Back to Journals » OncoTargets and Therapy » Volume 13
GCNT4 is Associated with Prognosis and Suppress Cell Proliferation in Gastric Cancer
Authors Sun H , Chang J , Ye M, Weng W, Zhang M, Ni S , Tan C, Huang D, Wang L, Du X, Xu M , Sheng W
Received 9 February 2020
Accepted for publication 3 August 2020
Published 25 August 2020 Volume 2020:13 Pages 8601—8613
DOI https://doi.org/10.2147/OTT.S248997
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Federico Perche
Hui Sun,1,2,* Jinjia Chang,3,4,* Min Ye,1,3,5,* Weiwei Weng,1,3,5 Meng Zhang,1,3,5 Shujuan Ni,1,3,5 Cong Tan,1,3,5 Dan Huang,1,3,5 Lei Wang,1,3,5 Xiang Du,1,3,5 Mi-die Xu,1,3,5 Weiqi Sheng1,3,5
1Department of Pathology, Fudan University Shanghai Cancer Center, Shanghai 200032, People’s Republic of China; 2Department of Pathology, Eye, Ear, Nose and Throat Hospital, Fudan University, Shanghai 200031, People’s Republic of China; 3Department of Oncology, Shanghai Medical College, Fudan University, Shanghai 200032, People’s Republic of China; 4Department of Medical Oncology, Fudan University Shanghai Cancer Center, Fudan University, Shanghai 200032, People’s Republic of China; 5Institute of Pathology, Fudan University, Shanghai 200032, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Mi-die Xu; Weiqi Sheng
Department of Pathology, Fudan University Shanghai Cancer Center, 270 Dong’an Road, Shanghai 200032, People’s Republic of China
Tel +86-21-64175590
Fax +86-21-64174774
Email [email protected]; [email protected]
Background: GCNT4 is a member of the glucosaminyl (N-acetyl) transferases family that has been implicated in multiple human malignancies. However, the role of GCNT4 in gastric cancer (GC) is unknown. In this present study, we aimed to explore the role and clinicopathological correlation of GCNT4 in GC.
Materials and Methods: We first evaluated the dysregulation of GCNT4 in The Cancer Genome Atlas (TCGA) and then we performed RT-qPCR and immunohistochemistry to validate the results in a cohort of in-house patients. The clinicopathological correlation and function of GCNT4 in GC were also analysed.
Results: GCNT4 was found to be significantly downregulated in GC. In addition, GCNT4 expression correlated with tumour depth, nervous invasion and pathological tumor-node-metastasis (pTNM) stage. Moreover, lower GCNT4 levels conferred poor overall survival (OS) and disease-free survival (DFS) to GC patients. Multivariate Cox regression analysis revealed that GCNT4 protein expression is an independent prognostic factor for OS in patients with GC. Further functional experimental results revealed that overexpression of GCNT4 appears to halt GC cell proliferation and the cell cycle.
Conclusion: Altogether, these findings indicated that GCNT4 regulates the GC cell cycle and have important implications for the selection of therapeutic targets to prevent tumour proliferation.
Keywords: GCNT4, gastric cancer, dysregulation, cell cycle, prognosis
Introduction
Currently, gastric cancer (GC) is one of the most common cancers, although, with a reducing incidence worldwide, its heavy mortality rate still threatens patients in China, with 498,000 deaths and 679,100 new cases in 2015.3,15 Surgery is the standard treatment for GC at an early stage, and although surgery followed by adjuvant chemotherapy is the first choice for advanced GC, it is still difficult to increase long-term survival for patients with GC. Recent results from many years of continuous research have yielded two molecules, HER2 and VEGFR2, that are clinically utilized as therapeutic targets for GC. These great achievements revealed the urgency and importance of improving optimal biomarkers for clinical prognosis and treatment of patients with GC.
