Back to Journals » Clinical Epidemiology » Volume 12
Epidemiology of Human Adenoviruses: A 20-Year Retrospective Observational Study in Hospitalized Patients in Bern, Switzerland
Authors Akello JO , Kamgang R, Barbani MT, Suter-Riniker F, Leib SL , Ramette A
Received 17 January 2020
Accepted for publication 20 March 2020
Published 5 April 2020 Volume 2020:12 Pages 353—366
DOI https://doi.org/10.2147/CLEP.S246352
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Henrik Sørensen
Joyce Odeke Akello,1– 3 Richard Kamgang,1 Maria Teresa Barbani,1 Franziska Suter-Riniker,1 Stephen L Leib,1 Alban Ramette1
1Institute for Infectious Diseases, University of Bern, Bern, Switzerland; 2Biology Division, Spiez Laboratory, Swiss Federal Office for Civil Protection, Spiez, Switzerland; 3Graduate School for Cellular and Biomedical Sciences, University of Bern, Bern, Switzerland
Correspondence: Alban Ramette
Institute for Infectious Diseases, University of Bern, Friedbühlstrasse 51, Bern 3001, Switzerland
Tel +41 31 632 9540
Email [email protected]
Background: Human adenovirus (HAdV) is an important pathogen seen in clinical practice. Long-term studies may help better understand epidemiological trends and changes in circulating genotypes over time.
Purpose: Using a large biobank of samples from hospitalized, adenovirus-positive patients over a 20-year period, we aimed to analyze long-term epidemiological trends and genotypic relatedness among circulating HAdV strains.
Methods: Based on samples from hospitalized patients confirmed to be HAdV positive in Bern, Switzerland, from 1998 to 2017, and on their associated demographic and clinical data, we identified epidemiological trends and risk factors associated with HAdV infection. HAdV genotyping was performed by PCR amplification and sequencing of the hypervariable hexon gene. The obtained sequences were phylogenetically compared with sequences from international HAdV strains.
Results: HAdV was identified in 1302 samples tested. Cases of HAdV infection were reported throughout the years with no clear seasonality. Upper respiratory tract samples, conjunctivitis swabs, and stool had the highest positivity rate (56.2%, 18.7%, and 14.2% of the cases, respectively). HAdV infection was highest among children ≤ 4 years old. Increased number of HAdV cases were observed in years 2009 (n = 110) and 2010 (n =112). HAdV8 was the predominant genotype among patients older than 20 years, and was mostly associated with ophthalmic infection. Predominant genotypes among children ≤ 4 years old were HAdV1, HAdV2, and HAdV3, which were mostly associated with respiratory tract infections. Recurring peaks of increased HAdV cases were evidenced every 4 years among children ≤ 4 years old.
Conclusion: Our study gives novel insights on long-term epidemiological trends and phylogenetic relatedness among circulating HAdV strains in Switzerland, country in which little data on HAdV prevalence and diversity was so far available.
Keywords: adenoviruses, human, molecular epidemiology, clinical infections, genotype
Introduction
Human adenovirus (HAdV) is one of the most important viruses seen in clinical and public health today. This DNA virus is associated with a wide range of illnesses ranging from benign colds to more serious conditions including gastroenteritis, acute respiratory infections, conjunctivitis, haemorrhagic cystitis, meningoencephalitis,1,2 and can cause severe disease in both immunocompetent3 and immunocompromised hosts, infections that sometimes result in death.2,4–7 HAdV is estimated to account for about 2% to 5% of all respiratory infections worldwide. This virus often causes epidemics and outbreaks, especially when associated with either ophthalmic infections or respiratory tract infections.8–10 It is also known to be causative of emerging viral respiratory tract infections in humans,8,11-14 along with Middle East respiratory syndrome coronavirus, H7N9 influenza A virus or swine-like influenza H3N2 variant virus.
HAdV comprises over 100 genotypes recognized by the Human Adenovirus Working Group, July 2019, Update (http://hadvwg.gmu.edu) classified into seven species (HAdV-A to HAdV-G). In clinical diagnostics laboratories, molecular typing of HAdV positive samples is not routinely performed, unless epidemiological analyses are conducted. PCR-based techniques targeting genes encoding for the hypervariable hexon, fiber, penton base or DNA polymerase, and/or sequencing of their PCR products, allow identification of HAdV species and genotypes.
