Back to Journals » Infection and Drug Resistance » Volume 14
Epidemiology of Community Origin Escherichia coli and Klebsiella pneumoniae Uropathogenic Strains Resistant to Antibiotics in Franceville, Gabon
Authors Mouanga Ndzime Y, Onanga R , Kassa Kassa RF, Bignoumba M , Mbehang Nguema PP , Gafou A, Lendamba RW, Mbombe Moghoa K, Bisseye C
Received 8 December 2020
Accepted for publication 6 January 2021
Published 16 February 2021 Volume 2021:14 Pages 585—594
DOI https://doi.org/10.2147/IDR.S296054
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Sahil Khanna
Yann Mouanga Ndzime,1,2 Richard Onanga,1 Roland Fabrice Kassa Kassa,1 Michelle Bignoumba,1 Pierre Philippe Mbehang Nguema,1,3 Amahani Gafou,1 Roméo Wenceslas Lendamba,1 Kelly Mbombe Moghoa,1 Cyrille Bisseye2
1Unité de Recherche et d’Analyses Médicales, Laboratoire de Bactériologie, Centre Interdisciplinaire de Recherches Médicales de Franceville, Franceville, BP 769, Gabon; 2Laboratoire de Biologie Moléculaire et Cellulaire (LABMC), Université des Sciences et Techniques de Masuku, Franceville, BP 943, Gabon; 3Institut de Recherche en Ecologie Tropicale, Centre National de la Recherche Scientifique et Technologique (CENAREST), Libreville, BP 13354, Gabon
Correspondence: Yann Mouanga Ndzime Tel +24177206217 Email [email protected]
Introduction: Urinary tract infection is one of the major causes of consultation, microbiologic exploration, intensive use of antibiotics worldwide, and the second leading cause of clinical consultation in community practice. Many bacteria play a role in the urinary tract infections etiology, including Enterobacteriaceae such as Escherichia coli (E. coli) and Klebsiella spp.
Objective: The study’s main objective was to examine the epidemiology of E. coli and Klebsiella pneumoniae (K. pneumoniae) uropathogenic strains resistant to antibiotics in Franceville.
Methodology: The study was carried out between January 2018 and June 2019 in Franceville South-East Gabon. We examined a total of 1086 cytobacteriological urine samples. The identification of E. coli and K. pneumoniae strains was carried out using the Vitek-2 compact automated system and the antibiogram with the disk diffusion method according to the European Committee on Antimicrobial Susceptibility Testing recommendations.
Results: The prevalence of urinary tract infections was 29.2% (317/1086), of which 25.1% and 4.1% were mono-infections and co-infections, respectively. The prevalence of UTIs with E. coli was 28.7% (91/317) with a predominance of isolation in women. K. pneumoniae was responsible for 16.2% (61/317) of UTIs. E. coli and K. pneumoniae Uropathogenic strains showed resistance to beta-lactams, quinolones and cotrimoxazole, whereas Nitrofurantoin, Amikacin, Imipenem and Ertapenem were the most active antibiotics against E. coli and K. pneumoniae uropathogenic strains.
Conclusion: This study showed a high prevalence of urinary tract infections with a major implication of E.coli and K. pneumoniae strains. E. coli and K. pneumoniae presented high frequency of resistance to antibiotics, highlighting the need to adapt their use accordingly at the local level.
Keywords: urinary tract infection, antibiotic resistance, E. coli, K. pneumoniae
Introduction
Urinary tract infections (UTIs) are the second most common ailment in community medical practice. Each year, around 150 million people are diagnosed worldwide with urinary tract infections costing more than 6 billion US dollars to the global economy.1 UTIs are a major global public health problem, because of the health costs they cause and the selection of multidrug-resistant strains both in hospitals and in community settings.2 These are common infections with an alarming increase in resistance to last-resort antibiotics.3 Many bacteria are involved in the etiology of urinary tract infections, including enterobacteria such as E. coli and Klebsiella spp.4 These bacteria are the most common pathogens in UTIs. However, less common agents may also cause UTIs, such as non-fermenters and fungi.5,6
Several studies conducted in Europe and the United States of America have shown a constant increase in the rate of resistance of uropathogenic bacteria to commonly prescribed antibiotics leading to reduced therapeutic efficacy.7
The main risk factor for resistance to an antibiotic is repeated previous exposure to the same antibiotic. Indeed, the misuse of an antibiotic or a class of antibiotics is the cause of development of bacterial resistance, possibly extending to other families of antibiotics because of the cross-transmission of mobile genetic elements carrying antibiotic resistance genes.8,9 These bacterial resistances develop more easily in the digestive microbiota due to a large number of bacteria (greater than 109 bacteria per gram of stool), promoting contact as well as the emergence of resistant mutants. UTIs are most often of ascending origin by contamination from the perineal flora, reflecting the digestive flora. Therefore this selection pressure has a definite clinical impact.8
Early treatment of UTIs decreases their severity rate, which, in most cases, involves antibiotic treatment prescribed empirically.10 In order to administer appropriate therapy, it is essential to identify the main bacteria commonly involved in urinary tract infection and their antibiotic resistance profile.11 In Gabon, the lack of treatment regimen standardization and clinical bacteriology laboratories make essential studies on uropathogenic germs and their susceptibility and/or resistance to antibiotics providing a local profile of bacterial resistance.
