Back to Journals » Infection and Drug Resistance » Volume 11
First molecular characterization of related cases of healthcare-associated infections involving multidrug-resistant Enterococcus faecium vanA in Algeria
Authors Benammar S , Pantel A , Aujoulat F, Benmehidi M , Courcol R , Lavigne JP , Romano-Bertrand S, Marchandin H
Received 3 February 2018
Accepted for publication 22 May 2018
Published 17 September 2018 Volume 2018:11 Pages 1483—1490
DOI https://doi.org/10.2147/IDR.S164487
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Professor Suresh Antony
Sonia Benammar,1,2 Alix Pantel,3,4 Fabien Aujoulat,5 Messaoud Benmehidi,1,2 René Courcol,6,7 Jean-Philippe Lavigne,3,4 Sara Romano-Bertrand,5,8 Hélène Marchandin3,5
1Department of Microbiology, University Hospital Center Touhami Benflis, Batna, Algeria; 2Department of Medicine, University Batna 2, Batna, Algeria; 3Department of Microbiology, Nîmes University Hospital, Nîmes, France; 4Faculty of Medicine, National Institute of Health and Medical Research, INSERM U1047, University of Montpellier, Nîmes, France; 5HydroSciences Montpellier, CNRS, IRD, University of Montpellier, University Hospital of Montpellier-Nîmes, Montpellier, France; 6Faculty of Medicine, University of Lille, Lille, France; 7Department of Bacteriology, Institute of Microbiology, Lille University Hospital, Lille, France; 8Department of Infection Control, Montpellier University Hospital, Montpellier, France
Purpose: Vancomycin-resistant Enterococcus (VRE) faecium (VREfm) are highly resistant bacteria emerging worldwide and rarely studied using molecular tools in Algeria since their first report in 2006. The aim of the study was to investigate healthcare-associated infections (HAIs) involving the first VRE in Batna University Hospital, Algeria, and characterize isolates using molecular tools.
Patients and methods: Medical charts were reviewed for patients with VREfm. van genes were detected by multiplex polymerase chain reaction (PCR), and strains were characterized by automated repetitive sequence-based PCR (rep-PCR), multiplex rep-PCR, pulsed-field gel electrophoresis (PFGE), and multilocus sequence typing (MLST).
Results: During a 6-month period, VREfm infections occurred in four patients hospitalized in three wards. The four isolates were E. faecium vanA belonging to the hospital-adapted clonal complex 17. PCR-based methods did not discriminate the isolates but MLST and PFGE delineated a subgroup of three VREfm of identical pulsotype and sequence type (ST) 80 (yet identified for five isolates in the international PubMLST database) while the fourth isolate was of ST789 (not previously identified for a VREfm) and displayed an unrelated pulsotype. The three genotypically related isolates were recovered in patients who underwent surgery in the same department, suggesting an outbreak for which the source and route of transmission remained unidentified.
Conclusion: This first molecular epidemiology study of VRE in Algeria was useful in delimiting an outbreak involving three of the four HAI cases and revealed rarely encountered genotypes. Considering the threat and burden of VRE infections worldwide, particularly in the USA, and the late emergence in Algeria, our study supports the urgent need for improved and early adequate infection control measures to avoid VRE spread in North African hospitals.
Keywords: outbreak, genotyping, MLST, rep-PCR, PFGE, molecular epidemiology
Introduction
Vancomycin-resistant enterococci (VRE) raise major concerns in medical practice due to the limited therapeutic options. They have emerged as an important cause of healthcare-associated infections (HAIs) and outbreaks worldwide, associated with a high mortality in patients with impaired host defenses.1–3 Nowadays, the most commonly reported clinical VRE isolates are multidrug-resistant (MDR) VanA-type Enterococcus faecium, displaying high-level resistances to both vancomycin and teicoplanin, for which a clonal spread has been demonstrated, especially through the worldwide dissemination of the hospital-adapted clonal complex 17 (CC17).4–6 Vancomycin-resistant E. faecium (VREfm) first emerged under the selective pressure of glycopeptides, then spread by cross-transmission through hands of healthcare workers (HCWs), with antibiotics facilitating VRE implantation and colonization in carrier patients. Hence, surveillance of VRE is crucial to limit their spread and the occurrence of outbreaks. This requires the accurate identification of infections, the development of screening strategies for patients, HCWs and the hospital environment, and finally, the implementation of investigations to trace the source and the transmission routes of VRE.7 Molecular typing techniques, including pulsed-field gel electrophoresis (PFGE), multilocus sequence typing (MLST), other polymerase chain reaction (PCR)-based methodologies, and more recently whole-genome sequencing, have substantially improved the ability to discriminate enterococcal isolates and provided critical insights into the epidemiology of VRE.8–11
In Algeria, sporadic cases are notified across the country since the first report of a VREfm in 2006.12 However, only few studies identified the genetic support for vancomycin resistance in VRE13 and there is no previously published molecular epidemiology study of VRE.14 Here, we report the clinical and bacteriological characteristics of the first cases of HAIs involving VRE in the Batna University Hospital (North-East Algeria) identified during a 6-month period. To our knowledge, here, we performed the third study with the molecular characterization of E. faecium vanA in Algeria and reported the first VRE of sequence type (ST) 789 while performing the first molecular characterization of related cases of HAIs involving VRE in Algeria.
