Back to Journals » Neuropsychiatric Disease and Treatment » Volume 16
Expression Profiles of Long Noncoding RNAs in Mice with High-Altitude Hypoxia-Induced Brain Injury Treated with Gymnadenia conopsea (L.) R. Br.
Authors Zhang Y, Liu L, Liang C, Zhou L, Tan L, Zong Y, Wu L, Liu T
Received 18 January 2020
Accepted for publication 15 April 2020
Published 12 May 2020 Volume 2020:16 Pages 1239—1248
DOI https://doi.org/10.2147/NDT.S246504
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Yuping Ning
Yongcang Zhang,1,2,* Lan Liu,2,3,* Cuiting Liang,2 Lingyu Zhou,2 Lixia Tan,2 Yonghua Zong,4 Lili Wu,5 Tonghua Liu4,5
1Department of Chemical Biology, School of Pharmaceutical Sciences, Peking University, Beijing 100191, People’s Republic of China; 2Medical College, Tibet University, Lhasa 850000, People’s Republic of China; 3West China Second University Hospital, Sichuan University, Chengdu, Sichuan 610041, People’s Republic of China; 4Tibet Traditional Medicine University, Lhasa 850000, People’s Republic of China; 5Key Laboratory of Health Cultivation of the Ministry of Education, Beijing University of Chinese Medicine, Beijing 100029, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Lili Wu
Key Laboratory of Health Cultivation of the Ministry of Education, Beijing University of Chinese Medicine, Beijing 100029, People’s Republic of China
Email [email protected]
Tonghua Liu
Tibet Traditional Medicine University, Lhasa 850000, People’s Republic of China
Key Laboratory of Health Cultivation of the Ministry of Education, Beijing University of Chinese Medicine, Beijing 100029, People’s Republic of China Email [email protected]
Background: The unique geographical environment at high altitudes may cause a series of diseases, such as acute altitude reaction, cerebral edema, and pulmonary edema. Gymnadenia conopsea (L.) R. Br. has been reported to have an effect on high-altitude hypoxia. However, the molecular mechanism, especially the expression of long noncoding RNAs (lncRNAs), is not yet clear.
Methods: The expression profiles of lncRNAs in high-altitude hypoxia-induced brain injury mice treated with Gymnadenia conopsea (L.) R. Br. by using a microarray method.
Results: A total of 226 differentially expressed lncRNAs, 126 significantly dysregulated mRNAs and 23 differentially expressed circRNAs were detected (> 2.0-fold, p< 0.05). The expression of selected lncRNAs, mRNAs and circRNAs was validated by qRT-PCR. KEGG analysis showed that the mRNAs coexpressed with lncRNAs were involved in inflammation and hypoxia pathways, including the HIF-1, PI3K-Akt, and NF-kappa B signaling pathways. The lncRNA-TF network analysis results indicated that the lncRNAs were regulated mostly by HMGA2, SRY, GATA4, SOX5, and ZBTB16.
Conclusion: This study is the first to report the expression profiles of lncRNAs, mRNAs and circRNAs in mice with high-altitude hypoxia-induced brain injury treated with Gymnadenia conopsea (L.) R. Br. and may improve the understanding of the molecular mechanism of Gymnadenia conopsea (L.) R. Br. in treating high altitude hypoxia-induced brain injury.