This study aimed to determine the epidemiology of Escherichia coli and Klebsiella pneumoniae uropathogenic strains isolated from patients in consultation at the Interdisciplinary Center for Medical Research of Franceville in South-East Gabon and to determine their resistance profile to common antibiotics.
Materials and Methods
Patients
The study was carried out between January 2018 and June 2019. It involved all patients from the community requesting a cyto-bacteriological examination of urine (CBEU) by the laboratory of medical analyzes, which is the only bacteriology laboratory in the city of Franceville, capital of the Haut-Ogooué province, the second-most populous province of Gabon bordering the Republic of Congo.
Patients of all ages and both sexes who presented for a CBEU were considered eligible for the study.
Sample Collection
Urine samples were preferably collected in the morning or after an absence of urination for about 4 hours, either in the laboratory or at home, under strict aseptic conditions. Urine was collected in a sterile, single-use urine container.
For children who could use the toilet on their own, urine samples were collected using a sterile adhesive collection bag under the supervision of a guardian. The urine vial was then sealed, identified, indicating the time of collection and then transported by the patient at room temperature to the laboratory. The socio-demographic and clinical data for each patient were collected through a structured questionnaire.
Culture and Identification of Bacterial Isolates
It consisted of inoculating ten (10) μL of total urine aseptically using a sterile single-use loop in the level 2 microbiological safety station. The inoculation was carried out systematically on agar media, CLED (Cystine-Lactose-Electrolytes-Deficient; Biomérieux, France), Mac Conkey (McC, Biomérieux, France), Eosin Methylene Blue (EMB, Biomérieux, France) after briefly homogenizing the urine. Urine samples were inoculated within two (2) hours of collection to avoid false positives. The inoculated media were systematically incubated aerobically in a bacteriological oven at 35°C for 18 to 24 hours. According to Kass criteria a number ≥ 105 colony forming units (CFU)/mL of urine was considered positive; a colony number < 105 CFU/mL or with more than two (2) types of bacterial colonies were considered contaminated.12
The identification of E. coli and K. pneumoniae strains was done after Gram stain, oxidase test and conventional biochemical tests (VITEK-2 automated system, Biomérieux, France). Only E. coli and K. pneumoniae strains isolated in monoinfected culture were retained in the study.
Antibiotic Sensitivity Test
All E. coli and K. pneumoniae isolates were tested for sensitivity to the following antibiotics: ampicillin, ticarcillin, amoxicillin-clavulanic acid, cefalotin, cefoxitin, cefotaxime, ceftazidime, cefepime, imipenem, ertapenem, gentamicin, tobramycin, amikacin, ofloxacin, ciprofloxacin, nalidixic acid, trimethoprim-sulfamethoxazole, nitrofurantoin according to the diffusion disc method (Kirby-Bauer) on Mueller-Hinton (MH) agar according to the recommendations of the European Committee on Antimicrobial Susceptibility Testing (EUCAST, V. 2.0 2018).13 Briefly, MH agars were seeded with a standardized suspension (0.5 McFarland) of each E. coli and K. pneumoniae isolate for a 24 hour MH agar culture. The antibiotic discs were firmly placed on the surface of the seed plates. The culture media were then incubated for 24 hours at 35°C. The diameters of antibiotics inhibition zones were interpreted according to the recommendations of the EUCAST.13 Multi-drug resistance (MDR) was defined as non-susceptibility to at least one agent in three or more antimicrobial categories.14
Statistical Analysis
All the data collected was entered in a Microsoft Excel 2013 file and then analyzed by the R software (version R × 64.3.4.3). The differences were considered significant for p <0.05.
Ethical Considerations
Informed and written consent was obtained from each adult patient prior to inclusion in the study. With regard to minors, consent has been obtained from their parents or legal guardians.