Patients and methods
Patients and bacterial isolates
Four enterococcal isolates recovered between November 2015 and April 2016 from clinical specimens in four patients hospitalized in the Batna University Hospital, a center of ~600 beds, were studied. Demographic data, clinical ward, isolation site, mono- or polymicrobial infection, underlying diseases, antimicrobial regimens, and clinical outcome were recorded.
Politic of infection control and prevention in the hospital
In case of MDR or emerging highly resistant (EHR) bacteria isolation, microbiologists alert the concerned ward and the infection prevention and control service, which then notify to the committee against nosocomial infections. According to the French health authority recommendations, standard precautions are reenforced, complementary precautions such as contact precaution are prescribed, and patient’s isolation in single room is implemented as far as possible, but rarely possible given the lack of single room in clinical wards.15 No dedicated team of HCWs is implemented, but care rounds are organized in order to take over patients colonized or infected by EHR bacteria in last. The screening of patients in contact with index cases is recommended but not always performed, depending on human and technical means. After patients’ discharge, their room is deeply cleaned and disinfected with Anios® products. Environmental investigation by surface sampling is not performed in routine.
Species identification, antimicrobial susceptibility testing, and resistance genotype determination
Strain identification was based on Gram staining, API 20 Strep system (bioMérieux, Marcy l’Etoile, France), and matrix-assisted laser desorption ionization-time of flight mass spectrometry (MALDI-TOF MS) (MALDI Biotyper Microflex™; Bruker Daltonics, Bremen, Germany), and confirmed by specific amplification of ddI genes encoding the d-Ala-d-Ala ligase as previously described.16 Antibiotic susceptibility testing was performed by using the disk diffusion method. For glycopeptides, minimum inhibitory concentrations (MICs) of vancomycin and teicoplanin were determined by the E-test method (bioMérieux). Results were interpreted according to the recommendations of the Clinical and Laboratory Standards Institute (CLSI) and MDR defined according to Magiorakos et al.17,18 Multiplex PCR assay allowing simultaneous and specific detection of vanA, vanB, vanC-1, and vanC-2/3 resistance genes was performed according to the method previously described with no previous DNA extraction and no further sequencing required.16
Molecular typing methods
Isolates were characterized by automated repetitive sequence-based PCR (rep-PCR) using the DiversiLab® Enterococcus kit and the DiversiLab® strain typing system (bioMérieux), multiplex rep-PCR, MLST (https://pubmlst.org/efaecium/), and PFGE.19–21 Multiplex PCR and MLST analyses were performed on bacterial genomic DNA extracted using the MasterPure™ kit as recommended by the manufacturer (Epicentre, Madison, WI, USA). For PFGE analysis, DNA was prepared in agarose plugs as previously described.22 Macrorestriction was then performed by 40 units of NotI endonuclease overnight at 37°C, and migration parameters were as follows: pulse ramp of 70–5 s during 40 h at 4.5 V/cm. Results were interpreted according to the method: strains were classified in the same rep-PCR cluster if the similarity score was >95%, multiplex rep-PCR and PFGE profiles were visually compared, and strain relatedness was estimated from the band difference between profiles.9,23 STs were identified in the E. faecium PubMLST database. Global optimal eBURST implemented by PHYLOViZ was used to construct a minimum-spanning tree based on allelic profiles of the four studied strains and those of all the VanA-type referenced strains deposited in the E. faecium PubMLST database (https://pubmlst.org/efaecium/) on September 5, 2017.24
Ethics statement
This study is an observational study that fell within routine practice with nonadditional diagnostic and monitoring procedures applied to the patient. Primary data derived from routine clinical care were retrospectively analyzed. Thus, research concerns only data, already collected with no intervention on the patient and therefore no risk for the patient. Ethics approval was obtained through the National Center for Biotechnology Research of Constantine, Algeria, and written informed consent for publication of clinical details was obtained from the patient or next of kin.
