Back to Journals » Cancer Management and Research » Volume 13
Exosomal circ_IFT80 Enhances Tumorigenesis and Suppresses Radiosensitivity in Colorectal Cancer by Regulating miR-296-5p/MSI1 Axis
Authors Li L, Jiang Z, Zou X, Hao T
Received 12 December 2020
Accepted for publication 2 February 2021
Published 24 February 2021 Volume 2021:13 Pages 1929—1941
DOI https://doi.org/10.2147/CMAR.S297123
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Antonella D'Anneo
Liang Li,1 Zhipeng Jiang,2 Xiangcai Zou,3 Tengfei Hao1
1Department of Digestive Medicine Center, The Seventh Affiliated Hospital, Sun Yat-Sen University, Shenzhen, People’s Republic of China; 2Department of Gastrointestinal Surgery, The Sixth Affiliated Hospital, Sun Yat-Sen University, Guangzhou, People’s Republic of China; 3Department of Gastrointestinal Surgery, The Second Affiliated Hospital of Guangzhou Medical University, Guangzhou, People’s Republic of China
Correspondence: Liang Li Tel +86-0755-81207006
Email [email protected]
Background: Exosomal circular RNAs (circRNAs) can act as biomarkers and play crucial roles in colorectal cancer (CRC) and radiosensitivity. The aim of this study was to explore the functions and regulatory mechanism of exosomal circRNA intraflagellar transport 80 (circ_IFT80) in tumorigenesis and radiosensitivity of CRC.
Methods: Exosomes were detected using transmission electron microscopy (TEM). Protein levels were determined by Western blot assay. The expression of circ_IFT80, microRNA-296-5p (miR-296-5p) and musashi1 (MSI1) was measured by quantitative real-time polymerase chain reaction (qRT-PCR). Cell cycle distribution, cell apoptosis, and cell proliferation were detected by flow cytometry and 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay, respectively. Colony formation assay was used to determine the radiosensitivity of cells. The interaction between miR-296-5p and circ_IFT80 or MSI1 was verified by dual-luciferase reporter assay. A xenograft tumor model was established to explore the role of exosomal circ_IFT80 in vivo.
Results: Circ_IFT80 was upregulated in exosomes derived from CRC patient serum and CRC cells. Exosomal circ_IFT80 or circ_IFT80 overexpression facilitated tumorigenesis by increasing cell proliferation and reducing apoptosis, and inhibited radiosensitivity via promoting colony formation and inhibiting apoptosis. Additionally, circ_IFT80 acted as a sponge of miR-296-5p, and miR-296-5p reversed the effects of circ_IFT80 on tumorigenesis and radiosensitivity. Moreover, MSI1 was a direct target of miR-296-5p. Furthermore, miR-296-5p overexpression inhibited tumorigenesis and promoted radiosensitivity by downregulating MSI1. Exosomal circ_IFT80 also accelerated tumor growth in vivo.
Conclusion: Exosomal circ_IFT80 promoted tumorigenesis and reduced radiosensitivity by regulating miR-296-5p/MSI1 axis, which might provide a novel avenue for treatment of CRC.
Keywords: colorectal cancer, exosomes, circ_IFT80, miR-296-5p, MSI1
Introduction
Colorectal cancer (CRC) is a common malignant tumor, ranking the third in incidence and the second in mortality worldwide.1 Despite great advances in treatment, the overall survival of CRC patients remains unsatisfactory.2 Radiation therapy is an important clinical treatment for CRC, but acquired radioresistance during radiotherapy is still a major clinical obstacle to achieve successful treatment.3,4 Hence, it is urgent to unveil the underlying mechanisms of CRC and find novel ways to improve the radiosensitivity of CRC.
Exosomes are a class of 30–100 nm sized extracellular vesicles that originate from endosomal multivesicular bodies.5 Almost all types of cells, including cancer cells, can generate exosomes.6 Exosomes have received widespread attention due to their multiple functions in modulating cell-to-cell communication. Exosomes exert their physiological roles through the exchange of biomolecules such as proteins, DNAs, lipids, circular RNAs (circRNAs), microRNAs (miRNAs) and mRNA among different cells.7,8 Exosomes transported from stroma or tumor cells have been shown to be indispensable in all stages of cancer progression and play a vital role in regulating radiosensitivity.9,10
CircRNAs are a special type of non-coding RNAs (ncRNAs) that form continuous covalently closed-loops without 5ʹ-end cup and 3ʹ-end ploy A tail.11 Currently, growing evidence has suggested that circRNAs are more stable and difficult to degrade in response to the RNA exonuclease due to their closed-loop structures.12 Currently, increasing evidence has indicated that circRNAs can function as pivotal mediators in regulating the progression of diverse cancers, including CRC.13 CircRNAs have been suggested to be stable in exosomes, and these circRNAs can act as messengers of intercellular communication, thereby regulating the function of target cells.14 It has been reported that exosomal circRNAs have potential applications as disease biomarkers and new therapeutic targets.14 CircRNA intraflagellar transport 80 (circ_IFT80) is derived from IFT80 gene and located at chr3:160073800–160099506, which acts as an oncogene in CRC.15 However, the roles of exosomal circ_IFT80 in tumorigenesis and radiosensitivity of CRC remain poorly understood.
MiRNAs are a group of small (about 22 nucleotides) ncRNAs that negatively regulate their target genes via directly combining with 3ʹuntranslated regions (3ʹUTR) of target mRNAs.16,17 Crucial roles of miRNAs in CRC progression and response to radiosensitivity have been proposed in recent years.18,19 Previous studies showed that miR-296-5p acted as a tumor suppressor in multiple cancers, including CRC.20–22 However, the roles and underlying mechanism of miR-296-5p in tumorigenesis and radiosensitivity of CRC are still largely unknown.