Members of the glucosaminyl (N-acetyl) transferase (GCNT) family are key mediators of mucin core structure synthesis, branching and oligomerization.13 Altered glycosylation is considered a universal cancer hallmark.17 Members of the GCNT family, including GCNT2, GCNT3 and GCNT4, have been previously identified as related to multiple human malignancies, such as colon,2,5,10,21 breast,9,24 prostate,8,11,18 hepatocellular,7 and pancreatic cancer.4,12,14 However, the relationship between the expression levels of GCNT family members and GC has not been fully investigated. The current study was aimed to explore the role of GCNT family in tumorigenesis and progression in gastric cancer.
To evaluate the dysregulation of GCNT family members in GC, we analysed the gene expression profile data in the online database—The Cancer Genome Atlas (TCGA) GC dataset. Then, we selected the most dysregulated gene, GCNT4, for further expression and function investigation.
Materials and Methods
Patients
The GC RNA-seq data from TCGA database (http://cancergenome.nih.gov/), which includes 360 GC and 37 unpaired normal gastric mucosa samples and the GEO GC datasets (GSE14210 and GSE15459, https://www.ncbi.nlm.nih.gov/geo) were used for the primary study. The GSE14210 data show gene expression data from human endoscopic biopsy samples of 145 metastatic gastric cancer patients prior to cisplatin and fluorouracil (CF) combination chemotherapy and 22 patients who initially responded to CF and then developed resistance to CF. The GSE15459 data show gene expression data from 200 primary gastric tumours from a Singapore patient cohort. The methylation and copy-number data of GCNT4 were download from TCGA GC dataset (http://www.cbioportal.org/).
A total of 194 patients analysed in this study underwent endoscopic biopsy (n = 29) of gastric intraepithelial neoplasia (GIN), or resection (n = 165) of primary GC at Fudan University Shanghai Cancer Center (FUSCC), and among these, 41 fresh frozen samples were obtained from the tissue bank of FUSCC, and the other 124 formalin-fixed paraffin-embedded (FFPE) blocks were enrolled from the archives of the Pathology Department of FUSCC. The GC diagnosis was histopathologically confirmed by two pathologists. The TNM classification system was used to evaluate the GC clinical stage.1 None of the patients underwent pre-operative treatment. Data included age, sex, and GC features and follow-up (eg, tumour size, histologic stage, depth of invasion, the status of vascular invasion, nervous invasion, lymphatic metastasis and peritoneal metastasis) were collected from all subjects as described previously.23 All patients provided written informed consent in this study, which was approved by The Research Ethics Committee of FUSCC. The study was conducted in compliance with the principles of the Helsinki Declaration.
Cell Lines and Culture Conditions
The human GC cell lines MKN-45, AGS, SGC-7901, MGC-803 and HGC-27, and the normal human gastric mucosa cell line GES-1 used in this study were cultured in 90% RMPI-1640 (Gibco) or 90% DMEM (Gibco Company, USA) supplemented with 10% foetal bovine serum (FBS, Gibco), 50 μg/mL streptomycin and 50 U/mL penicillin (Gibco), in a humidified atmosphere at 37°C and 5% CO2, as described previously23 The use of all the cell lines had approved by The Research Ethics Committee of FUSCC.
Transient Transfection
We amplified the full-length GCNT4 (NM_016591) sequence by PCR and then sub-cloned it into a pENTER vector (Transheep, Shanghai, China) using ClonExpress MultiS One Step Cloning Kit (Vazyme, Nanjing, China). The clone site was AsiSI/MluI. The primer of GCNT4 was shown as followed: GCNT4-AsiSI–F CTG CCG CCG CGA TCG CCG Cca cca tga agA TAT TCA AAT GTT ATT TTA AAC ATA C and GCNT4- MluI-R GCG GCC GCG TAC GCG ttg atg tgg taG TGA GAT TTC TA. GC cells (6 × 105) were seeded separately in 6-well plates for 24 h and transfected with pENTER- GCNT4, pENTER empty vector by lipofectamine 3000 reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions. Forty-eight hours after transfection, total RNA or protein was extracted from the harvested cells using Trizol reagent (Invitrogen, Carlsbad, CA, USA) or RIPA lysates buffer (Sigma, MO, U.S.A.), and GCNT4 expressions were measured using RT-qPCR or Western blotting as described below.