The research licence for this study was obtained from the Scientific Commission on Research Authorisations of the National Centre of Scientific and Technological Research (CENAREST) (permit 7 no. AR0033/17/MESRSFC/CENAREST/CG/CST/CSAR, dated 4 July 2017). This study was conducted in accordance with the Declaration of Helsinki.
Results
Characteristics of Patients
Of 1086 CBEUs performed, 61.7% (670/1086) were from female patients. The mean age of patients was 24.7 ± 18.9 years. More than half of these were aged 18 to 49 (53.7%). The vast majority of patients came from the city of Franceville (72.2%) and did not present any particular morbid ground (94.3%). Among these patients, 58.8% had urinary signs, and 86.8% of samples were collected mid-stream (Table 1).
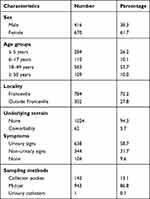 |
Table 1 Socioclinical Characteristics of the Study Patients |
Prevalence of Urinary Tract Infections Due to E. coli and K. pneumoniae
The overall prevalence of UTIs was 29.2% (317/1086), of which 25.1% (273/1086) were mono-infections while 4.1% (44/1086) were co-infections. Contaminations represented 23.2% (252/1086) of cultures, whereas 0.3% (3/1086) of CBEUs showed significant bacteriuria without leukocyturia.
The prevalence of UTIs due to E. coli was 28.7% (91/317) with 24.6% (78/317) of mono-infections and 4.1% (13/317) of co-infections. Regarding co-infections, E. coli was associated with K. pneumoniae (0.3%), K. oxytoca (0.3%), E. faecalis (0.9%), Staphylococcus spp. (1.3%), S. agalactiae (0.3%), E. aerogenes (0.3%), E. cloacae complex (0.63%) (Table 2).
 |
Table 2 Isolation of Uropathogenic Bacteria |
K. pneumoniae was found in 18.9% (60/317) of urinary tract infections with 14.2% (45/317) of the strains isolated in monoculture while 4.7% (15/317) were identified in association with K. oxytoca (0.3%), Enterococcus spp. (2.2%), Staphylococcus spp. (1.9%), or P. acidilactici (0.3%) (Table 2).
Distribution of Urinary Mono-Infections Due to E. coli and K. pneumoniae
A total of 78 E. coli urinary mono-infections were identified in the CBEUs of patients. These E.coli mono-infections were significantly more frequent in women compared to men (82% vs 18%; p <0.001) (Figure 1). The frequency of E. coli UTIs was significantly lower in patients aged 6–17 years compared to those aged ≤5 years (7.7% vs 25.6%; p = 0.03) and 18–49 years (7.7% vs 52.6%; p <0.001). In addition, E. coli was found predominantly in patients with urinary signs compared to those without urinary signs (68% vs 32%; p <0.001) (Figure 1).
 |
Figure 1 Distribution of E. coli and K. pneumoniae urinary mono-infections according to socioclinical parameters. |
Regarding the seasons, E. coli UTIs were significantly more frequent in the short rainy season compared to the short dry season (35.9% vs 17.95%; p = 0.012) or the long rainy season (35.9% vs 19.23%; p = 0.02) (Figure 1).
A total of 45 K. pneumoniae UTIs were identified in patients’ CBEUs. K. pneumoniae was found significantly more often in women than in men (66.7% vs 33.3%; p = 0.002). It was associated with patient age being significantly lower in the 6–17 years age group compared to the ≤5 years (4.4% vs 55.6%; p <0.001) and 18–49 years age groups (4.4% vs 28.9%; p = 0.002) (Figure 1). In addition, the distribution of UTIs due to K. pneumoniae was associated with seasonality and was more frequent during the short rainy season compared to the long dry season (37.8% vs 13.3%; p = 0.008) (Figure 1).
Antibiotic Resistance Assessment of E. coli and K. pneumoniae Isolates
Of the 123 strains of E. coli and K. pneumoniae isolated, the highest rates of antibiotic resistance were observed with Ampicillin (78%), Ticarcillin (75%), Trimethoprim-Sulfamethoxazole (63%), Cefalotin (47%), Amoxicillin-clavulanic acid (44%), Cefotaxime (41%), and Nalidixic acid (40%). The majority of bacterial strains isolated (56%) were multidrug resistant (MDR) (Table 3).
 |
Table 3 Resistance of E. coli and K. pneumoniae Strains to Antibiotics |
Overall, beta-lactam resistance was significantly more frequent in MDR strains compared to non-MDR strains for Ampicillin (p <0.001), Ticarcillin (p <0.001), Amoxicillin-clavulanic acid (p< 0.001), Cefalotin (p <0.001), Cefoxitin (p <0.001), Cefotaxime (p <0.001), Ceftazidime (p <0.001) and Cefepime (p = 0.026) (Table 3). This was also observed with quinolones including Nalidixic Acid (p <0.001), Ciprofloxacin (p <0.001) and Ofloxacin (p <0.001); and with aminoglycosides such as Gentamicin (p <0.001), Tobramycin (p <0.001) and Amikacin (p = 0.026) (Table 3). Furthermore, resistance to Trimethoprim-Sulfamethoxazole was more prevalent in MDR strains compared to non-MDR strains (p <0.001) (Table 3).
