Back to Journals » OncoTargets and Therapy » Volume 10
Downregulation of long noncoding RNA TUG1 inhibits proliferation and induces apoptosis through the TUG1/miR-142/ZEB2 axis in bladder cancer cells
Authors Liu Q, Liu H, Cheng HP, Li Y, Li XD, Zhu CY
Received 13 October 2016
Accepted for publication 7 March 2017
Published 5 May 2017 Volume 2017:10 Pages 2461—2471
DOI https://doi.org/10.2147/OTT.S124595
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 4
Editor who approved publication: Dr Tohru Yamada
Qian Liu,* Hui Liu,* Hepeng Cheng, Yang Li, Xiaodong Li, Chaoyang Zhu
Department of Urology Surgery, Huaihe Hospital of Henan University, Kaifeng, People’s Republic of China
*These authors contributed equally to this work
Background: Bladder cancer is a common serious disease around the world. Long noncoding RNAs (lncRNAs) have been demonstrated to participate in the development and progression of various cancers, including bladder cancer. The aim of this study was to investigate the effects of lncRNA taurine upregulated gene 1 (TUG1) on proliferation and apoptosis in bladder cancer cell lines and the underlying mechanism.
Methods: The levels of TUG1 were detected by quantitative real time polymerase chain reaction (qRT-PCR) in bladder cancer tissues and cells. The mRNA and protein levels of zinc finger E-box binding homeobox 2 (ZEB2) were measured by qRT-PCR and Western blot analysis, respectively. The functional targets of TUG1 were predicted by online softwares and confirmed by luciferase reporter assay. The effects of TUG1 on cell proliferation and apoptosis were examined by MTT and apoptosis assay, respectively. The expression levels of β-catenin, cyclinD1, and c-Myc in T24 cells were determined by Western blot analysis.
Results: The levels of TUG1 and ZEB2 were significantly increased in bladder cancer tissues and cells. Knockdown of either TUG1 or ZEB2 inhibited proliferation and induced apoptosis in bladder cancer cells. Interestingly, ZEB2 overexpression reversed the effects of TUG1 knockdown on cell proliferation and apoptosis. Moreover, ZEB2 was verified as a direct target of miR-142 and miR-142 could specially bind to TUG1. In addition, downregulation of TUG1 inhibited the Wnt/β-catenin pathway by regulating ZEB2 expression in bladder cancer cells.
Conclusion: Downregulation of TUG1 expression inhibited proliferation and induced apoptosis in bladder cancer cells by targeting ZEB2 mediated by miR-142 through the inactivation of Wnt/β-catenin pathway.
Keywords: bladder cancer, lncRNA TUG1, miR-142, ZEB2, Wnt/β-catenin pathway
Introduction
Bladder cancer ranks the ninth most common malignancy worldwide, and is the most common genitourinary malignant disease with estimated 140,430 new cases and 29,790 deaths in the United States.1,2 Despite much effort and progress in clinical therapy for bladder cancer, the overall survival time has not been improved markedly, and the 5-year survival rate is still only 50%–60%.3–5 Thus, searching new molecular targets and therapeutic strategies becomes increasingly important.
Long noncoding RNAs (lncRNAs) are ~200-nucleotide (nt)-long RNA molecules and stably exist in the plasma and urine, with disease and tissue specificity and without protein-coding potential.6 lncRNAs have been implied to be frequently aberrantly expressed in lots of cancers and involved in a large range of biological processes, such as transcriptional regulation, cell growth, and tumorigenesis.6,7 Taurine upregulated gene 1 (TUG1), a 7.1-kb lncRNA, was initially identified as a transcript upregulated by taurine.8 A previous report showed that TUG1 was upregulated in high-grade muscle-invasive bladder cancer and modulated proliferation and migration of bladder cancer cells.9 However, the precise mechanism by which TUG1 affects bladder cancer remains poorly defined.
MicroRNAs (miRNAs) are short noncoding RNA molecules with 19–22 nts and are implicated in multiple cellular processes at the posttranscriptional level by targeting the 3′ untranslated region of messenger RNA (mRNA), resulting in translation suppression or mRNAs degradation.10 A previous document showed that miR-142-3p expression was upregulated in bladder carcinoma cells.11 However, the function and mechanism of miR-142 in bladder cancer is not fully addressed.
