Back to Journals » Infection and Drug Resistance » Volume 11
Ertapenem non-susceptibility and independent predictors of the carbapenemase production among the Enterobacteriaceae isolates causing intra-abdominal infections in the Asia-Pacific region: results from the Study for Monitoring Antimicrobial Resistance Trends (SMART)
Authors Jean SS, Lee WS, Hsueh PR
Received 22 July 2018
Accepted for publication 29 August 2018
Published 17 October 2018 Volume 2018:11 Pages 1881—1891
DOI https://doi.org/10.2147/IDR.S181085
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Sahil Khanna
Shio-Shin Jean,1,2 Wen-Sen Lee,3,4 Po-Ren Hsueh5,6
on behalf of the SMART Asia-Pacific Group
1Department of Emergency, School of Medicine, College of Medicine, Taipei Medical University, Taipei, Taiwan; 2Department of Emergency Medicine, Department of Emergency and Critical Care Medicine, Wan Fang Hospital, Taipei Medical University, Taipei, Taiwan; 3Division of Infectious Diseases, Department of Internal Medicine, Wan Fang Hospital, Taipei Medical University, Taipei, Taiwan; 4Department of Internal Medicine, School of Medicine, College of Medicine, Taipei Medical University, Taipei, Taiwan; 5Departments of Laboratory Medicine, National Taiwan University Hospital, National Taiwan University College of Medicine, Taipei, Taiwan; 6Department of Internal Medicine, National Taiwan University Hospital, National Taiwan University College of Medicine, Taipei, Taiwan
Objectives: This study investigated the prevalence rates of carbapenemase positivity, antibiotic susceptibility, and independent predictors of carbapenemase producers among the Enterobacteriaceae isolates recovered from patients with intra-abdominal infections (IAI) in the Asia-Pacific region between 2008 and 2014.
Materials and methods: Multiplex PCR was used for the detection of specific β-lactamases, while the broth microdilution method was used to determine the minimum inhibitory concentrations (MICs) of antibiotics among the IAI-related Enterobacteriaceae isolates. We studied the abovementioned parameters in 484 ertapenem-non-susceptible (Erta-NS) isolates and explored the independent predictors of carbapenemase-producing Enterobacteriaceae (CPE) isolates.
Results: Eighty (16.5%) Erta-NS-IAI Enterobacteriaceae isolates were found to be CPE. Vietnam and the Philippines had the highest CPE prevalence rates. The IAI isolates of Enterobacter species and Klebsiella pneumoniae followed by Escherichia coli were the three major pathogens with 77.4%, 40.9%, and 11.7% Erta-NS prevalence rates, respectively. Furthermore, the highest CPE prevalence (35%) was noted among the Erta-NS-K. pneumoniae isolates. The CPE isolates harboring the blaNDM, blaKPC, or blaOXA-48-like alleles had higher imipenem MIC levels than those harboring the blaIMP alleles. Using multivariate logistic regression analysis, we concluded that Erta-NS-IAI isolates with an imipenem non-susceptible phenotype (OR, 56.4), with cefepime MIC >8 µg/mL (OR, 4.4), cultured from the peritoneal space samples (tissue or abscess; OR, 3.3), and harboring the extended-spectrum β-lactamase encoding allele (OR, 11.5) are independent predictors of CPE.
Conclusion: Imipenem non-susceptibility, cefepime MIC >8 μg/mL, and the peritoneal space as a culture site are independent clinical predictors of CPE among the Erta-NS-IAI Enterobacteriaceae isolates in the Asia-Pacific region.
Keywords: intra-abdominal infection, antimicrobial susceptibility, ertapenem-non-susceptible, carbapenemase-producing Enterobacteriaceae
Introduction
Infections with septicemia usually pose a threat to the survival of many inpatients. Among the diverse infection categories, intra-abdominal infections (IAI), mostly caused by Enterobacteriaceae species,1 are diseases with high fatality risks.2,3 In the last decade, resistance to carbapenem drug(s) among Enterobacteriaceae isolates has been gradually growing to be a critical concern for hospitalized patients worldwide.4–12 An Asia-Pacific IAI study conducted prior to 2010 by Sheng et al13 showed that the carbapenemase-producing Enterobacteriaceae (CPE) isolates accounted for up to 11.0% of the Enterobacteriaceae isolates having positive phenotypes for extended-spectrum β-lactamase (ESBL) production or ertapenem non-susceptibility (minimum inhibitory concentration [MIC] of ertapenem ≥1 µg/mL). In that study, the CPE predominantly harbored the alleles encoding New Delhi metallo-β-lactamases (NDM) and were mostly identified in India.13 However, this differs from the data in another recent study on Asia-Pacific IAI Enterobacteriaceae.14 We observed the emergence of NDM producing Enterobacteriaceae in many other Asian countries and Australia, as this worrisome situation challenges infection control.