Among 78 E. coli strains, the highest resistance to beta-lactams was observed with Ampicillin (65%), Ticarcillin (61%), Amoxicillin-clavulanic acid (45%), Cefalotin (41%) and Cefotaxime (33%) while the lowest frequency of resistance was found with Imipenem (2%), Ertapenem (2%) and Cefepime (7%) (Table 3). The majority of E. coli strains (55%) was MDR. The MDR phenotype Ampicillin/Nalidixic acid/Trimethoprim-Sulfamethoxazole was the most prevalent (70%) in E. coli strains. Resistance to Ampicillin (p <0.001), Ticarcillin (p <0.001), Amoxicillin-clavulanic acid (p <0.001), Cefalotin (p <0.001), Cefoxitin (p = 0.025), Cefotaxime (p <0.001), Ceftazidime (p <0.001), Nalidixic Acid (p <0.001), Ciprofloxacin (p = 0.002) and Ofloxacin (p <0.001) were more common in MDR-E. coli compared to non-MDR-E. coli-isolates (Table 3). Antibiotics multidrug-resistance has been observed more frequently in women compared to men (p <0.0001).
Of 45 K. pneumoniae strains tested, the highest rates of antibiotics resistance were observed with Cefalotin (58%), Cefotaxime (56%), Amoxicillin-clavulanic acid (46%) and Ceftazidime (47%) whereas the lowest frequency of resistance was obtained with Cefepime (4%), Imipenem (4%) and Ertapenem (4%) (Table 3). K. pneumoniae strains were 57% MDR, predominantly isolated from patients ≤ 5 years (p = 0.002). The MDR phenotype Amoxicillin-clavulanic acid/Cefotaxime/Trimethoprim-Sulfamethoxazole was most common (52%) in K. pneumoniae strains. Resistance to Amoxicillin-clavulanic acid, Cefalotin, Cefoxitin, Cefotaxime, Ceftazidime, nalidixic acid and Ofloxacin was significantly more frequently observed in MDR-K. pneumoniae compared to non-MDR-K. pneumoniae isolates (Table 3).
We also compared the resistance of E. coli and K. pneumoniae strains to antibiotics. K. pneumoniae strains of were more resistant to cefotaxime compared to those of E. coli (56% vs 33%; p = 0.024) (Table 3). Regarding quinolones, E. coli strains were more resistant to nalidixic acid (51% vs 22%; p = 0.002) and to Ciprofloxacin (34% vs 13%, p = 0.01) compared to K. pneumoniae strains while resistance to Ofloxacin was similar between the two types of bacteria (39% vs 28%; p = 0.226) (Table 3).
For aminoglycosides, the resistance of strains of E. coli and K. pneumoniae was evaluated with antibiotics such as Gentamicin (25% vs 28%), Tobramycin (28% vs 24%) and Amikacin (7% vs 4%). It was similar for the three antibiotics between E. coli and K. pneumoniae (Table 3). Resistance of K. pneumoniae to Cefotaxime (80% vs 43%; p = 0.044) and Trimethoprim-Sulfamethoxazole (73% vs 53%; p <0.0001) was significantly higher in men compared to women. Among MDR isolates, we identified one strain of E. coli and K. pneumoniae resistant to 14/18 (77%) and 13/18 (72%) of antibiotics tested.
Discussion
The overreaching goal of this study was to describe the epidemiology of E. coli and K. pneumoniae uropathogenic strains isolated in Franceville in South-eastern Gabon and to determine their resistance profiles to common antibiotics.