The recent emergence of the HAdV14 strains in the USA15 and Europe (Ireland)4 calls for a need to fill major gaps in understanding the molecular epidemiology and disease patterns of HAdVs. The predominant HAdV genotypes detected in association with clinical presentation differ among different countries and may change over time.16 Currently in Switzerland, the knowledge on the molecular epidemiology and clinical presentation of HAdV infection is lacking. This study presents the molecular epidemiology of HAdV and associated clinical findings based on a retrospective observational study from laboratory confirmed HAdV cases collected between 1998 and 2017, at the Institute for Infectious Disease (IFIK), following referral of specimens by clinicians from the largest hospitals and other regional clinics in Bern. Since outbreaks of HAdV have been reported in various countries,10,15,17-19 we therefore hypothesized that a thorough epidemiological analysis of the Bern HAdV clinical bio-bank, including genotyping and comparison to strains involved in international outbreaks, would provide information on potential outbreaks of HAdV that occurred in the last 20 years in Switzerland and how the epidemiological situation compares to that seen elsewhere in Europe during the studied period. To our knowledge, this is the first study to report HAdV epidemiology from a large clinical dataset over a 20-year period from patients of any age with positive HAdV infection. Overall, the study sheds light on epidemiological trends, risk factors for infection, and molecular epidemiology of HAdV circulating genotypes.
Materials and Methods
Laboratory Setting, Study Design and Ethical Approval
The institute for infectious diseases (IFIK), university of Bern, is a diagnostic, teaching and research institute in Switzerland. The institute covers the entire spectrum of microbiological diagnostics and also provides university education in clinical microbiology to students studying human medicine, dentistry, pharmacy, and cell biology. Samples used in this study are part of the IFIK clinical bio-bank and consisted of previously screened patient samples referred to the IFIK for diagnostics purposes by physicians from the Inselspital and other regional clinics in Bern. The clinical bio-bank consisted of patient samples and information on HAdV cases since 1998. The results of all cases were extracted from the electronic health-care records and information such as laboratory I.D., patient I.D., demographic data (age and sex), screening tests done, and sample type, which were available for most samples. Archived patients’ samples were approved for this research by the Swiss Ethics Committees on Research involving humans (BASEC-Nr: Req-2018-00158).
Study Population
HAdV positive infection was identified as sample(s) screened by either viral culture, immuno-chromatographic rapid test, immunofluorescence (IF), PCR (quantitative or qualitative real-time PCR), or all of the mentioned methods tested positive for HAdV by the department of clinical microbiology at IFIK. In instances in which more than one sample type was obtained from the same patient in a given year, multiple patient entries were removed and only the first screened positive sample type was kept for a given year. Similarly, subsequent HAdV infections from the same patient within a given year were excluded from epidemiological analyses.
Clinical Variables and Definitions
Patient age was defined at the date of sample collection. Clinical information and severity of HAdV positive cases were categorized as respiratory tract infection, fever only, ophthalmic infection, gastrointestinal infection, immunosuppressed, severe infection, co-infection, urinary tract infection, oncological disease, other manifestations, and unknown/no clinical information provided (Table 1).
 |
Table 1 Definition of Clinical Categories Used in the Study |
Statistical Analysis
The Chi-square test was used to test for significant differences between categorical variables (test of independence). Logistic regression was used to determine significant years associated with increased HAdV cases. Data analysis was performed with the statistical R environment, version 1.0.153.
DNA Extraction, PCR, and Sequencing
Viral DNA was extracted from 200 μL of either original sample or cell culture supernatant of HAdV positive samples with the Boom extraction method20 or the NUCLISENS easyMAG (bioMérieux, Geneva, Switzerland) extractor, as per manufactures instructions, and finally eluted in 50 μL. A PCR procedure that targeted the hypervariable regions 1–7 of the hexon gene was performed, according to Sarantis and colleagues.21 M13 universal priming tails (forward, 5′-TGTAAAACGACGGCCAGT-3′; and reverse, 5′-CAGGAAACAGCTATGACC-3′) were added to the Sarantis and colleagues primers (forward, 5′- CTGATGTACTACAACAGCACTGGCAACATGGG-3′; and reverse 5′- GCGTTGCGGTGGTGGTTAAATGGGTTTACGTTGTCCAT-3′). A standard protocol of the ZymoTaq Hotstart DNA PCR was applied, consisting of 2× Zymo reaction buffer, 0.2 mM dNTP mix, 0.5 µM of each primer, 5 U/µL ZymoTaq DNA Polymerase, 10 µL of DNA template and water up to 50 µL. Thermal cycling conditions were 95°C for 10 mins, 37 cycles at 95°C for 30 secs, 51°C for 30 secs, and 72°C for 1 min, with a final extension at 72°C for 7 mins. PCR products were resolved and visualized by electrophoresis on 1% agarose gel stained with RedSafe DNA Gel Stain (Invitrogen, Switzerland). Expected amplicon sizes ranged from 602 to 630 bp. Samples with PCR product band of the expected size were purified using 1.8× Agencourt Ampure XP beads (Beckman Coulter, Nyon, Switzerland), quantified by Qubit 3.0 fluorometer (ThermoFisher Scientific, Reinach, Switzerland), diluted to concentration of 180 ng in total reaction volume of 12 µL and then submitted to Microsynth AG (Balgach, Switzerland) for Sanger sequencing. All sequence data were deposited to the European Nucleotide Archive, under project reference PRJEB36200.