Zinc finger E-box binding homeobox 2 (ZEB2) has been revealed to be abnormally expressed in different tumors, including bladder cancer.12,13 A previous study indicated that ZEB2 was closely related to epithelial-to-mesenchymal transition (EMT) in hepatocellular carcinoma.14 Kurashige et al found that ZEB2 promoted cell proliferation, invasion, and migration in gastric carcinoma,15 which suggested that ZEB2 was a vital factor in promoting the initiation and development of cancer.
In this study, the expression levels of TUG1 and ZEB2 in bladder cancer tissue and cells were first detected. The role and regulatory mechanisms of TUG1 in bladder cancer cell lines were further investigated by gain- and loss-of-function experiments and bioinformatics analysis.
Materials and methods
Acquisition of tissue samples and cell culture
After approval from the Ethics Committee of Huaihe Hospital of Henan University and obtaining written informed consents from the patients, matching samples of bladder cancer tissues and adjacent normal tissues were collected from 36 patients undergoing surgical resection at the Huaihe Hosptital of Henan University. The specimens were immediately frozen and stored in liquid nitrogen. Pathological examination was performed by a pathologist to confirm bladder cancer.
Human urothelial cells HCV-29 and human bladder cancer cell lines (T24 and BIU-87) were purchased from American Type Culture Collection (ATCC, Manassas, VA, USA). All cell lines were cultured in RPMI 1640 medium (Invitrogen, Carlsbad, CA, USA) supplemented with 10% fetal bovine serum (GIBCO, Carlsbad, CA, USA) at 37°C in a humidified atmosphere containing 5% CO2.
Cell transfection
miR-124 mimics (miR-142), small interfering RNA (siRNA) against TUG1 (si-TUG1), TUG1 expression plasmid (pcDNA-TUG1), siRNA against ZEB2 (si-ZEB2) and ZEB2 expression plasmid (pcDNA-ZEB2) were purchased from Ribobio Co. (Guangzhou, China). siRNA sequences for TUG1 were 5′-GCU UGG CUU CUA UUC UGA AUC CUU U-3′ (sense) and 5′-AAA GGA UUC AGA AUA GAA GCC AAG C-3′ (antisense). siRNA sequences for ZEB2 were 5′-CAU CAG ACU UUG AGG AAU A-3′ (sense) and 5′-UAU UCC UCA AAG UCU GAU G-3′ (antisense). Cell transfection was performed by Lipofectamine 2000 reagent (Invitrogen) according to the manufacturer’s instructions. Cells were harvested for quantitative real time polymerase chain reaction (qRT-PCR), Western blot, MTT, or flow cytometry analysis at indicated time points.
RNA isolation and qRT-PCR
Total RNA was isolated from tissues and cells using TRIzol reagent (Thermo Fisher Scientific, Waltham, MA, USA) according to manufacturer’s instructions. Extracted RNA was used to synthesize complementary DNA (cDNA) using an All-in-One™ miRNA qRT-PCR Detection Kit (GeneCopoeia, FulenGen, China). To quantify mRNAs, the reverse-transcription reaction was performed using a Prime Script™ RT reagent kit (Takara, Shiga, Japan). Real-time quantitative reactions were performed in triplicate in a 384-well plate containing 1 μL of synthesized cDNA, Platinum SYBR Green qPCR SuperMix UDG (Invitrogen), and gene specific forward and reverse PCR primers (5′-CAA GAA ACA GCA ACA CCA GAA G-3′ and 5′-TAA GGT CCC CAT TCA AGT CAG T-3′ for TUG1; 5′-CCGTTGGACCTGTCATTACC-3′ and 5′-GACGATGAAGAAACACTGTTGTG-3′ for ZEB2; 5′-TGG ACT TCG AGC AGG AAA TGG-3′ and 5′-ACG TCG CAC TTC ATG ATC GAG-3′ for β-actin). The expression levels of miRNA and mRNA were normalized to endogenous U6 small nuclear RNA (U6-snRNA) and β-actin by the 2−ΔΔCt method.