Keywords: expression profile, noncoding RNAs, high-altitude hypoxia, Gymnadenia conopsea (L.) R. Br
Introduction
Acute high altitude sickness refers to a series of clinical syndromes caused by high altitude hypoxia occurring within hours or days upon exposure to high altitudes and includes acute mild altitude sickness, acute brain edema, acute high-altitude pulmonary edema, and high-altitude encephalopathy; among these conditions, high-altitude brain edema causes the most serious consequences.1–3 In recent years, researchers have focused on the pathogenesis of acute high-altitude hypoxia mainly from the perspectives of morphology, physiology, biochemistry and related molecular biology.4–6 High-altitude hypoxic brain injury is a complex pathophysiological process in which the cytokine network system participates in regulating and maintaining homeostasis in many aspects and stages, and the complex network system of cytokines is regulated by dynamic gene cascades.7,8 An increasing number of studies have shown that brain injury can induce changes in the expression of multiple genes in the central nervous system; in addition, there are inherent regulatory relations among some specific genes, and the differential expression of these genes directly or indirectly affects the expression levels of cytokines.9,10
Tibetan drugs, including Rhodiola rosea, Saussurea involucrate, Hippophae rhamnoides, Dracocephalum tanguticum, Potentilla anserina, and Gymnadenia conopsea (L.) R. Br. have been reported to have anti-hypoxia effects.11–16 Advanced research has shown that Gymnadenia conopsea (L.) R. Br. has anti-allergy and anti-silicosis fibrosis effects, promotes progenitor cell proliferation, scavenges free radicals, and inhibits hepatitis B virus surface antigen.17 No evidence indicates that gene expression profiles, especially ncRNA expression profiles, in mice with high-altitude hypoxia-induced brain injury treated with Tibetan drugs has been investigated. Long noncoding RNAs (lncRNAs) are newly identified ncRNA transcripts of more than 200 nucleotides. A recent study showed that lncRNAs play a wide range of regulatory roles in gene expression.18 lncRNAs were originally thought to be a byproduct of transcription. However, recent data suggest that lncRNAs can participate in gene expression by regulating transcription, posttranscriptional processing, chromatin remodeling, and the production of small ncRNAs.19–22 However, no research has been performed on the expression profiles of long noncoding RNAs in mice with high-altitude hypoxia-induced brain injury treated with Gymnadenia conopsea (L.) R. Br.
In the present study, we aimed to investigate differential expression profiles of lncRNAs, mRNAs and circRNAs in mice with high-altitude hypoxia-induced brain injury treated with Gymnadenia conopsea (L.) R. Br. using a DNA microarray.
Materials and Methods
Animals and Study Design
A total of 60 SPF, 6- to 8-week-old, male C57BL/6J mice weighing 22.5±5 g were purchased from Hunan Tianqin Biotechnology Co. Ltd. (Changsha, Hunan, China). Mice were housed at a constant temperature (21–23°C) and constant humidity (45–65%) on a 12-hour alternating light-dark cycle with free access to food and water. All animal experiments followed the guidelines of the National Institutes of Health Guide for the Care and Use of Laboratory Animals. All animal work was performed in the laboratory animal center of Tibet University. The research procedures were approved by the Ethics Committee of Tibet University (EC20190512), China.
Mice in the intervention group were given 750 mg/kg Gymnadenia conopsea (L.) R. Br. ethanol extract via intragastric administration once a day for 30 consecutive days. Mice in the control group were given distilled water in a similar manner. Then, mice in the intervention and control groups were housed in a decompression chamber and exposed to a simulated high altitude of 4000 m for 24 h.23
Sample Collection
Mice were anesthetized with pentobarbital sodium (2%, 0.1 mL/10 g) by intraperitoneal injection. Brain tissues were stored in 10% neutral formalin, and other tissues were stored at −80°C until further analysis.
RNA Extraction
Total RNA was isolated from each brain tissue sample by using an E.Z.N.A.® Total RNA Kit II (Omega, USA). Total RNA was quantified with a NanoDrop ND-2000 (Thermo Scientific, USA), and RNA integrity was assessed using an Agilent Bioanalyzer 2100 (Agilent Technologies, USA).
Microarray Analysis
Sample labeling, microarray hybridization and washing were performed based on the manufacturer’s standard protocols. Briefly, total RNA was transcribed to double-stranded cDNA, transcribed into cRNA and labeled with Cyanine-3-CTP. The labeled cRNAs were hybridized onto the microarray. After washing, the arrays were scanned with an Agilent Scanner G2505C (Agilent Technologies, USA). All experiments were carried out according to the manufacturers’ standard protocols. The experiments were performed by OE Biotechnology Co., Ltd. (Shanghai, China). An Agilent Mouse LncRNA Microarray V3 (4*180K, Design ID: 084388) was used in the present experiment.