Sequence and Phylogenetic Analysis
The forward and reverse sequences were trimmed and assembled using DNASTAR Lasergene SeqMan Pro version: 7.1.0 (44.1). The consensus sequence of each sample obtained was blasted against all HAdV sequences available in GenBank to identify the species and genotype. The phylogenetic trees were constructed using the Neighbor-Joining method based on evolutionary distances computed using the number of differences method. MEGA-X22 was used to generate phylogenetic trees and evaluate topologies with 1000 bootstrap replicates.
Results
Descriptive Epidemiology of HAdV Cases in Bern
We analysed samples that were referred to the IFIK for HAdV screening from January 1998 to December 2017. Of the 43,562 samples screened for HAdV from suspected patients, 34,556 (79.3%) samples were identified as unique patient cases based on patient I.D. and year of entry. Based on the laboratory confirmed results, 1302 (3.8%) were identified as positive. Of the 1302 HAdV positive cases, one case had information missing on age, gender, and clinical information, and was thus excluded from further analysis. The median age of HAdV positive patients was 3 years (range 0 months – 100 years; Table 2). A significant difference in the number of HAdV positive cases among males and females was observed (P < 0.001) with females accounting for 42.4% (552/1301) of the cases. A significant difference in the frequency of positive HAdV cases among the different age groups was also observed (P < 0.001), with the highest number of HAdV positive cases being found among young children aged ≤4 years (744; 57.2%), followed by those in the age group 20–44 years old (163; 12.5%). The least age group (59; 4.5%) was for children between 5 and 9 years. Logistic regression analyses confirmed that the overall effect of age category was statistically significant in determining positivity for HAdV infection as compared to those aged ≤4 years (P < 0.001).
 |
Table 2 Characteristics of the Study Population |
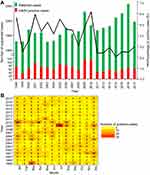
The yearly positive rate for HAdV ranged from 2.0% to 6.1%, with an average rate of 3.9%. A peak in proportion of HAdV positive cases was observed in years 1998 (5.6%), 2001 (5.9%), 2003 (5.2%), 2005 (5.5%), 2009 (4.8%), and 2010 (6.1%). An increased number of HAdV positive cases occurred in years 2009 and 2010, with 110 and 112 cases, respectively (Figure 1A). A heat map of the monthly incidence showed that HAdV appeared to occur throughout the year. Although the data showed no clear seasonality, a pattern reminiscent of HAdV outbreaks was observed particularly in January 2010 (n = 24), July 2009 (n = 33), and March 1998 (n = 24). The mean expected incidence of HAdV cases per month from 1998 to 2017 was 5.43 cases (95% CI, 1.86–12.26 cases). Therefore, HAdV incidence of ≥13 cases was defined as significantly higher number than the mean expected incidence (Figure 1B).
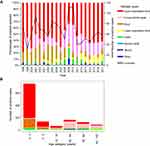
Upper respiratory tract samples were the most common positive sample types (N = 731; 56.2%), followed by conjunctivitis swab (243; 18.7%), and stool (184; 14.1%).
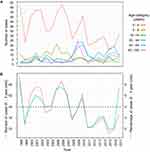
HAdV was least detected from EDTA blood and genital swab samples. Higher numbers of positive conjunctivitis swab samples were observed in years 2009 and 2010 than in any other year (Figure 2A). Among young children, upper respiratory tract and stool samples were associated with the largest number of positive cases, whilst conjunctivitis swab samples were the most positive among young adults, middle aged and elderly (Figure 2B). A peak in HAdV cases was also observed between the years 2009 and 2010, as compared to other years in the study period, across all age categories. The most unexpected finding was a peak in the number of HAdV cases reoccurring every 4 years among children ≤4 years old, with increased incidences during the years 1998, 2001, 2002, 2006, 2010, and 2014, with an average rate of 3.91% (Figure 3A and 3B).