Analysis of Cell Proliferation and Cell Cycle
The ability of cell proliferation was evaluated using Cell Counting Kit-8 (CCK-8) (Dojindo, Kumamoto, Japan), and flow cytometry assays were performed to analyse cell cycle progression as described previously.20 Briefly, for the CCK-8 assay, 2 × 103 cells/well were incubated in 100 μL culture medium in 96-well plates. The cells were transfected with vector or pENTER–GCNT4 and incubated for 1, 2, 3, or 4 days before 10 μL CCK-8 (5 mg/mL) was added to the culture medium in each well. After 1 h incubation at 37°C, the absorbance at 450 nm was measured in a ThermoMax Microplate Reader (Molecular Devices); for cell cycle analysis, after 48-h post-transfection, the cells were collected and fixed with ethanol for 24 h. The cells were then washed by PBS twice and then stained with propidium iodide (PI, Calbiochem) for 20 min and analysed by flow cytometry analysis. Cells were stained with fixed with Ethanol overnight at −20°C followed by subsequent propidium iodide (PI, Calbiochem) staining for DNA content (cell cycle) analysis. Representative data from one of the three independent experiments are shown.
RNA Extraction and qRT-PCR Analysis
Total RNA was extracted with TRIzol (Invitrogen) according to the manufacturer’s instructions. Real-time RT-PCR was performed as previously described.23 The primers for RT-PCR analysis were purchased from Transheep (Shanghai, China). Primers were as follows: β-actin-F, 5ʹ- AGTCATTCCAAATATGAGATGCGTT-3ʹ, β-actin-R, 5ʹ- TGCTATCACCTCCCCTGTGT-3ʹ; and GCNT4-F, 5ʹ-GTTGTGGCAATGACCAGTGAT-3ʹ, GCNT4-R, 5ʹ- AGCATGGATAAGCCTTTCAACC-3ʹ. The relative expression of GCNT4 was calculated using the comparative cycle threshold (CT) (2−ΔΔCT) method with β-actin as the endogenous control for data normalization. The range of the obtained Ct values was 10–34. Each sample was analyzed in triplicate.
Immunohistochemistry (IHC) Staining
A total of 95 GC sample pairs and 29 gastric intraepithelial neoplasia (GIN) samples were studied. The 10 × 12 tissue microarray (TMA) was made by FUSCC Tissue Bank. IHC was performed on 7-μm-thick TMA sections using the antibody against GCNT4 (HPA 037431, rabbit polyclonal antibody; Sigma, Darmstadt, Germany; 1:200 dilution). Each case had two cores made from separate sources to preclude the heterogeneity of tumours. IHC staining was performed as described previously.22 PBS was used as a negative control. Each section was evaluated and scored independently by two pathologists as previously described.16
Western Blotting
Antibodies against P53 (#10442) were purchased from Protein Tech (Chicago, IL, USA). Antibodies against P57 (#DF2993), and P21 (#AF6290) were purchased from Affinity Bioscience (USA). Antibodies against GCNT4 (#138788) were purchased from Sigma-Aldrich (Darmstadt, Germany). Antibodies against cyclin D1 (#2978), cyclin B1 (#4138), cdc25B (#9525), and GAPDH (#2118) were bought from Cell Signalling Technology (Cambridge, MA, USA).