Results
Description of the four cases of VREfm infections and investigations
The four VREfm isolates were recovered during the first report of cases of infections involving VRE in the Batna University Hospital. As summarized in the timeline in Figure 1, the four patients were considered to have an HAI because of their length of hospital stay before VRE isolation. Patients 1, 3, and 4 experienced surgery in adjoining surgical rooms in the Unit for Medical and Surgical Emergencies (UMSE) over a 6-month period (related patients), patients 3 and 4 hospitalized in the Nephrology department, and patient 2 remained throughout his hospitalization in the Haematology ward having no contact with the three other patients (unrelated patient).
The index case (patient 1) was admitted to the Neurosurgery department on October 20, 2015 for the management of spina bifida hydrocephalus, and placement of a ventriculoperitoneal shunt was performed 1 week later. The patient received an antimicrobial therapy associating ceftriaxone, gentamicin, and metronidazole. While the neurological evolution was favorable, the patient presented with peritonitis requiring surgery performed in the UMSE on November 5, and treatment switch to imipenem and vancomycin.
Surgery was performed again on November 12 and 17 for recurrent peritonitis. Glycopeptide-susceptible E. faecium and Escherichia coli were identified from the first two samples of peritoneal pus sent to the laboratory (November 5 and 12), and VREfm was isolated from the peritoneal pus sampled on November 17 (Figure 1). No care sectoring of patient was performed in the Neurosurgery department, and the rectal screening of contact patients hospitalized in the ward was negative for VRE. Unfortunately, no screening was performed for patients in the UMSE.
Patient 2 was admitted to the Haematology department on November 5, 2015 for an acute myeloid leukemia treated with a standard immunosuppressive chemotherapy. Twenty days later, a blood culture was performed due to fever and an antimicrobial therapy with cefotaxime, gentamicin, and metronidazole was initiated. The laboratory identified a Streptococcus oralis resistant to cefotaxime, and the patient was given vancomycin on November 28. The second blood culture was sampled on December 3 due to persisting fever, and the treatment was changed to cefotaxime, amikacin, and vancomycin. This second blood culture and the third blood culture performed on December 5 were positive for VREfm. The patient died on December 7, as there is no effective alternative antimicrobial therapy available in Algeria (Figure 1). All rectal screening of contact patients hospitalized in the ward showed negative.
Patient 3 was admitted to the Nephrology department and then transferred to the UMSE for renal transplantation performed on December 20, 2015. The patient received ceftriaxone for 10 days. In the postoperative course, the patient was feverish, but all samples were negative. Vancomycin and later imipenem and ciprofloxacin were added to the treatment. On January 20, 2016, parietal infection was diagnosed on the basis of VREfm identification in a surgical wound fluid sample (Figure 1). The patient did not receive any antimicrobial agent because the VREfm was only susceptible to antibiotics without effectiveness at the infectious site. This patient was transferred in a local hospital and was alive in May 2018. Rectal samples were requested for patient 3 and contact cases in the Nephrology department and the UMSE but have not been performed.
Patient 4 with chronic renal failure and undergoing peritoneal dialysis was hospitalized in the Department of Nephrology for peritonitis in March 15, 2016, and antibiotic treatment (cefotaxime+gentamicin and then vancomycin+imipenem) was started. Peritoneal drainage was performed in the UMSE twice in April (22 and 24), and the patient also received colistin. A polymicrobial infection was identified as VREfm, Klebsiella pneumoniae, and Morganella morganii were identified from the peritoneal pus sampled during both surgical procedures. The patient died on April 25 (Figure 1). No rectal screening was performed either for the patient or for other patients.
Characterization of the four VREfm isolates
The four isolates were identified as E. faecium by biochemical method and MALDI-TOF MS (score >2.2) and confirmed by the specific amplification of the E. faecium ddI gene (unique amplification product of 550 bp). They displayed MDR to amoxicillin, gentamicin (high-level resistance), rifampin, levofloxacin, ciprofloxacin, vancomycin (MIC >256 mg/L), and teicoplanin (MICs 16–24 mg/L) while being susceptible to chloramphenicol, tigecyclin, fosfomycin, and nitrofurantoin. Variable susceptibilities were observed for erythromycin (isolate 1148 being the sole resistant isolate) and tetracycline (isolate 1148 being the sole-susceptible isolate), resulting in distinct antimicrobial susceptibility pattern A2 for isolate 1148 compared with isolates 90, 439, and 1043 (Figure 1). vanA gene was amplified from DNA extracted for the four isolates.
