Back to Journals » Cancer Management and Research » Volume 10
Deletion of 3p13 is a late event linked to progression of TMPRSS2:ERG fusion prostate cancer
Authors Kluth M, Volta H, Hussein M, Taskin B, Frogh S, Möller-Koop C, Büscheck F, Jacobsen F, Tsourlakis MC, Lübke AM, Hinsch A, Clauditz T, Graefen M, Heinzer H , Huland H , Minner S, Sauter G, Wilczak W, Schlomm T, Simon R
Received 29 April 2018
Accepted for publication 28 June 2018
Published 19 November 2018 Volume 2018:10 Pages 5909—5917
DOI https://doi.org/10.2147/CMAR.S172637
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Antonella D'Anneo
Martina Kluth,1,* Heinke Volta,1,* Mohammad Hussein,1 Billurvan Taskin,1 Sohall Frogh,1 Christina Möller-Koop,1 Franziska Büscheck,1 Frank Jacobsen,1 Maria Christina Tsourlakis,1 Andreas M Lübke,1 Andrea Hinsch,1 Till Clauditz,1 Markus Graefen,2 Hans Heinzer,2 Hartwig Huland,2 Sarah Minner,1 Guido Sauter,1 Waldemar Wilczak,1 Thorsten Schlomm,2–4 Ronald Simon1
1Institute of Pathology, University Medical Center Hamburg-Eppendorf, Hamburg, Germany; 2Martini-Clinic, Prostate Cancer Center, University Medical Center Hamburg-Eppendorf, Hamburg, Germany; 3Department of Urology, Section for Prostate Cancer Research, University Medical Center Hamburg-Eppendorf, Hamburg, Germany; 4Department of Urology, Charitè Universitätsmedizin Berlin, Berlin, Germany
*These authors contributed equally to this work
Introduction: Deletion of 3p13 is one of the most common alterations in prostate cancer preferentially occurring in tumors with TMPRSS2:ERG fusion. The cause for the striking association between 3p13 loss and ERG fusion is unknown.
Methods: Here, we made use of a preexisting heterogeneity prostate cancer tissue microarray including ten tissue spots from ten different tumor areas of 317 cancers to examine the spatial distribution of 3p13 deletions (determined by fluorescence in situ hybridization) in prostate cancer areas with and without ERG overexpression (determined by immunohistochemistry).
Results: 3p13 deletions were found in 61 of 299 (20.4%) and ERG positivity in 174 of 317 (54.9%) interpretable cancers. The likelihood of 3p13 loss was twice as high in ERG-positive cancers (39/152, 25.7%) than in ERG-negative cancers (17/124, 13.7%, P=0.010). At least three tissue spots were interpretable for 3p13 deletion status in 279 cancers: only these were used for heterogeneity assessment. Among these tumors, 58 (20.8%) had a 3p13 deletion and 221 (79.2%) were undeleted. The majority of 3p13-deleted cancers showed marked intratumoral heterogeneity. Areas with and without 3p13 loss were found in 50 (18%) of 279 cancers with three or more interpretable tissue spots, while only eight (3%) tumors had a homogeneous 3p13 loss. Comparison with ERG data revealed that ERG fusion usually precede 3p13 deletions. In total, 26 (66.7%) of 39 cancers with ERG and 3p13 alteration had only focal 3p13 deletions in an otherwise ERG-positive background. In contrast, none of the cancers showed a pattern that would be consistent with 3p13 deletion preceding ERG fusion.
Conclusion: Our study identifies 3p13 deletion as a highly heterogeneous alteration in prostate cancer preferentially developing at rather late stages of progression in TMPRSS2:ERG fusion-positive tumors.