Among all carbapenem agents, ertapenem is most vulnerable to hydrolysis of the advanced β-lactamases (including ESBL plus AmpC or most carbapenemases) in carbapenem-resistant Enterobacteriaceae (CRE) isolates. Therefore, non-susceptibility to ertapenem is deemed a sensitive indicator for the initial screening for potential carbapenemase production in the Enterobacteriaceae isolates.15 However, when compared to the high case fatality rates (44%–67%) in patients with septicemia due to CRE (producers of Klebsiella pneumoniae carbapenemase [KPC]) showing in vitro non-susceptibility to imipenem,10,16 patients infected with Enterobacteriaceae isolates having ertapenem resistance (MIC >2 µg/mL, in accordance with the criteria in 2009) but relative susceptibility to imipenem correlated with lower mortality rates (10%–31%).17,18 Therefore, the infection entities related to ertapenem-non-susceptible (Erta-NS) Enterobacteriaceae with or without susceptibility to imipenem may have different clinical outcomes. In addition, in a study by Tamma et al,19 the MIC values for imipenem and meropenem in the subgroup of CPE isolates (>90% were KPC producers) were significantly higher than those of the non-CPE subgroup. Furthermore, the overall 14-day mortality rate was four-fold higher in patients with CPE bacteremia than that seen in the non-CPE group. Consequently, discrimination of clinical infections caused by carbapenem-NS Enterobacteriaceae isolates by non-CPE and CPE is useful in predicting the outcomes of hospitalized patients. Using our existing database documenting the demographic factors, abdominal sampling sites for culture, health care settings for sample collection, as well as the detailed in vitro susceptibility testing results on bacterial isolates from Asia-Pacific IAI patients, we attempted to identify the independent predictors of CPE among the clinical Erta-NS-IAI Enterobacteriaceae isolates that produced important plasmidic β-lactamase(s) (including AmpC, ESBL, and carbapenemases). In addition, because new susceptibility break points of cefepime against Enterobacteriaceae species were proposed in Clinical and Laboratory Standards Institute (CLSI),20 we used different MIC levels for cefepime (8 and 16 µg/mL) to explore the independent predictors of CPE among the Erta-NS-IAI Enterobacteriaceae strains.
Patients and methods
Ethical considerations
In this survey, all Enterobacteriaceae isolates were collected as part of the routine hospital laboratory procedure and were not specially isolated for this study from the participating hospitals. This study was approved by the Institutional Review Boards and Ethical Committees of the participating centers, including the National Taiwan University Hospital, Taipei, Taiwan (NTUH 9561709108). The Ethical Committees waived the need for informed consent because limited private health information was collected and this research involved minimal risk to the subjects.
Study countries, isolates, and definitions of demographic variables
Thirty-seven medical centers from 12 countries/regions across the Asia-Pacific region participated in the Study for Monitoring Antimicrobial Resistance Trends (SMART) program during 2008–2014,14 which included Australia (n=5), the Hong Kong Special Administrative Region of China (n=2), Japan (n=3), Kazakhstan (n=1), Malaysia (n=2), New Zealand (n=4), Singapore (n=2), South Korea (n=2), Taiwan (n=8), Thailand (n=2), the Philippines (n=2), and Vietnam (n=4). Isolates of Enterobacteriaceae that were cultured from the first clinical samples of respective patients with IAI were obtained. Clinical samples from intra-abdominal sites related to IAI comprised tissue, fluid, or deep wound cultures obtained intraoperatively, or fluid obtained from percutaneous aspiration of abscesses. Duplicate isolates (the same species from the same patient within 30 days of the first positive culture) were excluded. Aside from the gender of IAI patients and the hospital settings where the IAI isolates were collected (intensive care units [ICUs] vs non-ICU, medical general ward vs surgical general ward), we also classified the ages of patients into three subgroups for analysis, including pediatric (age ≤14 years), adult (age between 15 and 64 years), and geriatric (age ≥65 years) subgroups.
Antimicrobial susceptibility testing and molecular detection of ESBLs, AmpC β-lactamases, and carbapenemases
Antibiotic susceptibility testing (MIC of the following antibiotics: cefoxitin, ceftriaxone, piperacillin/tazobactam, cefepime, ertapenem, imipenem, amikacin, ciprofloxacin, and levofloxacin) determined by the broth microdilution method, quality control testing, and molecular analyses (by multiplex PCR) for all Enterobacteriaceae isolates were performed at the Central Laboratory (International Health Management Associates, Inc., Schaumburg, IL, USA). Antibiotic susceptibility results were interpreted according to the MIC break points recommended by the CLSI 2015.20 In this study, we excluded the IAI Enterobacteriaceae strains from the analysis if they did not harbor β-lactamase encoding genes based on the multiplex PCR results (see below).