Differential Expression Analysis
Raw data in the array images were analyzed with Feature Extraction software (version 10.7.1.1, Agilent Technologies). The basic analysis was performed using GeneSpring (version 14.8, Agilent Technologies, USA) with the raw data. First, the quintile algorithm was used to normalize the raw data. Fold change values (≥ 2.0), as well as t-test-based P values (≤ 0.05), were calculated to identify the differentially expressed genes. Then, gene ontology (GO) analysis (http://www.geneontology.org) and kyoto encyclopedia of genes and genomes (KEGG) analysis (http://www.genome.jp/kegg/) were applied to determine the roles of these differentially expressed mRNAs, as well as circRNA host mRNAs. Finally, hierarchical clustering of differentially expressed lncRNAs, circRNAs and mRNAs between brain samples from the intervention and control groups was performed to visualize the distinguishable gene expression patterns.
Co-Expression Network Analysis
A coexpression network was constructed using the top 20 differentially expressed lncRNAs and mRNAs. Coexpression of lncRNAs and mRNAs was identified by the Pearson correlation coefficients (PCC) (≥0.7). Values of p < 0.05 indicated a statistically significant correlation. Additionally, via coexpression analysis of the differentially expressed lncRNAs and mRNAs, the transcription data of the top 40 lncRNAs were extracted, and a network map (lncRNAs- Transcription factors (TFs)) was drawn according to the top 5 lncRNAs enriched on the TFs. The top 20 lncRNAs were sorted according to the lncRNAs enriched on the TFs, and the top 2 lncRNAs were used to draw a network map (lncRNA-target mRNA-TF). Cytoscape software (The Cytoscape Consortium, San Diego, CA, USA) was used to construct the networks.
Quantitative Real-Time-PCR (qRT-PCR)
qRT-PCR was performed using a HiScript II One Step qRT-PCR SYBR Green Kit (Vazyme Biotech Co., Ltd., Nanjing, China) in a QuantStudio 3 Flex Real-Time PCR System (Thermo Scientific, USA). The 2−ΔΔCt method was used to evaluate the relative expression levels of lncRNAs, circRNAs and mRNAs. The significance of the differences was assessed by Student’s t-test. Repeated measures ANOVA was performed. Values of p < 0.05 were considered statistically significant. SPSS (version 19.0, SPSS, Inc., Chicago, IL, USA) was used to perform the statistical tests.
Results
Differentially Expressed LncRNAs, CircRNAs and mRNAs in Mice with High-Altitude Hypoxia-Induced Brain Injury Mice Treated with Gymnadenia conopsea (L.) R. BR
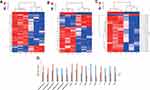
A total of 226 differentially expressed lncRNAs were identified in mice with high-altitude hypoxia-induced brain injury mice treated with Gymnadenia conopsea (L.) R. Br. compared with normal controls. Among these differentially expressed lncRNAs, 186 were upregulated and 40 were downregulated. NONMMUT033643.2, on chr19, was the most downregulated lncRNA, with a FC of 4.46. NONMMUT117581.1, on chr4, was the most upregulated lncRNA, with a FC of 8.01. In addition, 126 significantly dysregulated mRNAs (85 upregulated and 41 downregulated) were detected. Adamts4 (FC= 7.42) and Lcn2 (FC= 15.67) were the most downregulated and upregulated mRNAs, respectively. Furthermore, 23 differentially expressed circRNAs were identified. The most upregulated circRNA was Pum2 (FC= 4.71). The most downregulated circRNA was Cit (FC= 3.68) (Figure 1A–C). Six lncRNAs and mRNAs and four circRNAs were randomly selected and verified by using qRT-PCR. The results were consistent with those of the microarray chip data analysis (Figure 1D).
Identification of Coexpression Networks and Potential Functions
The correlations between the top 20 dysregulated lncRNAs and mRNAs were predicted. The p values of each lncRNA-mRNA correlation were ranked. The coexpression network was constructed with the selected lncRNA-mRNA pairs with the highest Pearson correlation coefficient. A total of 265 connections (168 positive connections and 97 negative connections) were included in the network (Figure 2).