Keywords: 3p13 deletion, prostate cancer, heterogeneity, TMPRSS2:ERG fusion, tissue microarray
Introduction
Chromosomal deletions are a hallmark of prostate cancer. Partial or complete losses involving chromosomes 2q, 3p, 5q, 6q, 8p, 10q, 12, 13q, 16q, 17, 18q, and 21q occur in up to 60% of tumors.1–3 Deletions of 3p and 10q are of particular interest because they are typically small as compared to all other deletion loci.2,4–6 The 10q23 deletion locus has been extensively analyzed. A magnitude of studies have demonstrated that the PTEN tumor suppressor is targeted by 10q23 deletion and that PTEN loss is linked to aggressive tumor features and poor patient prognosis.3,7,16 Much less is known about the 3p deletion. Global copy number screening studies demonstrated that 3p deletions are strongly linked to the subset of prostate cancers harboring the TMPRSS2:ERG fusion and that they typically do not exceed two megabases in size.2,5 In a previous study, we found 3p13 deletion in 15% of prostate cancers, where it was linked to early biochemical recurrence and adverse tumor phenotype.5 Moreover, we assessed the tumor-suppressive properties of three candidate genes, including the SHQ1 H/ACA ribonucleoprotein assembly factor, the FOXP1 transcription factor, and the RYBP polycomb suppressor, in our earlier study.5 The striking association between ERG fusion and 3p13 deletion strongly suggests cooperative effects driving tumor growth. However, it is currently unknown whether development of these two alterations follows a specific sequence and whether 3p13 deletion is an early or late event during prostate cancer progression.
Intratumoral heterogeneity can serve as an excellent tool to unravel the chronology of molecular alterations arising during tumor growth and progression. It can be hypothesized that early tumor-initiating molecular alterations are present in virtually all cancer cells, whereas events occurring at later stages of tumor progression will be detectable only in a subset of cancer cells. We have previously developed a prostate cancer heterogeneity tissue microarray (TMA) platform.8,9 This TMA contains multiple samples from the same prostate cancer, thus enabling a high-throughput mapping of molecular features across the entire tumors. We have successfully used this approach to dissect the chronology of key molecular alterations of prostate cancers, including activation of the ETS transcription factor ERG and recurrent deletions of the PTEN tumor suppressor or deletions at chromosome 6q15.8–11
In the present study, we took advantage of our heterogeneity TMA approach in order to evaluate the degree of heterogeneity of 3p13 deletions in 317 prostate cancers and to use this information in order to determine the chronology of TMPRSS2:ERG fusion and 3p13 deletion development.
Materials and methods
Patient samples and TMA construction
All patients were treated by radical prostatectomy at our center between January and March 2010. Removed prostates were macroscopically dissected in a standardized way. All prostatectomy specimens were completely paraffin-embedded and processed according to a modified Stanford protocol as previously described.12,13 The prostates were fixed in 4% buffered formalin, serially blocked at 3 mm intervals in transverse planes perpendicular to the rectal surface, and embedded in paraffin. For each cancer, the number of independent tumor foci was determined according to Wise et al.14 In brief, tumor areas were defined as part of a single focus if they were within 3 mm of each other in any section or within 4 mm in adjacent sections. A total of 317 cancers were considered unifocal according to this definition and were included in the study. From each of these cancers, ten different tumor-containing tissue blocks were selected for TMA construction. From each block, one 0.6 mm punch core was removed, and the ten cores were assembled side by side in a TMA. If less than ten blocks were available, multiple cores were taken from one or several blocks in order to obtain the total number of ten punches per tumor focus. If more than ten blocks were available, blocks were selected to obtain an optimal representation of the entire tumor mass (ie, blocks were selected that enabled maximal distances between selected tumor areas). This approach resulted in TMA containing a total of 3,170 cores obtained from 317 cancer foci. Histopathological data of all arrayed tumors are given in Table 1. Presence of cancer was histologically confirmed in each tissue spot. Normal prostate glands were immunohistochemically identified using the antibody 34BE12 (clone MA903, 1:12.5, pH 7.8; Dako, Glostrup, Denmark) for basal cell detection. The usage of archived anonymized diagnostic leftover tissues for manufacturing of TMAs without written consent and their analysis for research purposes as well as patient data analysis has been approved by local laws (HmbKHG, §12,1) and by the local ethics committee (Ethics Commission Hamburg, WF-049/09 and PV3652). All work was carried out in compliance with the Declaration of Helsinki.