Musashi1 (MSI1) is a neural RNA-binding protein that plays a critical role in regulating cell differentiation.23 Previous studies have shown that MSI1 acts as a tumor promoter in multiple cancers.24,25 In addition, MSI1 knockdown can sensitize colon cancer cells to radiation therapy.26 Furthermore, MSI1 was found to be overexpressed in CRC.27 Nevertheless, there is no report on the connections among exosomal circ_IFT80, miR-296-5p and MSI1 in tumorigenesis and radiosensitivity of CRC.
In this research, we separated exosomes from CRC patient serum. Circ_IFT80 and negative control (Vector) were loaded into exosomes by electroporation and then co-cultured with CRC cells to investigate the biological functions of exosomal circ_IFT80 in cell proliferation, cell cycle, apoptosis, and radiosensitivity. Moreover, the relationships among circ_IFT80, miR-296-5p and MSI1 were also studied in CRC cells.
Materials and Methods
Serum Collection
Blood samples were obtained from 15 CRC patients and 9 healthy control subjects at The Seventh Affiliated Hospital, Sun Yat-sen University. Blood samples were collected and centrifuged at 1600 g for 10 min at room temperature, followed by a second centrifugation at 12,000 g for 10 min at 4°C to remove cell debris. After that, the serum supernatant was transferred to RNase free tube, and then stored in −80°C. This research has been authorized by the ethics committee of The Seventh Affiliated Hospital, Sun Yat-sen University, which was in accordance with the Declaration of Helsinki Principles. Informed consent was acquired from each participant.
Cell Culture and Transfection
Human colonic epithelial cells (FHC) and CRC cells (SW480 and SW620) were purchased from Bena Culture Collection (Beijing, China). These cells were cultured in Dulbecco’s modified eagle medium (DMEM; Invitrogen, Carlsbad, CA, USA) containing 10% fetal bovine serum (FBS, Invitrogen). All cells were maintained in constant temperature incubator with 5% CO2 at 37°C.
Circ_IFT80 overexpression vector (circ_IFT80), miR-296-5p mimic (miR-296-5p), miR-296-5p inhibitor (anti-miR-296-5p), MSI1-overexpressing plasmid (MSI1), and their negative controls (Vector, miR-NC, anti-miR-NC, and pcDNA) were obtained from Genechem (Shanghai, China). Transient transfection was performed by Lipofectamine 3000 Reagent (Invitrogen).
Exosome Isolation and Incubation with CRC Cells
The collected serum samples were subjected to centrifugation at 3000g for 15 min to eliminate cells and cellular debris. Exosomes from serum samples were isolated using ExoQuick precipitation kit (SBI, System Biosciences, Mountain View, CA) following the manufacturer’s instructions. Gene Pulser X Electroporator (Bio-Rad, Hercules, CA, USA) was used to electroporate circ_IFT80 or Vector into exosomes as previously described.28 Briefly, exosomes (3 μg) and circ_IFT80 or Vector were mixed in electroporation buffer (400 μL) at 4°C. After electroporation at 350 V and 150 μF in a 4-mm cuvette, the mixture was then incubated for 30 min at 37°C to ensure the exosome membrane was fully recovered. For incubation of exosomes with SW480 and SW620 cells, these cells were co-inoculated into a 12-well plate and exosomes containing circ_IFT80 or Vector were added to per well. SW480 and SW620 cells were collected after incubation for indicated time for further analysis.
Transmission Electron Microscopy (TEM)
The exosomes were resuspended in phosphate-buffered saline (PBS; Beyotime, Shanghai, China) and then placed on a sheet of parafilm. After that, a carbon-coated copper grid was floated on the drop at room temperature for 5 min. Next, copper grid with exosomes adsorbed was then suspended in 2% phosphotungstic acid (pH 7.0, Sangon Biotech, Shanghai, China) for 5 seconds, and then excess liquid was carefully drained off. The morphology and ultrastructure of exosomes were subsequently observed with JEM-1400 (JEOL, Akishima, Japan) at 100 kV.
Western Blot Assay
Total protein was extracted by RIPA lysis buffer (Beyotime). The protein samples were denatured by heating at 100°C for 3–5 min, and then quantified by BCA protein assay kit (Abcam, Cambridge, UK). After that, protein samples (about 30 μg/lane) were loaded onto sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE). Next, the gels were transferred to polyvinylidene fluoride membranes (PVDF; Invitrogen). After blocking with 5% non-fat milk (Sangon Biotech), these membranes were then incubated for 12 h at 4°C with primary antibodies. The primary antibodies including TSG101 (1:500, ab30871), CD63 (1:1000, ab118307), CD9 (1:500, ab223052) B-cell lymphoma-2 (Bcl-2; 1:1000, ab59348), BCL2-associated X protein (Bax; 1:500, ab53154), MSI1 (1:1000, ab154497), and GAPGH (1:2000, ab37168) were purchased from Abcam. Subsequently, these membranes were continuously probed with secondary antibody (ab205718, 1:4000, Abcam). At last, protein bands were detected using enhanced chemiluminescence reagent (Tanon, Shanghai, China).
Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR)
Total exosomal RNAs were obtained using the exoRNeasy Midi Kit (Qiagen, Valencia, CA, USA). Total RNA from cells was isolated by TRIzol reagent (Invitrogen). Next, complementary DNA (cDNA) was synthesized by One Step PrimeScript cDNA Synthesis Kit (Takara, Tokyo, Japan) or miRNA cDNA synthesis kit (Ribobio, Guangzhou, China). Then, qRT-PCR reactions were performed on ABI Prism 7900HT (Applied Biosystems, Foster city, CA, USA) using a SYBR Premix Ex Taq II (Takara). In this study, primers used for amplification were listed as follows: circ_IFT80, 5ʹ-TATGGGATAGTTACGGCCGC-3ʹ (sense) and 5ʹ-TGGGATGCTCATGAGGTTGT-3ʹ (anti-sense); IFT80, 5ʹ-AGCAGGCAAGCAGCTAATCA-3ʹ (sense) and 5ʹ-CTCCAGCAATCTGAGTGCCA-3ʹ (anti-sense); miR-296-5p, 5ʹ-GTATCCAGTGCAGGGTCCGA-3ʹ (sense) and 5ʹ-CGACGAGGGCCCCCCCT-3ʹ (anti-sense); MSI1, 5ʹ-GCTCGACTCCAAAACAATTGACC-3ʹ (sense) and 5ʹ-GGCTGAGCTTTCTTACATTCCAC-3ʹ (anti-sense); U6, 5ʹ-CTCGCTTCGGCAGCACATATACT-3ʹ (sense) and 5ʹ-ACGCTTCACGAATTTGCGTGTC-3ʹ (anti-sense); glyceraldehyde-3-phosphate dehydrogenase (GAPDH), 5ʹ-GGAGAAGCTGAGTCATGGGT-3ʹ; (sense) and 5ʹ- GGTGGGAGATCTGGTTTCCG-3ʹ (anti-sense). The expression of genes was calculated by 2-ΔΔCt method. U6 or GAPDH was used as an internal reference.
Actinomycin D Treatment
Actinomycin D (2 mg/mL, Sigma-Aldrich, St. Louis, MO, USA) was added into cells to block transcription, followed by RNA extraction. At last, the expression levels of circ_IFT80 and IFT80 mRNA were tested using qRT-PCR analysis.
Flow Cytometry
For analysis of cell cycle progression, cells were collected after treatment for 48 h, and then fixed with 70% ice-cold ethanol (Beyotime) at −20°C for 12 h. Next, the fixed cells were subsequently incubated with propidium iodide (PI; Sangon Biotech) and RNase A (Sangon Biotech) in the darkness for 15 min. At last, cell cycle distribution was determined by flow cytometer (Partec AG, Arlesheim, Switzerland). For the cell apoptosis analysis, cells were collected at 48 h post-treatment, and resuspended in 400 μL of binding buffer. Next, the cells were then incubated with 5 µL of Annexin V-fluorescein isothiocyanate (FITC) and 10 µL of PI (Sangon Biotech) in the darkness for 20 min. The apoptotic cells were analyzed by flow cytometry.
Cell Proliferation Assay
Cell proliferation was measured by 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT; Beyotime). In brief, 100 μL of cell suspension was seeded into 96-well plates. Twenty microliters of MTT was added to per well at different time points. After incubation for 3–4 h, the supernatant was discarded and then 150 μL of methyl sulfoxide solution (DMSO; Beyotime) was added to dissolve formazan product. The absorbance at 570 nm was determined by microplate reader (Bio-Rad).
Irradiation Treatment
Cells (SW480 and SW620) were treated with different doses of radiation (0, 2, 4, and 6Gy) by a linear accelerator (Varian, Palo Alto, CA, USA) at a dose rate of 3.5 Gy/min.
Colony Formation Assay
Treated cells (SW480 and SW620) were seeded into six-well plates and the medium was refreshed every 3 days. After 2 weeks of incubation, the colonies were fixed with 4% paraformaldehyde (Beyotime). After that, the colonies were stained with crystal 0.1% violet (Beyotime). The number of colonies (one colony > 50 cells) was counted using a microscope (Leica, Wetzlar, Germany). Cell survival fraction was calculated as previously described.29
Bioinformatics Prediction and Dual-Luciferase Reporter Assay
The putative binding sequence of miR-296-5p and circ_IFT80 or MSI1 was predicted by circinteractome or starBase. The circ_IFT80 or MSI1 3ʹUTR fragments containing binding sequence for miR-296-5p were synthesized and inserted into pmirGlO luciferase reporter vector (Promega, Madison, WI, USA), thereby generating wild-type luciferase reporter plasmid (circ_IFT80-WT or MSI1-WT). Meanwhile, the mutated binding sites were used for the mutant reporter (circ_IFT80-MUT or MSI1-MUT) in the same way. Afterward, SW480 and SW620 cells were co-transfected with miR-296-5p or miR-NC and the constructed reporter plasmid for 48 h. At last, the luciferase activity was examined by Dual-Luciferase Reporter assay system (Promega), followed by normalization to the Renilla luciferase.
Tumor Formation Assay in vivo
A total of 12 BALB/c nude mice (female, 5 weeks, weighing 18 ± 2 g) were purchased from Vital River Laboratory Animal Technology Co., Ltd (Beijing, China). SW480 cells were subcutaneously injected into the right flank of nude mice. These mice were randomly separated into two groups (6 mice per group). One week later, the collected exosomes containing circ_IFT80 or Vector was subsequently injected into mice via the tail vein (5 μg/mouse) every 3 days. The volume of tumor was monitored with calipers every week, followed by calculation of tumor volume with the following equation: length × width2 × 0.5. All mice were sacrificed 4 weeks later, and tumor samples were resected, weighted and collected for further analysis. The animal experiments were approved by the Animal Care and Use Committee of The Seventh Affiliated Hospital, Sun Yat-sen University, and performed according to the guidelines for laboratory animal welfare (GB/T 35892–2018).
