Back to Journals » Infection and Drug Resistance » Volume 11
Detection and characterization of a clinical Escherichia coli ST3204 strain coproducing NDM-16 and MCR-1
Authors Li X , Mu X, Zhang P, Zhao D , Ji J, Quan J, Zhu Y, Yu Y
Received 23 May 2018
Accepted for publication 13 June 2018
Published 15 August 2018 Volume 2018:11 Pages 1189—1195
DOI https://doi.org/10.2147/IDR.S175041
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Suresh Antony
Xi Li,1,* Xinli Mu,2,* Ping Zhang,2 Dongdong Zhao,2 Jingshu Ji,3 Jingjing Quan,2 Yongze Zhu,1 Yunsong Yu2
1Centre of Laboratory Medicine, Zhejiang Provincial People’s Hospital, People’s Hospital of Hangzhou Medical College, Hangzhou, Zhejiang 310014, China; 2Department of Infectious Diseases, Sir Run Run Shaw Hospital, College of Medicine, Zhejiang University, Hangzhou, Zhejiang 310016, China; 3Department of Clinical Laboratory, Sir Run Run Shaw Hospital, College of Medicine, Zhejiang University, Hangzhou, Zhejiang 310016, China
*These authors contributed equally to this work
Objectives: A plasmid-mediated colistin resistance gene, mcr-1, has been reported worldwide and has caused concern regarding a major therapeutic challenge. Alarmingly, mcr-1 has spread into clinical carbapenem-resistant Enterobacteriaceae isolates, resulting in extensively drug-resistant and even pan drug-resistant isolates that can cause untreatable infections. In this study, we report isolation of an extensively drug-resistant Escherichia coli strain EC1188 that coproduces NDM-16 and MCR-1 from a urine sample taken from a patient with craniocerebral injury.
Materials and methods: E. coli strain EC1188 was identified and subjected to genotyping, susceptibility testing and conjugation experiments. The genetic locations of blaNDM-16 and mcr-1 were established with southern blot hybridization. The complete genome sequence of this strain was obtained and the genetic characteristics of the mcr-1- and blaNDM-16-harboring plasmids were analyzed. In addition, comparative genetic analyses of mcr-1 and blaNDM-16 with closely related plasmids were also carried out.
Results: Whole-genome sequencing revealed that strain EC1188 possess various resistance genes and virulence genes. S1-pulsed-field gel electrophoresis and southern blot suggested that the blaNDM-16 and mcr-1 genes were located on an ~65 kb plasmid and an ~80 kb plasmid, respectively. Moreover, the two genes could successfully transfer their resistance phenotype to E. coli strain C600. Sequence analysis showed that these two plasmids possessed high sequence similarity to previously reported blaNDM-5-harboring and mcr-1-harboring plasmids in China.
Conclusion: To the best of our knowledge, this is the first report to isolate an E. coli strain that coproduces NDM-16 and MCR-1. In addition, we characterized the blaNDM-16-harboring plasmid for the first time. Our study further emphasizes that the co-occurrence of the two prevalent transferrable resistance plasmids in a single isolate is highly significant because infections caused by MCR-1–producing carbapenem-resistant Enterobacteriaceae isolates are increasing each year. It is imperative to perform active surveillance to prevent further dissemination of MCR-1–producing CRE isolates.
Keywords: E. coli, mcr-1, NDM-16, CRE, colistin resistance, extensively drug-resistant bacteria
Corrigendum for this paper has been published
Introduction
Polymyxins are regarded as last-line therapeutic options to treat the growing number of infections caused by multidrug-resistant Gram-negative bacteria, especially carbapenem-resistant Enterobacteriaceae (CRE).1 Meanwhile, since the initial discovery of the plasmid-mediated colistin resistance gene, mcr-1, this gene has been reported in Enterobacteriaceae isolates from the environment, food, animals and humans worldwide.2 Even more worrisome, mcr-1 has spread into clinical CRE isolates, resulting in extensively drug-resistant isolates that can cause untreatable infections. Our research group and others have reported on a number of clinical isolates that coproduce MCR-1 and carbapenemase. Among these clinical MCR-1–producing CRE isolates, the NDM-type carbapenemases are highly represented, including NDM-13–5 and NDM-5.6–11 Other carbapenemases have also been identified, such as KPC-2,12,13 VIM-114 and OXA-48.15,16 Such strains have the potential to seriously compromise current treatment options for CRE infections. Moreover, these resistance genes are located on different plasmids, allowing their further spread. It is clear that MCR-1–producing CRE isolates result in so-called “superbug” isolates, accelerating the entry into a “post-antibiotic” era.17
In the present study, we report the isolation of an extensively drug-resistant Escherichia coli strain that coproduces NDM-16 and MCR-1 from a urine sample taken from a patient with a craniocerebral injury. The genetic characteristics of the mcr-1- and blaNDM-16-harboring plasmids were analyzed. In addition, comparative genetic analyses of mcr-1 and blaNDM-16 with closely related plasmids were also carried out. To the best of our knowledge, this is the first report describing the isolation of an isolate E. coli strain coproducing NDM-16 and MCR-1.