Molecular Typing of Cases Associated with Increased Incidence in Years 2009 and 2010
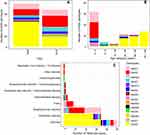
Of the 222 HAdV positive samples, 154 (69.4%) samples were available for HAdV molecular typing. Out of the 154 samples subjected to typing, HAdV genotypes were identified in 145 samples, including 63 (43.4%) samples from patients aged 0–4 years, 8 (5.5%) from 5 to 9-year-olds, 5 (3.4%) from 10 to 19-year-olds, 21 (14.5%) from 20 to 44-year-olds, 21 (14.5%) from 45 to 64-year-olds, and 27 (18.6%) from 65 to 100-year-olds. Fourteen HAdV genotypes were identified to be circulating during this time period (Figure 4A), with 10 and 11 different HAdV genotypes identified in the years 2009 and 2010, respectively. The predominant circulating types in these 2 years were HAdV8 (63; 43.4%), HAdV2 (31; 21.4%), HAdV1 (21; 14.5%), and HAdV3 (11; 7.6%), with HAdV species D genotypes accounting for the majority (48.3%) of the cases. However, in the year 2010 only, HAdV2 belonging to HAdV species C became the predominant genotype accounting for 29.9% (20/67) of the cases, followed by HAdV8 with 26.9%, and HAdV1 with 16.4% of the cases. Genotypes HAdV41, HAdV56, and HAdV64 were detected in the year 2009 only, whilst HAdV6, HAdV7, HAdV28, and HAdV53 were detected in the year 2010 only. Among samples from those aged 0–4 years (n = 63), HAdV2 (46.0%), HAdV1 (30.2%), HAdV3 (7.9%), and HAdV5 (6.3%) were the main genotypes. Among samples from 5 to 9-year-olds (n = 8), HAdV3 (50.0%) was the main genotype. HAdV8 was the predominant genotype, accounting for 40.0%, 71.4%, 85.7%, and 100% of the cases, among samples from 10–19, 20–44, 45–64, and 65–100-year-olds, respectively (Figure 4B).
HAdV cases with ophthalmic infection (n = 36), respiratory tract infection (n = 36), and with unknown/missing clinical information (n = 51), accounted for the majority of the genotyped samples (Figure 4C). Most cases presenting with ophthalmic infection (88.8%) and those with unknown/missing clinical information (74.5%) were identified to be infected with HAdV species D genotypes, particularly HAdV8. HAdV cases presenting with respiratory tract infection were associated with HAdV species C genotypes (HAdV1, HAdV2, and HAdV5), and HAdV species B genotype (HAdV3). Similarly, the former HAdV species C genotypes were also found in patients presenting with fever. Among patients presenting with gastrointestinal infection, HAdV2, HAdV3 and HAdV41 were identified. HAdV6 was identified in a case presenting with immune suppression. Among those who had more than one clinical presentation and comorbidity (ie, respiratory tract infection and gastrointestinal infection, respiratory tract infection and immune suppression, respiratory tract infection and co-infection), HAdV2 and HAdV3 were identified.
Molecular Epidemiology of HAdV Between 1998 and 2017 in Young Children
Out of the 745 identified positive samples from young children (≤4 years old) during years 1998 −2017, 198 (26.6%) were available for typing. We identified a wide variety of HAdV genotypes circulating among young children during this period (Figure 5A), with HAdV1 (n = 49), HAdV2 (n = 93), HAdV3 (n = 17), and HAdV41 (n = 13) being the most common HAdV genotypes accounting for 24.7%, 47.0%, 8.6%, and 6.6% of the cases, respectively. Most typed samples were from the years 2010 (n = 38), 2009 (n = 25), 2006 (n = 24), 2002 (n = 15), 2001 (n = 17), and 2000 (n = 15). We did not identify instances of co-infection with different genotypes of HAdV. The majority of HAdV genotypes were identified among those presenting with respiratory tract infections, fever, gastrointestinal infection, and missing clinical information (Figure 5B). HAdV2, HAdV5 and HAdV1 were identified in three dead young children (≤4 years old) in years 2006, 2012, and 2014, respectively. All three cases were associated with sudden infant death syndrome.
 |
Figure 5 (A) Distribution of circulating HAdV genotypes among children ≤4 years old from 1998 to 2017. The x-axis labels indicate isolation year, and in parentheses, the number of genotyped cases as compared to the total number of positive HAdV cases for that year. (B) Distribution of HAdV genotypes among children ≤4 years old with various clinical presentations and comorbidities. Detail on the definition of the categories is provided in Table 1. The same legend for the colour code applies for the two panels. |
Phylogenetic Analysis
The nucleotide sequences of the partial hypervariable hexon gene 1–7 from HAdV genotypes identified in this study were aligned and analysed with sequences from circulating HAdV genotypes identified in China including Taiwan, Egypt, France, Germany, Japan, Korea, Russia, Sweden, and the USA (Figure 6). The phylogenetic analysis showed that the sequences of the HAdV genotypes (HAdV1, HAdV2, HAdV3, HAdV4, HAdV5, HAdV6, HAdV7, HAdV8, HAdV28, HAdV37, HAdV41, HAdV53, HAdV56, and HAdV64) identified in the years 2009 and 2010 in this study from patients of all ages clustered with other international sequences of similar HAdV genotypes and exhibited similarity >98% at the nucleotide level. Similarly, phylogenetic analysis of the 14 HAdV genotypes identified among young children ≤4 years infected with HAdV from years 1998 to 2017 in our study clustered with international sequences of the same HAdV genotypes (Figures S1A and B).