The epidemiological profile of uropathogenic bacteria varies from one region to another.15 As a result, the knowledge of the local epidemiology, as well as its evolution is crucial in the selection of an effective first-line antibiotic therapy adapted to each region.15 Of the 1086 patients’ CBEUs, the prevalence of urinary tract infections was 29.2%. This prevalence is lower than 59.8% reported in a previous study in Cameroon.16 E. coli and K. pneumoniae represented 44.9% of the germs involved in the etiology of urinary tract infections with 28.7% and 16.2% respectively. Such prevalence is similar to 24.5% for E. coli and 18.4% for K. pneumoniae reported in an earlier study in Nigeria.17 However, the prevalence of E. coli found in UTIs is significantly lower than those between 60% and 90% reported in Chad;18 Madagascar;19 Rwanda20 and Morocco.21 We found that urinary tract infections due to E. coli mainly affected women, which is in agreement with similar studies carried out in Cameroon22 and Nigeria.23 This could be explained on the one hand by the proximity of the urethral, anal and vaginal orifices in women and on the other hand by poor hygiene practices as well as pregnancies responsible for a biased immune response favorable to microbial agents development in pregnant women.22,24 We also observed that the urinary tract infection with K. pneumoniae was more frequent among young patients (≤ 5 years) and the older patients (≥ 50 years). Our results are in agreement with those reported in a previous study.22 These two groups of patients are at higher risk.22 Indeed, in the elderly, the risk of urinary tract infection increases due to the decline of the immune system efficiency and decreased functional autonomy.8 In children aged ≤ 5 years, UTI incidence is about 5% in girls and 20% in uncircumcised boys. For febrile infants in the first two months of life and during the first 6 months, the risk of developing an UTI is 10 to 12 times higher in uncircumcised boys.25 This susceptibility of developing UTIs could be attributed to a less robust immune system.25
E. coli and K. pneumoniae isolated strains have shown strong resistance to several antibiotics. The resistance of E. coli to Ampicillin and Ticarcillin was 65% and 61%, respectively. Similar resistance rates have been reported on an outpatient basis in West Africa,26 Madagascar,19 the Central African Republic27 and Chad.18 These high resistance rates could explain why aminopenicillins and carboxypenicillins are no longer recommended for the probabilistic treatment of UTIs.28
The resistance rates of E. coli and K. pneumoniae to amoxicillin-clavulanic acid were 45% and 47%, respectively. These prevalence are higher than those ranging between 25 and 35% reported in France.8 However, this resistance rate is similar to that of 50% reported in Rwanda in patients of community origin.20 The resistance of these two bacteria to amoxicillin-clavulanic acid reported in this study is much lower than that observed in Senegal, 60.17% for E. coli and 73.18% for K. pneumoniae, respectively.29
K. pneumoniae strains have shown a higher resistance rate of 76.09% to amoxicillin-clavulanic acid in Pakistan.30 These high rates of antibiotic resistance could be explained by the broad probabilistic prescription of these antibiotics, particularly in outpatient medicine in the absence of CBEUs results.22
The resistance rate of E. coli to 3rd generation cephalosporins was 33% and 30% for cefotaxime and ceftazidime, respectively. These resistance rates were higher than those ranging from 0 to 5% observed in the Central African Republic,27 Sudan,31 and Madagascar.19 The resistance of E. coli to ceftazidime was similar to that of 29.1% obtained in Rwanda.20 On the other hand, the resistance of the strains of K. pneumoniae to 3rd generation cephalosporins was 56% for Cefotaxime and 47% for Ceftazidime. These resistance rates are in line with those reported by studies in Pakistan and Senegal, which were respectively 54.35% for 3rd generation cephalosporins30 and 57.85% for Cefotaxime.29 A very high 84% resistance rate of K. pneumoniae to Ceftazidime has been reported in India.32
This E. coli strains resistant to third-generation cephalosporins (C3G) could be explained by their acquisition of a plasmid-extended-spectrum beta-lactamase (ESBL).8,24 Previous studies carried out in Gabon reported high prevalence of bacterial strains producing ESBLs.33,34 In addition, the Global Antimicrobial Resistance Surveillance Report found that five out of six (5/6) and six out of six regions (6/6) reported E. coli and K. pneumoniae resistance > 50% to C3Gs, respectively.35 These findings were in agreement with the results of this study.