Western blot analysis
Total proteins were isolated from tissues and cells by RIPA buffer. Protein quantification was performed by a NanoDrop 2000 spectrophotometer (Thermo Fisher Scientific). A total of 50 μg proteins were separated by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and then transferred to polyvinylidene fluoride membranes (Millipore, Bradford, MA, USA). The membranes were blocked in buffer (Tris-buffered saline and 0.1% Tween 20 (TBST) buffer with 5% bovine serum albumin) for 2 h at room temperature and then were incubated with primary antibodies overnight at 4°C. The membranes were washed twice in TBST and incubated with secondary antibody labeled with horse radish peroxidase for 1 h at 37°C before detected by an enhanced chemiluminescence system (Pierce, Rockford, IL, USA). Finally, the signal intensity of the interest proteins was determined by ImageJ software (NIH, Bethesda, MD, USA). The primary antibodies used in this study are anti-ZEB2 (1:1,000; Santa Cruz Biotechnology, Santa Cruz, CA, USA), anti-β-catenin (1:1,000; Cell Signaling Technology, Beverly, MA, USA), anti-cyclinD1 (1:1,000; Cell Signaling Technology), anti-c-Myc (1:1,000; Cell Signaling Technology), and anti-β-actin (1:5,000; Sigma, St Louis, MO, USA).
MTT assay
Cells were seeded in 96-well plates (Corning Costar, Corning, NY, USA) at a density of 2.0×103 cells per well and incubated for 24 h prior to transfection. Then, 20 μL of MTT (Sigma) solution (5 mg/mL) was added to each well at indicated times, and incubation was continued for further 4 h at 37°C. After discarding supernatant, intracellular formazan crystals were dissolved by addition of 200 μL of dimethyl sulfoxide (DMSO; Sigma) in each well. Cell proliferation was determined by measuring the absorbance at 490 nm on a Multiskan Ascent 354 microplate reader (Thermo Labsystems, Waltham, MA, USA).
Flow cytometry
Cultured cells were collected and washed with phosphate-buffered saline (PBS). Cells (1×106) from each sample were resuspended in binding buffer and stained with the AnnexinV-fluorescein isothiocyanate (AnnexinV-FITC)/propidium iodide kit (BD Pharmingen, San Diego, CA, USA) following the manufacturer’s instructions. Positive cells were detected and quantified by flow cytometry (BD FACS Aria; BD Biosciences, Franklin Lakes, NJ, USA) 48 h posttransfection.
Luciferase assay
The binding sequences of TUG1 and miR-142 were predicted by online softwares starBase v2.0 and TargetScan. TUG1 targeted miR-142 and ZEB2 was confirmed to be a direct target of miR-142. The sequences of wild type TUG1 (TUG1-WT), mutant TUG1 (TUG1-MUT), wild type ZEB2 (ZEB2-WT), and mutant ZEB2 (ZEB2-MUT) containing the putative binding sites of miR-142 were amplified and cloned into the pGL3-control luciferase reporter vectors (Promega, Madison, WI, USA). The reporter vectors with miR-142 or miR-control were cotransfected into T24 and BIU-87 cells using Lipofectamine 2000 (Invitrogen). After 36 h of incubation, cells were lysed and luciferase activities were detected with Dual-Luciferase Reporter Assay System (Promega).
Statistical analysis
Data were presented as mean ± standard deviation (SD) of at least 3 independent experiments. The difference between 2 groups was examined by the Student’s t-test, while one-way analysis of variance was employed to compare difference among 3 or more groups using SPSS 19.0. A P-value of <0.05 was considered statistically significant.
Results
TUG1 and ZEB2 were upregulated in bladder cancer tissues
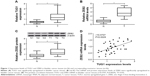
To begin with, qRT-PCR and Western blot was performed to detect the expression of TUG1 and ZEB2 in 39 pairs of bladder cancer tissue and normal counterparts. Analysis of qRT-PCR showed that expression levels of both TUG1 and ZEB2 were significantly increased in bladder cancer tissues, compared with the corresponding normal tissues (Figure 1A and B). Western bolt analysis indicated that expression levels of ZEB2 protein were significantly elevated in bladder cancer tissues compared with the adjacent normal tissues (Figure 1C). We also analyzed the relationships between TUG1 and ZEB2, and found that TUG1 expression was positively correlated with ZEB2 mRNA in bladder cancer tissues (Figure 1D). From these data, it is speculated that TUG1 and ZEB2 might play an oncogenic role in bladder cancer.