Statistical Analysis
All data from at least 3 independent experiments were shown as the mean ± standard deviation (SD) and analyzed by Graphpad Prism version 6.0 software (GraphPad Software, La Jolla, CA, USA). The statistical differences between groups two or more groups were analyzed by Student’s t-test and one-way analysis of variance (ANOVA). Statistical significance was considered when P<0.05.
Results
Circ_IFT80 Was Upregulated in Exosomes from CRC Patient Serum and CRC Cells
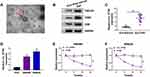
Exosomes were extracted from the serum of CRC patients. As shown in Figure 1A, serum-derived exosomes were observed under a TEM and the diameter of vesicles was about 40–60 nm, which was in consistent with the morphological characteristics of exosomes. The levels of exosome protein markers (TSG101, CD63 and CD9) were detected by Western blot. Results showed that TSG101, CD63 and CD9 were all highly enriched in the isolated exosomes (Figure 1B). In addition, the results of qRT-PCR demonstrated that circ_IFT80 expression was significantly increased in serum-derived exosomes from CRC patients compared to those derived from healthy individuals (Figure 1C). Moreover, circ_IFT80 level was also enhanced in SW480 and SW620 cells compared to that in FHC cells (Figure 1D). Actinomycin D assay revealed that the half-life of circ_IFT80 transcript exceeded 24 h, while that of the IFT80 mRNA displayed about 8 h (Figure 1E and F), suggesting that circ_IFT80 transcript was more stable than linear IFT80 mRNA transcript in SW480 and SW620 cells. These results suggested that exosomes were successfully isolated from serum of CRC patients and circ_IFT80 was overexpressed in CRC.
Exosomal circ_IFT80 Promoted Tumorigenesis and Decreased Radiosensitivity of CRC Cells
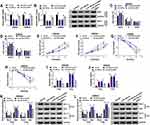
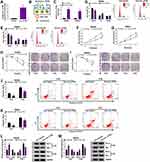
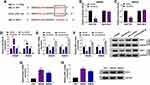
Circ_IFT80 or Vector was electroporated into exosomes to determine whether circ_IFT80 was successfully overexpressed. The data revealed that circ_IFT80 was upregulated in exosomes electroporated with circ_IFT80 (Exo-circ_IFT80) compared to exosomes electroporated with Vector (Exo-Vector) (Figure 2A). As shown in Figure 2B, circ_IFT80 and Vector could be delivered into CRC cells through exosomes. The expression of circ_IFT80 in CRC cells co-cultured with Exo-circ_IFT80 or Exo-Vector was detected by qRT-PCR. Results showed that Exo-circ_IFT80 obviously increased the expression of circ_IFT80 in SW480 and SW620 cells compared to Exo-Vector group (Figure 2C). Flow cytometry analysis showed that fewer cells were distributed in the G0/G1 phase in Exo-circ_IFT80 group, while more cells were distributed in S phase (Figure 2D and E). MTT assay indicated that Exo-circ_IFT80 increased SW480 and SW620 cell proliferation (Figure 2F and G), indicating that Exo-circ_IFT80 promoted the proliferation by blocking the CRC cell cycle to S phase. SW480 and SW620 cells were incubated with Exo-circ_IFT80 or Vector and then irradiated with 0 Gy to 6 Gy to explore the effect of Exo-circ_IFT80 on radiosensitivity. It was also showed that Exo-circ_IFT80 increased the survival rate of SW480 and SW620 cells exposed to radiation (Figure 2H and I). Moreover, radiation-induced apoptosis was decreased by Exo-circ_IFT80 in SW480 and SW620 cells (Figure 2J and K). Furthermore, the expression levels of apoptosis-related proteins Bcl-2 (anti-apoptotic molecule) and Bax (pro-apoptotic molecule) were analyzed by Western blot assay. As displayed in Figure 2L and M, Exo-circ_IFT80 increased the protein level of Bcl-2 while decreased the protein expression of Bax with or without irradiation. Taken together, these findings indicated that Exo-circ_IFT80 increased tumorigenesis and decreased radiosensitivity of CRC cells.
MiR-269-5p Was a Direct Target of circ_IFT80
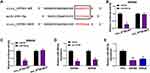
It is well known that circRNAs can act as ceRNAs to upregulate mRNAs through binding with miRNAs.30 To investigate the underlying molecular mechanisms of circ_IFT80 in CRC. Target miRNAs of circ_IFT80 were screened by circinteractome. As presented in Figure 3A, binding sites were observed between circ_IFT80 and miR-296-5p, which indicated an interaction relationship. To verify the targeting relationship between circ_IFT80 and miR-296-5p, dual-luciferase reporter assay was conducted. As illustrated Figure 3B and C, the luciferase activity of circ_IFT80-WT was markedly decreased after transfected with miR-296-5p, but transfection of miR-296-5p did not affect the luciferase activity of circ_IFT80-MUT. Moreover, we found that miR-296-5p expression was reduced by overexpression of circ_IFT80 (Figure 3D), suggesting that miR-296-5p was negatively regulated by circ_IFT80. Furthermore, it was observed that miR-296-5p was lowly expressed in CRC cells (SW480 and SW620) compared to that in FHC cells (Figure 3E). Collectively, these data indicated that circ_IFT80 acted as a sponge of miR-296-5p.