Materials and methods
Patient and isolate data
A carbapenem-resistant strain of E. coli EC1188 was isolated from a 40-year-old female patient who was admitted to a hospital for a craniocerebral injury caused by a traffic accident in Hangzhou, Zhejiang, China in September 2017. The strain was cultured from urine during hospitalization of the patient and was preliminarily identified by the VITEK 2 system (Sysmex-bioMérieux, Marcy l’Etoile, France) and further confirmed by 16S rRNA sequencing. Multilocus sequence typing (MLST) of resistance genes and the Inc-type plasmid of the strain were performed using the MLST 1.8 server, ResFinder 3.0, Virulence Finder 1.5 and Plasmid Finder 1.3, which are available at the Center for Genomic Epidemiology (http://www.genomicepidemiology.org/). Antibiotic susceptibilities were determined using the broth microdilution method and the results were interpreted according to the Clinical and Laboratory Standard Institute guidelines,18 except for tigecycline and colistin, which were interpreted according to the European Committee on Antimicrobial Susceptibility Testing breakpoints for Enterobacteriaceae (http://www.eucast.org/clinical_breakpoints). E. coli ATCC 25922 was used as a quality control strain of antibiotic susceptibility test.
S1-pulsed-field gel electrophoresis (PFGE) and southern blot analysis
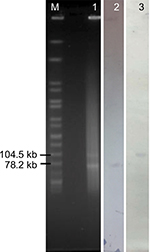
To ascertain the plasmids location of the blaNDM-16 and mcr-1 genes, genomic DNA digested with S1-nuclease (TaKaRa, Osaka, Japan) was electrophoresed on a CHEF-mapper XA PFGE system (Bio-Rad Laboratories Inc., Hercules, CA, USA) for 22 hours at 14°C with run conditions of 6 V/cm and pulse times from 2.16 to 63.8 seconds. The DNA fragments were transferred to a positive-charged nylon membrane (EMD Millipore, Billerica, MA, USA) and then hybridized with a digoxigenin-labeled blaNDM-16- and mcr-1-specific probe. An NBT/BCIP color detection kit (Hoffman-La Roche Ltd., Basel, Switzerland) was then used to detect the fragments. The Salmonella enterica serotype Braenderup H9812 was used as the size marker.
Conjugation experiments
A filter-mating experiment was carried out between the isolate and rifampicin-resistant E. coli C600 as the recipient strain.8 Transconjugants were selected for Mueller–Hinton (MH) agar plates containing 500 mg/L rifampicin and 100 mg/L ampicillin or 500 mg/L rifampicin and 2 mg/L colistin. Polymerase chain reaction sequencing and antimicrobial susceptibility testing of the transconjugants were subsequently performed to confirm whether the plasmid was successfully transferred to the recipient.
Genomic DNA extraction and analysis
Total genomic DNA extraction and analysis was performed as previously described.12 Briefly, the genomic DNA of E. coli strain EC1188 was extracted using a QIAamp DNA MiniKit (Qiagen, Valencia, CA, USA) following the manufacturer’s recommendations. Sequencing was performed via a single molecule real-time technique using a PacBio RS II platform and the resulting sequences were assembled de novo using the hierarchical genome assembly process with the default settings of the single molecule real-time Analysis v2.3.0 software package. Next, the Rapid Annotation using Subsystems Technology annotation website server (http://rast.nmpdr.org/rast/cgi) was used to annotate the genomes. A comparison of pEC1188-NDM16 and pEC1188-MCR and their related plasmids was performed with EasyFig 2.2.2.19
Plasmid stability
Plasmid stability was determined according to a previous study.20 Briefly, the isolate was individually streaked out on the MH agar, incubated at 37°C for 24 hours and then transferred to a fresh MH agar plate. After repeating this procedure for 10 days, 12 individual colonies were randomly selected. Subsequently, the mcr-1 and blaNDM-16 genes were screened by polymerase chain reaction and sequenced.
