with growth defects in MTB and M. smegmatis,30,34-36 while MTB strains harboring mutations in rplC are unimpaired.35 As with the results of the growth characteristics of the representative strains in vitro, there were no distinct differences in the total lung or spleen viable CFU counts in vivo after 4 weeks post-infection. This implies that the LPDR strains replicate normally in mice.
Despite recent studies that have reported various virulence factors of MTB, relatively few studies have provided information on the difference in virulence among MTB drug-resistant strains.20,37-39 We compared the virulence of the six representative strains mainly based on the survival time in the mice, and results of which were also consistent with results from organ coefficient. Actually, we had chosen three mice in each group for histopathology, but there were no distinct differences between groups in pathology (results not shown). One possible reason may be that commonly used BALB/c mice do not exhibit caseous pathology after infection with MTB.29
There are some limitations in our study. First, because this study used H37Rv as a parental strain, there might be some differences between the induced LPDR MTB strains in vitro and the clinical LPDR MTB isolates from patients; hence these mutants cannot completely cover the drug resistance of LZD and PA-824 in clinical practice. Second, the mechanisms of drug resistance of the representative strains obtained in current study have not been fully clarified. To better construct a more standardized LPDR-TB model for further research, such as utilizing them to screen for new compounds, the new mechanisms of both LZD and PA-824 resistant to LPDR strains should be better understood. Third, although our findings in the present study may have certain guiding significance for BPaL regimen in clinical settings, further studies should be performed to shed light on the evolution of the LZD- or PA-824-resistant mutants to the LPDR strains. We will explore the virulence factors and drug resistance mechanisms of the LPDR strains through whole genome-wide research at a later stage, which we hope will help us to adopt new strategies against MDR-TB, lay the foundation for rapid molecular target detection of LPDR strains, and promote new anti-TB drug target research.
In conclusion, we induced 38 mutants of LZD and PA-824 mono- or dual-resistant MTB strains from H37Rv in vitro and, for first time, used a mouse model to evaluate growth viability (replication or propagation) and virulence properties in six representative strains, including LZD-resistant strains, PA-824-resistant strains, and LPDR strains. Our study showed that rplC (C154R) gene had a high mutation frequency in LPDR strains; the virulence of LPDR strains was similar to PA-824-resistant strains, and the virulence of the LZD-resistant strains was weaker than PA-824-resistant strains. Our findings provided insights on the genetic and virulence properties of LPDR strains, which may have important implications for early detection of LPDR strains in clinical settings and help to predict their replication and virulence in TB patients. Moreover, the LPDR strain model can be used to screen for new compounds and provide the possibility for the exploration of new mechanisms of LZD and PA-824 dual resistance.
Acknowledgments
This work was supported by the National Mega-project for Innovative Drugs of China under Grant [2019ZX09721001-007-003].
Disclosure
The authors report no conflicts of interest in this work.
References
1. World Health Organization (WHO). Global tuberculosis report 2019. WHO CDS/TB/2019.20. Geneva, Switzerland: World Health Organization; 2019.
2. Conradie F, Diacon AH, Ngubane N, et al. Treatment of highly drug-resistant pulmonary tuberculosis. N Engl J Med. 2020;382(10):893–902. doi:10.1056/NEJMoa1901814
3. Dawson R, Diacon AH, Everitt D, et al. Efficiency and safety of the combination of moxifloxacin, pretomanid (PA-824), and pyrazinamide during the first 8 weeks of antituberculosis treatment: a phase 2b, open-label, partly randomised trial in patients with drug-susceptible or drug-resistant pulmonary tuberculosis. Lancet. 2015;385(9979):1738–1747. doi:10.1016/S0140-6736(14)62002-X
4. Lee M, Lee J, Carroll MW, et al. Linezolid for treatment of chronic extensively drug-resistant tuberculosis. N Engl J Med. 2012;367(16):1508–1518. doi:10.1056/NEJMoa1201964
5. Singh R, Manjunatha U, Boshoff HI, et al. PA-824 kills nonreplicating Mycobacterium tuberculosis by intracellular NO release. Science. 2008;322(5906):1392–1395. doi:10.1126/science.1164571
6. Sotgiu G, Pontali E, Migliori GB. Linezolid to treat MDR-/XDR-tuberculosis: available evidence and future scenarios. Eur Respir J. 2015;45(1):25–29. doi:10.1183/09031936.00145014
7. World Health Organization. Rapid Communication: Key changes to treatment of multidrug- and rifampicin-resistant tuberculosis (MDR/RR-TB). World Health Organization Document (2018).WHO/CDS/TB/2018.18.