Discussion
Our 20-year retrospective observational study from patients across all age categories underlines the need to carry epidemiological studies and molecular typing of HAdV among patients with HAdV infection. Although long-term epidemiological trends demonstrated no clear seasonality, a pattern reminiscent of recurring outbreaks was evidenced with high number of HAdV cases observed in January 2010, July 2009 and March 1998 as compared to other months or years during the study period. Genotyping of samples from years 2010 and 2009 identified HAdV8 as predominant genotype causing HAdV infection among young adults, middle-aged and elderly, and HAdV1 and HAdV2 as predominant genotypes causing HAdV infection among young children. Although HAdV infections are known to fluctuate over time, increased cases of HAdV infections were evidenced among young children every 4 years. This study is the first to report such recurring pattern. Previously, Heo and colleagues23 showed that cases of HAdV respiratory infection are common during winter and spring. A study by Yao and colleagues concluded that HAdV prevalence may be related to age and season.24 Our findings suggest that increased HAdV infection in Switzerland was related to age, but to a lesser extent to a particular season, as HAdV infections occurred throughout the year. Also when detected, HAdV was predominantly found among those aged ≤4 years. HAdV is therefore confirmed to be an important pathogen in clinical practice, mainly in the paediatric population.
The high incidence of HAdV8 associated with ophthalmic infections among young adults, middle-aged and elderly in the years 2009 and 2010 is consistent with what was seen in Germany for instance.25 Our finding is also consistent with other studies that showed HAdV8 to be a major culprit of adenovirus-associated conjunctivitis.9,26,27 In addition, the predominance of HAdV species C genotypes, particularly HAdV1 and HAdV2, found among young children is also supported by other studies.28 As young children have developing immune system, they may be more prone to infections. In the same line of evidence, most HAdV infections in immunocompromised hosts are known to be associated with HAdV species C genotypes 1 and 2.14,29 Our study focused on molecular typing of HAdV positive samples collected between 1998 and 2017 among young children ≤4 years in order to identify the genotypes associated with an increase of cases of HAdV infection among this age category every 4 years. Various genotypes were identified to be causing HAdV infections, among which HAdV1 and HAdV2 genotypes were predominantly detected in the years with reoccurring peaks. These genotypes were also observed in the non-reoccurring peaks, but to a lesser extent. These typing results give insight into circulating HAdV genotypes causing disease mainly among young children who are known to be at high risk of HAdV infection.
Although patients from all age categories may be susceptible to HAdV infections, more cases of HAdV infections in this study were observed among children ≤4 years old. HAdV species C genotypes (HAdV1, HAdV2, and HAdV5) and HAdV species B genotype (HAdV3) were the most commonly identified genotypes, indicating that young children may be particularly sensitive to these genotypes. Previous studies have shown that HAdV infections in young children may be severe, including manifestations ranging from meningitis, intussusception, subacute focal encephalitis and myocarditis, to disseminated disease, or even death.30–33 Of particular note to this study are the five patients with HAdV infection who died in the 20-year period. Four patients who died were aged ≤4 years (three males and one female) and one patient was a 6-year-old male. Our data showed that, of the three patients who died and whose samples were available for typing, one had been infected with HAdV1, one had been infected with HAdV2, and one had been infected with HAdV5. All three patients presented with sudden infant death syndrome. The case of another patient aged ≤4 years who died suffering from disseminated adenoviral infection with hepatitis, but whose sample was not available in this study for typing, was reported by Steiner and colleagues.34 Although HAdV infections are usually self-limited and mild, several studies implicating HAdV causing fatal cases have been reported.5,6,35
The association between HAdV genotypes and clinical presentations and comorbidities in infected patients have been examined in several studies: Wurzel and colleagues found that HAdV species C genotypes (HAdV1 and HAdV2) are predominant in the airways of young children and are therefore a likely etiological agent of the disease.36 Wang and colleagues showed that only HAdV B, C and E were identified among children with significant respiratory symptoms.37 Furthermore, Liu and colleagues reported that HAdV species B genotypes (HAdV3 and HAdV7) are predominant genotypes among paediatric population presenting with acute lower respiratory tract infections.38 A strong association of HAdV40 and HAdV41 with gastroenteritis is known. In our study, we noticed that, besides these genotypes, other HAdV genotypes were identified in those presenting with gastrointestinal infection (Figure 5B), a finding that is similar to what was reported by Kumthip and colleagues.39 In addition, it is known that specific HAdV species have distinct tissue tropism which may determine the clinical presentation.40
The presence of HAdV in respiratory tract and stool samples from patients not presenting with any clinical symptoms might be detected incidentally, thus making interpretation of the clinical significance of a positive result difficult. Yet, in our study, samples of the patients who had any clinical presentation associated with HAdV infection were referred for laboratory diagnostics testing to our institute upon the request of the seeing physician when an adenovirus infection was suspected. Therefore, majority of the HAdV positive, hospitalized patients in this study had an adenovirus associated clinical presentation. Persistence of HAdV in a latent state among young children has been described.41–43 Reports of asymptomatic individuals shedding virus into the stool was also described.41,44,45 In our study, 86.1% of young children ≤4 years old whose stool sample tested positive for HAdV had a clinical presentation, with gastrointestinal infection, fever, and respiratory tract infection being the most predominant clinical presentations.