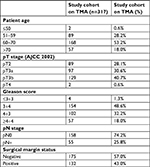  | Table 1 Clinicopathological features of the 317 arrayed prostate cancers Abbreviations: AJCC, American Joint Committee on Cancer; p, pathological; TMA, tissue microarray. |
Immunohistochemistry
Immunohistochemical analysis of ERG was performed as previously described.8,15 Freshly cut TMA sections were analyzed in 1 day and in one experiment. Slides were deparaffinized and exposed to heat-induced antigen retrieval for 5 minutes in an autoclave at 121°C in pH 7.8 citrate buffer. The primary antibody (clone EPR3864; Epitomics) was diluted at 1:450. Bound primary antibody was visualized using the Dako EnVision Kit. Only nuclear ERG staining was scored. Any detectable staining was considered ERG positive.
Fluorescence in situ hybridization (FISH)
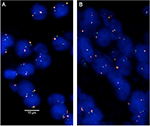
Four-micrometer TMA sections were used for FISH. For proteolytic slide pretreatment, a commercial kit was used (paraffin pretreatment reagent kit; Abbott, Chicago, IL, USA). TMA sections were deparaffinized, air-dried, and dehydrated in 70%, 85%, and 100% ethanol, followed by denaturation for 5 minutes at 74°C in 70% formamid 2X SSC solution. The FISH probe set consisted of a SpectrumGreen-labeled 3p13 (FOXP1 locus) probe (made from a mixture of RP11-154H23 and RP11-49E03) and a SpectrumOrange-labeled commercial centromere 3 probe (#06J36-003; Abbott) as a reference. Hybridization was performed overnight at 37°C in a humidified chamber. Slides were subsequently washed and counterstained with 0.2 µmol/L DAPI in antifade solution. Stained slides were manually interpreted with an epifluorescence microscope, and the predominant FISH signal numbers were recorded in each tissue spot. Homozygous deletion of 3p13 was defined as complete absence of 3p13 FISH probe signals in ≥60% of tumor nuclei, with the presence of one or two 3p13 FISH signals in adjacent normal cells. Heterozygous deletion of 3p13 was defined as the presence of fewer 3p13 signals than centromere 3 probe signals in ≥60% of tumor nuclei (Figure 1). This threshold was based on a previous validation study comparing PTEN copy number results analyzed by FISH and array comparative genomic hybridization in prostate cancer.16 Tissue spots with a lack of 3p13 signals in all (tumor and normal cells) or lack of any normal cells as an internal control for successful hybridization of the 3p13 probe were excluded from analysis.
Results
Technical issues
Interpretable results for 3p13 were found in 2,195 (69.2%) of the 3,170 tissue spots from 317 different prostate cancers included in the heterogeneity TMA. Out of the 317 cancers, 279 (88.0%) had at least three TMA spots that were interpretable for 3p13, including 55 tumors with ten interpretable spots, 66 cancers with nine interpretable spots, 52 cancers with eight interpretable spots, 42 cancers with seven interpretable spots, 26 cancers with six interpretable spots, 17 cancers with five interpretable spots, nine cancers with four interpretable spots, and 12 cancers with three interpretable spots for 3p13. The remaining 38 cancers had only less than three interpretable tissue spots for 3p13 FISH. In summary, there were 6.9±2.9 (average) and eight (median) analyzable tissue spots per cancer. All data are summarized in Figures 2 and S1.
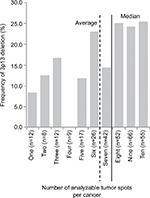  | Figure 2 Frequency of 3p13 deletion in dependence of the number of analyzable tissue spots per cancer. |
Impact of the number of tissue spots on 3p13 deletion frequency
The 3p13 deletion rate increased with the number of analyzable tumor spots per cancer. 3p13 deletion was found in six (10.3%) of 58 cancers with 1–5 analyzable tumor spots, but in 43 (24.9%) of 143 cancers with 8–10 analyzable tumor spots. All data are summarized in Figure 2.