Nucleotide sequence accession numbers
The complete nucleotide sequences of the plasmids pEC1188-NDM16 and pEC1188-MCR reported in the present study have been deposited in the GenBank nucleotide database under accession numbers MH213345 and MH213346, respectively.
Ethical approval
The clinical isolate E. coli EC1188 was part of the routine hospital laboratory procedure.
The Ethics Committee of the Zhejiang Provincial People’s Hospital exempted this study from review because the present study only focused on bacteria.
Results and discussion
Isolate characteristics
The antimicrobial susceptibility testing results showed that strain EC1188 was resistant to the majority of the antimicrobial agents tested (Table 1) and was classified as an extensively drug-resistant bacterium according to the recently proposed international classification scheme.21 Acquired resistance to antimicrobial agents was further identified, including beta-lactams (blaTEB-1B, blaCTX-199 and blaNDM-16), aminoglycoside (aadA5, rmtB and aadA2), colistin (mcr-1), macrolide (mph(A)), sulfonamide (sul1), tetracycline (tetA) and trimethoprim (dfrA12 and dfrA17). MLST analysis showed the isolate belonged to ST3204.
To date, some clinical isolates coproducing MCR-1 and various carbapenemases have been identified since an MCR-1 and NDM-5 coproducing isolate was first reported from a duck sample.22 However, strains coproducing NDM-16 and MCR-1 have not been reported. At present, E. coli is the primary host among strains coproducing MCR-1 and carbapenemase (Table 2). According to the MLST results (Table 2), these isolates have various sequence types, indicating that these isolates are not clonally related to each other, even though some of the strains were collected from the same hospital.
In addition, the type one fimbriae genes fimABCDEFGHI, which encode virulence determinants, were identified by the Rapid Annotation using Subsystems Technology annotation. Furthermore, the type IV fimbrial genes pilABC were also identified. Other virulence genes for increased serum survival (iss), glutamate decarboxylase (gad), enterobactin siderophore receptor protein (iron), colicin M (cma) and several putative virulence factors were also identified in the genome of this strain. The identified virulence determinants may have contributed to the infection or colonization of the strain.
Plasmid location
S1-PFGE followed by southern blotting showed that the mcr-1 and blaNDM-16 genes were located on an ~65 kb IncHI2 plasmid and an ~80 kb IncF plasmid, respectively (Figure 1). Filter mating experiments were then performed to confirm their transferability and the two genes could successfully transfer their resistance phenotype to E. coli strain C600 (Table 1). In addition, after ten rounds of subculture on MH agar without the addition of antibiotics, randomly selected colonies were confirmed to carry mcr-1 and blaNDM-16 genes and plasmids identical to their parental isolate in size, indicating that the plasmids harboring the blaNDM-16 or mcr-1 genes may be stably maintained in the hosts.
Genetic context of plasmids bearing mcr-1
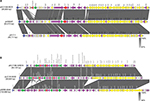
The whole genome sequence of strain EC1188 was obtained to better characterize the mcr-1- and blaNDM-16-positive plasmids. The mcr-1 gene was located in a plasmid named pEC118-MCR. Sequence analysis showed that the plasmid was 65,806 bp in length (Figure 2), belonged to IncI2 incompatibility group and had a G+C content of 42.5%.
Sequence analysis showed that the genes coding for plasmid transfer functions, such as tra and pil, and the genes responsible for plasmid stability functions, such as mok and parA, were identified. BLASTn search results showed that pEC1188-MCR exhibited 91% query coverage and 99% identity to the originally described mcr-1-encoding plasmid pHNSHP45.23 The pEC118-MCR plasmid sequence was nearly identical to the plasmid pHNSHP45, primarily differing by the following: 1) the lack of a 2,704 bp insertion of IS683 downstream of the repA gene, which is consistent with another plasmid (pH17-1) carrying the mcr-1 gene but lacking the inserted sequence;12 2) an ISApl1 transposable element was not observed in this plasmid, but an insertion sequence ISEcp1was present (Figure 2) and 3) an extended-spectrum β-lactamase gene blaCTX-M-199 was identified in this plasmid. Note that the insertion sequence ISApl1 gene was not found upstream of mcr-1, but the pap2 gene encoding the PAP2-family protein was identified downstream of the mcr-1 gene. The insertion sequence of ISApl1 is not always associated with mcr-1 and it is absent in some mcr-1-positive plasmids, implying a relic for frequent conjugation of ISApl1-mcr-1-pap2.24
So far, mcr-1 has been reported to be carried in different plasmids incompatibility such as IncI2, IncX4, IncHI2 and IncP.24 The IncI2-type plasmid was the first reported mcr-1–harboring plasmid. In addition, in our previous study, of 21 mcr-1–positive isolates recovered from bloodstream infections in China, 15 isolates contained mcr-1 on the IncI2-type plasmid.8 Furthermore, incompatibility of strains coproducing MCR-1 and carbapenemases isolated from humans being was analyzed, indicating that IncI2-type plasmid was also prevalent (Table 2). Overall, IncI2-type plasmid played an important role in the dissemination of mcr-1 in E. coli, even in strains coproducing MCR-1 and carbapenemases.