The phylogenetic analysis showed that the sequences of HAdV genotypes identified in our study displayed great similarity with nucleotide sequences from HAdV genotypes isolated in other countries. Although this finding may indicate that the same circulating genotypes may be found at the global scale, we cannot exclude that the partial sequences of the hypervariable hexon gene does not allow enough resolution among genotypes. As such, targeting other regions of the HAdV genomes such as fiber, penton base, and/or DNA polymerase genes may provide further information on differences among circulating HAdV genotypes, may help clarify the HAdV genotypes and also identify recombination events.
Our study has a number of limitations. First, our reported trends likely underestimate the true burden of HAdV infections in both the hospitalized patients and in the community, as samples sent for laboratory diagnostic testing to the IFIK rely on request by physicians, when an adenovirus infection is suspected. Second, samples were tested using different diagnostic methods for HAdV and some methods are known to be less sensitive for HAdV detection than others.46 Therefore, there is a chance that this could have also affected the results of our study. Third, some of the clinical information was missing, hence restricting interpretation of trends in HAdV clinical presentation associated with circulating genotypes. In addition, clinical information was obtained from electronic clinical records provided to the IFIK diagnostic department and could not be improved in most of the cases. Fourth, several HAdV positive samples could not be genotyped due to unavailability/lack of sample material. Although the missing information was not systematic for a particular age group, sample type or year, the unavailable information might have biased our results and therefore, the reported HAdV genotypes might not be fully representative of circulating HAdV genotypes in a given year. In addition, a complete quantitative description of the main genotype (s) causing an increase in HAdV infection every 4 years among young children could not be reached due to this lack of samples.
In conclusion, our study gives insight on HAdV infections from the largest hospital in Bern and other regional clinics serving the Swiss population. Patients of various age categories were identified to be infected, with peak years in 2009 and 2010. In these 2 years, HAdV8 was identified as the predominant genotypes and was associated with HAdV infections among young adults, middle aged and the elderly, whilst HAdV2 and HAdV1 genotypes were identified to be predominantly causing HAdV infection among young children. The findings support that variation in the clinical presentation and epidemiology of HAdV genotypes exists, and may present unexpected patterns when long-term records are examined. By uncovering HAdV molecular epidemiology and its clinical presentations in our 20-year observation study, we have highlighted the existence of differences in HAdV infections among various age categories and circulating HAdV genotypes. Future work will be performed to sequence the whole genomes of representative HAdV isolates to assess the recombination potential in our strain collection.
Acknowledgments
This work was supported by the European Union’s Horizon 2020 research and innovation programme, under the Marie Skkłodowska-Curie Actions grant agreement no. 721367 (HONOURs). The authors thank the diagnostics department of Institute for Infectious Diseases, University of Bern for their contribution to this study. We would also like to thank Prof. Daniel Paris from the Swiss Tropical and Public Health Institute for useful discussions.
Disclosure
The authors report no potential conflicts of interest in this work.
References
1. Kondo M, Kojima S, Kato K, et al. Late-onset hemorrhagic cystitis after hematopoietic stem cell transplantation in children. Bone Marrow Transplant. 1998;22(10):995–998. doi:10.1038/sj.bmt.1701482
2. Lion T. Adenovirus infections in immunocompetent and immunocompromised patients. Clin Microbiol Rev. 2014;27(3):441–462. doi:10.1128/CMR.00116-13
3. Hakim FA, Tleyjeh IM. Severe adenovirus pneumonia in immunocompetent adults: a case report and review of the literature. Eur J Clin Microbiol Infect Dis. 2007;27(2):153–158. doi:10.1007/s10096-007-0416-z
4. Carr MJ, Kajon AE, Lu X, et al. Deaths associated with human adenovirus-14p1 infections, Europe, 2009–2010. Emerg Infect Dis. 2011;17(8):1402–1408. doi:10.3201/1708.101760
5. Ahmad NMW, Weinstein MP, Boruchoff SE. Life-threatening adenovirus pneumonia in an immunocompetent civilian adult. Infect Dis Clin Pract. 2005;13(1):39–41. doi:10.1097/01.idc.0000152475.55049.cd
6. Ryan MA, Gray GC, Malasig MD, et al. Two fatal cases of adenovirus-related illness in previously healthy young adults––Illinois, 2000. MMWR. 2001;50:5.