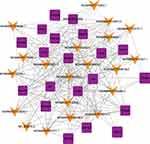 |
Figure 2 lncRNA-mRNA-network analysis. Purple squares represent dysregulated mRNAs, green arrows represent dysregulated lncRNAs. |
KEGG and GO Analyses
As presented in Figure 3A, GO enrichment analysis indicated that the differentially expressed lncRNAs were mostly enriched in the hemoglobin complex cellular component, the peroxidase activity molecular function, and the branching involved in blood vessel morphogenesis biological process. The differentially expressed circRNAs were mostly enriched in the protein complex cellular component, the ATP binding molecular function, and the dendrite development biological process. Furthermore, KEGG analysis indicated that the lncRNAs were involved in the following pathways: IL-17 signaling pathway, HIF-1 signaling pathway, PI3K-Akt signaling pathway, NF-kappa B signaling pathway, TNF signaling pathway, Apoptosis, and VEGF signaling pathway. The circRNAs were involved in the following pathways: TNF signaling pathway, Apoptosis, Neurotrophin signaling, and MAPK signaling pathway (Figure 3B).
LncRNA-TF Network Analysis
Transcription data for the top 40 lncRNAs were extracted. Then, the top 5 enriched lncRNAs on TFs were used to construct the lncRNA-TF network. The results showed that a total of 87 TFs with 1354 lncRNA-TF pairs were predicted. The top 5 TFs enriched with lncRNAs were HMGA2, SRY, GATA4, SOX5, and ZBTB16 (Figure 4).
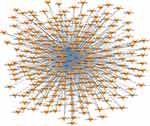 |
Figure 4 Network of the top 40 most related LncRNA-Transcription factors (TFs) pairs (the most 5 related lncRNA-TFs pairs according to the P value). Orange arrow: TFs; Blue diamonds: lncRNAs. |
LncRNA-Target-TF Network Analysis
Transcription data for the top 20 lncRNAs were extracted. Then, analysis was conducted on the top 2 related lncRNA-mRNA and lncRNA-TF pairs ranked according to the p values. A total of 80 TFs and 68 mRNAs were predicted to regulate or be the target of these lncRNAs, respectively. Among these TFs, HMGA2 and SRY were shown to regulate most of the lncRNAs. For example, HMGA2 and SRY were predicted to regulate NONMMUT033643.2 and NONMMUT115383.1 (Figure 5).
 |
Figure 5 lncRNA-target-Transcription factors (TFs) network of 20 most differentially expressed lncRNAs. Red arrow: lncRNAs; Green round: target mRNAs; Blue diamond: TFs. |
Discussion
In this study, we explored the expression profiles of lncRNAs, mRNAs and circRNAs in mice with high-altitude hypoxia-induced brain injury treated with Gymnadenia conopsea (L.) R. Br. In total, our study identified 226 lncRNAs, 126 mRNAs and 23 circRNAs. The differentially expressed lncRNAs were involved in several hypoxia- and inflammation-related pathways, including the HIF-1 signaling pathway, PI3K-Akt signaling pathway, and NF-kappa B signaling pathway.
Initially, 186 lncRNAs upregulated and 40 lncRNAs downregulated relative to their control expression levels were detected. To the best of our knowledge, this is the first identification of differentially expressed lncRNAs in mice with high-altitude hypoxia-induced brain injury treated with Gymnadenia conopsea (L.) R. Br. Accumulating evidence has indicated that lncRNAs may participate in multiple pathological processes, such as the occurrence of cancer,24 neurodegenerative lesions,25 and cardiovascular diseases.26 Recently, two lncRNAs, nuclear-enriched abundant/autosomal transcript 1 (NEAT1) and metastasis-associated lung adenocarcinoma transcript 1 (MALAT1), were found to be strongly induced by hypoxia in cultured cells.27 According to a study by Jiménez et al,28 hypoxia could contribute to the metastatic spread of breast cancer via HIF-mediated induction of EFNA3 lncRNAs and subsequent Ephrin-A3 protein accumulation in human breast cancer cells. The above conclusions may support the hypothesis that lncRNAs play an important role in hypoxia. However, there has been no research on the relationship between the expression of lncRNAs and high-altitude hypoxia. Although the results were identified randomly by RT-PCR, we should perform more studies to obtain more concise results in the future.