Multiplex PCR was used to detect genes encoding ESBLs, AmpC β-lactamase, and carbapenemases in all isolates with ertapenem MIC >0.5 µg/mL. Whole genomic DNA of the isolates was extracted using the QIAamp DNA Mini kit and the QIAcube instrument (Qiagen NV, Venlo, the Netherlands) from the colonies grown overnight in blood agar plates (Thermo Fisher Scientific, Waltham, MA, USA). Specific primers for detection of the ESBL alleles (blaCTX-M, blaTEM, blaSHV, blaVEB, blaGES, and blaPER), plasmid-borne AmpC genes (blaACC, blaCMY, blaMOX, blaFOX, blaDHA, blaACT, and blaMIR), and carbapenemase encoding genes (including blaSPM, blaGIM, blaKPC, blaVIM, blaNDM, blaIMP, and blaOXA-48-like) were used as previously described.21 If the isolates tested positive for the abovementioned allele(s) encoding carbapenemase, they were categorized as CPE.
Statistical analysis
Continuous variables were presented as mean ± SD or median with an IQR and were compared using Student’s t-test or Wilcoxon’s rank sum test for the two indicated groups depending on the normality of distribution. In addition, the Kruskal–Wallis test was employed for more than two subsets if the assumption of normality was invalid. By contrast, the categorical variables were expressed as absolute numbers and their respective percentages in a subgroup were analyzed and evaluated for differences by using Pearson chi-squared test or Fisher’s exact test as appropriate. To explore the independent predictors of CPE among the Erta-NS-IAI isolates, all variables with P<0.10 between different subgroups in the univariate analysis were included in a multivariate logistic regression model with a backward conditional method. Overall goodness of fit was analyzed by the Hosmer and Lemeshow test with Nagelkerke’s R-square. Data with a P-value <0.05 were considered statistically significant. All the tests were two-tailed and performed analyses using the SPSS statistical software for Windows (version 17.0; SPSS Inc., Chicago, IL, USA).
Results
Distribution of IAI isolates, including those with an Erta-NS phenotype
In this Asia-Pacific Enterobacteriaceae survey from 2008 through 2014, a total of 1,564 isolates harboring alleles encoding β-lactamase(s) were collected from abdomen. There were 484 (30.9%) IAI Enterobacteriaceae isolates with in vitro non-susceptibility to ertapenem. These 484 IAI Enterobacteriaceae isolates accounted for ~2.6% of the overall 18,689 Asia-Pacific IAI Enterobacteriaceae isolates over the 7-year period. Of note, the isolates of Enterobacter species (n=223), followed by K. pneumoniae (n=140) and Escherichia coli (n=98) accounted for the majority (95.2%) of the Erta-NS-IAI isolates. The remainder were 23 Erta-NS isolates that comprised Citrobacter species (n=12), Serratia species (n=8), Klebsiella oxytoca (n=1), Cronobacter sakazakii (n=1), and Pantoea agglomerans (n=1). Among the Erta-NS-IAI isolates, 80 (16.5%) isolates were confirmed to harbor genes encoding carbapenemases, while the remaining 404 Erta-NS isolates did not. All IAI-CPE isolates harbored not more than one carbapenemase encoding gene.
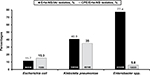
High Erta-NS prevalence rates among the IAI isolates of K. pneumoniae as well as Enterobacter species (40.9% and 77.4%, respectively), and a notably higher CPE prevalence rate (35%) among the Erta-NS-K. pneumoniae isolates (the NDM producers predominantly, followed by the imipenemase [IMP] producers) were noted relative to the Erta-NS group of E. coli and Enterobacter species (P<0.001; Figure 1). In addition, the NDM producers were also the main CPE among the isolates of E. coli and Enterobacter spp.
Distribution of Erta-NS-IAI and CPE-IAI isolates collected by geographic regions, patients’ age, and hospital settings
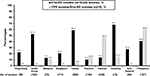
Except for Japan and Kazakhstan with <10 Erta-NS-IAI isolates (Figure 2), the highest Erta-NS prevalence rates were observed in samples obtained from Australia, Taiwan, and South Korea, followed by the Philippines, New Zealand, the Hong Kong region, Singapore, and Vietnam during the 2008–2014 period. In stark contrast, samples from the Philippines and Vietnam had the highest CPE prevalence rates (58.9% and 45.8%) among their Erta-NS-IAI isolates, respectively. Among the CPE isolates, IMP-26 producers (n=14) and NDM-1 producers (n=12) accounted for the majority of the CPE isolates in the Philippines, while NDM-1 producers (n=16) followed by OXA-48-like (n=7) dominated the CPE isolates collected from Vietnam.
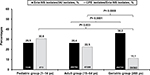
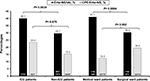
When the IAI patients were stratified by age, a trend toward higher Erta-NS prevalence was observed in the IAI isolates from the geriatric (≥65 years old) subgroup (P<0.0001). Nevertheless, the pediatric subgroup had the highest CPE prevalence among the Erta-NS isolates followed by the adult and geriatric subgroups (Figure 3). Additionally, significantly higher Erta-NS prevalence rates among the IAI isolates identified by culturing samples from patients hospitalized in the ICU (40.1%) or medical general ward (41.1%) were noted when compared with that of the non-ICU (29.7%) or surgical general ward (30.4%) settings, respectively (Figure 4).