Carbapenems (Imipenem and Ertapenem) had good efficacy on E. coli and K. pneumoniae strains with resistance rates of 2% and 4%, which corroborated the results obtained by Leopold et al.36
Quinolones retained acceptable activity against K. pneumoniae strains with resistance rates of 13% to 28%. However, this resistance was higher in strains of E. coli ranging from 34% to 51%. An earlier study conducted in sub-Saharan Africa reported E. coli strain resistance to quinolones ranging between 0 and 98%.36 The first-line use of fluoroquinolones as a probabilistic treatment against these bacteria could explain the emergence of their resistance to quinolones.37 Aminoglycoside activity was conserved across all E. coli and K. pneumoniae strains with resistance rates ranging between 4% and 28%. Amikacin has been the most active antibiotic among the aminoglycosides. Resistance to aminoglycosides observed in this study was similar to that of 16.7% and 21.8% reported in Iran38 and between 8% and 14% observed in Morocco.21 The apparently preserved efficacy of aminoglycosides could be explained by their frequent parenteral administration, limiting use. In contrast, the resistance of E. coli and K. pneumoniae to Cotrimoxazole were 65% and 60%, respectively. The resistance rates to Cotrimoxazole of 80% in the Central African Republic; 69.5% in Madagascar and 55% in Morocco have been reported in previous studies.19,21,27 This high resistance of E. coli and K. pneumoniae to Cotrimoxazole could explain its exclusion in the probabilistic treatment of uncomplicated urinary tract infections. Nitrofurantoin appeared to be the most active antibiotic against E. coli and K. pneumoniae uropathogenic strains.
We found that 57% of E. coli and K. pneumoniae isolated were MDR. The presence of these MDR strains suggests that inappropriate use of antibiotics without medical prescription is taking place. This is mainly due to the lack of clinical microbiology laboratory,39,40 which tends to promote self-medication, increasing the risk of selection of resistant bacteria leading to the emergence of multidrug resistant bacteria.41,42
Conclusion
This study showed a high prevalence of urinary tract infections with great involvement of E. coli and K. pneumoniae strains. These bacteria have shown high levels of resistance to beta-lactams, quinolones and cotrimoxazole, while Nitrofurantoin, Amikacin, Imipenem and Ertapenem remain the most active antibiotics against E. coli and K. pneumoniae uropathogenic strains. The levels of bacterial resistance observed in this study requires the use of antibiotics adapted to the local epidemiology and the promotion of the antimicrobial resistance surveillance by the health authorities.
Abbreviations
CBEU, Cytobacteriological examination of urine; CFU, Colony forming unit; CLED, Cystine-Lactose-Electrolytes-Deficient; C3G, third generation cephalosporins; EMB, Eosine Methylene Blue; ESBL, Extended-spectrum-beta-lactamase; EUCAST, European Committee on Antimicrobial Susceptibility Testing; GNB, Gram negative bacilli; McC, Mac Conkey; MDR, Multidrug resistance; MH, Mueller-Hinton; UTI, Urinary tract infection.
Data Sharing Statement
Data supporting the conclusions of this study will be made available on request to the corresponding author.
Acknowledgments
The authors thank all the staff of the Medical Research Unit. We also thank the study participants for their kind participation.
Author Contributions
All authors made a significant contribution to the work reported, whether that is in the conception, study design, execution, acquisition of data, analysis and interpretation, or in drafting, revising or critically reviewing the article; gave final approval of the version to be published; have agreed on the journal to which the article has been submitted; and agree to be accountable for all aspects of the work.
Funding
There is no funding to report.
Disclosure
The authors declare that they have no conflicts of interest for this work.
References
1. Gonzalez CM, Schaeffer AJ. Treatment of urinary tract infection: what’s old, what’s new, and what works. World J Urol. 1999;17(6):372–382. doi:10.1007/s003450050163
2. Gajdacs M, Batori Z, Abrok M, Lazar A, Burian K. Characterization of resistance in gram-negative urinary isolates using existing and novel indicators of clinical relevance: a 10-year data analysis. Life. 2020;10(2):16. doi:10.3390/life10020016
3. WHO. Système mondial de surveillance de la résistance aux antimicrobiens: Manual de mise en oeuvre initiale. Zurich: World Health Organization; 2016.
4. Linhares I, Raposo T, Rodrigues A, Almeida A. Frequency and antimicrobial resistance patterns of bacteria implicated in community urinary tract infections: a ten-year surveillance study (2000–2009). BMC Infect Dis. 2013;13:19. doi:10.1186/1471-2334-13-19
5. Gajdacs M, Burian K, Terhes G. Resistance levels and epidemiology of non-fermenting gram-negative bacteria in urinary tract infections of inpatients and outpatients (RENFUTI): a 10-year epidemiological snapshot. Antibiotics. 2019;8(3).
6. Gajdacs M, Doczi I, Abrok M, Lazar A, Burian K. Epidemiology of candiduria and Candida urinary tract infections in inpatients and outpatients: results from a 10-year retrospective survey. Cent Eur J Urol. 2019;72(2):209–214. doi:10.5173/ceju.2019.1909
7. Sire JM, Nabeth P, Perrier-Gros-Claude JD, et al. Antimicrobial resistance in outpatient Escherichia coli urinary isolates in Dakar, Senegal. J Infect Dev Ctries. 2007;1(3):263–268. doi:10.3855/jidc.362
8. SPILF. Diagnostic et antibiothérapie des infections urinaires bactériennes communautaires de l’adulte. Paris: SPILF; 2015:1–43.