Cells were lysed in RIPA buffer (Sigma-Aldrich, Darmstadt, Germany), which contain a phosphatase inhibitor (Roche, Basel, Switzerland) and a protease inhibitor (Roche, Basel, Switzerland). As mentioned above, the BCA Protein Assay Kit (Thermo Scientific, USA) was selected to measure protein concentration.19 The cell lysates were added with sample buffer and boiled at 95°C for 5 min. The samples were transferred to 12% SDS-PAGE at 80 V for 3 h and then electrophoretically transferred to PVDF membranes (Millipore, Billerica, MA, USA) for another 2 h. The membranes then were blocked by 5% BSA for 30 min and incubated with primary antibodies at 4°C overnight and washed by 1% TBST for three times followed by incubated with secondary antibodies (1:2000, Cell Signaling Technology) for 1 h. The final detection of the substrates was performed with the ECL system (#32209, Thermo Fisher, USA). Protein expression levels were normalized to that of GAPDH.
Localization
Nuclear and cytoplasmic RNA of AGS and MGC-803 cells were separately extracted by using the PARIS (PARIS) kit (Life Technologies, USA), and then submitted to qRT-PCR analysis as mentioned previously.22 Of the control genes, MT RNR1 as the mitochondrial gene was expressed in the cytoplasm, whereas U6 as the nuclear transcript was expressed in the nucleus. β-actin was used for normalization.
Statistical Analysis and Gene Set Enrichment Analysis (GSEA)
Statistical analysis was carried out using R Studio (R version 3.5.1), SPSS 22.0 or GraphPad Prism EdgeR from R Studio was used to identify different expression of GCNT4 in TCGA GC dataset, the P values were determined by the Benjamini-Hochberg method. Comparisons between groups were determined by a Student’s t-test, Wilcoxon rank-sum test or a one-way ANOVA followed by Tukey’s multiple comparison tests. A two-tailed χ2 test was used to evaluate the relationship between the clinicopathological features and GCNT4 expression. The survival data of GSE14210 and GSE15459 data were download from http://kmplot.com. The survival curves of FUSCC IHC cohort were estimated by Kaplan-Meier analysis, and P values were calculated by Log rank test. Univariate and multivariate Cox proportional hazard regressions analysis curves were used to estimate the individual hazard ratio (HR) for OS and DFS. HR with a 95% confidence interval (CI) was measured to estimate the hazard risk of individual factors. P < 0.05 was considered statistically significant, and all P values were two-sided.
GSEA was carried out on TCGA dataset using the java implementation available from www.broadinstitute.org/gsea. We divided GCNT4 expression into terciles and compared samples in the third tercile against samples in the first tercile, and then we carried out GSEA analysis using H hallmarks sets.6 The cell cycle KEGG pathway map was downloaded from https://www.kegg.jp.
Results
GCNT4 mRNA and Protein Expression in GC Tissues
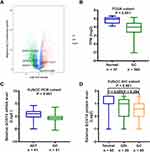
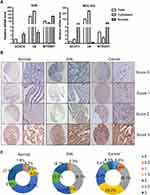
We first analysed the GCNT family (GCNT1, GCNT2, GCNT3, GCNT4, and GCNT7) gene expression levels in the TCGA GC dataset. GCNT4 was the most significantly dysregulated gene in GC (Figure 1A). Therefore, we focused further on the expression and clinicopathologic involvement of GCNT4 in GC. By analysing the GC RNA-seq data from TCGA, we found that the levels of GCNT4 mRNA were significantly higher in normal gastric mucous than those in tumour samples (P < 0.001, with DESeq2 test, Figure 1B).
We then validated the GCNT4 transcripts in 41 pairs of the primary GC tissues and adjacent normal mucosa. It turned out that the GCNT4 mRNA level was significantly decreased in cancer tissues compared to that in adjacent normal tissues (ANT; P = 0.039, with Wilcoxon rank-sum test, Figure 1C). To detect the expression of GCNT4 protein in GC, we performed GCNT4 IHC staining on a tissue microarray with 29 cases of GIN, 95 pairs of GC and normal mucosa samples which showed that GC and normal gastric mucosa expressed significantly different levels of GCNT4; (P < 0.001, with one-way ANOVA); however, there was no significant difference in GCNT4 expression between tumour tissue and GIN, or between GIN and ANT (P = 0.059 and P = 0.384, with one-way ANOVA, respectively; Figure 1D).