In addition, 85 upregulated and 41 downregulated mRNAs were detected. We identified Adamts4 and Lcn2 as the most differentially expressed mRNAs in the treated group compared with the control group. ADAMTS-4 is one of the members of the polyprotein polysaccharide family and has been reported to cleave aggrecan, which may contribute to cartilage breakdown during arthritis.29,30 In addition, a previous study clarified that hypoxia may inhibit the expression of ADAMTS-4 and ADAMTS-5 through HIF-1α, maintain the homeostasis of the intervertebral disc environment, and delay the occurrence of disc degeneration.31 ADAMTS-4 is also present in the central nervous system (CNS), including the cortex, hippocampus, striatum, and spinal cord.32 Cross et al revealed that the expression of ADAMTS-4 was significantly increased in human astrocyte cultures induced by TNF-a.33 Ashlin et al34 provided evidence that the expression of ADAMTS-4 in macrophages increased after treatment with TNF-α, IFN-g, and TGF-β, which may represent an external source of ADAMTS-4 in CNS injury. Additionally, ADAMTS-4 was shown to reduce neuroinflammation in vitro in microglia or astrocyte cultures and in vivo in mice after ischemic stroke.35 Collectively, these studies suggest that the increased expression of endogenous ADAMTS-4 during neuronal injury may be an adaptive change to protect the brain against neuroinflammation. Interestingly, our findings indicated that ADAMTS-4 mRNA expression was upregulated in brain tissue from mice with high-altitude hypoxia-induced brain injury treated with Gymnadenia conopsea (L.) R. Br. compared with brain tissue from control mice, suggesting that ADAMTS-4 may play an important role in the neuroprotective process during high altitude hypoxia.
Lipid transport protein-2 (LCN-2) is also called neutrophil gelatinase-related lipid transport protein (neutrophelase neutrophelase).36 It is a secretory protein that is related to inflammation, the apoptosis of immune response cells, and the development of various tumors.37 In the central nervous system, the LCN-2 protein is secreted by small glial cells and astrocytes and is related to the inflammatory response of the central nervous system.38 As Niu et al reported, the release of LCN-2 has an obvious correlation with cell injury caused by oxygen-sugar deprivation/poly-oxy sugar.39 In addition, as an anti-inflammatory mediator, adenosine can significantly ameliorate the cell damage caused by LCN-2.39 Moreover, LCN2 can promote the inflammatory activation of astrocytes via the RAS homologous gene-Rho related spiral curling protein kinase (Rho-ROCK)-GFAP pathway, and inflammatory activated astrocytes can selectively promote neuronal apoptosis.40 Therefore, LCN2 plays an important role in the inflammatory activation and neuronal apoptosis of glial cells under various pathological conditions in the CNS. We found that the expression level of LCN2 was significantly decreased in the group treated with Gymnadenia conopsea (L.) R. Br. compared to the control group, which may indicate that Gymnadenia conopsea (L.) R. Br. may provide neuroprotection by inhibiting LCN2 expression in brain tissue. However, the mechanism requires further research.
According to KEGG analysis, the dysregulated lncRNAs were involved in several pathways, such as the IL-17 signaling pathway, HIF-1 signaling pathway, PI3K-Akt signaling pathway, NF-kappa B signaling pathway, TNF signaling pathway, apoptosis, and VEGF signaling pathway. HIF-1 is an important transcription factor involved in the hypoxia regulation process. Zhang et al demonstrated that the relative expression of HIF-1α mRNA increased significantly in the hypoxia group compared to the normoxia group.41 Furthermore, Cao et al reported that Twenty flavor Shen Xiang Pill can protect against hypoxia by interfering with the protein levels of HIF-1α and EPO.42 In the oxygenated state, HIF-1α expression is maintained at a low level mainly through the activity of proteasome degradation pathways. However, the HIF-α degradation pathway is inhibited in hypoxia.43 In hypoxia, HIF-α also interacts with HIF-1β to form a heterodimer and is transferred to the nucleus; is the heterodimer then interacts with hypoxia response elements (HREs) and activates the transcription of downstream target genes such as heme oxygenase-1 (HO-1), erythropoietin (EPO), vascular endothelial growth factor (VEGF), and a variety of glycolysis enzymes.44 Recent studies have found that the expression of HIF-1α under normoxic or hypoxic conditions can be mediated by phosphatidylinositol 3-kinase (PI3 kinase, PI3K) and regulated by the Akt signaling pathway.45,46 Su et al reported that rosamultin can ameliorate functional damage to vascular endothelial cells induced by acute hypoxia by activating the PI3K/Akt signaling pathway and then increasing HIF-α expression.47 Thus, Gymnadenia conopsea (L.) R. Br. may protect against high-altitude hypoxia via these signaling pathways.