Knockdown of TUG1 suppressed proliferation and induced apoptosis in bladder cancer cells
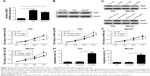
To investigate the functional effects of TUG1 in bladder cancer in vitro, the expression levels of TUG1 were determined in human urothelial cells HCV-29 and human bladder cancer cell lines (T24 and BIU-87). The result showed that expression levels of TUG1 were significantly higher in T24 and BIU-87 cells than those in HCV-29 cells (Figure 2A). To study the possible role of TUG1 in bladder cancer cell proliferation and apoptosis, T24 and BIU-87 cells were transfected with si-TUG1. MTT assay indicated that proliferation of T24 and BIU-87 cells were both significantly inhibited in the si-TUG1 group compared with the si-control group (Figure 2B and C). Flow cytometry analysis revealed that transfection of si-TUG1 led to an obvious increase of apoptosis rates in both T24 (Figure 2D and E) and BIU-87 (Figure 2F and G) cells. These data suggested that TUG1 knockdown inhibited cell proliferation and promoted apoptosis in bladder cancer in vitro.
Downregulation of ZEB2 inhibited proliferation and promoted apoptosis in bladder cancer cells
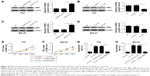
To explore the functional effects of ZEB2 in bladder cancer in vitro, mRNA and protein levels of ZEB2 in HCV-29, T24, and BIU-87 cells were evaluated by using qRT-PCR and Western blot analysis, respectively. The results indicated that ZEB2 was significantly upregulated in mRNA (Figure 3A) and protein levels (Figure 3B) in T24 and BIU-87 cells compared with HCV-29 cells. To observe the possible roles of ZEB2 in bladder cancer cell proliferation and apoptosis, T24 and BIU-87 cells were transfected with si-ZEB2 and pcDNA-ZEB2 plasmids, respectively. As presented in Figure 3C, knockdown of ZEB2 dramatically repressed expression of ZEB2 and transfection of pcDNA-ZEB2 evidently improved expression of ZEB2. MTT assay demonstrated that proliferation of si-ZEB2-transfected T24 cells was significantly inhibited, whereas ZEB2 upregulation promoted growth of T24 cells (Figure 3D and E). Similarly, ZEB2 downregulation impeded proliferation of BIU-87 cells, whereas ZEB2 overexpression enhanced cell proliferation (Figure 3F and G). In addition, flow cytometry analysis implied that the apoptosis rates of both T24 and BIU-87 cells transfected with si-ZEB2 were dramatically increased compared with control groups (Figure 3H and I). These data suggested that ZEB2 knockdown inhibited cell proliferation and induced apoptosis in bladder cancer in vitro.
ZEB2 overexpression reversed effects of TUG1 knockdown on proliferation and apoptosis of bladder cancer cells
To further explore the relationship between TUG1 and ZEB2 in bladder cancer cells, the protein level of ZEB2 were measured in cells transfected with si-TUG1 and pcDNA-TUG1, respectively. The results showed that enforced expression of TUG1 triggered a significant promotion in ZEB2 expression in both T24 and BIU-87 cells (Figure 4A and C), whereas ZEB2 expression level was markedly reduced following inhibition of TUG1 (Figure 4B and D). Furthermore, cells were cotransfected with si-TUG1 and pcDNA-ZEB2 to investigate their effects on cell proliferation and apoptosis. The data indicated that knockdown of TUG1 significantly suppressed proliferation (Figure 4E and F) and promoted apoptosis (Figure 4G and H) of T24 and BIU-87 cells, whereas ectopic expression of ZEB2 abolished these effects.
TUG1 regulates ZEB2 by sponging miR-142 in bladder cancer cells
TUG1 was predicted to contain binding sites of miR-142 by bioinformatic software starBase v2.0 (Figure 5A). Then pGL3 luciferase reporter plasmid containing the 3′-UTR sequences of wild type TUG1 (TUG1-WT) and mutant type TUG1 (TUG1-MUT) were constructed. The results showed that mature miR-142 strongly decreased the luciferase expression of TUG1-WT in both T24 and BIU-87 cells, whereas luciferase expression of TUG1-MUT was not significantly affected after transfection with miR-142 mimic (Figure 5B and C). Then, the expression level of miR-142 was detected in T24 and BIU-87 cells transfected with si-TUG1 and pcDNA-TUG1, respectively. The results confirmed that knockdown of TUG1 significantly upregulated the expression levels of miR-142, whereas overexpression of TUG1 significantly reduced the expression levels of miR-142 in T24 and BIU-87 cells (Figure 5D and E). All data indicated that TUG1 targetedly suppressed miR-142 expression.