In vitro susceptibility and MIC distributions of the Erta-NS- and CPE-IAI isolates
The majority (>80%) of the Erta-NS-non-CPE isolates exhibited in vitro susceptibility to imipenem. Furthermore, a significantly lower rate of non-susceptibility to imipenem was observed in the non-CP subgroup than in the CP subgroup among the Erta-NS-IAI isolates of Enterobacter species (4.3% vs 61.5%; P<0.001). A similar scenario was also detected among the isolates of Erta-NS-E. coli as well as Erta-NS-K. pneumoniae. Geographically, among the Erta-NS-IAI isolates, 56 isolates collected from the Philippines as well as 59 isolates collected from Vietnam had the highest MICs (average in distributions) for imipenem (median [IQR] MIC value, 2 [1–8] and 1 [0.25–8] µg/mL; the MIC50 and MIC90 values of imipenem, 2 and 32 µg/mL, as well as 1 and 32 µg/mL, respectively) when the imipenem MIC data of all of the 10 countries were evaluated by the Kruskal–Wallis test. Nevertheless, the Erta-NS-IAI isolates from New Zealand (n=19), Taiwan (n=166), Australia (n=49), and Thailand (n=16) had relatively higher imipenem MIC ranges (overall median [IQR] MIC value, 0.5 [0.5–1.0] µg/mL, with a MIC50 and MIC90 of 0.5 and 4.0 µg/mL for imipenem in 250 IAI isolates) than did the following four (Hong Kong region, South Korea, Malaysia, and Singapore) as seen from the nonparametric analysis.
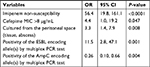
As for the CPE isolates, 95% harbored the ESBL encoding allele(s). The imipenem MICs (average in the distributions) among those harboring NDM-, KPC-, or OXA-48-like encoding genes were notably higher than those of the IMP producing CPE according to the Wilcoxon’s rank sum test (Table 1).
Univariate and multivariate logistic analyses of CPE in the Erta-NS-IAI isolates
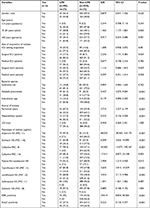
The differences among the various demographic factors, hospital settings of collection, intra-abdominal cultured sites, types of leading species, and the non-susceptibility percentages against the evaluated antimicrobial agents in the CP and non-CP subgroups of Erta-NS-IAI Enterobacteriaceae isolates, are illustrated in Table 2. As all CPE isolates had cefepime MICs >8 µg/mL, we chose the cefepime MIC of >8 µg/mL as the threshold for the univariate analysis of CPE. According to the univariate analysis, isolates of K. pneumoniae, IAI isolates from the peritoneal cavity (identified by culturing), patients aged >15 years old, IAI isolates acquired from the overall ICU, surgical ICU alone, or the medical general ward settings, and those showing non-susceptibility to the antibiotics except cefoxitin, the molecular confirmation for ESBL positive and plasmidic AmpC negative were likely to be CPE, whereas the reverse was true for the Erta-NS isolates of Enterobacter spp., or isolates identified by culturing samples from the hepatobiliary system. Furthermore, the multivariate analysis for Erta-NS-CPE revealed that the Erta-NS strains with imipenem non-susceptibility (OR, 56.4), with cefepime MIC >8 µg/mL (OR, 4.4), identified by culturing of samples from the peritoneal space (tissue or abscess; OR, 3.3), or carrying the ESBL encoding gene(s) according to PCR test (OR, 11.5) were found to be independent factors predicting CPE in the Erta-NS-IAI isolates, whereas the Erta-NS-IAI isolates that lacked the gene(s) encoding plasmid-borne AmpC enzyme(s) were an independent predictor of non-CPE (OR, 0.26; Table 3). This model has an acceptable goodness of fit (P=0.363) with a pseudo-R-square (Nagelkerke) of 0.731.
Discussion
Until recently, only few studies have specifically addressed IAI related to carbapenem-NS Enterobacteriaceae species in the PubMed literature.22,23 In addition, with a significant difference to previous surveys pertaining to carbapenem-NS Enterobacteriaceae isolates related to KPC production,8–10,19,24 the Erta-NS-CPE isolates in this Asia-Pacific IAI investigation mostly harbored the blaNDM or blaIMP alleles. Among the NDM-1 producers of IAI-CPE in this survey, 38.7% and 51.6% were collected from the Philippines and Vietnam, respectively. Therefore, the Philippines and Vietnam will likely face a huge therapeutic challenge in treating substantial CPE burden among clinical IAI isolates in the future. Of the five independent CPE predictors for the Erta-NS-IAI Enterobacteriaceae isolates (not including samples collected in ICU settings), imipenem non-susceptibility, cefepime MIC >8 µg/mL, and Erta-NS-IAI strains detected by culturing samples from the peritoneal space could be conveniently applied to identify isolates likely to be CPE in the Asia-Pacific region.