Genetic characteristics of plasmids bearing blaNDM-16
The blaNDM-16 gene is located in a plasmid named pEC1188-NDM16. Sequence analysis showed that the plasmid was 80,215 bp in length (Figure 2), belonged to the IncF incompatibility group and had a G+C content of 51.8%. The plasmid was further analyzed in the nr/nt database and showed an overall 99% nucleotide identity and 100% query coverage to the blaNDM-5-positive plasmids pLZ135-NDM (GenBank Accession Number MF353156) and pNDM-d2e9 (GenBank Accession Number CP026201), as shown in Figure 2. Interestingly, analysis of the typical segment flanking blaNDM-16 on pEC1188-NDM16 revealed that the blaNDM-16 gene was likely to be transferred by two flanking IS15DIV elements, which share the same flanking sequence (IS15DIV-blaNDM-16-ble-dsbD-IS91-dfrA12-IS15DIV) with the two blaNDM-5 gene-bearing IncX3 plasmids.
In addition, NDM-5 is the primary type of carbapenemase among MCR-1–producing CRE isolates (Table 2). Our recent study also found that blaNDM-5 was able to coexist in the MCR-1–producing strain isolated from bloodstream infections.8 Furthermore, NDM-16, which differs from NDM-5, has aspartate to valine amino acid substitution at position 233. Note that dissemination of the blaNDM-5 gene via a plasmid among non-clonal E. coli isolates has been observed in our hospital.25 Although no genetic association was observed among MCR-1–producing CRE isolates, the widespread dissemination of blaNDM-5 in recent years in Enterobacteriaceae highlights the need for active surveillance to prevent further dissemination of MCR-1–producing isolates.
Conclusion
In summary, to the best of our knowledge, this is the first report of a clinical E. coli strain coproducing MCR-1 and NDM-16. In addition, this study also characterizes the blaNDM-16-harboring plasmid for the first time. Our study further underscores that the co-occurrence of the two prevalent transferrable resistance plasmids in a single isolate raises extensive concern, because infections caused by MCR-1–producing CRE isolates increase each year. It is imperative to perform active surveillance to prevent further dissemination of MCR-1–producing CRE isolates.
Acknowledgments
This study was supported by the National Natural Science Foundation of China (31700125), the Natural Science Young Foundation of Zhejiang Province, China (LQ17H190006), the Medical and Health Research Project of Zhejiang Province, China (2017KY224, 2018KY229, and 2018RC011). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author contributions
YY and YZ conceived and designed the experiments; XL, PZ, and XM performed the experiments; DZ, JJ, and JQ analyzed the data; XL and XM wrote the manuscript. All authors read and approved the final manuscript. All authors contributed toward data analysis, drafting and revising the paper and agree to be accountable for all aspects of the work.
Disclosure
The authors report no conflicts of interest in this work.