GCNT4 Subcellular Localization and Immunostaining
We used nuclear separation methods to extract nuclear and plasma RNA, and then detected the subcellular localization of GCNT4 mRNA in AGS and MGC-803 cells by RT-qPCR. The results indicated that GCNT4 mRNA is mainly located in the nucleus of GC cells (Figure 2A). In contrast, the positive immunostaining signals of GCNT4 were observed in the cell membrane and cytoplasm of benign and malignant gastric mucosa (Figure 2B). Low expression (score 0–1.5) was observed in 64 (67.3%) GC cases, 14 (46.7%) GIN cases and 20 (20.6%) normal cases; high expression (score 2–3) was observed in 31 (32.7%) GC cases, 16 (53.3%) GIN cases and 75 (79.4%) ANT cases (Figure 2C). In proportion, 27.4% normal and 16.7% of GIN samples scored 3, while only 6.5% gastric cancer samples scored 3.
GCNT4 Expression and Clinicopathologic Factors of GC
We divided 41 GC tissue samples into high GCNT4 mRNA level group and low GCNT4 mRNA level group according to the mean value of GCNT4 mRNA expression, and then analysed the correlation between the expression level of GCNT4 mRNA and the clinicopathological parameters of GC patients. However, we did not find any positive correlation (all P > 0.05; Table 1). The relationship between the expression levels of GCNT4 protein and the clinicopathological parameters of 95 cases GC patients were then analysed by Chi-square analysis, which demonstrated that the expression of GCNT4 protein was correlated with nervous invasion (P = 0.016, withχ2 test), tumour depth (P = 0.013, with χ2 test) and pTNM stage (P = 0.008, with χ2 test); whereas there was no significant relationship between the expression of GCNT4 protein and any other clinicopathologic features (Table 1).
 |
Table 1 Clinical Characteristics of Gastric Cancer Patients (χ2 Test) |
Association Between GCNT4 Expression and GC Patient Prognosis
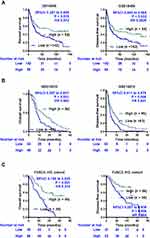
The overall survival (OS) and disease-free survival (DFS) curves reference to the expression levels of GCNT4 mRNA in the GSE14210 and GSE15459 data were plotted by using the Kaplan–Meier method. Compared with high GCNT4 expression group, GC patients with lower GCNT4 mRNA expression showed significantly shorter survival times in the GSE15459 (P = 0.015 for OS, and P = 0.032 for DFS, Log rank test, Figure 3A) and GSE14210 (P = 0.001 for OS, and P = 0.006 for DFS, Log rank test, Figure 3B) datasets. Next, we analysed the prognostic value of GCNT4 protein in the FUSCC IHC cohort. Analogously, patients with higher GCNT4 protein expression showed an obviously better OS (P = 0.006) and DFS (P = 0.002) than those of GC patients with lower GCNT4 expression levels (Figure 3C). Univariate analysis results of OS showed that the expression level of GCNT4 (P < 0.001), tumour depth (P = 0.008), tumour size (P = 0.020), nervous invasion (P = 0.021), tumour stage (P < 0.001), lymphatic metastasis (P = 0.017) and peritoneal metastasis (P < 0.001) were prognostic indicators (Table 2). Multivariate analysis results of OS showed that the peritoneal metastasis (P = 0.006), pTNM stage (P = 0.022) and GCNT4 expression (P < 0.001), all were independent prognostic indicator for OS in GC patients (Table 2). However, GCNT4 expression was not an independent prognostic indicator for DFS in patients with GC (data not shown).