Molecular epidemiology study of the four E. faecium vanA isolates
Three isolates (1043, 90, and 439 recovered in patients 1, 3, and 4, respectively) showed identical molecular characteristics (Figure 1). Genotyping also revealed that MLST and PFGE were the most discriminatory methods showing strains of different STs, ST80 and ST789 (differing by 20 nt of the 556 analyzed for the atpA locus), and pulsotypes (unrelated pulsotypes P1 and P2 showing more than seven DNA fragments of difference) while rep-PCR did not clearly distinguish the four isolates (>95% of pattern similarity for semiautomated rep-PCR using the DiversiLab® system: pattern R1, one faint band of difference for multiplex rep-PCR: closely related patterns M1 and M1′) (Figure 1). Finally, both identified STs belonged to the CC17 and their position within CC17 is illustrated in Figure 2A. They were however only rarely identified to date, and the analysis of the PubMLST database found 1) five strains of ST80 E. faecium vanA (Figure 2B) isolated between 1997 and 2014 from blood cultures of four patients in Israel, Germany, and Russia, and the fifth isolate was from skin origin during a hospital survey in 2005 in Italy, and 2) a unique strain of ST789 E. faecium isolated from a blood culture in an hospitalized patient from South Korea in 2012 (data not shown), the strain was not registered as a vancomycin-resistant one, and we therefore identified the first VanA-type ST789 E. faecium (Figure 2C). The four isolates were therefore deposited in the E. faecium PubMLST database (https://pubmlst.org/efaecium/). All the molecular typing results supported the epidemiologic link between three of the four clinical isolates and a VREfm cross-transmission between patients 1, 3, and 4 for whom strains displayed both the same ST and pulsotype.
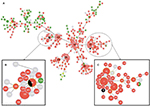  | Figure 2 (A) Minimum spanning tree generated using the goeBURST algorithm implemented in the PHYLOViZ software showing the position of the four studied strains (black) among the whole population of VanA-type strains (n=1,064) deposited in the Enterococcus faecium PubMLST database49 on September 5, 2016. Each circle corresponds to an ST. The number given in the circle corresponds to the ST designation. The size of each circle is proportional to the number of isolates of each ST. Red indicates strains of human origin, dark green indicates strains of animal origin, light green indicates strains of environmental origin, and gray indicates strains with no associated information on their origin. (B) Focus on cluster including strains of ST80. (C) Focus on cluster including the ST789 isolate. Abbreviation: ST, sequence type. |
Discussion
We reported the first VRE in Batna and the first E. faecium vanA in Algeria, consisting in four nonrepeat and consecutive strains isolated from clinical specimens of infected patients presenting with peritonitis, postoperatively infected wound, and bacteremia. The main risk factors for VRE nosocomial acquisition, such as prolonged hospital stay, use of broad-spectrum antimicrobials such as third-generation cephalosporins known to select enterococci due to the intrinsic resistance of enterococci to these antibiotics, severity of underlying diseases, and prior surgery, were found in the four patients included in this study.2,25–27 As observed in our study, VRE have been previously isolated from recurrent peritonitis, reported in patients undergoing chronically peritoneal dialysis or ventriculoperitoneal shunt, identified during healthcare-associated bacteremia, particularly in neutropenic patients and elderly, with a high risk of death, and are becoming important pathogens in several infections complicating solid organ transplantation.25,28–33 The four VREfm isolates characterized herein were MDR. This result is of great concern and consistent with other studies, which show a high-level resistance among enterococci, especially in E. faecium, and even more in VREfm.14,25,34 The treatment of these enterococcal infections in Algeria is challenging because tigecyclin, daptomycin, linezolid, and quinupristin–dalfopristin are not yet available, and effective therapy for patients infected by MDR VRE is consequently limited, as in our study for patients 2 and 3.