Overexpressed circ_IFT80 Facilitated CRC Cell Progression and Reduced Radiosensitivity by Downregulating miR-296-5p
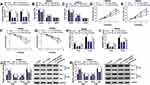
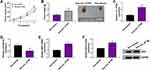
Given that circ_IFT80 acted as a sponge of miR-296-5p, rescue experiments were performed to explore whether miR-296-5p was involved in circ_IFT80-mediated cellular behaviors and radiosensitivity. The results showed that miR-296-5p expression was blocked by circ_IFT80 transfection, which was restored by miR-296-5p upregulation (Figure 4A). Flow cytometry analysis indicated that circ_IFT80 upregulation blocked SW480 and SW620 cells at G1/G0 phase and increased the proportions of cells in S phase, which was reversed by overexpression of miR-296-5p (Figure 4B and C). MTT assay demonstrated that enforced expression of miR-296-5p abated the promotive effect of circ_IFT80 overexpression on cell proliferation (Figure 4D and E). Next, we explored the effect of circ_IFT80 and miR-296-5p on radiosensitivity. As shown in Figure 4F and G, survival rate was significantly higher in circ_IFT80-overexpressing cells than that in Vector group after irradiation treatment, which could be abolished by miR-296-5p upregulation. Moreover, elevation of circ_IFT80 could reduce the apoptotic rate of SW480 and SW620 cells without or with radiation, which could be reversed by miR-296-5p upregulation (Figure 4H and I). Furthermore, upregulated circ_IFT80 increased the expression of Bcl-2 protein and decreased the expression of Bax protein in SW480 and SW620 cells with or without radiation, while these effects were reversed by upregulated miR-296-5p (Figure 4J and K). Taken together, these results revealed that circ_IFT80 could regulate tumorigenesis and radiosensitivity by sponging miR-296-5p in CRC cells.
MSI1 Was a Direct Target of miR-296-5p in CRC Cells
Growing evidence has shown that miRNAs exert their functions through binding to 3ʹUTR of target mRNAs.31 The possible target genes of miR-296-5p were predicted by starBase. As shown in Figure 5A, 3ʹUTR of MSI1 had potential binding sites for miR-296-5p, suggesting that MSI1 might be a target of miR-296-5p. Next, the dual-luciferase reporter assay was performed to verify whether miR-296-5p could directly target bind to MSI1. The results showed that miR-296-5p overexpression led to an obvious decrease in the luciferase activity of MSI1-WT, but it did not affect the luciferase activity of MSI1-MUT (Figure 5B and C). The expression of miR-296-5p was increased by miR-296-5p transfection and decreased by anti-miR-296-5p (Figure 5D), indicating that miR-296-5p and anti-miR-296-5p were successfully transfected. Next, we explored the effect of miR-296-5p on the expression of MSI1. The results of qRT-PCR and Western blot demonstrated that overexpression of miR-296-5p inhibited the mRNA and protein expression of MSI1, which was reversed by anti-miR-296-5p (Figure 5E and F). In addition, it was found that MSI1 mRNA and protein levels were both increased in SW480 and SW620 cells compared to that in FHC cells (Figure 5G and H). These results indicated that miR-296-5p directly targeted MSI1 and negatively regulated MSI1 expression.
MiR-296-5p Overexpression Inhibited Tumorigenesis and Improved Radiosensitivity by Downregulating MSI1 in CRC Cells
To further explore whether miR-296-5p functions were regulated by MSI1, rescue assays were performed. The results of qRT-PCR and Western blot showed that overexpression of miR-296-5p reduced the expression of MSI1, which was restored by addition of MSI1 (Figure 6A and B). Moreover, miR-296-5p upregulation induced cell cycle arrest at G1/G0 phase and inhibited cell proliferation, which could be abolished by upregulating MSI1 (Figure 6C–F). Moreover, the inhibitory effect of miR-296-5p on survival rate was overturned by overexpression of MSI1 in SW480 and SW620 cells treated with radiation (Figure 6G and H). Furthermore, upregulation of miR-296-5p increased apoptosis, miR-296-5p overexpression and radiation further promoted apoptosis in SW480 and SW620 cells, which could be abated by overexpressed MSI1 (Figure 6I and J). Besides, MSI1 elevation also abolished the effects of miR-296-5p on Bcl-2 and Bax (Figure 6K and L). These above findings indicated that miR-296-5p exerted its functions by targeting MSI1.
Exo-circ_IFT80 Promoted Tumor Growth in vivo
Lastly, we attempted to unveil the impact of Exo-circ_IFT80 on CRC tumor formation in nude mice. SW480 cells were injected subcutaneously into nude mice to form tumor masses. One week later, Exo-circ_IFT80 or Exo-Vector was subsequently injected into the mice every 3 days. Similar to the in vitro analysis, Exo-circ_IFT80 enhanced tumor volume and weight (Figure 7A and B). Moreover, the expression of circ_IFT80 was increased and miR-296-5p expression was decreased in tumor tissues from Exo-circ_IFT80 group compared to control group (Figure 7C and D). In addition, Exo-circ_IFT80 increased the mRNA and protein expression of MSI1 in tumor tissues (Figure 7E and F). All findings indicated that Exo-circ_IFT80 also accelerated tumor growth by regulating miR-296-5p and MSI1 in vivo.
Discussion
CRC is an aggressive malignant tumor with the high recurrence rate, which is increasingly threatening people’s health.32 Radiotherapy is one of the effective strategies for CRC treatment.33 Nevertheless, the effects of exosomal circRNAs on radiosensitivity of CRC cells have not been fully elucidated. The aim of our study was to investigate the influence of exosomal circ_IFT80 on tumorigenesis and radiosensitivity of CRC.