Conclusion
We identified a number of dysregulated lncRNAs, mRNAs and circRNAs that might be potential biomarkers or targets in mice with high-altitude hypoxia-induced brain injury treated with Gymnadenia conopsea (L.) R. Br. Further investigation is needed to elucidate the detailed mechanisms underlying the regulation of these differentially expressed lncRNAs.
Disclosure
The authors declare no conflicts of interest.
References
1. Schneider M, Bernasch D, Weymann J, et al. Acute mountain sickness: influence of susceptibility, preexposure, and ascent rate. Med Sci Sports Exerc. 2002;34(12):1886–1891. doi:10.1097/00005768-200212000-00005
2. Dehnert C, Bärtsch P. Acute mountain sickness and high-altitude cerebral edema. N Engl J Med. 2017;74(10):535–541.
3. Garlick V, O’Connor A, Shubkin CD. High-altitude illness in the pediatric population: a review of the literature on prevention and treatment. Curr Opin Pediatr. 2017;29(4):503–509. doi:10.1097/MOP.0000000000000519
4. Richalet JP, Hornych A, Rathat C, et al. Plasma prostaglandins, leukotrienes and thromboxane in acute high altitude hypoxia. Respir Physiol. 1991;85(2):205–215. doi:10.1016/0034-5687(91)90062-N
5. Luks AM, Levett D, Martin DS. Changes in acute pulmonary vascular responsiveness to hypoxia during a progressive ascent to high altitude (5300m). Exp Physiol. 2017;102(6):711–724. doi:10.1113/EP086083
6. Baze MM. Effects of hypoxia and high altitude on gene expression, energetics, and immune function. Dissertations & Theses – Gradworks. 2011; 3457511.
7. Severinghaus JW. Hypothesis: angiogenesis cytokines in high altitude cerebral oedema. Acta Anaesthesiol Scand. 2008;39(s107):177–178. doi:10.1111/j.1399-6576.1995.tb04354.x
8. Jenny L, Feiner John R, Andrew S. Increased cytokines at high altitude: lack of effect of ibuprofen on acute mountain sickness, physiological variables, or cytokine levels. High Alt Med Biol. 2018;19(3):249–258. doi:10.1089/ham.2017.0144
9. Li YH, Tana WR, Bai ZH. Transcriptomic profiling reveals gene expression kinetics in patients with hypoxia and high altitude pulmonary edema. Gene. 2018;651:200–205. doi:10.1016/j.gene.2018.01.052
10. MacInnis MJ, Koehle MS. Evidence for and against genetic predispositions to acute and chronic altitude illnesses. High Alt Med Biol. 2016;17(4):281–293. doi:10.1089/ham.2016.0024
11. Qiang W, Xiao R, Yan QC. Study on the effect of plant hormones and pre-chilled treatment to break dormancy and germination of Rhodiola rosea seeds. J Zhejiang Univ. 2005;31:113–118.
12. Ma HP, Fan PC, Jing LL, et al. Anti-hypoxic activity at simulated high altitude was isolated in petroleum ether extract of Saussurea involucrata. J Ethnopharmacol. 2011;137(3):1510–1515. doi:10.1016/j.jep.2011.08.037
13. Tulsawani R, Gupta R, Misra K. Efficacy of aqueous extract of Hippophae rhamnoides and its bio-active flavonoids against hypoxia-induced cell death. Indian J Pharmacol. 2013;45(3):258–263. doi:10.4103/0253-7613.111943
14. Maimaitiyiming D, Hu G, Aikemu A, et al. The treatment of uygur medicine dracocephalum moldavica L on chronic mountain sickness rat model. Pharmacogn Mag. 2014;10(40):477–482. doi:10.4103/0973-1296.141817
15. Li LZ, Wang LJ, Wang Y, et al. Effect of n-butanol extract from potentilla anserina on hypoxia-induced calcium overload and SERCA2 expression of rat cardiomyocytes. Chin Herb Med. 2012;4(2):142–149.