Initially, the major reservoir of VRE in Europe was the community healthy population contaminated from an animal source attributed to the widespread veterinary use of avoparcin during 1980s.35 Endemic in the USA since 1990s because of the overuse of vancomycin for methicillin-resistant Staphylococcus aureus and Clostridium difficile infections, VREfm was spread throughout the world and emerged in the European hospitals since 2004 with the report of outbreaks.5,6,35–39 In Batna hospital, the first VREfm was isolated at the end of 2015 and we assumed that this late emergence, compared to Europe and the USA, may be related to the never use of avoparcin in Algeria, the increased prescription of vancomycin since very few years, and the lowest level of enterococcal infections. As an example, 30 E. faecium strains were isolated during 2015 in our institution, all from patients with HAIs. As in Batna, very few reports have described VRE in other Algerian (http://www.sante.dz/aarn/alerte.htm) or North Africa hospitals.12,14,40,41 In Algeria, the first VRE identified in Algiers University Hospital was an E. faecalis isolate from a urine sample in 2006.12 More recently, single-center and multicenter epidemiological studies conducted on Enterococcus spp. isolated from human infections in three other cities located in north-eastern Algeria between 2010 and 2013 did not find any VRE while a study on MDR bacteria isolated from patients hospitalized in the Intensive Care Unit of the University Hospital of Constantine, the main regional hospital of North-East Algeria, between 2011 and 2015, found 10 VREfm, with no VRE identified during the first 2 years, and two, six, and two VREfm identified in 2013, 2014, and 2015, respectively.40,42,43 Finally, sporadic or small-scale outbreaks of VREfm were reported according to the Algerian Network for Monitoring the Resistance of Bacteria to Antibiotics depending on the year considered.44 However, only two previous reports included the identification of the genetic support for vancomycin resistance, the isolates were VanA-type E. faecium recovered from a blood culture in a 47-year-old severely burned patient in 201013 and from a surgical wound in a patient who carried the strain in his digestive tract in 2011, respectively.14 Our work is therefore the third one in which the vancomycin-resistant genetic support was identified in a VRE. It is also noteworthy that none of the previous study reporting VRE included genotyping. In this study, we showed that the four VREfm belonged to CC17, consistent with the literature reporting the currently worldwide dissemination of this high-risk enterococcal clonal complex, well adapted to the hospital.4–6 Genotyping helped us to delineate a subgroup of three patients infected by ST80 VREfm while the fourth patient was infected by an unrelated strain of ST789. PFGE confirmed that the three strains of ST80 were clonally related as they displayed indistinguishable fingerprints unrelated to that observed for the ST789 VREfm. As in previous studies, we showed that PCR-based methods had lower discriminative power than PFGE being unable to distinguish the four strains and thereby overestimating clonal spread.8,9 To the best of our knowledge, this is the first VRE outbreak in Algeria that has been investigated using molecular epidemiology tools. Interestingly, the four VREfm belonged to rarely described genotypes, ST789 being identified for the first time for a VRE while ST80 isolates previously described in Algeria were vancomycin susceptible and represented a minor genotype beside the major ST78 or ST17.42,43 These results emphasize the importance of genotyping the VRE isolated in Algeria with the aim to increase our knowledge on national epidemiology of the circulating clones and of performing molecular epidemiology to assess strain relatedness in case of clustered isolation of such EHR bacteria.
ST80 VREfm persistence in our institution is highly probable, as three clonally related strains have been identified during a 6-month period. Regarding investigations and infection control measures, Algeria had no specific recommendations for the management of EHR bacteria infections. In Batna, in the absence of infection preventionists and of a hospital hygiene laboratory specifically dedicated to the investigations and management of outbreaks, an alert is given by the Microbiology team to the ward concerned and to the infection prevention and control service and notified to the committee against nosocomial infections in case of MDR or EHR bacteria isolation. While the basic and specific hygiene procedures are reenforced, the French recommendations for the management of MDR or EHR cases are implemented as far as possible.15 However, during the outbreak period, detection of contact cases was incomplete despite requested in all the units concerned. These partial investigations did not find any patient carrying a VRE in both the Neurosurgery and the Haematology units, and it is noteworthy that no other VRE infection case was diagnosed in our institution during and since the 6-month outbreak period, despite the UMSE hosted a large number of patients and made thousand surgical interventions. Altogether, we were unable to find the source and the route of ST80 VREfm transmission between the index case and patients 2 and 4, but ST80 VREfm persistence in hospital environment in the UMSE is highly suspected given the ability of VRE to persist in hospital environment, even in nonoutbreak settings, and knowing that hospital environment plays an important role in the transmission of VRE.45–48
Conclusion
Considering the threat and burden of VRE infections worldwide, particularly in the USA, outbreak cases of HAIs involving multidrug VanA-type VREfm in Batna are worrisome and our study, revealing some defaults in the current management of such infections in Algeria, supports the urgent need for improved and early adequate infection control measures to avoid VRE spread in North African hospitals. This is particularly reinforced by the fact that antimicrobials potentially still efficient on MDR VRE are not available in Algeria, which greatly complicated the treatment of these severe and sometimes lethal bacterial infections. Therefore, implemented procedures and actions in the event of a VRE alert are urgently required, including edition of specific recommendations that could be applied in Algerian healthcare structures regarding labeling of patients, patient isolation, dedicated material use, care sectoring to limit contact patients, and ward disinfection/decontamination to eliminate a potential environmental reservoir. Regarding investigations, rectal screening of contact cases should be more systematically performed and requires a higher compliance of the medical staffs; environmental contamination detection should be promoted to identify the source and route of dissemination of VRE and the availability of rapid, affordable, and reliable techniques of molecular epidemiology is required for early diagnosis, surveillance, and investigation of potential outbreaks.