 |
Table 2 Univariate and Multivariate Analyses of Clinicopathological Factors for Overall Survival in Gastric Cancer Patients (IHC Cohort, Cox Proportional Hazard Regressions Analysis) |
GCNT4 Halts GC Cell Proliferation and Cell Cycle in GC
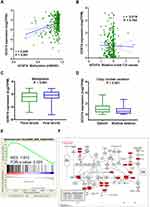
To investigate the possible mechanisms underlying GCNT4 downregulation in gastric cancer, we analysed the methylation and copy-number data of GCNT4 from TCGA GC dataset, which showed a large fraction of samples suffering methylation and copy-number alteration (shallow deletion) in GC (n = 369, Figure 4A and B). We further assessed the effect of copy number variation and methylation on GCNT4 expression. For methylation, we divided samples into terciles based on methylation and compared GCNT4 expression in hyper-methylated samples (third tercile) against rather unmethylated samples (first tercile), which showed that hyper-methylated samples exhibited lower GCNT4 expression (P < 0.001, Figure 4C). For copy number alteration, we compared GCNT4 expression in shallow deletion samples to diploid samples, which showed that shallow deletion samples exhibited lower GCNT4 expression than diploid samples (P < 0.001, Figure 4D).
We further applied GSEA to detect coherent changes of genes annotated to specific biological processes: we divided GCNT4 expression into terciles and compared samples in the third tercile against samples in the first tercile, and then we carried out GSEA analysis using H hallmarks sets (Figure 4C and Supplementary Table 1) and found the G2/M checkpoint of cell cycle enriched (Figure 4E and F).
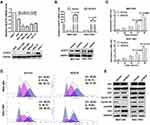
We then detected the baseline expression levels of GCNT4 in 5 GC cell lines (MGC-803, SGC-7901, AGS, HGC-27and MKN-45) and one normal gastric mucosal epithelial cell line (GES-1). The level of GCNT4 in GES-1 cell was observed to be significantly higher than that in gastric cancer cells (all P < 0.001, Figure 5A). Then, we selected SGC-7901 and MGC-803 cells for overexpression experiments; the efficiency of overexpression was verified by qRT–PCR and Western blotting (both P < 0.001, Student’s t-test, Figure 5B). Further cell function experiments in vitro showed that the up-regulation of GCNT4 suppressed GC cell proliferation (all P < 0.05, Student’s t-test, Figure 5C), inhibited cell cycle arrest at the G2/M phase (P < 0.05; Figure 5D). Western blotting showed that GCNT4 overexpression suppressed the G2/M phase by attenuating the expression of cdc25B and CyclinB1. However, GCNT4 has no effect on the G1/S phase-related molecules including P21, P27, P57 and CyclinD1 (Figure 5E).
Discussion
GCNT4 is a key mediator of mucin core structure synthesis, branching and oligomerization.13.Low expression of GCNT4 has been reported in colon21.and breast9.cancer, and its high expression conferred a poorer prognosis in breast cancer.9.However, the relationship between GCNT4 expression and GC had not yet been fully investigated.
In the present study, we examined the expression of GCNT4 in the TCGA GC datasets and found that GCNT4 mRNA was downregulated in GC, which was confirmed by using samples from the FUSCC cohort. There was no significant difference in GCNT4 expression between tumour tissue and GIN or between GIN and normal gastric mucous; these results suggest that GCNT4 may not be downregulated during the development of GC. However, the significant difference between the results for GC and those of normal gastric mucous still suggests a mechanism and clinical implication of GCNT4 in GC.
The correlation of GCNT4 expression with survival time in patients with GC in GSE14210 and GSE15459 datasets showed that GCNT4 is a positive tumour prognostic marker. As we were limited by the sample size, we did not validate the survival curve in our FUSCC PCR cohort. Specifically, the clinicopathological correlation in the FUSCC IHC cohort implied that GCNT4 expression was associated with GC carcinogenesis and progression. To reach a conclusion, we further analysed the prognostic potential of GCNT4 in the FUSCC IHC cohort; the results demonstrated that a lower GCNT4 expression was significantly correlated with a shorter OS and DFS in GC. Given that the primary GC tissue samples in GSE 14210 and 15459 were collected from patients in America and Singapore, respectively, the prognostic significance of GCNT4 in these two GEO datasets and our in-house cohort provide an extensive potential for GCNT4 to be utilized as a universally applicable biomarker in GC. Future studies should be conducted to validate the clinical significance of GCNT4 protein expression in GC samples from multiple centres to confirm its prognostic value for GC patients.