16. Zhang JY, Men LC, Wu XJ, et al. Research progress of tibetan medicine in protection of plateau hypoxia. Chin J Ethnomed Ethnopharm. 2017;26(22):42–45.
17. Shang XF, Guo X, Liu Y, et al. Gymnadenia conopsea (L.) R. Br.: a systemic review of the ethnobotany, phytochemistry, and pharmacology of an important Asian folk medicine. Front Pharmacol. 2017;8:24. doi:10.3389/fphar.2017.00024
18. Gudenas BL, Wang J, Kuang SZ. Genomic data mining for functional annotation of human long noncoding RNAs. J Zhejiang Univ Sci B. 2019;20(6):476–487. doi:10.1631/jzus.B1900162
19. Kanduri C. Long noncoding RNAs: lessons from genomic imprinting. Biochim Biophys Acta. 2016;1859(1):102–111. doi:10.1016/j.bbagrm.2015.05.006
20. Chen YG, Satpathy AT, Chang HY. Gene regulation in the immune system by long noncoding RNAs. Nat Immunol. 2017;18(9):962–972. doi:10.1038/ni.3771
21. Xue M, Zhuo Y, Shan B. MicroRNAs, long noncoding RNAs, and their functions in human disease. Methods Mol Biol. 2017;1–25.
22. Hui KM, Xia HP. Mechanism of cancer drug resistance and the involvement of noncoding RNAs. Curr Med Chem. 2014;21(26):3029–3041. doi:10.2174/0929867321666140414101939
23. Ma HP, Wu JH, Gao RM, et al. Establishment of an animal model for acute mountain sickness with a decompression chamber. Pharm J Chin PLA. 2013;29(4):301–304.
24. Camacho CV, Choudhari R, Gadad SS. Long noncoding RNAs and cancer, an overview. Steroids. 2018;133:1–12. doi:10.1016/j.steroids.2017.12.012
25. Wu P, Zuo XL, Deng HL. Roles of long noncoding RNAs in brain development, functional diversification and neurodegenerative diseases. Brain Res Bull. 2013;97(Complete):69–80. doi:10.1016/j.brainresbull.2013.06.001
26. Haemmig S, Simion V, Yang D, et al. Long noncoding RNAs in cardiovascular disease, diagnosis, and therapy. Curr Opin Cardiol. 2017;32(6):776–783. doi:10.1097/HCO.0000000000000454
27. Lelli A, Nolan KA, Santambrogio S, et al. Induction of long noncoding RNA MALAT1 in hypoxic mice. Hypoxia. 2015;8(3):45–52.
28. Jiménez B, Tiana M, Del PL. lnc RNAs, hypoxia and metastasis. Oncoscience. 2015;2(10):795–796. doi:10.18632/oncoscience.247
29. Fontanil T, ÁlvarezTeijeiro S, Ángeles VM. Cleavage of Fibulin-2 by the aggrecanases ADAMTS-4 and ADAMTS-5 contributes to the tumorigenic potential of breast cancer cells. Oncotarget. 2017;8(8):13716–13729. doi:10.18632/oncotarget.14627
30. Song RH, Tortorella MD, Malfait AM, et al. Aggrecan degradation in human articular cartilage explants is mediated by both ADAMTS-4 and ADAMTS-5. Arthritis Rheum. 2014;56(2):575–585. doi:10.1002/art.22334
31. Hua DF, Xu MM, Yang C, et al. Regulation of HIF-1αdependent hypoxia on expression of ADAMTS-4 and ADAMTS-5 in nucleus pulposus cells. J Spina Surg. 2017;15(4):233–238.