In this Asia-Pacific Enterobacteriaceae IAI study, a significantly higher CPE percentage (35%) was observed in the Erta-NS-K. pneumoniae isolates than the other two species (E. coli [15.3%] and Enterobacter species [5.8%]; P-values, 0.0007 and <0.0001, respectively; Figure 1). The diverse carbapenemases on CP K. pneumoniae isolates were dominated by NDM (42.9%; especially NDM-1), followed by IMP (38.8%; especially IMP-26) and OXA-48-like (8.2%). Spread of the mobile NDM-1 encoding genes which are primarily located on an easily transferable, 180-kb plasmid (with high horizontal transfer potential) in K. pneumoniae is facilitated by population mobility.12,25 In addition, in a CRE study reported by Gupta et al26 issuing the clinical US cases with IMP or NDM producing Enterobacteriaceae infections (reported to the Centers for Disease Control and Prevention, USA), K. pneumoniae isolates accounted for the majority (70%) of implicated Enterobacteriaceae strains. Other surveys also showed that clinical isolates of K. pneumoniae predominated the NDM producing Enterobacteriaceae strains.27,28 The findings of the above studies suggest that clinical isolates of K. pneumoniae indeed have a higher likelihood of harboring plasmid-mediated carbapenemases than the other Enterobacteriaceae spp. These results corresponded with our Erta-NS-IAI Enterobacteriaceae study as well. Nevertheless, isolates of carbapenem-resistant K. pneumoniae were surprisingly found not to be an independent predictor of CPE in the multivariate logistic regression analysis in this IAI study. The reason why IAI-Erta-NS-K. pneumoniae was not an independent CPE predictor might be related to the possibility that the CPE proportion among Erta-NS-K. pneumoniae isolates was not high enough.
Prior research into CPE infections has indicated that most (>75%) of these patients are overexposed to some antimicrobials.8,9 We did not investigate this aspect in our IAI study. Of note, this Asia-Pacific IAI Enterobacteriaceae study indicates that the Erta-NS strains isolated from the peritoneal cavity were independently predicted to be CPE. In addition, the molecular confirmation of ESBL encoding genetic determinants is another predictor of Erta-NS-CPE among the IAI-cultured isolates. With respect to these associations, we consider that extended-spectrum cephalosporin in conjunction with metronidazole is often prescribed as the first-line treatment for most mild-to-moderate IAI episodes during the initial therapy.29 Usually a prolonged duration of treatment with antimicrobials targeting the bacteria in the peritoneal space is needed, especially for patients who are too vulnerable to undergo surgery. For these debilitated patients, such a therapeutic maneuver is highly likely to predispose them to acquiring clinical CPE infections in nosocomial settings.25,30,31 In addition, Ben-David et al suggested that such a situation may simultaneously cause the originally susceptible Enterobacteriaceae isolates to acquire plasmidic ESBL encoding gene(s) from other bacteria residing in the same microbial environment.32 As for the higher Erta-NS prevalence rates among the IAI Enterobacteriaceae isolates for two age groups, whether there were significant differences in the past antimicrobial exposure30 between them also needs to be clarified.
In this Asia-Pacific IAI study, the majority of Erta-NS-non-CP Enterobacteriaceae isolates virtually displayed in vitro non-susceptibility to imipenem. A UK study conducted by Doumith et al investigated 16 Erta-NS-non-CPE isolates (including 10 K. pneumoniae and six Enterobacter species) that were mostly non-susceptible to imipenem (MICs range, 2–8 µg/mL among 11 strains). Apart from ESBL and/or AmpC production, they found that most of them lacked or had a significantly poor expression of major porin(s) encoding genes as well (OmpK35 and OmpK36 for K. pneumoniae; OmpC and OmpF for Enterobacter species).33 The isolates of Enterobacter species are known to be constitutive hyperproducers of inducible AmpC β-lactamase(s). Furthermore, in this Asia-Pacific IAI survey, ~70% of the 223 isolates of Erta-NS Enterobacter species (more than three-fourths of which were Enterobacter cloacae; of note, the overall imipenem non-susceptibility rate was only 7.6%) were proven to be plasmidic AmpC producers, whereas only 16% and 5.8% of them were ESBL producers and CPE, respectively. In addition, among the Erta-NS-non-CP K. pneumoniae isolates in our IAI survey, the prevalence rate of ESBL plus concomitantly plasmidic AmpC was 44.0% (not shown in the “Results” section), disproportionally higher than the imipenem non-susceptibility rate (7.7%; P<0.01). These results imply that most Erta-NS-non-CPE IAI isolates in the Asia-Pacific region likely have intact outer-membrane porins. The above mechanisms might plausibly explain why imipenem non-susceptibility (MIC >2 µg/mL) is identified as an independent predictor of CPE among these Asia-Pacific Erta-NS-IAI Enterobacteriaceae isolates.