References
Kaye KS, Pogue JM, Tran TB, Nation RL, Li J. Agents of last resort: polymyxin resistance. Infect Dis Clin North Am. 2016;30(2):391–414. | ||
Schwarz S, Johnson AP. Transferable resistance to colistin: a new but old threat. J Antimicrob Chemother. 2016;71(8):2066–2070. | ||
Delgado-Blas JF, Ovejero CM, Abadia-Patiño L, Gonzalez-Zorn B. Coexistence of mcr-1 and blaNDM-1 in Escherichia coli from Venezuela. Antimicrob Agents Chemother. 2016;60(10):6356–6358. | ||
Zheng B, Dong H, Xu H, et al. Coexistence of MCR-1 and NDM-1 in clinical Escherichia coli isolates. Clin Infect Dis. 2016;63(10):1393–1395. | ||
Zhong LL, Zhang YF, Doi Y, et al. Coproduction of MCR-1 and NDM-1 by colistin-resistant Escherichia coli isolated from a healthy individual. Antimicrob Agents Chemother. 2017;61(1):e01962–16. | ||
Mediavilla JR, Patrawalla A, Chen L, et al. Colistin- and carbapenem-resistant Escherichia coli harboring mcr-1 and blaNDM-5, causing a complicated urinary tract infection in a patient from the United States. MBio. 2016;7(4):e01191-16. | ||
Zheng B, Lv T, Xu H, et al. Discovery and characterisation of an Escherichia coli ST206 strain producing NDM-5 and MCR-1 from a patient with acute diarrhoea in China. Int J Antimicrob Agents. 2018;51(2):273–275. | ||
Quan J, Li X, Chen Y, et al. Prevalence of mcr-1 in Escherichia coli and Klebsiella pneumoniae recovered from bloodstream infections in China: a multicentre longitudinal study. Lancet Infect Dis. 2017;17(4):400–410. | ||
Yu H, Qu F, Shan B, et al. Detection of the mcr-1 colistin resistance gene in carbapenem-resistant Enterobacteriaceae from different hospitals in China. Antimicrob Agents Chemother. 2016;60(8):5033–5035. | ||
Li A, Yang Y, Miao M, et al. Complete sequences of mcr-1-harboring plasmids from extended-spectrum-β-lactamase- and carbapenemase-producing Enterobacteriaceae. Antimicrob Agents Chemother. 2016;60(7):4351–4354. | ||
Zhang Y, Liao K, Gao H, et al. Decreased fitness and virulence in ST10 Escherichia coli harboring blaNDM-5 and mcr-1 against a ST4981 strain with blaNDM-5. Front Cell Infect Microbiol. 2017;7:242. | ||
Zhao D, Zhou Z, Hua X, et al. Coexistence of mcr-1, blaKPC-2 and two copies of fosA3 in a clinical Escherichia coli strain isolated from urine. Infect Genet Evol. 2018;60:77–79. | ||
Falgenhauer L, Waezsada SE, Yao Y, et al. Colistin resistance gene mcr-1 in extended-spectrum β-lactamase-producing and carbapenemase-producing Gram-negative bacteria in Germany. Lancet Infect Dis. 2016;16(3):282–283. | ||
Poirel L, Kieffer N, Liassine N, Thanh D, Nordmann P. Plasmid-mediated carbapenem and colistin resistance in a clinical isolate of Escherichia coli. Lancet Infect Dis. 2016;16(3):281. | ||
Mulvey MR, Mataseje LF, Robertson J, et al. Dissemination of the mcr-1 colistin resistance gene. Lancet Infect Dis. 2016;16(3):289–290. | ||
Lu H, Zhang X, Zhang Y, et al. First report of multidrug-resistant Escherichia coli isolates co-harbouring mcr-1 and blaOXA-48 from clinical patients in China. J Glob Antimicrob Resist. 2018;13:94–95. | ||
Bulman ZP, Chen L, Walsh TJ, et al. Polymyxin combinations combat Escherichia coli harboring mcr-1 and blaNDM-5: preparation for a postantibiotic era. MBio. 2017;8(4):e00540-17. | ||
CLSI. Performance Standards for Antimicrobial Susceptibility Testing. CLSI Supplement M100. 27th ed. Wayne, PA: Clinical and Laboratory Standards Institute; 2017. | ||
Sullivan MJ, Petty NK, Beatson SA. Easyfig: a genome comparison visualizer. Bioinformatics. 2011;27(7):1009–1010. | ||
Zhu YQ, Zhao JY, Xu C, et al. Identification of an NDM-5-producing Escherichia coli sequence type 167 in a neonatal patient in China. Sci Rep. 2016;6:29934. | ||
Magiorakos AP, Srinivasan A, Carey RB, et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect. 2012;18(3):268–281. | ||
Yang RS, Feng Y, Lv XY, et al. Emergence of NDM-5- and MCR-1-producing Escherichia coli clones ST648 and ST156 from a single Muscovy duck (Cairina moschata). Antimicrob Agents Chemother. 2016;60(11):6899–6902. | ||
Liu YY, Wang Y, Walsh TR, et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect Dis. 2016;16(2):161–168. | ||
Wang Q, Sun J, Li J, et al. Expanding landscapes of the diversified mcr-1-bearing plasmid reservoirs. Microbiome. 2017;5(1):70. | ||
Li X, Fu Y, Shen M, et al. Dissemination of blaNDM-5 gene via an IncX3-type plasmid among non-clonal Escherichia coli in China. Antimicrob Resist Infect Control. 2018;7:59. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.