Back to Journals » OncoTargets and Therapy » Volume 8
Expression of CPEB4 in invasive ductal breast carcinoma and its prognostic significance
Authors Sun H, Wen X, Han T, Liu Z, Li S, Wang J, Liu X
Received 29 April 2015
Accepted for publication 4 July 2015
Published 26 November 2015 Volume 2015:8 Pages 3499—3506
DOI https://doi.org/10.2147/OTT.S87587
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Faris Farassati
Hao-Ting Sun,1,2,* Xin Wen,3,* Tian Han,4,* Zhen-Hua Liu,5 Shao-Bo Li,1 Ji-Gang Wang,1 Xiu-Ping Liu6
1Department of Pathology, School of Basic Medical Sciences, Fudan University, Shanghai, 2Department of General Surgery, Huashan Hospital, Fudan University, Shanghai, 3Sun Yat-sen University Cancer Center, State Key Laboratory of Oncology in South China, Collaborative Innovation Center for Cancer Medicine, Canton, Guangdong Province, 4Key Lab of Myopia, Ministry of Health, Department of Ophthalmology, Eye & ENT Hospital of Fudan University, Shanghai, 5Urology Department and Institute of Urology, Peking University First Hospital, Peking University, Beijing, 6Department of Pathology, The Fifth People’s Hospital of Shanghai, Fudan University, Shanghai, People’s Republic of China
*These authors contributed equally to this work
Aims: Cytoplasmic polyadenylation element binding proteins (CPEBs) are RNA-binding proteins that regulate translation by inducing cytoplasmic polyadenylation. CPEB4 has been reported in association with tumor growth, vascularization, and invasion in several cancers. To date, the expression of CPEB4 with clinical prognosis of breast cancer was never reported before. We aim to investigate the expression of CPEB4 and its prognostic significance in invasive ductal breast carcinoma.
Methods: Immunohistochemical staining of CPEB4 and estrogen receptor, progesterone receptor, and human epidermal growth factor receptor was performed in 107 invasive ductal carcinoma (IDC) samples, and prognostic significance was evaluated.
Results: High expression of CPEB4 was observed in 48.6% of IDC samples. Elevated CPEB4 expression was possibly related to increased histological grading (P=0.037) and N stage (P<0.001). Patients with high expression of CPEB4 showed shorter overall survival (P=0.001). High CPEB4 expression was an independent prognostic factor for overall survival (P=0.022, hazard ratio =4.344, 95% confidence interval =1.235–15.283).
Conclusion: High CPEB4 expression is associated with increased histological grading and N stage, and it can serve as an independent prognostic factor in IDC.
Keywords: cytoplasmic polyadenylation element binding protein 4, invasive ductal carcinoma, immunohistochemistry, prognosis
Introduction
Invasive breast cancer is the most frequent cancer and the leading cause of cancer death in young women, in which the invasive ductal carcinoma (IDC) is the most common pathologic type.1,2 Several biomarkers have been routinely tested clinically to evaluate the prognosis and establish the treatment strategy. For example, estrogen receptor (ER) and progesterone receptor (PR) have served as predictors to patient’s suitability for endocrine therapy.3,4 The human epidermal growth factor receptor-2 (HER2/neu) has also been used as a valuable prognostic and treatment biomarker.5,6 Trastuzumab and lapatinib have been included in clinical practice for HER2-postive breast cancer patients.7–9 However, drug resistance is quite common,10,11 and the clinical outcome remains hard to predict for individual patients. Therefore, there is a continual drive to find new biomarkers as reliable prognostic indicators and treatment targets.12
Cytoplasmic polyadenylation element binding protein (CPEB) is a combination of a sequence-specific RNA-binding protein with a RNA-recognition motif and a zinc-finger.13,14 CPEBs specifically target a sequence with a cis-acting sequence in their 3′-untranslated region (UTR) and contribute to polyadenylation, resulting in translation termination.15–17 CPEB4, one of the most important subtypes that affects cell proliferation and differentiation, can bind to a distinct loop-forming U-rich motif.18,19 Till now, the specific CPEB4 binding sequence overlapped with the cytoplasmic polyadenylation element is unclear.18,20,21 According to previous studies, elevated CPEB4 expression seems to contribute to tumor growth, vascularization, migration, invasion, and metastasis.22–24 However, the expression characteristics of CPEB4 in breast cancer have not been reported yet.
In this study, we performed an immunohistochemical study on 107 cases of IDC. The aim of this study was to investigate the clinicopathologic significance of CPEB4 expression in IDC and evaluate its potential value when served as a prognostic indicator.