Acknowledgments
The authors warmly thank Professor Ammar Azioune, Professor Abdellatif Niar (CRBt of Constantine), and Professor Mourad Aribi (University of Tlemcen) for invaluable assistance with ethical issues; Pr Yamina Ouaghlent, Dr Ahmed Bougroura, Dr Mohamed Amrane, and Dr Salima Ait Madjber for authorizing access to patient and clinical data; and Dr Faiza Bouziane, France Joullié, and Agnès Masnou for laboratory assistance.
Disclosure
The authors report no conflicts of interest in this work.
References
Arias CA, Murray BE. The rise of the Enterococcus: beyond vancomycin resistance. Nat Rev Microbiol. 2012;10(4):266–278. | ||
O’Driscoll T, Crank CW. Vancomycin-resistant enterococcal infections: epidemiology, clinical manifestations, and optimal management. Infect Drug Resist. 2015;8:217–230. | ||
Reyes K, Bardossy AC, Zervos M. Vancomycin-resistant enterococci: epidemiology, infection prevention, and control. Infect Dis Clin North Am. 2016;30(4):953–965. | ||
Klare I, Konstabel C, Mueller-Bertling S, et al. Spread of ampicillin/vancomycin-resistant Enterococcus faecium of the epidemic-virulent clonal complex-17 carrying the genes esp and hyl in German hospitals. Eur J Clin Microbiol Infect Dis. 2005;24(12):815–825. | ||
Leavis HL, Bonten MJM, Willems RJL. Identification of high-risk enterococcal clonal complexes: global dispersion and antibiotic resistance. Curr Opin Microbiol. 2006;9(5):454–460. | ||
Top J, Willems R, Bonten M. Emergence of CC17 Enterococcus faecium: from commensal to hospital-adapted pathogen. FEMS Immunol Med Microbiol. 2008;52(3):297–308. | ||
Cattoir V, Leclercq R. Twenty-five years of shared life with vancomycin-resistant enterococci: is it time to divorce? J Antimicrob Chemother. 2013;68(4):731–742. | ||
Chuang YC, Wang JT, Chen ML, Chen YC. Comparison of an automated repetitive-sequence-based PCR microbial typing system with pulsed-field gel electrophoresis for molecular typing of vancomycin-resistant Enterococcus faecium. J Clin Microbiol. 2010;48(8):2897–2901. | ||
Werner G, Fleige C, Neumann B, Bender JK, Layer F, Klare I. Evaluation of DiversiLab®, MLST and PFGE typing for discriminating clinical Enterococcus faecium isolates. J Microbiol Methods. 2015;118:81–84. | ||
Freitas AR, Tedim AP, Francia MV, et al. Multilevel population genetic analysis of vanA and vanB Enterococcus faecium causing nosocomial outbreaks in 27 countries (1986–2012). J Antimicrob Chemother. 2016;71(12):3351–3366. | ||
Lytsy B, Engstrand L, Gustafsson Å, Kaden R. Time to review the gold standard for genotyping vancomycin-resistant enterococci in epidemiology: comparing whole-genome sequencing with PFGE and MLST in three suspected outbreaks in Sweden during 2013–2015. Infect Genet Evol. 2017;54:74–80. | ||
Aggoune N, Chabani A, Tiouit D, Naim M, Rahal K. Premier cas d’Enterococcus faecalis résistant à la vancomycine en Algérie [First case of vancomycin-resistant Enterococcus faecalis in Algeria]. Med Mal Infect. 2008;38(10):557–558. | ||
sante.dz/aarn [homepage on the Internet] Alertes du Réseau Algérien de la Surveillance de la Résistance des Bactéries aux Antibiotiques (A.A.R.N.): alerte du 29 mars 2011; alerte du 03 novembre 2010 [updated June 17, 2018]. Available from: http://www.sante.dz/aarn/alerte.htm. Accessed July 3, 2018. | ||
Hamidi M, Ammari H, Ghaffor M, et al. Emergence of glycopeptide resistant Enterococcus faecium in Algeria: a case report. Ann Biol Clin (Paris). 2013;71(1):104–106. | ||