There are some limitations to this study. Firstly, although we designed the study to collect clinical IAI isolates from many Asia-Pacific countries for 7 years, the total number of Erta-NS-CPE isolates is relatively small (n=80). Secondly, with respect to the detection of accurate variants among the diverse blaOXA-48-like alleles, the PCR test failed to differentiate them among the Enterobacteriaceae isolates. The OXA-163 hydrolyzes cephalosporins as well as aztreonam more effectively than carbapenems, while the OXA-232 enzyme also hydrolyzes the carbapenem agents less efficiently than the other OXA-48-like carbapenemases.34 Thirdly, the genetic determinants encoding carbapenemases among the Erta-NS-CPE isolates of this Asia-Pacific IAI study are much different from those in most Western countries where KPC producers are endemic.25 Consequently, the conclusions of our IAI survey cannot be generalized to other geographic regions. Fourthly, the genetic relatedness and clones of special sequence types (eg, ST131 E. coli and ST11 K. pneumoniae) of the enrolled IAI isolates in this study were not determined.25,35,36 Lastly, we did not analyze the major porin(s) encoding genes and emergence of efflux pump(s) in the Erta-NS-IAI Enterobacteriaceae isolates investigated in this study.
Conclusion
Many countries in the Asia-Pacific region have high (>20%) Erta-NS prevalence rates among the IAI isolates. The isolates of Enterobacter species followed by K. pneumoniae have higher prevalence rates of the Erta-NS phenotype (77.4%, 40.9%) than E. coli. Of note, the Erta-NS-IAI K. pneumoniae isolates have an alarmingly high prevalence rate (35%) of CPE. The Erta-NS Enterobacteriaceae isolates detected by culturing samples from the peritoneal space (fluid, tissue), with in vitro cefepime MIC >8 µg/mL, with imipenem non-susceptibility, or carrying the ESBL encoding gene(s) proven by the PCR test, are independent predictors of CPE.
Acknowledgments
We thank all the investigators in the Asia-Pacific region for their participation in the SMART program. This study was supported by Merck Sharp & Dohme.
Other investigators from the SMART Asia-Pacific Group include Tony Korman (Monash Medical Centre, Clayton, VIC, Australia), Justin Ellem (Westmead Hospital, Westmead, NSW, Australia), Narelle George (Royal Brisbane Hospital, Brisbane, QLD, Australia), Geoffrey Coombs (Royal Perth Hospital, Perth, WA, Australia), Thomas Ling (Prince of Wales Hospital, Shatin, New Territories, Hong Kong), Owen Tsang (Princess Margaret Hospital, Hong Kong), V Balaji (Christian Medical College, Vellore, India), Hiroshige Mikamo (Aichi Medical University Hospital, Nagakute, Japan), Shinya Kusachi (Toho University, Tokyo, Japan), Tetsu Mizutani (Osaka Police Hospital, Osaka, Japan), Min-Ja Kim (Korea University Anam Hospital, Seoul, South Korea), In-Gyu Bae (Gyeongsang National University Hospital, Jinju, South Korea), Nurulhuda Binti Umur (Hospital Kuala Lumpur, Kuala Lumpur, Malaysia), Datin Ganeswrie Rajasekaram (Hospital Sultanah Aminah, Johor Bahru, Malaysia), Susan Taylor (Middlemore Hospital at Counties Manukau District, Otahuhu, New Zealand), Sally Roberts (Auckland City Hospital, Grafton, New Zealand), Koen van der Werff (Wellington Hospital, Wellington, New Zealand), Dragana Drinkovic (North Shore Hospital, Auckland, New Zealand), Evelina Lagamayo (St Luke’s Medical Centre, Quezon, the Philippines), Myrna Mendoza (Philippine General Hospital, Manila, the Philippines), Thean Yen Tan (Changi General Hospital, Singapore), Prabha Krishnan (Tan Tock Seng Hospital, Singapore), Ellie Wang (National Cheng Kung University Hospital, Tainan, Taiwan), Po-Liang Lu (Kaohsiung Medical University Hospital, Kaohsiung, Taiwan), Chun-Eng Liu (Changhua Christian Hospital, Changhua, Taiwan), Kenneth Yin-Ching Chuang (Chi-Mei Medical Centre, Tainan, Taiwan), Kwok-Woon Yu (Taipei Veterans General Hospital, Taipei, Taiwan), Yao-Shen Chen (Kaohsiung Veterans General Hospital, Kaohsiung, Taiwan), Min-Chi Lu (Chung Shan Medical University Hospital, Taichung, Taiwan), Siripen Panthuwong (Songklanakarin Hospital, Songkhla, Thailand), Pattarachai Kiratisin (Siriraj Hospital, Bangkok-Noi, Thailand), Nguyen Tran My Phoung (Binh Dan Hospital, Ho Chi Minh City, Vietnam), Doan Mai Phuong (Bach Mai Hospital, Hanoi, Vietnam), Nguyen Thi Van (Benh Vien Viet Duc Hospital, Hanoi, Vietnam), and Tran Thi Thanh Nga (Cho Ray Hospital, Ho Chi Minh City, Vietnam).