8. Chiang CY, Van Deun A, Rieder HL. Gatifloxacin for short, effective treatment of multidrug-resistant tuberculosis. Int J Tuberc Lung Dis. 2016;20(9):1143–1147. doi:10.5588/ijtld.15.0884
9. Ndjeka N, Schnippel K, Master I, et al. High treatment success rate for multidrug-resistant and extensively drug-resistant tuberculosis using a bedaquiline-containing treatment regimen. Eur Respir J. 2018;52(6):6. doi:10.1183/13993003.01528-2018
10. Stover CK, Warrener P, VanDevanter DR, et al. A small-molecule nitroimidazopyran drug candidate for the treatment of tuberculosis. Nature. 2000;405(6789):962–966. doi:10.1038/35016103
11. Haver HL, Chua A, Ghode P, et al. Mutations in genes for the F420 biosynthetic pathway and a nitroreductase enzyme are the primary resistance determinants in spontaneous in vitro-selected PA-824-resistant mutants of Mycobacterium tuberculosis. Antimicrob Agents Chemother. 2015;59(9):5316–5323. doi:10.1128/AAC.00308-15
12. Manjunatha UH, Boshoff H, Dowd CS, et al. Identification of a nitroimidazo-oxazine-specific protein involved in PA-824 resistance in Mycobacterium tuberculosis. Proc Natl Acad Sci U S A. 2006;103(2):431–436. doi:10.1073/pnas.0508392103
13. Beckert P, Hillemann D, Kohl TA, et al. rplC T460C identified as a dominant mutation in linezolid-resistant Mycobacterium tuberculosis strains. Antimicrob Agents Chemother. 2012;56(5):2743–2745. doi:10.1128/AAC.06227-11
14. Richter E, Rusch-Gerdes S, Hillemann D. First linezolid-resistant clinical isolates of Mycobacterium tuberculosis. Antimicrob Agents Chemother. 2007;51(4):1534–1536. doi:10.1128/AAC.01113-06
15. Zhang Z, Pang Y, Wang Y, Liu C, Zhao Y. Beijing genotype of Mycobacterium tuberculosis is significantly associated with linezolid resistance in multidrug-resistant and extensively drug-resistant tuberculosis in China. Int J Antimicrob Agents. 2014;43(3):231–235. doi:10.1016/j.ijantimicag.2013.12.007
16. Pang Y, Zong Z, Huo F, et al. In vitro drug susceptibility of bedaquiline, delamanid, linezolid, clofazimine, moxifloxacin, and gatifloxacin against extensively drug-resistant tuberculosis in Beijing, China. Antimicrob Agents Chemother. 2017;61(10):10. doi:10.1128/AAC.00900-17
17. Bloemberg GV, Keller PM, Stucki D, et al. Acquired resistance to bedaquiline and delamanid in therapy for tuberculosis. N Engl J Med. 2015;373(20):1986–1988. doi:10.1056/NEJMc1505196
18. Zimenkov DV, Nosova EY, Kulagina EV, et al. Examination of bedaquiline- and linezolid-resistant Mycobacterium tuberculosis isolates from the Moscow region. J Antimicrob Chemother. 2017;72(7):1901–1906. doi:10.1093/jac/dkx094
19. Gygli SM, Borrell S, Trauner A, Gagneux S. Antimicrobial resistance in Mycobacterium tuberculosis: mechanistic and evolutionary perspectives. FEMS Microbiol Rev. 2017;41(3):354–373. doi:10.1093/femsre/fux011
20. Smith KL, Saini D, Bardarov S, et al. Reduced virulence of an extensively drug-resistant outbreak strain of Mycobacterium tuberculosis in a murine model. PLoS One. 2014;9(4):e94953. doi:10.1371/journal.pone.0094953
21. De Majumdar S, Sikri K, Ghosh P, et al. Genome analysis identifies a spontaneous nonsense mutation in ppsD leading to attenuation of virulence in laboratory-manipulated Mycobacterium tuberculosis. BMC Genomics. 2019;20(1):129. doi:10.1186/s12864-019-5482-y
22. Smith I. Mycobacterium tuberculosis pathogenesis and molecular determinants of virulence. Clin Microbiol Rev. 2003;16(3):463–496. doi:10.1128/CMR.16.3.463-496.2003
23. Ismail N, Omar SV, Ismail NA, Peters RPH. In vitro approaches for generation of Mycobacterium tuberculosis mutants resistant to bedaquiline, clofazimine or linezolid and identification of associated genetic variants. J Microbiol Methods. 2018;153:1–9. doi:10.1016/j.mimet.2018.08.011