Materials and methods
Patient population and clinical data
One hundred and seven patients with primary IDC underwent curative surgery at the Huashan Hospital of Fudan University between January 1999 and December 2002. None of the patients in this study received preoperative neoadjuvant chemotherapy and/or radiotherapy, and all patients received four cycles of cyclophosphamide, methotrexate, and 5-fluorouracil after surgery. All patients were women aging from 34 years to 87 years, with a mean age of 53 years. In addition, as controls, normal breast tissues were taken from randomly selected tissues of breast IDC patients who received operation over the same period. The study was approved by the Ethical Committee for Clinical Research of Fudan University, and informed consent was obtained from all subjects.
All pathologic slides were reevaluated by two independent pathologists. The pathologic diagnosis was made according to the WHO classification of breast tumors, and histological grading was assessed according to the Nottingham modification of the Bloom and Richardson grading criteria.25 The American Joint Committee on Cancer (AJCC)/International Union for Cancer Control (UICC) tumor, node, metastasis (TNM) classification and stage grouping system was used to evaluate the clinical stage.26 Patients’ characteristics are listed in Table 1.
  | Table 1 Clinicopathological characteristics and follow-up data of 107 IDC patients |
The follow-up started postoperatively and ended on December 31, 2008. The follow-up time ranged from 3.5 months to 119.6 months, with a median time of 81.6 months. At the end of the follow-up period, 89 patients were still alive and 18 patients had died of the disease.
Antibodies
Polyclonal anti-CPEB4 antibody was purchased from Abcam (Cat ab83009, Cambridge, UK); mouse monoclonal antibodies anti-HER2/neu/c-erbb-2 (Cat M-0196), ER (Cat M-00241), and PR (Cat M-0448) were all purchased from Shanghai Long Island Biotech Co., Ltd. (Shanghai, People’s Republic of China).
Immunohistochemistry
Tissue samples were fixed in 10% formalin, embedded with paraffin, and cut into sections of 4–5 μm. After that, all slides were dehydrated with xylene and graded alcohol/water mixtures. Antigen retrieval was performed with 0.01 M citrate buffer (pH =6.0) at 95°C for 20 minutes. Then slides were incubated with diluted primary antibodies (anti-CPEB4, 1:200 dilution; anti-ER, anti-PR, and anti-HER2, 1:100 dilution) at 4°C for 12 hours, followed by incubations with biotinylated secondary antibody for 1 hour and peroxidase-labeled streptavidin (Shanghai Long Island Biotech Co., Ltd.) for 15 minutes. The color was developed by reacting with 3,3-diaminobenzidine for 1 minute. Slides were again counterstained with Mayer’s hematoxylin. The primary antibody was omitted as a negative control, being replaced by phosphate-buffered solution. The reproducibility of CPEB4 staining was examined by two independent pathologists.
Semi-quantitative analysis
The immunohistochemical results were evaluated by two pathologists. Ten visual fields at a high power (×400) were observed in each slide by a light microscope (Carl Zeiss Meditec AG, Jena, Germany). For CPEB4 expression, the staining intensity was observed (score 0, negative staining; score 1, pale yellow; score 2, dark yellow; score 3, brown), and the percentage of positive cells was calculated (score 0, <25% positive cells/field; score 1, 25%–50% positive cells/field; score 2, 50%–75% positive cells/field; and score 3, >75% positive cells/field). Based on the product of the two scores, the staining grades were classified into low (<4) and high (≥4).
Scoring of ER, PR, and HER2 expressions was also reevaluated by two independent pathologists. Based on the percentage of positive cells and the intensity of the staining, if there were no reactivity or nuclear (ER or PR)/membranous (HER2) reactivity in <1% of tumor cells, the samples will be regarded as negative stains, otherwise positive stains.
Statistical analysis
Statistical analyses were performed using the Statistical Package for the Social Sciences, Version 20 (IBM Corporation, Armonk, NY, USA) and Prism® 5.0 software (GraphPad Software, Inc., La Jolla, CA, USA). Pearson’s correlation coefficients were used to determine the relationship between the CPEB4 expression and the clinicopathologic parameters, including ER, PR, and HER2 expressions. Kaplan–Meier survival analysis was used to estimate the prognostic value of CPEB4, and the log-rank test was used to assess the survival differences between different groups. Univariate and multivariate Cox regression analyses were performed to evaluate differences of all possible factors in the risk of death. For all tests, a P-value <0.05 was defined as statistically significant.
Results
Expression of CPEB4 in normal breast and IDC tissues
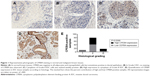
We analyzed CPEB4 expression in 107 primary IDC samples. Immunoreactivity of CPEB4 was detected only in cytoplasm. In normal breast tissues, CPEB4 was negative in all adipocytes and myoepithelial cells but sometimes positive in ductal epithelium (Figure 1). High expression of CPEB4 expression was observed in 48.6% (52/107) of IDC samples.