Exosomes can be excreted into peripheral blood and widely spread in human fluids such as saliva, urine, blood, and breast milk.34 Many studies have suggested that exosomes are involved in regulating the tumor-normal communication in the tumor microenvironment.35,36 Exosomes have been suggested to play critical roles in the initiation, development and progression of cancer, as well as chemosensitivity and radiosensitivity.37,38 The interest in using exosomes as a therapeutic carrier for the in vivo delivery of circRNA has increased dramatically over the past few years, due to its lowered immunogenicity.39 Considered about their availability and stability, exosomal circRNAs have been proposed as a novel cancer diagnostic tool and have prognostic value.14,40 Moreover, exosomal circRNAs also play essential roles in CRC. For instance, exosomal circPACRGL contributed to the progression of CRC by regulating miR-142-3p/miR-506-3p-TGF-β1 axis.41 Exosome-delivered hsa_circ_0005963 facilitated glycolysis to induced chemoresistance by the miR-122/PKM2 axis in CRC.42 A previous report showed that circ_IFT80 expression was upregulated in CRC serum exosomes, CRC tissues and CRC cells, circ_IFT80 silencing repressed cell proliferation while accelerated apoptosis through miR-1236-3p/HOXB7 axis in CRC.15 However, the role of exosomal circ_IFT80 in tumorigenesis and radiosensitivity of CRC have not been reported. In this research, we found that circ_IFT80 was upregulated in serum-derived exosomes from CRC patients and CRC cells. Functionally, exosomal circ_IFT80 or circ_IFT80 upregulation increased the proliferation of CRC cells via promoting cell cycle, and exosomal circ_IFT80 or circ_IFT80 overexpression reduced radiosensitivity by promoting survival and decreasing apoptosis, indicating that exosomal circ_IFT80 could promote tumorigenesis and suppress radiosensitivity in CRC cells.
Mechanistically, circRNAs play a role in human malignant tumors through competitively binding to miRNAs and regulating the expression of target genes.43 Subsequently, the underlying mechanism of how circ_IFT80 exerted its functions was explored. Through bioinformatics analysis (circinteractome) and dual-luciferase reporter assay, miR-296-5p was found to be a target of circ_IFT80. MiR-296-5p has been suggested to play a tumor-suppressive role in many cancers. For example, Lee et al revealed that miR-296-5p functioned as a tumor suppressor via targeting PIK3R1 in prostate cancer.44 Moreover, Xu et al proved that miR-296-5p exerted the anti-tumor role in lung cancer by targeting PLK1.45 Furthermore, Zhang et al. pointed out that miR-296 suppressed cell proliferation and facilitated cell apoptosis in CRC via regulating ARRB1-mediated AKT activation.46 MiR-296-5p has also been reported to be associated with the resistance of laryngeal carcinoma to radiotherapy.47 In this study, miR-296-5p was lowly expressed in CRC cells. Additionally, miR-296-5p overexpression reversed the effects of circ_IFT80 on promoting cell proliferation and reducing radiosensitivity in CRC cell, suggesting circ_IFT80 exerted its functions by regulating miR-296-5p.
Previous studies have demonstrated that miRNAs participate in regulating cancer progression and radiosensitivity by binding to their target mRNAs.48,49 Next, the possible targets of miR-296-5p were predicted by starBase. Due to its tumor-promotive effect, MSI1 was selected as a candidate target gene of miR-296-5p. Then, we demonstrated that MSI1 was directly targeted by miR-296-5p. MSI1 has been reported to act as a diagnostic marker and potential therapeutic target in some cancers.50 Moreover, MSI1 is a critical regulator in tumorigenesis and progression of several cancers. For instance, Yang et al pointed out that upregulation of MSI1 weakened the anti-cancer role of miR-331 in gastric cancer.51 Gao et al stated that MSI1 was highly expressed in colon cancer tissues, and its downregulation inhibited the growth of colon cancer by targeting p21cip1.52 Moreover, Araujo et al revealed that MSI1 could affect radioresistance via controlling DNA-protein kinase catalytic subunit in glioblastoma.53 Smith et al illustrated that MSI1 was upregulated in CRC tissue samples, miR-137 acted as a tumor-suppressive miRNA by targeting MSI1.27 In this research, MSI1 was found to be upregulated in CRC cells. Rescue assays indicated MSI1 upregulation abolished the effects of miR-296-5p on reduction of tumorigenesis and enhancement of radiosensitivity in CRC cells. In addition, in vivo experiments demonstrated that exosomal circ_IFT80 promoted tumor growth by inhibiting miR-296-5p and increasing MSI1 expression.
In conclusion, our study revealed that exosomal circ_IFT80 cells could promote cell proliferation and cell cycle progression as well as decrease apoptosis and radiosensitivity in CRC cells via regulating miR-296-5p/MSI1 axis. Our study not only provided novel insight into the roles of exosomes in CRC but also offered a promising biomarker for the diagnosis or treatment for CRC.
Highlights
- Exosomal circ_IFT80 was upregulated in CRC.
- Exosomal circ_IFT80 enhanced tumorigenesis and inhibited radiosensitivity by downregulating miR-296-5p.
- MiR-296-5p suppressed tumorigenesis and improved radiosensitivity by targeting MSI1.
- MiR-296-5p could directly bind to circ_IFT80, and MSI1 was a target of miR-296-5p.
Data Sharing Statement
The analyzed data sets generated during the present study are available from the corresponding author on reasonable request.
Ethics Approval and Consent to Participate
The present study was approved by the ethical review committee of The Seventh Affiliated Hospital, Sun Yat-sen University.
Author Contributions
Conceptualization and Methodology: Zhipeng Jiang and Xiangcai Zou; Formal analysis and Data curation: Xiangcai Zou and Tengfei Hao; Validation and Investigation: Liang Li and Tengfei Hao; Writing – original draft preparation and Writing – review and editing: Liang Li, Zhipeng Jiang and Xiangcai Zou; Approval of final manuscript: all authors. All authors contributed to data analysis, drafting or revising the article, have agreed on the journal to which the article will be submitted, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Funding
No funding was received.