Lepelletier D, Berthelot P, Lucet JC, et al. French recommendations for the prevention of ‘emerging extensively drug-resistant bacteria’ (eXDR) cross-transmission. J Hosp Infect. 2015;90(3):186–195. | ||
Dutka-Malen S, Evers S, Courvalin P. Detection of glycopeptide resistance genotypes and identification to the species level of clinically relevant enterococci by PCR. J Clin Microbiol. 1995;33(1):24–27. | ||
Clinical and Laboratory Standards Institute (CLSI). Performance Standards for Antimicrobial Susceptibility Testing. 26th ed. Wayne, PA: CLSI; 2016. CLSI supplement M100S. | ||
Magiorakos AP, Srinivasan A, Carey RB, et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect. 2012;18(3):268–281. | ||
Dupont C, Michon A-L, Jumas-Bilak E, Nørskov-Lauritsen N, Chiron R, Marchandin H. Intrapatient diversity of Achromobacter spp. involved in chronic colonization of cystic fibrosis airways. Infect Genet Evol. 2015;32:214–223. | ||
Homan WL, Tribe D, Poznanski S, et al. Multilocus sequence typing scheme for Enterococcus faecium. J Clin Microbiol. 2002;40(6):1963–1971. | ||
Jolley KA, Maiden MC. BIGSdb: scalable analysis of bacterial genome variation at the population level. BMC Bioinformatics. 2010;11:595. | ||
Murray BE, Singh KV, Heath JD, Sharma BR, Weinstock GM. Comparison of genomic DNAs of different enterococcal isolates using restriction endonucleases with infrequent recognition sites. J Clin Microbiol. 1990;28(9):2059–2063. | ||
Malathum K, Singh KV, Weinstock GM, Murray BE. Repetitive sequence-based PCR versus pulsed-field gel electrophoresis for typing of Enterococcus faecalis at the subspecies level. J Clin Microbiol. 1998;36(1):211–215. | ||
Francisco AP, Vaz C, Monteiro PT, Melo-Cristino J, Ramirez M, Carriço JA. PHYLOViZ: phylogenetic inference and data visualization for sequence based typing methods. BMC Bioinformatics. 2012;13:87. | ||
Cetinkaya Y, Falk P, Mayhall CG. Vancomycin-resistant enterococci. Clin Microbiol Rev. 2000;13(4):686–707. | ||
Bonten MJ, Willems R, Weinstein RA. Vancomycin-resistant enterococci: why are they here, and where do they come from? Lancet Infect Dis. 2001;1(5):314–325. | ||
Fossi Djembi L, Hodille E, Chomat-Jaboulay S, et al. Factors associated with vancomycin-resistant Enterococcus acquisition during a large outbreak. J Infect Public Health. 2017;10(2):185–190. | ||
Agudelo Higuita NI, Huycke MM [webpage on the internet]. Enterococcal disease, epidemiology, and implications for treatment. In: Gilmore MS, Clewell DB, Ike Y, et al., editors. Enterococci: From Commensals to Leading Causes of Drug Resistant Infection. Boston: Massachusetts Eye and Ear Infirmary; 2014. Available from: https://www.ncbi.nlm.nih.gov/books/NBK190429/. Accessed July 3, 2018. | ||
Lai KK. Treatment of vancomycin-resistant Enterococcus faecium infections. Arch Intern Med. 1996;156(22):2579–2584. | ||
Alatorre-Fernández P, Mayoral-Terán C, Velázquez-Acosta C, et al. A polyclonal outbreak of bloodstream infections by Enterococcus faecium in patients with hematologic malignancies. Am J Infect Control. 2017;45(3):260–266. | ||
Satlin MJ, Walsh TJ. Multidrug-resistant Enterobacteriaceae, Pseudomonas aeruginosa, and vancomycin-resistant Enterococcus: three major threats to hematopoietic stem cell transplant recipients. Transpl Infect Dis. Epub 2017 Oct 25;19(6):doi: 10.1111/tid.12762. | ||
Patel G, Snydman DR. AST infectious diseases community of practice. Vancomycin-resistant Enterococcus infections in solid organ transplantation. Am J Transplant. 2013;13(Suppl 4):59–67. | ||
Zirakzadeh A, Patel R. Vancomycin-resistant enterococci: colonization, infection, detection, and treatment. Mayo Clin Proc. 2006;81(4):529–536. | ||