Interestingly, an online software TargetScan also showed that miR-142 may bind to 3′-UTR of ZEB2 mRNA (Figure 5F). The reporter plasmids harboring the predicted miR-142 binding region sites (ZEB2-WT) and its mutant (ZEB2-MUT) were both established. These results showed that cotransfection of mature miR-142 and ZEB2-WT significantly limited the luciferase activity in both T24 and BIU-87 cells, which was abated by transfection of TUG1 overexpression (Figure 5G and H). However, for cells with ZEB2-MUT, there was no significant difference of luciferase activity in miR-control, miR-142, and miR-142 + pcDNA-TUG1. Then the expression levels of ZEB2 protein in T24 and BIU-87 cells transfected with miR-142 or miR-142 + pcDNA-TUG1 were determined using Western blot analysis. The results implicated that miR-142 resulted in a significant suppression in ZEB2 expression in T24 and BIU-87 cells, which was abrogated following transfection of pcDNA-TUG1 (Figure 5I and J). These data suggested that TUG1 regulated ZEB2 expression by targeting miR-142.
TUG1 regulated the Wnt/β-catenin pathway by affecting ZEB2 expression
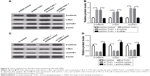
To further explore the molecular mechanism of TUG1 regulating the biological processes in bladder cancer cells, Western blot analysis was used to detect the expression levels of proteins related to the Wnt/β-catenin signaling pathway (β-catenin, cyclinD1, and c-Myc) in T24 cells treated with pcDNA-TUG1, pcDNA-TUG1 + si-ZEB2, si-TUG1, or si-TUG1 + pcDNA-ZEB2. The results suggested that TUG1 upregulation significantly increased the expression levels of β-catenin, cyclinD1, and c-Myc in T24 cells, whereas knockdown of ZEB2 reversed the positive effects of TUG1 upregulation on these proteins (Figure 6A and B). Conversely, knockdown of TUG1 generated a significant reduction in the expression levels of β-catenin, cyclinD1, and c-Myc in T24 cells, which was eliminated after the transfection of pcDNA-ZEB2 (Figure 6C and D). These data suggested that knockdown of TUG1 downregulated ZEB2 to further block Wnt/β-catenin signaling pathway, which has a vital role in bladder cancer cells growth and apoptosis.
Discussion
Bladder cancer is a common serious disease around the world. Recent studies showed that the dysregulation of lncRNAs were involved in several human cancers, including bladder cancer.16,17 It is of great significance to illuminate the molecular mechanism of lncRNAs for the treatment of bladder cancer.
Previous studies indicated lncRNA TUG1 was upregulated in bladder cancer, gastric cancer, and osteosarcoma.18–20 Consistent with this study, the levels of TUG1 in bladder cancer tissues were significantly higher than those in paired nontumor tissues. These data suggested that TUG1 may exhibit vital roles in bladder cancer development and progression. In this study, loss of function method was used to evaluate the potential effects of TUG1 on bladder cancer cells. The results demonstrated that TUG1 downregulation exhibited cell proliferation inhibition and apoptosis promotion in T24 and BIU-87 cells, which was consistent with Zhang’s results.20
lncRNAs have been elaborated to function as endogenous miRNA sponges to bind to miRNAs and modulate their biological function.21,22 To explore whether TUG1 acts as a miRNA sponge, bioinformatics analysis was carried out and the results showed that TUG1 contained binding sites for miR-142. A great many miRNAs are dysregulated in cancers and may play essential roles in tumorigenesis.23 A recent study reported that TUG1 decreased the expression of miR-145 and knocking TUG1 inhibited EMT and radioresistance through the miR-145/ZEB2 axis in human bladder cancer cells.24 However, the potential molecular mechanism of miR-142 during cancer progression is unclear. In this study, it was confirmed that miR-142 could target directly ZEB2 mRNA and inhibited expression of ZEB2 protein. Thus, it was speculated that TUG1 exerted its function via the miR-142/ZEB2 axis.
ZEB2 is a member of the Zfh1 family of 2-handed zinc finger/homeodomain proteins, and can bind to paired ZEB-type E boxes within target gene promoters to inhibit their transcription.25 A previous document indicated that knockdown of ZEB2 inhibited cell proliferation, migration, invasion, and promoted cell apoptosis in glioma cells.26 In agreement with these findings, the data in this study showed that ZEB2 expression was significantly upregulated in bladder cancer tissues and cells, and inhibited cell proliferation and induced apoptosis in T24 and BIU-87 cells.