9. Gajdacs M, Albericio F. Antibiotic resistance: from the bench to patients. Antibiotics. 2019;8(3):129. doi:10.3390/antibiotics8030129
10. De Francesco MA, Ravizzola G, Peroni L, Negrini R, Manca N. Urinary tract infections in Brescia, Italy: etiology of uropathogens and antimicrobial resistance of common uropathogens. Med Sci Monit. 2007;13(6):BR136–44.
11. Neto JAD, Martins ACP, Silva LDMD. Community acquired urinary tract infection: etiology and bacterial susceptibility. Acta Cir Bras. 2003;18(suppl 5):33–36. doi:10.1590/S0102-86502003001200012
12. Hay AD, Birnie K, Busby J, Delaney B, Downing H, Dudley J. The diagnosis of urinary tract infection in young children (DUTY): a diagnostic prospective observational study to derive and validate a clinical algorithm for the diagnosis of urinary tract infection in children presenting to primary care with an acute illness. Health Technol Assess (Rockv). 2016;20(51):1–294. doi:10.3310/hta20510
13. SFM/EUCAST. Comité de l’Antibiogramme de la Sociéte Française de Microbiologie: recommandations. Paris; 2018. Available from: https://www.sfm-microbiologie.org/wp-content/uploads/2018/12/CASFMV2_SEPTEMBRE2018.pdf.
14. Magiorakos AP, Srinivasan A, Carey RB, et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect. 2012;18(3):268–281. doi:10.1111/j.1469-0691.2011.03570.x
15. El Bouamri MC, Arsalane L, Kamouni Y, et al. Current antibiotic resistance profile of uropathogenic Escherichia coli strains and therapeutic consequences. Prog Urol. 2014;24(16):1058–1062.
16. Nzalie RN, Gonsu HK, Koulla-Shiro S. Bacterial etiology and antibiotic resistance profile of community-acquired urinary tract infections in a cameroonian city. Int J Microbiol. 2016;2016:3240268. doi:10.1155/2016/3240268
17. El-Mahmood MA. Antimicrobial susceptibility pattern of pathogenic bacteria causing urinary tract infections at the Specialist Hospital, Yola, Adamawa state, Nigeria. J Clin Med Res. 2009;1(1):1–8.
18. Yandai FH, Ndoutamia G, Nadlaou B, Barro N. Prevalence and resistance profile of Escherichia coli and Klebsiella pneumoniae isolated from urinary tract infections in N’Djamena, Tchad. Int J Biol Chem Sci. 2019;13(4):2065–2073. doi:10.4314/ijbcs.v13i4.13
19. Randrianirina F, Soares JL, Carod JF, et al. Antimicrobial resistance among uropathogens that cause community-acquired urinary tract infections in Antananarivo, Madagascar. J Antimicrob Chemother. 2007;59(2):309–312. doi:10.1093/jac/dkl466
20. Muvunyi CM, Masaisa F, Bayingana C, et al. Decreased susceptibility to commonly used antimicrobial agents in bacterial pathogens isolated from urinary tract infections in Rwanda: need for new antimicrobial guidelines. Am J Trop Med Hyg. 2011;84(6):923–928. doi:10.4269/ajtmh.2011.11-0057
21. Arsalane L, Kamouni Y, Yahyaoui H, Bennouar N, Berraha M, Zouhair S. Profil actuel de résistance aux antibiotiques des souches d’Escherichia coli uropathogènes et conséquences thérapeutiques. Prog Urol. 2014;24(16):1058–1062. doi:10.1016/j.purol.2014.09.035
22. Gonsu Kamga H, Nzengang R, Toukam M, Sando Z, Koulla Shiro S. Phénotypes de résistance des souches d’Escherichia coli responsables des infections urinaires communautaires dans la ville de Yaoundé (Cameroun). Afr J Pathol Microbiol. 2014;3:1–4. doi:10.4303/ajpm/235891
23. Kolawole AS, Kolawole OM, Kandaki-Olukemi YT, Babatunde SK, Durowade KA, Kolawole CF. Prevalence of urinary tract infections (UTI) among patients attending Dalhatu Araf Specialist Hospital, Lafia, Nasarawa state, Nigeria. Int J Med Sci. 2010;1(5):163–167.