Kristich CJ, Rice LB, Arias CA [webpage on the Internet]. Enterococcal infection – treatment and antibiotic resistance. In: Gilmore MS, Clewell DB, Ike Y, et al., editors. Enterococci: From Commensals to Leading Causes of Drug Resistant Infection. Boston: Massachusetts Eye and Ear Infirmary; 2014. Available from: https://www.ncbi.nlm.nih.gov/books/NBK190420/. Accessed July 3, 2018. | ||
Marcel JP, Alfa M, Baquero F, et al. Healthcare-associated infections: think globally, act locally. Clin Microbiol Infect. 2008;14(10):895–907. | ||
Treitman AN, Yarnold PR, Warren J, Noskin GA. Emerging incidence of Enterococcus faecium among hospital isolates (1993 to 2002). J Clin Microbiol. 2005;43(1):462–463. | ||
Top J, Willems R, Blok H, et al. Ecological replacement of Enterococcus faecalis by multiresistant clonal complex 17 Enterococcus faecium. Clin Microbiol Infect. 2007;13(3):316–319. | ||
Leclercq R. Epidemiological and resistance issues in multidrug-resistant staphylococci and enterococci. Clin Microbiol Infect. 2009;15(3):224–231. | ||
Werner G, Coque T, Hammerum A, et al. Emergence and spread of vancomycin resistance among enterococci in Europe. Euro Surveill. 2008;13(47):ii=19046. | ||
Hecini-Hannachi A, Bentchouala C, Lezzar A, Laouar H, Benlabed K, Smati F. Multidrug-resistant bacteria isolated from patients hospitalized in Intensive Care Unit in University Hospital of Constantine, Algeria (2011–2015). Afr J Microbiol Res. 2016;10(33):1328–1336. | ||
Abbassi MS, Znazen A, Mahjoubi F, Hammami A, Ben Hassen A. Emergence of vancomycin-resistant Enterococcus faecium in Sfax: clinical features and molecular typing. Med Mal Infect. 2007;37(4):240–241. | ||
Djahmi N, Boutet-Dubois A, Nedjai S, Dekhil M, Sotto A, Lavigne J-P. Molecular epidemiology of Enterococcus sp. isolated in a university hospital in Algeria. Scand J Infect Dis. 2012;44(9):656–662. | ||
Bourafa N, Abat C, Loucif L, et al. Identification of vancomycin-susceptible major clones of clinical Enterococcus from Algeria. J Glob Antimicrob Resist. 2016;6:78–83. | ||
sante.dz/aarn [homepage on the Internet]. Rapports du Réseau Algérien de la Surveillance de la Résistance des Bactéries aux Antibiotiques (A.A.R.N.) [updated January 1, 2018]. Available from: http://www.sante.dz/aarn/rapports.htm. Accessed July 3, 2018. | ||
Elhani D, Klibi N, Dziri R, et al. vanA-containing E. faecium isolates of clonal complex CC17 in clinical and environmental samples in a Tunisian hospital. Diagn Microbiol Infect Dis. 2014;79(1):60–63. | ||
Dziri R, Lozano C, Ben Said L, et al. Multidrug-resistant enterococci in the hospital environment: detection of novel vancomycin-resistant E. faecium clone ST910. J Infect Dev Ctries. 2016;10(8):799–806. | ||
McDermott H, Skally M, O’Rourke J, Humphreys H, Fitzgerald-Hughes D. Vancomycin-Resistant Enterococci (VRE) in the intensive care unit in a nonoutbreak setting: identification of potential reservoirs and epidemiological associations between patient and environmental VRE. Infect Control Hosp Epidemiol. 2018;39(1):40–45. | ||
Hota B. Contamination, disinfection, and cross-colonization: are hospital surfaces reservoirs for nosocomial infection? Clin Infect Dis. 2004;39(8):1182–1189. | ||
Enterococcus faecium MLST Databases [homepage on the Internet]. Website sited at the University of Oxford (Jolley & Maiden 2010, BMC Bioinformatics, 11:595) [updated July 16, 2018]. Available from: https://pubmlst.org/efaecium. Accessed July 3, 2018. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.