32. Lemarchant S, Wojciechowski S, Vivien D, et al. ADAMTS-4 in central nervous system pathologies. J Neurosci Res. 2017;95(9):1703–1711. doi:10.1002/jnr.24021
33. Cross AK, Haddock G, Stock CJ, et al. ADAMTS-1 and −4 are up-regulated following transient middle cerebral artery occlusion in the rat and their expression is modulated by TNF in cultured astrocytes. Brain Res. 2006;1088:19–30. doi:10.1016/j.brainres.2006.02.136
34. Ashlin TG, Kwan AP, Ramji DP. Regulation of ADAMTS-1, −4 and −5 expression in human macrophages: differential regulation by key cytokines implicated in atherosclerosis and novel synergism between TL1A and IL-17. Cytokine. 2013;64:234–242. doi:10.1016/j.cyto.2013.06.315
35. Lemarchant S, Dunghana H, Pomeshchik Y, et al. Anti-inflammatory effects of ADAMTS-4 in a mouse model of ischemic stroke. Glia. 2016;64(9):1492–1507. doi:10.1002/glia.23017
36. Jin M, Kim JH, Jang E, et al. Lipocalin-2 deficiency attenuates neuroinflammation and brain injury after transient middle cerebral artery occlusion in mice. J Cereb Blood Flow Metab. 2014;34(8):1306–1314. doi:10.1038/jcbfm.2014.83
37. Wang G, Weng YC, Han X, et al. Lipocalin-2 released in response to cerebral ischaemia mediates reperfusion injury in mice. J Cell Mol Med. 2015;19(7):1637–1645. doi:10.1111/jcmm.12538
38. Lee S, Kim JH, Kim JH. Lipocalin-2 is a Chemokine Inducer in the Central Nervous System: role of chemokine ligand 10(CXCL10) in lipocalin-2-induced cell migration. J Biol Chem. 2011;286(51):43855–43870. doi:10.1074/jbc.M111.299248
39. Niu CY, Hu YN, Wang J, et al. Effects of adenosine pretreatment on astrocytes and the expression of lipocalin-2 under oxygen-glucose deprivation and reoxygenation. Chin J Anat. 2016;39(2):192–195.
40. Bi F, Huang C, Tong J, et al. Reactive astrocytes secrete lcn2 to promote neuron death. Proc Natl Acad Sci USA. 2013;110(10):4069–4074. doi:10.1073/pnas.1218497110
41. Zhang WB, Su C, Zhang YL, et al. Temporal changes of HIF-1α and rat brain hippocampus injury induced by acute hypobaric hypoxia. Shandong Med. 2016;56(4):8–11.
42. Cao CZ. Effect of 20-flavor Chenxiang pill on hypoxia-inducible factor-1 in rats exposed to hypoxia. J Qinghai Med Coll. 2012;2:177–179.
43. Huang LE, Gu J, Schau M, et al. Regulation of hypoxia-inducible factor 1alpha is mediated by an O2-dependent degradation domain via the ubiquitin-proteasome pathway. Proc Natl Acad Sci U S A. 1998;95(14):7987–7992. doi:10.1073/pnas.95.14.7987
44. Pagé EL, Robitaille GA, Jacques P, et al. Induction of hypoxia inducible factor-1α by transcriptional and translational mechanisms. J Biol Chem. 2002;277(50):48403–48409. doi:10.1074/jbc.M209114200
45. Pore N, Jiang Z, Shu HK, et al. Akt1 activation can augment hypoxia-inducible factor-1α expression by increasing protein translation through a mammalian target of rapamycin-independent pathway. Mol Cancer Res. 2006;4(7):471–479. doi:10.1158/1541-7786.MCR-05-0234
46. Yao D, He X, Wang JH. Effects of PI3K/Akt signaling pathway on learning and memory abilities in neonatal rats with hypoxic-ischemic brain damage. Zhongguo Dang Dai Er Ke Za Zhi. 2011;13(5):424–427.
47. Zhang L, Liu Y, Li JY. Protective effect of rosamultin against H 2 O 2 -induced oxidative stress and apoptosis in H9c2 Cardiomyocytes. Oxid Med Cell Longev. 2018;2018(6):1–13. doi:10.1155/2018/4296985
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.