Expressions of ER, PR, and HER2 in IDC tissues
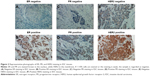
For ER, PR, and HER2, typical patterns of positive and negative immunohistochemical staining are shown in Figure 2. ER and PR were stained brown in the nucleus, while HER2 in the membrane. Positive rates for ER, PR, and HER2 were 49.5%, 42.1%, and 64.5%, respectively, in our 107 IDCs.
Correlations between CPEB4, ER, PR, and HER2 expressions and clinicopathologic parameters
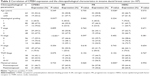
Statistical analysis showed that CPEB4 expression was positively correlated with the histological grading (P=0.037) and N stage (lymph node status, P<0.001) of IDC, and it was not statistically related to patients’ age, T stage (tumor size), M stage (metastasis), TNM stage, or menopausal status. PR expression was significantly associated with age (P=0.028) and menopausal status (P=0.005), while no significant association was found between ER and menstrual status. No significant relationship was found between HER2 expression and age, histological grading, T stage, N stage, M stage, TNM stage, and menstrual status. Detailed information is listed in Table 2.
Correlations between CPEB4 and ER, PR, and HER2 expressions
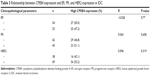
The correlations between CPEB4 and ER, PR, and HER2 expressions were evaluated. No significant relationship was observed between CPEB4 and ER (r=-0.028, P=0.770), PR (r=0.043, P=0.658), and HER2 (r=0.096, P=0.319) (Table 3).
Survival analysis
The average follow-up time for the 107 IDC cases was 83.6 months. Event was defined as death from any disease. No patient was excluded from the analysis. A total of 18 patients died (16.8%) during the follow-up period.
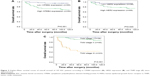
Correlation between higher CPEB4 expression and shorter overall survival times was revealed by Kaplan–Meier survival analysis (P=0.001, log-rank test) (Figure 3A). High HER2 expression and advanced TNM stage were both negatively correlated to survival time (P=0.020, P<0.001, log-rank test) (Figure 3B and C).
Univariate analysis regarding age, menstrual status, histological grading, TNM stage, ER, PR, HER2, and CPEB4 expression showed that the positive HER2 expression (P=0.036), high TNM stage (P<0.001), and high CPEB4 expression (P=0.005) were risk factors for IDC. Multivariate analysis using the Cox model demonstrated that the HER2 expression (P=0.026, hazard ratio [HR] =5.439, 95% confidence interval [CI] =1.227–24.114) was an independent risk factor, and high CPEB4 expression (P=0.022, HR =4.344, 95% CI =1.235–15.283) and high TNM stage (P<0.001, HR =13.804, 95% CI =4.769–39.935) were also independent risk factors (Table 4).
Discussion
In this study, we analyzed CPEB4 expression in 107 IDC tissues and evaluated its value as a potential prognostic indicator when compared with commonly used biomarkers: ER, PR, and HER2. Our data showed that high CPEB4 overexpression was observed in 48.6% of IDC samples, and its expression level was related to the histological grading and N stage. Patients with higher CPEB4 expression appeared to have poorer prognosis. Multivariate analysis showed that high CPEB4 expression was an independent prognostic factor for overall survival.
CPEB4 protein is overexpressed in a large variety of tumors (17 out of a total of 20 tumor types listed at www.proteinatlas.org/ENSG00000113742-CPEB4/cancer/tissue), and CPEB4 mRNA has been also confirmed in many tumor cells.27 Rhodes et al carried out a meta-analysis of 42 studies comparing global gene expression in 92 human cancers with matched normal tissue using oncomine.28 At this cutoff of 1.5-fold and a P-value of <0.05, 90 of 245 analyses showed a change in CPEB4. In the IDC, the expression of CPEB4 mRNA was upregulated with the score of 2.2, which was consistent with our result.