24. Lu Y, Zheng M, Wang B, et al. Clofazimine analogs with efficacy against experimental tuberculosis and reduced potential for accumulation. Antimicrob Agents Chemother. 2011;55(11):5185–5193. doi:10.1128/AAC.00699-11
25. Xu J, Wang B, Hu M, et al. Primary clofazimine and bedaquiline resistance among isolates from patients with multidrug-resistant tuberculosis. Antimicrob Agents Chemother. 2017;61(6):6. doi:10.1128/AAC.00239-17
26. Zhang S, Chen J, Cui P, et al. Mycobacterium tuberculosis mutations associated with reduced susceptibility to linezolid. Antimicrob Agents Chemother. 2016;60(4):2542–2544. doi:10.1128/AAC.02941-15
27. Feuerriegel S, Koser CU, Bau D, et al. Impact of Fgd1 and ddn diversity in Mycobacterium tuberculosis complex on in vitro susceptibility to PA-824. Antimicrob Agents Chemother. 2011;55(12):5718–5722. doi:10.1128/AAC.05500-11
28. Cheng G, Hussain T, Sabir N, et al. Comparative study of the molecular basis of pathogenicity of M. bovis strains in a mouse model. Int J Mol Sci. 2018;20(1):1. doi:10.3390/ijms20010005
29. Ordonez AA, Tasneen R, Pokkali S, et al. Mouse model of pulmonary cavitary tuberculosis and expression of matrix metalloproteinase-9. Dis Model Mech. 2016;9(7):779–788. doi:10.1242/dmm.025643
30. Hillemann D, Rusch-Gerdes S, Richter E. In vitro-selected linezolid-resistant Mycobacterium tuberculosis mutants. Antimicrob Agents Chemother. 2008;52(2):800–801. doi:10.1128/AAC.01189-07
31. Makafe GG, Cao Y, Tan Y, et al. Role of the Cys154Arg substitution in ribosomal protein L3 in oxazolidinone resistance in mycobacterium tuberculosis. Antimicrob Agents Chemother. 2016;60(5):3202–3206. doi:10.1128/AAC.00152-16
32. Dye C, Espinal MA. Will tuberculosis become resistant to all antibiotics? Proc Biol Sci. 2001;268(1462):45–52. doi:10.1098/rspb.2000.1328
33. Koch A, Mizrahi V, Warner DF. The impact of drug resistance on Mycobacterium tuberculosis physiology: what can we learn from rifampicin? Emerg Microbes Infect. 2014;3(3):e17. doi:10.1038/emi.2014.17
34. Besier S, Ludwig A, Zander J, Brade V, Wichelhaus TA. Linezolid resistance in Staphylococcus aureus: gene dosage effect, stability, fitness costs, and cross-resistances. Antimicrob Agents Chemother. 2008;52(4):1570–1572. doi:10.1128/AAC.01098-07
35. McNeil MB, Dennison DD, Shelton CD, Parish T. In vitro isolation and characterization of oxazolidinone-resistant mycobacterium tuberculosis. Antimicrob Agents Chemother. 2017;61(10):10. doi:10.1128/AAC.01296-17
36. Sander P, Belova L, Kidan YG, Pfister P, Mankin AS, Bottger EC. Ribosomal and non-ribosomal resistance to oxazolidinones: species-specific idiosyncrasy of ribosomal alterations. Mol Microbiol. 2002;46(5):1295–1304. doi:10.1046/j.1365-2958.2002.03242.x
37. Montoya-Rosales A, Provvedi R, Torres-Juarez F, et al. lysX gene is differentially expressed among Mycobacterium tuberculosis strains with different levels of virulence. Tuberculosis (Edinb). 2017;106:106–117. doi:10.1016/j.tube.2017.07.005
38. Paolino M, Brindisi M, Vallone A, et al. Development of potent inhibitors of the Mycobacterium tuberculosis virulence factor zmp1 and evaluation of their effect on Mycobacterial survival inside macrophages. ChemMedChem. 2018;13(5):422–430. doi:10.1002/cmdc.201700759
39. Rifat D, Campodonico VL, Tao J, et al. In vitro and in vivo fitness costs associated with Mycobacterium tuberculosis RpoB mutation H526D. Future Microbiol. 2017;12(9):753–765. doi:10.2217/fmb-2017-0022
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.