Activation of the Wnt/β-catenin signaling pathway was observed in many human cancers, including bladder cancer.27 Previous reports revealed that β-catenin, a vital regulator of Wnt pathway, can transcriptionally regulate c-Myc and cyclinD1 of Wnt target genes, which participate in cell survival and proliferation.28,29 As is well-known, c-Myc and cyclinD1 are 2 proto-oncogenes and are vital regulators of cellular growth and proliferation.30 In this study, knockdown of TUG1 significantly downregulated the protein levels of β-catenin, c-Myc, and cyclinD1 in T24 cells, while ZEB2 overexpression reversed the inhibitory effects. Thus, these data suggested that knockdown of TUG1 inhibited the Wnt/β-catenin pathway through affecting ZEB2 expressions.
Conclusion
TUG1 was upregulated in bladder cancer tissues and cells and TUG1 knockdown suppressed proliferation and promoted apoptosis of bladder cancer cells. ZEB2 improved cell proliferation and inhibited apoptosis in bladder cancer cells. TUG1 could bind to miR-142 to regulate ZEB2 expression. Overexpression of ZEB2 overturned the effects of TUG1 knockdown on proliferation and apoptosis in bladder cancer cells. Knockdown of TUG1 suppressed the activation of the Wnt/β-catenin pathway by affecting ZEB2 expression. Collectively, this study demonstrated that TUG1 knockdown inhibited proliferation and induced apoptosis through suppressing ZEB2 mediated by miR-142 via the inactivation of the Wnt/β-catenin pathway in bladder cancer. This TUG1/miR-142/ZEB2 regulatory axis and the possible molecular mechanism provide a theoretical basis for the research on lncRNA-directed therapeutics in bladder cancer.
Disclosure
The authors report no conflicts of interest in this work.
References
Siegel R, Naishadham D, Jemal A. Cancer statistics, 2013. CA Cancer J Clin. 2013;63(1):11–30. | ||
Witjes JA, Comperat E, Cowan NC, et al; European Association of Urology. EAU guidelines on muscle-invasive and metastatic bladder cancer: summary of the 2013 guidelines. Eur Urol. 2014;65(4):778–792. | ||
Marta GN, Hanna SA, Gadia R, Correa SF, Silva JL, Carvalho Hde A. The role of radiotherapy in urinary bladder cancer: current status. Int Braz J Urol. 2012;38(2):144–153. | ||
Racioppi M, D’Agostino D, Totaro A, et al. Value of current chemotherapy and surgery in advanced and metastatic bladder cancer. Urol Int. 2012;88(3):249–258. | ||
Chen J, Wang L, Tang Y, et al. Maspin enhances cisplatin chemosensitivity in bladder cancer T24 and 5637 cells and correlates with prognosis of muscle-invasive bladder cancer patients receiving cisplatin based neoadjuvant chemotherapy. J Exp Clin Cancer Res. 2016;35:2. | ||
Yang F, Xue X, Bi J, et al. Long noncoding RNA CCAT1, which could be activated by c-Myc, promotes the progression of gastric carcinoma. J Cancer Res Clin Oncol. 2013;139(3):437–445. | ||
Yuan JH, Yang F, Wang F, et al. A long noncoding RNA activated by TGF-beta promotes the invasion-metastasis cascade in hepatocellular carcinoma. Cancer Cell. 2014;25(5):666–681. | ||
Khalil AM, Guttman M, Huarte M, et al. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc Natl Acad Sci U S A. 2009;106(28):11667–11672. | ||
Iliev R, Kleinova R, Juracek J, et al. Overexpression of long non-coding RNA TUG1 predicts poor prognosis and promotes cancer cell proliferation and migration in high-grade muscle-invasive bladder cancer. Tumor Biol. 2016;37(10):13385–13390. | ||
Hua Y, Duan S, Murmann AE, et al. miRConnect: identifying effector genes of miRNAs and miRNA families in cancer cells. PLoS One. 2011;6(10):e26521. | ||
Jia AY, Castillo-Martin M, Domingo-Domenech J, et al. A common MicroRNA signature consisting of miR-133a, miR-139-3p, and miR-142-3p clusters bladder carcinoma in situ with normal umbrella cells. Am J Pathol. 2013;182(4):1171–1179. | ||