The relationship between CPEB4 expression and IDC progression together with poor survival has never been reported before, although it was demonstrated in other cancers. Tian et al showed that CPEB4 was commonly suppressed in hepatocellular carcinoma (HCC), and its expression was correlated with HCC prognosis. CPEB4 was directly targeted by miR-550a, which was frequently upregulated in HCC and facilitated HCC cell migration and invasion.24 Ortiz-Zapater et al demonstrated that the overexpression of CPEB4 regulated tPA expression to contributing tumor growth and angiogenesis in pancreatic ductal adenocarcinoma and glioblastomas.22 Xu and Liu suggested that CPEB4 was a candidate biomarker for defining metastatic cancers and promoted invasion and metastasis through TGF-beta signaling pathway.23
It is well established that the function of a protein depends on its location and is affected by normal or abnormal expression.29–31 We found that immunoreactivity of CPEB4 was detected only in cytoplasm. It supports the opinion that CPEB4 is associated with specific sequences in mRNA 3′-UTR, influencing translation by inducing cytoplasmic polyadenylation,32 which may help to explain how CPEB4 functions in the development of IDC. The present data supported that CPEB4-mediated regulation of gene expression might be a more general mechanism in cancer. It is obvious that factors in translation can influence cancer development. The relationship between CPEB4 expression and many clinical prognosis of cancer still needs further investigation.
In the present study, ER, PR, and HER2-positive rates were 49.5%, 42.1%, and 64.5%, respectively. PR expression was correlated with age and menopausal status (P=0.005). Multivariate Cox regression analysis showed that HER2 was associated with poor prognosis (P=0.026), correspondent with previous studies.27,28,33 By comparing commonly used biomarkers to CPEB4, no correlations between CPEB4 and ER, PR, and HER2 were observed.
The current study has several limitations. First, our findings should be replicated in other populations and larger cohorts to further validate our results. Second, further studies are needed to delineate the mechanisms behind this association between CPEB4 and IDC. Third, how to make best use of CPEB4 to stratify cancer patients for personalized treatment remains a critical goal.
In conclusion, high CPEB4 expression is associated with increased histological grading and N stage. Our study suggested that CPEB4 could serve as a useful prognostic indicator for IDC.
Acknowledgments
This work was supported by grants from the National Natural Science Foundation of China (81272387, 81470857, and 81502272), and the Shanghai Natural Science Foundation (134119b1100). We would like to thank the Department of Pathology of Huashan Hospital for supporting the research. We would also like to express our gratitude to all the reviewers for their precious advice and all the editors for their hard work.
Disclosure
The authors state that they have no conflicts of interest in relation to this article or the funding bodies.
References
Siegel R, Ma J, Zou Z, Jemal A. Cancer statistics, 2014. CA Cancer J Clin. 2014;64:9–29. | ||
Lebeau A, Kriegsmann M, Burandt E, Sinn HP. Invasive Mammakarzinome: Die aktuelle WHO-Klassifikation. [Invasive breast cancer: the current WHO classification]. Pathologe. 2014;35:7–17. German. | ||
Osborne CK, Yochmowitz MG, Knight WA 3rd, McGuire WL. The value of estrogen and progesterone receptors in the treatment of breast cancer. Cancer. 1980;46:2884–2888. | ||
Hammond ME, Hayes DF, Dowsett M, et al. American Society of Clinical Oncology/College Of American Pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer. J Clin Oncol. 2010;28:2784–2795. | ||
Wolff AC, Hammond ME, Hicks DG, et al. Recommendations for human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists clinical practice guideline update. Arch Pathol Lab Med. 2014;138: 241–256. | ||
Shaaban AM, Purdie CA, Bartlett JM, et al. HER2 testing for breast carcinoma: recommendations for rapid diagnostic pathways in clinical practice. J Clin Pathol. 2014;67:161–167. | ||
Piccart-Gebhart MJ, Procter M, Leyland-Jones B, et al. Trastuzumab after adjuvant chemotherapy in HER2-positive breast cancer. N Engl J Med. 2005;353:1659–1672. | ||
Payne SJ, Bowen RL, Jones JL, Wells CA. Predictive markers in breast cancer – the present. Histopathology. 2008;52:82–90. | ||