7. Schilham MW, Claas EC, van Zaane W, et al. High levels of adenovirus DNA in serum correlate with fatal outcome of adenovirus infection in children after allogeneic stem-cell transplantation. Clin Infect Dis. 2002;35(5):526–532. doi:10.1086/341770
8. Tate JE, Bunning ML, Lott L, et al. Outbreak of severe respiratory disease associated with emergent human adenovirus serotype 14 at a US air force training facility in 2007. J Infect Dis. 2010;202(2):214–222.
9. Li D, Zhou J-N, Li H, et al. An outbreak of epidemic keratoconjunctivitis caused by human adenovirus type 8 in primary school, southwest China. BMC Infect Dis. 2019;19(1):624. doi:10.1186/s12879-019-4232-8
10. Lewis PF, Schmidt M, Lu X, et al. A community-based outbreak of severe respiratory illness caused by human adenovirus serotype 14. J Infect Dis. 2009;199(10):1427–1434.
11. Barraza EM, Ludwig SL, Gaydos JC, Brundage JF. Reemergence of adenovirus type 4 acute respiratory disease in military trainees: report of an outbreak during a lapse in vaccination. J Infect Dis. 1999;179(6):1531–1533. doi:10.1086/jid.1999.179.issue-6
12. Metzgar D, Osuna M, Kajon A, et al. Abrupt emergence of diverse species B adenoviruses at US military recruit training centers. J Infect Dis. 2007;196(10):1465–1473. doi:10.1086/525005
13. Gray GC, Chorazy ML, Kondo M, Kojima S, Kato K, Matsuyama T. Human adenovirus 14a: a new epidemic threat. J Infect Dis. 2009;199(10):1413–1415.
14. Leen AM, Rooney CM. Adenovirus as an emerging pathogen in immunocompromised patients. Br J Haematol. 2005;128(2):135–144. doi:10.1111/bjh.2005.128.issue-2
15. Esposito DH, Gardner T, Schneider E, et al. Outbreak of pneumonia associated with emergent human adenovirus serotype 14–Southeast Alaska, 2008. J Infect Dis. 2010;202(2):214–222. doi:10.1086/653014
16. Lynch JP, Kajon AE. Adenovirus: epidemiology, global spread of novel serotypes, and advances in treatment and prevention. Semin Respir Crit Care Med. 2016;37(04):586–602. doi:10.1055/s-0036-1584923
17. Cassir N, Hraiech S, Nougairede A, Zandotti C, Fournier PE, Papazian L. Outbreak of adenovirus type 1 severe pneumonia in a French intensive care unit. Euro Surveill. 2014;19(39):20914. doi:10.2807/1560-7917.ES2014.19.39.20914
18. Zhao S, Wan C, Ke C, et al. Re-emergent human adenovirus genome type 7d caused an acute respiratory disease outbreak in Southern China after a twenty-one year absence. Sci Rep. 2014;4:7365. doi:10.1038/srep07365
19. James L, Vernon MO, Jones RC, et al. Outbreak of human adenovirus type 3 infection in a pediatric long-term care facility–Illinois, 2005. Clin Infect Dis. 2007;45(4):416–420. doi:10.1086/519938
20. Boom R, Sol CJ, Salimans MM, et al. Rapid and simple method for purification of nucleic acids. J Clin Microbiol. 1990;28(3):495–503. doi:10.1128/JCM.28.3.495-503.1990
21. Sarantis H, Johnson G, Brown M, et al. Comprehensive detection and serotyping of human adenoviruses by PCR and sequencing. J Clin Microbiol. 2004;42(9):3963–3969. doi:10.1128/JCM.42.9.3963-3969.2004
22. Kumar S, Stecher G, Li M, et al. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol. 2018;35(6):1547–1549. doi:10.1093/molbev/msy096
23. Heo JY, Noh JY, Jeong HW, et al. Molecular epidemiology of human adenovirus–associated febrile respiratory illness in Soldiers, South Korea(1). Emerg Infect Dis. 2018;24(7):1221–1227. doi:10.3201/eid2407.171222
24. Yao LH, Wang C, Wei T-L, et al. Human adenovirus among hospitalized children with respiratory tract infections in Beijing, China, 2017–2018. Virol J. 2019;16(1):78. doi:10.1186/s12985-019-1185-x
25. Adlhoch C, Schöneberg I, Fell G, Brandau D, Benzler J. Increasing case numbers of adenovirus conjunctivitis in Germany. Euro Surveill. 2010;15(45).