Disclosure
The authors declare that they have no competing interests.
References
1. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. doi:10.3322/caac.21492
2. Siegel R, Naishadham D, Jemal A. Cancer statistics, 2013. CA Cancer J Clin. 2013;63(1):11–30.
3. Dell’Acqua V, Surgo A, Kraja F, et al. Stereotactic radiation therapy in oligometastatic colorectal cancer: outcome of 102 patients and 150 lesions. Clin Exp Metastasis. 2019;36(4):331–342.
4. Park SY, Lee CJ, Choi JH, et al. The JAK2/STAT3/CCND2 Axis promotes colorectal Cancer stem cell persistence and radioresistance. J Exp Clin Cancer Res. 2019;38(1):399.
5. Cocucci E, Meldolesi J. Ectosomes and exosomes: shedding the confusion between extracellular vesicles. Trends Cell Biol. 2015;25(6):364–372.
6. Théry C, Zitvogel L, Amigorena S. Exosomes: composition, biogenesis and function. Nat Rev Immunol. 2002;2(8):569–579.
7. Vlassov AV, Magdaleno S, Setterquist R, Conrad R. Exosomes: current knowledge of their composition, biological functions, and diagnostic and therapeutic potentials. Biochim Biophys Acta. 2012;1820(7):940–948.
8. Ge R, Tan E, Sharghi-Namini S, Asada HH. Exosomes in Cancer Microenvironment and Beyond: have we Overlooked these Extracellular Messengers? Cancer Microenviron. 2012;5(3):323–332.
9. Malla B, Zaugg K, Vassella E, Aebersold DM, Dal Pra A. Exosomes and Exosomal MicroRNAs in Prostate Cancer Radiation Therapy. Int J Radiat Oncol Biol Phys. 2017;98(5):982–995.
10. Boelens MC, Wu TJ, Nabet BY, et al. Exosome transfer from stromal to breast cancer cells regulates therapy resistance pathways. Cell. 2014;159(3):499–513. doi:10.1016/j.cell.2014.09.051
11. Memczak S, Jens M, Elefsinioti A, et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013;495(7441):333. doi:10.1038/nature11928
12. Chen -L-L, Yang L. Regulation of circRNA biogenesis. RNA Biol. 2015;12(4):381–388. doi:10.1080/15476286.2015.1020271
13. Patop IL, Kadener S. circRNAs in Cancer. Curr Opin Genet Dev. 2018;48:121–127. doi:10.1016/j.gde.2017.11.007
14. Wang Y, Liu J, Ma J, et al. Exosomal circRNAs: biogenesis, effect and application in human diseases. Mol Cancer. 2019;18(1):116. doi:10.1186/s12943-019-1041-z
15. Feng W, Gong H, Wang Y, et al. circIFT80 Functions as a ceRNA of miR-1236-3p to Promote Colorectal Cancer Progression. Mol Ther Nucleic Acids. 2019;18:375–387. doi:10.1016/j.omtn.2019.08.024
16. Ardekani AM, Naeini MM. The role of microRNAs in human diseases. Avicenna J Med Biotechnol. 2010;2(4):161.
17. Jansson MD, Lund AH. MicroRNA and cancer. Mol Oncol. 2012;6(6):590–610. doi:10.1016/j.molonc.2012.09.006
18. Afshar S, Najafi R, Sedighi Pashaki A, et al. MiR-185 enhances radiosensitivity of colorectal cancer cells by targeting IGF1R and IGF2. Biomed Pharmacother. 2018;106:763–769. doi:10.1016/j.biopha.2018.07.002
19. Samadi P, Afshar S, Amini R, et al. Let-7e enhances the radiosensitivity of colorectal cancer cells by directly targeting insulin-like growth factor 1 receptor. J Cell Physiol. 2019;234(7):10718–10725.
20. He Z, Yu L, Luo S, et al. miR-296 inhibits the metastasis and epithelial-mesenchymal transition of colorectal cancer by targeting S100A4. BMC Cancer. 2017;17(1):140.
21. Shi DM, Li LX, Bian XY, et al. miR-296-5p suppresses EMT of hepatocellular carcinoma via attenuating NRG1/ERBB2/ERBB3 signaling. J Exp Clin Cancer Res. 2018;37(1):294.
22. Zhou SL, Tang QL, Zhou SX, Ren RZ. MiR-296-5p suppresses papillary thyroid carcinoma cell growth via targeting PLK1. Eur Rev Med Pharmacol Sci. 2019;23(5):2084–2091.
23. Li D, Peng X, Yan D, et al. Msi-1 is a predictor of survival and a novel therapeutic target in colon cancer. Ann Surg Oncol. 2011;18(7):2074–2083.
24. Nikpour P, Emadi-Baygi M, Mohhamad-Hashem F, Maracy MR, Haghjooy-Javanmard S. MSI1 overexpression in diffuse type of gastric cancer. Pathol Res Pract. 2013;209(1):10–13.
25. Gong P, Wang Y, Gao Y, et al. Msi1 promotes tumor progression by epithelial-to-mesenchymal transition in cervical cancer. Hum Pathol. 2017;65:53–61.
26. Sureban SM, May R, George RJ, et al. Knockdown of RNA binding protein musashi-1 leads to tumor regression in vivo. Gastroenterology. 2008;134(5):1448–1458.