Disclosure
Dr Hsueh received a grant from Merck Sharp & Dohme. The authors report no other conflicts of interest in this work.
References
Ko WC, Hsueh PR. Increasing extended-spectrum beta-lactamase production and quinolone resistance among Gram-negative bacilli causing intra-abdominal infections in the Asia/Pacific region: data from the Smart Study 2002–2006. J Infect. 2009;59(2):95–103. | ||
Sartelli M, Catena F, Ansaloni L, et al. Complicated intra-abdominal infections observational European study (CIAO Study). World J Emerg Surg. 2011;6(1):40. | ||
Tan TY, Ong M, Cheng Y, Ng LSY. Hypermucoviscosity, rmpA, and aerobactin are associated with community-acquired Klebsiella pneumoniae bacteremic isolates causing liver abscess in Singapore. J Microbiol Immunol Infect. Epub 2017 Jul 14. | ||
Chiu CC, Lin TC, Wu RX, et al. Etiologies of community-onset urinary tract infections requiring hospitalization and antimicrobial susceptibilities of causative microorganisms. J Microbiol Immunol Infect. 2017;50(6):879–885. | ||
Ku YH, Chen CC, Lee MF, Chuang YC, Tang HJ, Yu WL. Comparison of synergism between colistin, fosfomycin and tigecycline against extended-spectrum β-lactamase-producing Klebsiella pneumoniae isolates or with carbapenem resistance. J Microbiol Immunol Infect. 2017;50(6):931–939. | ||
Tseng SP, Wang SF, Ma L, et al. The plasmid-mediated fosfomycin resistance determinants and synergy of fosfomycin and meropenem in carbapenem-resistant Klebsiella pneumoniae isolates in Taiwan. J Microbiol Immunol Infect. 2017;50(5):653–661. | ||
Jao YT, Wang WH, Wang A, Siu LK, Lu PL. First report of OXA-48 carbapenemase-producing Escherichia coli in Taiwan. J Microbiol Immunol Infect. 2017;50(3):403–404. | ||
Tumbarello M, Trecarichi EM, De Rosa FG, et al. Infections caused by KPC-producing Klebsiella pneumoniae: differences in therapy and mortality in a multicentre study. J Antimicrob Chemother. 2015;70(7):2133–2143. | ||
Qureshi ZA, Paterson DL, Potoski BA, et al. Treatment outcome of bacteremia due to KPC-producing Klebsiella pneumoniae: superiority of combination antimicrobial regimens. Antimicrob Agents Chemother. 2012;56(4):2108–2113. | ||
Fraenkel-Wandel Y, Raveh-Brawer D, Wiener-Well Y, Yinnon AM, Assous MV. Mortality due to blaKPC Klebsiella pneumoniae bacteraemia. J Antimicrob Chemother. 2016;71(4):1083–1087. | ||
Woodford N, Tierno PM Jr, Young K, et al. Outbreak of Klebsiella pneumoniae producing a new carbapenem-hydrolyzing class A beta-lactamase, KPC-3, in a New York Medical Center. Antimicrob Agents Chemother. 2004;48(12):4793–4799. | ||
Rolain JM, Parola P, Cornaglia G. New Delhi metallo-beta-lactamase (NDM-1): towards a new pandemia? Clin Microbiol Infect. 2010;16(12):1699–1701. | ||
Sheng WH, Badal RE, Hsueh PR; SMART Program. Distribution of extended-spectrum β-lactamases, AmpC β-lactamases, and carbapenemases among Enterobacteriaceae isolates causing intra-abdominal infections in the Asia-Pacific region: results of the study for Monitoring Antimicrobial Resistance Trends (SMART). Antimicrob Agents Chemother. 2013;57(7):2981–2988. | ||
Jean SS, Hsueh PR; SMART Asia-Pacific Group. Distribution of ESBLs, AmpC β-lactamases and carbapenemases among Enterobacteriaceae isolates causing intra-abdominal and urinary tract infections in the Asia-Pacific region during 2008-14: results from the Study for Monitoring Antimicrobial Resistance Trends (SMART). J Antimicrob Chemother. 2017;72(1):166–171. | ||
Anderson KF, Lonsway DR, Rasheed JK, et al. Evaluation of methods to identify the Klebsiella pneumoniae carbapenemase in Enterobacteriaceae. J Clin Microbiol. 2007;45(8):2723–2725. | ||
Zarkotou O, Pournaras S, Voulgari E, et al. Risk factors and outcomes associated with acquisition of colistin-resistant KPC-producing Klebsiella pneumoniae: a matched case-control study. J Clin Microbiol. 2010;48(6):2271–2274. | ||
Hyle EP, Ferraro MJ, Silver M, Lee H, Hooper DC. Ertapenem-resistant Enterobacteriaceae: risk factors for acquisition and outcomes. Infect Control Hosp Epidemiol. 2010;31(12):1242–1249. | ||