From coherently changed pathways detected with GSEA in the GCNT4 low expression phenotype, we speculated that overexpression of GCNT4 might halt the G2/M checkpoint in GC cells. The relationship of GCNT4 with gastric cancer progression was further confirmed by our study using cell cycle and Western blotting experiments, which indicated that GCNT4 can act as a tumour suppressor and induce tumour growth arrest. Further endeavours should focus on the role and underlying mechanism of GCNT4 in GC cell growth of GC cells.
The major strength of this study was that the information regarding GCNT4 expression in GC was obtained from four independent populations (TCGA, GSE14210, GSE15459 datasets and the FUSCC cohort) and that, for the first time, we illustrated the role of GCNT4 in GC. Although two GEO databases showed a significant prognostic correlation of GCNT4 expression and GC, we did not obtain a consistent result from the TCGA cohort (data not shown). As there were many factors such as patients’ race, social status, sampling techniques, as well as the response to adjuvant therapy affecting GC patients’ overall survival, the biomarkers from a single centre or region are not sufficient. Second, further in vivo functional studies are warranted to confirm our in vitro findings. Third, we mainly focused on the clinical significance of GCNT4 in GC tissues; further studies should focus on the expression and early diagnostic significance of GCNT4 in the peripheral blood from patients with GC.
In summary, we presented evidence that low expression of GCNT4 at both the mRNA and protein levels is associated with in GC. Thus, that GCNT4 might serve as a promising prognostic biomarker and therapeutic target for GC.
Disclosure
The authors declare no conflicts of interest.
References
1. Amin MB, Greene FL, Edge SB, et al. The eighth edition AJCC cancer staging manual: continuing to build a bridge from a population-based to a more “personalized” approach to cancer staging. CA Cancer J Clin. 2017;67:93–99. doi:10.3322/caac.21388
2. Chao CC, Wu PH, Huang HC, et al. Downregulation of miR-199a/b-5p is associated with GCNT2 induction upon epithelial-mesenchymal transition in colon cancer. FEBS Lett. 2017;591:1902–1917.
3. Chen W, Zheng R, Baade PD, et al. Cancer statistics in China, 2015. CA Cancer J Clin. 2016;66:115–132. doi:10.3322/caac.21338
4. Gonzálezvallinas M, Molina S, Vicente G, et al. Expression of microRNA-15b and the glycosyltransferase GCNT3 correlates with antitumor efficacy of rosemary diterpenes in colon and pancreatic cancer. PLoS One. 2014;9:e98556. doi:10.1371/journal.pone.0098556
5. Gonzálezvallinas M, Vargas T, Morenorubio J, et al. Clinical relevance of the differential expression of the glycosyltransferase gene GCNT3 in colon cancer. Eur J Cancer. 2015;51:1–8. doi:10.1016/j.ejca.2014.10.021
6. Kim M, Kim JH, Jang HR, et al. LRRC3B, encoding a leucine-rich repeat-containing protein, is a putative tumor suppressor gene in gastric cancer. Cancer Res. 2008;68:7147–7155. doi:10.1158/0008-5472.CAN-08-0667
7. Liu T, Zhang S, Chen J, et al. The transcriptional profiling of glycogenes associated with hepatocellular carcinoma metastasis. PLoS One. 2014;9:e107941. doi:10.1371/journal.pone.0107941
8. Jotaro M, Yuki T, Tohru Y, et al. I‐branching N‐acetylglucosaminyltransferase regulates prostate cancer invasiveness by enhancing α5β1 integrin signaling. Cancer Sci. 2015;107:359–368.