27. Smith AR, Marquez RT, Tsao WC, et al. Tumor suppressive microRNA-137 negatively regulates Musashi-1 and colorectal cancer progression. Oncotarget. 2015;6(14):12558–12573.
28. Qu L, Ding J, Chen C, et al. Exosome-Transmitted lncARSR Promotes Sunitinib Resistance in Renal Cancer by Acting as a Competing Endogenous RNA. Cancer Cell. 2016;29(5):653–668.
29. Zhou S, Ye W, Ren J, et al. MicroRNA-381 increases radiosensitivity in esophageal squamous cell carcinoma. Am J Cancer Res. 2015;5(1):267.
30. Kulcheski FR, Christoff AP, Margis R. Circular RNAs are miRNA sponges and can be used as a new class of biomarker. J Biotechnol. 2016;238:42–51.
31. Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell. 2009;136(2):215–233.
32. Siegel RL, Miller KD, Fedewa SA, et al. Colorectal cancer statistics, 2017. CA Cancer J Clin. 2017;67(3):177–193.
33. Tam SY, Wu VWC A. Review on the Special Radiotherapy Techniques of Colorectal Cancer. Front Oncol. 2019;9:208.
34. Cappello F, Logozzi M, Campanella C, et al. Exosome levels in human body fluids: a tumor marker by themselves? Eur J Pharm Sci. 2017;96:93–98.
35. Takahashi K, Yan IK, Kogure T, Haga H, Patel T. Extracellular vesicle‐mediated transfer of long non‐coding RNA ROR modulates chemosensitivity in human hepatocellular cancer. FEBS Open Bio. 2014;4(1):458–467.
36. Jella KK, Nasti TH, Li Z, Malla SR, Buchwald ZS, Khan MK. Exosomes, Their Biogenesis and Role in Inter-Cellular Communication, Tumor Microenvironment and Cancer Immunotherapy. Vaccines. 2018;6(4):212.
37. Bach DH, Hong JY, Park HJ, Lee SK. The role of exosomes and miRNAs in drug-resistance of cancer cells. Int J Cancer. 2017;141(2):220–230.
38. Han F, Huang D, Huang X, Wang W, Yang S, Chen S. Exosomal microRNA-26b-5p down-regulates ATF2 to enhance radiosensitivity of lung adenocarcinoma cells. J Cell Mol Med. 2020;24(14):7730–7742.
39. Jiang L, Gu Y, Du Y, Liu J. Exosomes: diagnostic Biomarkers and Therapeutic Delivery Vehicles for Cancer. Mol Pharm. 2019;16(8):3333–3349.
40. Bai H, Lei K, Huang F, Jiang Z, Zhou X. Exo-circRNAs: a new paradigm for anticancer therapy. Mol Cancer. 2019;18(1):56.
41. Shang A, Gu C, Wang W, et al. Exosomal circPACRGL promotes progression of colorectal cancer via the miR-142-3p/miR-506-3p- TGF-β1 axis. Mol Cancer. 2020;19(1):117.
42. Wang X, Zhang H, Yang H, et al. Exosome-delivered circRNA promotes glycolysis to induce chemoresistance through the miR-122-PKM2 axis in colorectal cancer. Mol Oncol. 2020;14(3):539–555.
43. Greene J, Baird AM, Brady L, et al. Circular RNAs: biogenesis, Function and Role in Human Diseases. Front Mol Biosci. 2017;4:38.
44. Lee KH, Lin FC, Hsu TI, et al. MicroRNA-296-5p (miR-296-5p) functions as a tumor suppressor in prostate cancer by directly targeting Pin1. Biochim Biophys Acta. 2014;1843(9):2055–2066.
45. Xu C, Li S, Chen T, et al. miR-296-5p suppresses cell viability by directly targeting PLK1 in non-small cell lung cancer. Oncol Rep. 2016;35(1):497–503.
46. Zhang Z, Zhong X, Xiao Y, Chen C. MicroRNA-296 inhibits colorectal cancer cell growth and enhances apoptosis by targeting ARRB1-mediated AKT activation. Oncol Rep. 2019;41(1):619–629.
47. Maia D, de Carvalho AC, Horst MA, Carvalho AL, Scapulatempo-Neto C, Vettore AL. Expression of miR-296-5p as predictive marker for radiotherapy resistance in early-stage laryngeal carcinoma. J Transl Med. 2015;13:262.
48. Lu HJ, Jin PY, Tang Y, et al. microRNA-136 inhibits proliferation and promotes apoptosis and radiosensitivity of cervical carcinoma through the NF-κB pathway by targeting E2F1. Life Sci. 2018;199:167–178.
49. Su H, Wu Y, Fang Y, et al. MicroRNA‑301a targets WNT1 to suppress cell proliferation and migration and enhance radiosensitivity in esophageal cancer cells. Oncol Rep. 2019;41(1):599–607.
50. Wang XY, Penalva LO, Yuan H, et al. Musashi1 regulates breast tumor cell proliferation and is a prognostic indicator of poor survival. Mol Cancer. 2010;9:221.
51. Yang LY, Song GL, Zhai XQ, Wang L, Liu QL, Zhou MS. MicroRNA-331 inhibits development of gastric cancer through targeting musashi1. World J Gastrointest Oncol. 2019;11(9):705–716.
52. Gao C, Han C, Yu Q, et al. Downregulation of Msi1 suppresses the growth of human colon cancer by targeting p21cip1. Int J Oncol. 2015;46(2):732–740.
53. de Araujo PR, Gorthi A, da Silva AE, et al. Musashi1 impacts radio-resistance in glioblastoma by controlling DNA-protein kinase catalytic subunit. Am J Pathol. 2016;186(9):2271–2278.
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.