Scaltriti M, Chandarlapaty S, Prudkin L, et al. Clinical benefit of lapatinib-based therapy in patients with human epidermal growth factor receptor 2-positive breast tumors coexpressing the truncated p95HER2 receptor. Clin Cancer Res. 2010;16:2688–2695. | ||
Gianni L, Dafni U, Gelber RD, et al. Treatment with trastuzumab for 1 year after adjuvant chemotherapy in patients with HER2-positive early breast cancer: a 4-year follow-up of a randomised controlled trial. Lancet Oncol. 2011;12:236–244. | ||
Wang Q, Quan H, Zhao J, Xie C, Wang L, Lou L. RON confers lapatinib resistance in HER2-positive breast cancer cells. Cancer Lett. 2013;340:43–50. | ||
Maric P, Ozretic P, Levanat S, Oreskovic S, Antunac K, Beketic-Oreskovic L. Tumor markers in breast cancer – evaluation of their clinical usefulness. Coll Antropol. 2011;35:241–247. | ||
Hake LE, Richter JD. CPEB is a specificity factor that mediates cytoplasmic polyadenylation during Xenopus oocyte maturation. Cell. 1994;79:617–627. | ||
Stebbins-Boaz B, Hake LE, Richter JD. CPEB controls the cytoplasmic polyadenylation of cyclin, Cdk2 and c-mos mRNAs and is necessary for oocyte maturation in Xenopus. EMBO J. 1996;15:2582–2592. | ||
Richter JD. CPEB: a life in translation. Trends Biochem Sci. 2007;32:279–285. | ||
Wells DG, Dong X, Quinlan EM, et al. A role for the cytoplasmic polyadenylation element in NMDA receptor-regulated mRNA translation in neurons. J Neurosci. 2001;21:9541–9548. | ||
McGrew LL, Dworkin-Rastl E, Dworkin MB, Richter JD. Poly(A) elongation during Xenopus oocyte maturation is required for translational recruitment and is mediated by a short sequence element. Genes Dev. 1989;3:803–815. | ||
Huang YS, Kan MC, Lin CL, Richter JD. CPEB3 and CPEB4 in neurons: analysis of RNA-binding specificity and translational control of AMPA receptor GluR2 mRNA. EMBO J. 2006;25:4865–4876. | ||
Wang XP, Cooper NG. Comparative in silico analyses of cpeb1-4 with functional predictions. Bioinform Biol Insights. 2010;4:61–83. | ||
Igea A, Mendez R. Meiosis requires a translational positive loop where CPEB1 ensues its replacement by CPEB4. EMBO J. 2010;29:2182–2193. | ||
Afroz T, Skrisovska L, Belloc E, Guillen-Boixet J, Mendez R, Allain FH.A fly trap mechanism provides sequence-specific RNA recognition by CPEB proteins. Genes Dev. 2014;28:1498–1514. | ||
Ortiz-Zapater E, Pineda D, Martinez-Bosch N, et al. Key contribution of CPEB4-mediated translational control to cancer progression. Nat Med. 2012;18:83–90. | ||
Xu H, Liu B. CPEB4 is a candidate biomarker for defining metastatic cancers and directing personalized therapies. Med Hypotheses. 2013;81:875–877. | ||
Tian Q, Liang L, Ding J, et al. MicroRNA-550a acts as a pro-metastatic gene and directly targets cytoplasmic polyadenylation element-binding protein 4 in hepatocellular carcinoma. PLoS One. 2012;7:e48958. | ||
Bloom HJ, Richardson WW. Histological grading and prognosis in breast cancer; a study of 1,409 cases of which 359 have been followed for 15 years. Br J Cancer. 1957;11:359–377. | ||
Sobin LH, Fleming ID. TNM Classification of Malignant Tumors, fifth edition. Union Internationale Contre le Cancer and the American Joint Committee on Cancer. Cancer. 1997;80:1803–1804. | ||
Maric P, Ozretic P, Levanat S, Oreskovic S, Antunac K, Beketic-Oreskovic L. Tumor markers in breast cancer-evaluation of their clinical usefulness. Coll Antropol. 2011;35(1):241–247. | ||
Rhodes DR, Kalyana-Sundaram S, Mahavisno V, et al. Oncomine 3.0: genes, pathways, and networks in a collection of 18,000 cancer gene expression profiles. Neoplasia. 2007;9:166–180. | ||
Fanelli MA, Montt-Guevara M, Diblasi AM, et al. P-cadherin and beta-catenin are useful prognostic markers in breast cancer patients; beta-catenin interacts with heat shock protein Hsp27. Cell Stress Chaperones. 2008;13:207–220. | ||
Cuello-Carrion FD, Shortrede JE, Alvarez-Olmedo D, et al. HER2 and beta-catenin protein location: importance in the prognosis of breast cancer patients and their correlation when breast cancer cells suffer stressful situations. Clin Exp Metastasis. 2015;32:151–168. | ||
Sloan EK, Ciocca DR, Pouliot N, et al. Stromal cell expression of caveolin-1 predicts outcome in breast cancer. Am J Pathol. 2009;174:2035–2043. | ||
D’Ambrogio A, Nagaoka K, Richter JD. Translational control of cell growth and malignancy by the CPEBs. Nat Rev Cancer. 2013;13:283–290. | ||
Krop IE, Kim SB, Gonzalez-Martin A, et al. Trastuzumab emtansine versus treatment of physician’s choice for pretreated HER2-positive advanced breast cancer (TH3RESA): a randomised, open-label, phase 3 trial. Lancet Oncol. 2014;15:689–699. |
 © 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.