9. Mildelangosch K, Karn T, Schmidt M, et al. Prognostic relevance of glycosylation-associated genes in breast cancer. Breast Cancer Res Treat. 2014;145:295–305. doi:10.1007/s10549-014-2949-z
10. Nakamura K, Yamashita K, Sawaki H, et al. Aberrant methylation of GCNT2 is tightly related to lymph node metastasis of primary CRC. Anticancer Res. 2015;35:1411–1421.
11. Petrosyan A, Holzapfel MS, Muirhead DE, et al. Restoration of compact golgi morphology in advanced prostate cancer enhances susceptibility to galectin-1-induced apoptosis by modifying mucin O-glycan synthesis. Mol Cancer Res. 2014;12:1704–1716. doi:10.1158/1541-7786.MCR-14-0291-T
12. Rao CV, Janakiram NB, Madka V, et al. Small molecule inhibition of GCNT3 disrupts mucin biosynthesis and malignant cellular behaviors in pancreatic cancer. Cancer Res. 2016;76:1965. doi:10.1158/0008-5472.CAN-15-2820
13. Rinaldi M, Dreesen L, Hoorens PR, et al. Infection with the gastrointestinal nematode Ostertagia ostertagi in cattle affects mucus biosynthesis in the abomasum. Vet Res. 2011;42:61. doi:10.1186/1297-9716-42-61
14. Satomura Y, Sawabu N, Takemori Y, et al. Expression of various sialylated carbohydrate antigens in malignant and nonmalignant pancreatic tissues. Pancreas. 1991;6:448. doi:10.1097/00006676-199107000-00012
15. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2018. CA Cancer J Clin. 2018;68:7–30. doi:10.3322/caac.21442
16. Sun H, Ni SJ, Ye M, et al. Hedgehog interacting protein 1 is a prognostic marker and suppresses cell metastasis in gastric cancer. J Cancer. 2018;9:4642–4649. doi:10.7150/jca.27686
17. Vajaria BN, Patel PS. Glycosylation: a hallmark of cancer? Glycoconj J. 2017;34:147–156. doi:10.1007/s10719-016-9755-2
18. Wang L, Mitoma J, Tsuchiya N, et al. An A/G polymorphism of core 2 branching enzyme gene is associated with prostate cancer. Biochem Biophys Res Commun. 2005;331:958–963. doi:10.1016/j.bbrc.2005.04.022
19. Wang YQ, Xu MD, Weng WW, et al. BCL6 is a negative prognostic factor and exhibits pro-oncogenic activity in ovarian cancer. Am J Cancer Res. 2015;5:255–266.
20. Weng W, Ni S, Wang Y, et al. PTTG3P promotes gastric tumour cell proliferation and invasion and is an indicator of poor prognosis. J Cell Mol Med. 2017;21:3360–3371. doi:10.1111/jcmm.13239
21. Wu F, Yuan G, Chen J, et al. Network analysis based on TCGA reveals hub genes in colon cancer. Contemp Oncol. 2017;21:136–144. doi:10.5114/wo.2017.68622
22. Xu MD, Dong L, Qi P, et al. Pituitary tumor-transforming gene-1 serves as an independent prognostic biomarker for gastric cancer. Gastric Cancer. 2016;19:107–115. doi:10.1007/s10120-015-0459-2
23. Xu MD, Wang Y, Weng W, et al. A positive feedback loop of lncRNA-PVT1 and FOXM1 facilitates gastric cancer growth and invasion. Clin Cancer Res. 2017;23:2071–2080. doi:10.1158/1078-0432.CCR-16-0742
24. Zhang H, Meng F, Wu S, et al. Engagement of I-branching {beta}-1, 6-N-acetylglucosaminyltransferase 2 in breast cancer metastasis and TGF-{beta} signaling. Cancer Res. 2011;71:4846. doi:10.1158/0008-5472.CAN-11-0414
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.