Teo J, Cai Y, Tang S, et al. Risk factors, molecular epidemiology and outcomes of ertapenem-resistant, carbapenem-susceptible Enterobacteriaceae: a case-case-control study. PLoS One. 2012;7(3):e34254. | ||
Tamma PD, Goodman KE, Harris AD, et al. Comparing the outcomes of patients with carbapenemase-producing and non-carbapenemase-producing carbapenem-resistant Enterobacteriaceae bacteremia. Clin Infect Dis. 2017;64(3):257–264. | ||
Clinical and Laboratory Standards Institute (CLSI). Performance Standards for Antimicrobial Susceptibility Testing: Twenty-fifth Informational Supplement CLSI document M100-S25. Wayne, PA: CLSI; 2015. | ||
Lob SH, Kazmierczak KM, Badal RE, et al. Trends in susceptibility of Escherichia coli from intra-abdominal infections to ertapenem and comparators in the United States according to data from the SMART program, 2009 to 2013. Antimicrob Agents Chemother. 2015;59(6):3606–3610. | ||
Di Carlo P, Pantuso G, Cusimano A, et al. Two cases of monomicrobial intraabdominal abscesses due to KPC--3 Klebsiella pneumoniae ST258 clone. BMC Gastroenterol. 2011;11:103. | ||
Oteo J, Domingo-García D, Fernández-Romero S, et al. Abdominal abscess due to NDM-1-producing Klebsiella pneumoniae in Spain. J Med Microbiol. 2012;61(Pt 6):864–867. | ||
Weisenberg SA, Morgan DJ, Espinal-Witter R, Larone DH. Clinical outcomes of patients with Klebsiella pneumoniae carbapenemase-producing K. pneumoniae after treatment with imipenem or meropenem. Diagn Microbiol Infect Dis. 2009;64(2):233–235. | ||
Jean SS, Lee WS, Lam C, Hsu CW, Chen RJ, Hsueh PR. Carbapenemase-producing Gram-negative bacteria: current epidemics, antimicrobial susceptibility and treatment options. Future Microbiol. 2015;10(3):407–425. | ||
Gupta N, Limbago BM, Patel JB, Kallen AJ. Carbapenem-resistant Enterobacteriaceae: epidemiology and prevention. Clin Infect Dis. 2011;53(1):60–67. | ||
Kumarasamy KK, Toleman MA, Walsh TR, et al. Emergence of a new antibiotic resistance mechanism in India, Pakistan, and the UK: a molecular, biological, and epidemiological study. Lancet Infect Dis. 2010;10(9):597–602. | ||
Jain A, Hopkins KL, Turton J, et al. NDM carbapenemases in the United Kingdom: an analysis of the first 250 cases. J Antimicrob Chemother. 2014;69(7):1777–1784. | ||
Solomkin JS, Mazuski JE, Bradley JS, et al. Diagnosis and management of complicated intra-abdominal infection in adults and children: guidelines by the Surgical Infection Society and the Infectious Diseases Society of America. Clin Infect Dis. 2010;50(2):133–164. | ||
Bratu S, Landman D, Haag R, et al. Rapid spread of carbapenem-resistant Klebsiella pneumoniae in New York City: a new threat to our antibiotic armamentarium. Arch Intern Med. 2005;165(12):1430–1435. | ||
Tzouvelekis LS, Markogiannakis A, Psichogiou M, Tassios PT, Daikos GL. Carbapenemases in Klebsiella pneumoniae and other Enterobacteriaceae: an evolving crisis of global dimensions. Clin Microbiol Rev. 2012;25(4):682–707. | ||
Ben-David D, Kordevani R, Keller N, et al. Outcome of carbapenem resistant Klebsiella pneumoniae bloodstream infections. Clin Microbiol Infect. 2012;18(1):54–60. | ||
Doumith M, Ellington MJ, Livermore DM, Woodford N. Molecular mechanisms disrupting porin expression in ertapenem-resistant Klebsiella and Enterobacter spp. clinical isolates from the UK. J Antimicrob Chemother. 2009;63(4):659–667. | ||
Bakthavatchalam YD, Anandan S, Veeraraghavan B. Laboratory detection and clinical implication of oxacillinase-48 like carbapenemase: the hidden threat. J Glob Infect Dis. 2016;8(1):41–50. | ||
Netikul T, Sidjabat HE, Paterson DL, et al. Characterization of an IncN2-type blaNDM-1-carrying plasmid in Escherichia coli ST131 and Klebsiella pneumoniae ST11 and ST15 isolates in Thailand. J Antimicrob Chemother. 2014;69(11):3161–3163. | ||
Ma L, Siu LK, Lin JC, et al. Updated molecular epidemiology of carbapenem-non-susceptible Escherichia coli in Taiwan: first identification of KPC-2 or NDM-1-producing E. coli in Taiwan. BMC Infect Dis. 2013;13:599. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.