24. Behzadi P, Urban E, Matuz M, Benko R, Gajdacs M. The role of gram-negative bacteria in urinary tract infections: current concepts and therapeutic options. Adv Exp Med Biol. 2020.
25. Chang SL, Shortliffe LD. Pediatric urinary tract infections. Pediatr Clin North Am. 2006;53(3):379–400. doi:10.1016/j.pcl.2006.02.011
26. Bernabe KJ, Langendorf C, Ford N, Ronat J-B, Murphy RA. Antimicrobial resistance in West Africa: a systematic review and meta-analysis. Int J Antimicrob Agents. 2017;50(5):629–639. doi:10.1016/j.ijantimicag.2017.07.002
27. Hima-Lerible H, Menard D, Talarmin A. Antimicrobial resistance among uropathogens that cause community-acquired urinary tract infections in Bangui, Central African Republic. J Antimicrob Chemother. 2003;51(1):192–194. doi:10.1093/jac/dkg053
28. Benhiba I, Bouzekraoui T, Zahidi J. Epidémiologie et antibio-résistance des infections urinaires à entérobactéries chez l’adulte dans le CHU de Marrakech et implication thérapeutique. Revue Africaine. 2015;1(4).
29. Dia ML, Chabouny H, Diagne R. Profil antibiotypique des bactéries uropathogènes isolées au CHU de Dakar. Revue Africaine. 2015;1(4).
30. Ullah F, Malik SA, Ahmed J. Antimicrobial susceptibility pattern and ESBL prevalence in Klebsiella pneumoniae from urinary tract infections in the North-West of Pakistan. Afr J Microbiol Res. 2009;3(11):676–680.
31. Ahmed AA, Osman H, Mansour AM, et al. Antimicrobial agent resistance in bacterial isolates from patients with diarrhea and urinary tract infection in the Sudan. Am J Trop Med Hyg. 2000;63(5–6):259–263. doi:10.4269/ajtmh.2000.63.259
32. Singh NP, Goyal R, Manchanda V, Das S, Kaur I, Talwar V. Changing trends in bacteriology of burns in the burns unit, Delhi, India. Burns. 2003;29(2):129–132. doi:10.1016/S0305-4179(02)00249-8
33. Schaumburg F, Alabi A, Kokou C, et al. High burden of extended-spectrum beta-lactamase-producing Enterobacteriaceae in Gabon. J Antimicrob Chemother. 2013;68(9):2140–2143. doi:10.1093/jac/dkt164
34. Yala J-F, Mabika Mabika R, Bisseye C, et al. Phenotypic and genotypic characterization of extended-spectrum-beta-lactamases producing-enterobacteriaceae (ESBLE) in patients attending Omar Bongo Ondimba military hospital at Libreville (Gabon). J Mol Microbiol Biotechnol. 2016;4(6):944–949.
35. WHO. Antimicrobial Resistance: Global Report on Surveillance: World Health Organization. 2014.
36. Leopold SJ, van Leth F, Tarekegn H, Schultsz C. Antimicrobial drug resistance among clinically relevant bacterial isolates in sub-Saharan Africa: a systematic review. J Antimicrob Chemother. 2014;69(9):2337–2353. doi:10.1093/jac/dku176
37. Dromigny JA, Nabeth P. Distribution and susceptibility of bacterial urinary tract infections in Dakar, Senegal. Int J Antimicrob Agents. 2002;20(5):339–347. doi:10.1016/S0924-8579(02)00196-6
38. Naziri Z, Derakhshandeh A, Soltani Borchaloee A, Poormaleknia M, Azimzadeh N. Treatment failure in urinary tract infections: a warning witness for virulent multi-drug resistant ESBL- producing escherichia coli. Infect Drug Resist. 2020;13:1839–1850. doi:10.2147/IDR.S256131
39. Radyowijati A, Haak H. Improving antibiotic use in low-income countries: an overview of evidence on determinants. Soc Sci Med. 2003;57(4):733–744. doi:10.1016/S0277-9536(02)00422-7
40. Tadesse BT, Ashley EA, Ongarello S, et al. Antimicrobial resistance in Africa: a systematic review. BMC Infect Dis. 2017;17(1):616. doi:10.1186/s12879-017-2713-1
41. Grigoryan L, Burgerhof JG, Haaijer-Ruskamp FM, et al. Is self-medication with antibiotics in Europe driven by prescribed use? J Antimicrob Chemother. 2007;59(1):152–156. doi:10.1093/jac/dkl457
42. Planta MB. The role of poverty in antimicrobial resistance. J Am Board Fam Med. 2007;20(6):533–539. doi:10.3122/jabfm.2007j.06.070019
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