Lee H, Jun SY, Lee YS, Lee HJ, Lee WS, Park CS. Expression of miRNAs and ZEB1 and ZEB2 correlates with histopathological grade in papillary urothelial tumors of the urinary bladder. Virchows Arch. 2014;464(2):213–220. | ||
Prislei S, Martinelli E, Zannoni GF, et al. Role and prognostic significance of the epithelial-mesenchymal transition factor ZEB2 in ovarian cancer. Oncotarget. 2015;6(22):18966–18979. | ||
Yang Z, Sun B, Li Y, et al. ZEB2 promotes vasculogenic mimicry by TGF-beta1 induced epithelial-to-mesenchymal transition in hepatocellular carcinoma. Exp Mol Pathol. 2015;98(3):352–359. | ||
Kurashige J, Kamohara H, Watanabe M, et al. MicroRNA-200b regulates cell proliferation, invasion, and migration by directly targeting ZEB2 in gastric carcinoma. Ann Surg Oncol. 2012;19 (Suppl 3):S656–S664. | ||
Xue Y, Ma G, Zhang Z, et al. A novel antisense long noncoding RNA regulates the expression of MDC1 in bladder cancer. Oncotarget. 2015;6(1):484–493. | ||
Peter S, Borkowska E, Drayton RM, et al. Identification of differentially expressed long noncoding RNAs in bladder cancer. Clin Cancer Res. 2014;20(20):5311–5321. | ||
Cao WJ, Wu HL, He BS, Zhang YS, Zhang ZY. Analysis of long non-coding RNA expression profiles in gastric cancer. World J Gastroenterol. 2013;19(23):3658–3664. | ||
Han Y, Liu Y, Gui Y, Cai Z. Long intergenic non-coding RNA TUG1 is overexpressed in urothelial carcinoma of the bladder. J Surg Oncol. 2013;107(5):555–559. | ||
Zhang Q, Geng PL, Yin P, Wang XL, Jia JP, Yao J. Down-regulation of long non-coding RNA TUG1 inhibits osteosarcoma cell proliferation and promotes apoptosis. Asian Pac J Cancer Prev. 2013;14(4):2311–2315. | ||
Sen R, Ghosal S, Das S, Balti S, Chakrabarti J. Competing endogenous RNA: the key to posttranscriptional regulation. ScientificWorldJournal. 2014;2014:896206. | ||
Wang P, Ning S, Zhang Y, et al. Identification of lncRNA-associated competing triplets reveals global patterns and prognostic markers for cancer. Nucleic Acids Res. 2015;43(7):3478–3489. | ||
Cong N, Du P, Zhang A, et al. Downregulated microRNA-200a promotes EMT and tumor growth through the wnt/β-catenin pathway by targeting the E-cadherin repressors ZEB1/ZEB2 in gastric adenocarcinoma. Oncol Rep. 2013;29(4):1579–1587. | ||
Tan J, Qiu K, Li M, Liang Y. Double-negative feedback loop between long non-coding RNA TUG1 and miR-145 promotes epithelial to mesenchymal transition and radioresistance in human bladder cancer cells. FEBS Lett. 2015;589(20 Pt B):3175–3181. | ||
Vandewalle C, Comijn J, De Craene B, et al. SIP1/ZEB2 induces EMT by repressing genes of different epithelial cell-cell junctions. Nucleic Acids Res. 2005;33(20):6566–6578. | ||
Qi S, Song Y, Peng Y, et al. ZEB2 mediates multiple pathways regulating cell proliferation, migration, invasion, and apoptosis in glioma. PLoS One. 2012;7(6):e38842. | ||
Du HF, Ou LP, Lv CK, et al. Expression of hepaCAM inhibits bladder cancer cell proliferation via a Wnt/beta-catenin-dependent pathway in vitro and in vivo. Cancer Biol Ther. 2015;16(10):1502–1513. | ||
He TC, Sparks AB, Rago C, et al. Identification of c-MYC as a target of the APC pathway. Science. 1998;281(5382):1509–1512. | ||
Tetsu O, McCormick F. Beta-catenin regulates expression of cyclin D1 in colon carcinoma cells. Nature. 1999;398(6726):422–426. | ||
Li YJ, Wei ZM, Meng YX, Ji XR. Beta-catenin up-regulates the expression of cyclinD1, c-myc and MMP-7 in human pancreatic cancer: relationships with carcinogenesis and metastasis. World J Gastroenterol. 2005;11(14):2117–2123. |
 © 2017 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2017 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.