26. Hage E, Espelage W, Eckmanns T, et al. Molecular phylogeny of a novel human adenovirus type 8 strain causing a prolonged, multi-state keratoconjunctivitis epidemic in Germany. Sci Rep. 2017;7(1):40680. doi:10.1038/srep40680
27. Rutala WA, Peacock JE, Gergen MF, et al. Efficacy of hospital germicides against adenovirus 8, a common cause of epidemic keratoconjunctivitis in health care facilities. Antimicrob Agents Chemother. 2006;50(4):1419–1424. doi:10.1128/AAC.50.4.1419-1424.2006
28. Chehadeh W, Al-Adwani A, John SE, et al. Adenovirus types associated with severe respiratory diseases: A retrospective 4‐year study in Kuwait. J Med Virol. 2018;90(6):1033–1039. doi:10.1002/jmv.v90.6
29. Kajon AE, Suárez MV, Avendano LF, et al. Genome type analysis of South American adenoviruses of subgenus C collected over a 7-year period. Arch Virol. 1993;132(1):29–35. doi:10.1007/BF01309841
30. Pham TTN, Burchette JL, Hale LP. Fatal disseminated adenovirus infections in immunocompromised patients. Am J Clin Pathol. 2003;120(4):575–583. doi:10.1309/AWXDGNC5D70EN7YT
31. Toivanen AL, Valanne L, Tatlisumak T. Acute disseminated encephalomyelitis following nephropathia epidemica. Acta Neurol Scand. 2002;105(4):333–336. doi:10.1034/j.1600-0404.2002.1c168.x
32. Chou SM, Roos R, Burrell R, et al. Subacute focal adenovirus encephalitis*. J Neuropathol Exp Neuro. 1973;32(1):34–50. doi:10.1097/00005072-197301000-00003
33. Frange P, Peffault de Latour R, Arnaud C, et al. Adenoviral infection presenting as an isolated central nervous system disease without detectable viremia in two children after stem cell transplantation. J Clin Microbiol. 2011;49(6):2361–2364. doi:10.1128/JCM.00080-11
34. Steiner I, Aebi C, Ridolfi Lüthy A, et al. Fatal adenovirus hepatitis during maintenance therapy for childhood acute lymphoblastic leukemia. Pediatr Blood Cancer. 2008;50(3):647–649. doi:10.1002/(ISSN)1545-5017
35. Awosika OO, Lyons JL, Ciarlini P, et al. Fatal adenovirus encephalomyeloradiculitis in an umbilical cord stem cell transplant recipient. Neurology. 2013;80(18):1715–1717. doi:10.1212/WNL.0b013e3182904f96
36. Wurzel DF, Mackay IM, Marchant JM, et al. Adenovirus species C is associated with chronic suppurative lung diseases in children. Clin Infect Dis. 2014;59(1):34–40. doi:10.1093/cid/ciu225
37. Wang Y-F, Shen F-C, Wang S-L, et al. Molecular epidemiology and clinical manifestations of adenovirus respiratory infections in taiwanese children. Medicine (Baltimore). 2016;95(18):e3577. doi:10.1097/MD.0000000000003577
38. Liu C, Xiao Y, Zhang J, et al. Adenovirus infection in children with acute lower respiratory tract infections in Beijing, China, 2007 to 2012. BMC Infect Dis. 2015;15(1):408. doi:10.1186/s12879-015-1126-2
39. Kumthip K, Khamrin P, Ushijima H, et al. Enteric and non-enteric adenoviruses associated with acute gastroenteritis in pediatric patients in Thailand, 2011 to 2017. PLoS One. 2019;14(8):e0220263. doi:10.1371/journal.pone.0220263
40. Kosulin K. Intestinal HAdV infection: tissue specificity, persistence, and implications for antiviral therapy. Viruses. 2019;11(9):804. doi:10.3390/v11090804
41. Wold W, Horwitz M. Adenoviruses. In: Knipe DM, Howley PM, editors. Fields Virology. Philadelphia (PA): Lippincott Williams & Wilkins; 2007.
42. Kosulin K, Geiger E, Vécsei A, et al. Persistence and reactivation of human adenoviruses in the gastrointestinal tract. Clin Microbiol Infect. 2016;22(4):381 e1–381 e8. doi:10.1016/j.cmi.2015.12.013
43. Garnett CT, Talekar G, Mahr JA, et al. Latent species C adenoviruses in human tonsil tissues. J Virol. 2009;83(6):2417–2428. doi:10.1128/JVI.02392-08
44. Roy S, Calcedo R, Medina-Jaszek A, et al. Adenoviruses in lymphocytes of the human gastro-intestinal tract. PLoS One. 2011;6(9):e24859. doi:10.1371/journal.pone.0024859
45. Adrian T, Schäfer G, Cooney MK, et al. Persistent enteral infections with adenovirus types 1 and 2 in infants. Epidemiol Infect. 1988;101:503–509. doi:10.1017/S0950268800029393
46. Stroparo E, Cruz CR, Debur MDC, et al. Adenovirus respiratory infection: significant increase in diagnosis using PCR comparing with antigen detection and culture methods. Rev Inst Med Trop São Paulo. 2010;52(6):317–321. doi:10.1590/S0036-46652010000600006
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.