Heterogeneity of 3p13 deletion
Only the subset of 279 cancers with at least three tissue spots interpretable for 3p13 FISH analysis was included in this evaluation. Deletion of 3p13 was found in 58 (20.8%) of these cancers. There was a high degree of intratumoral heterogeneity: only eight (13.8%) of 58 tumors were homogenously 3p13 deleted (ie, 100% of all interpretable cancer spots showed a 3p13 deletion), while 50 (86.2%) cancers were heterogeneously 3p13 deleted (ie, at least one tissue spot had no 3p13 deletion). All data are summarized in Figure 3.
Association of 3p13 deletion and ERG expression
In order to estimate the patterns of ERG expression and 3p13 deletion on a spot-by-spot basis, a subset of 2,085 tissue spots that were interpretable for both ERG and 3p13 was analyzed. Deletion of 3p13 was strongly linked to an ERG-positive cancer phenotype: 3p13 loss was seen in 199 (19.3%) of 1,032 ERG-positive cancer spots but only in 69 (6.6%) of 1,053 ERG-negative cancer spots (P<0.0001). To study whether this association also held true on the cancer level, we further studied the subset of 276 tumors that had at least three informative tissue spots for both 3p13 deletion and ERG expression status. Tissue spots that yielded results only for either ERG or 3p13 were excluded from this analysis. In total, this analysis included 2,039 tissue spots from 276 cancers (range: 3–10 spots per cancer, average: 7.34±1.98 spots per cancer). This analysis revealed that the likelihood for 3p13 deletion was 1.88-fold higher in ERG-positive cancers (39/152, 25.7%) as compared to ERG-negative cancers (17/124, 13.7%, P=0.010).
Chronology of 3p13 deletion and ERG expression
To determine whether the presence of ERG facilitates 3p13 deletion or vice versa, we sought to determine the chronological sequence in 39 cancers with 3p13 deletion and ERG expression. All of these cancers showed a pattern consistent with ERG rearrangement preceding 3p deletions. This includes 28 cancers (71.8%) with focal 3p13 deletions in an otherwise ERG-positive background as well as eleven (28.2%) cancers with homogeneous positivity for both alterations. All data are summarized in Figure 4.
Discussion
The average 3p13 deletion rate of 21% in our current study on 279 cancers with 3–10 different analyzable tissue spots each was significantly higher than in our previous study analyzing only a single 0.6 mm TMA spot per cancer (14.5% in 1,300 cancers; P=0.006).5 This discrepancy is obviously due to the higher likelihood of detecting deletions in heterogeneous tumors if multiple samples are analyzed. That the maximal deletion rate (26%) was reached when at least eight spots per cancer were analyzed demonstrates that the ten tissue spots per cancer included in our study was sufficient to reliably detect the vast majority of 3p13-deleted cancers. It is, however, noteworthy that the FOXP1 FISH probe selected for this project does not identify all 3p13 deletions. We previously found about 4%–5% of 3p13-deleted prostate cancers with alternative deletions not including FOXP1 centering around the RYBP gene located 790 kilobases centromeric to FOXP1.5 Accordingly, we estimate that more than 30% of prostate cancers may have at least some focal 3p13 deletions.
The degree of heterogeneity of a molecular event may represent a parameter for its early or late occurrence. Homogeneous molecular features are likely to have developed early in tumor evolution. This is especially true for “low-malignant” alterations that do not trigger rapid outgrowth of a subclone. ERG overexpression is an example for a molecular aberration that is often homogeneous in prostate cancer and does not confer a marked proliferative advantage to cancer cells.8,15 ERG overexpression can therefore be considered an early event based on our heterogeneity analysis, which is in line with other data from the literature.17–20 If a molecular cancer feature is limited to a small cancer area only, this feature must have occurred rather late in tumor evolution. That the vast majority (86%) of 3p13-deleted cancers were heterogeneously deleted (ie, contained tumor areas with normal 3p13 copy numbers) fits thus with the notion of 3p13 loss representing a relevant event during late tumor progression.5
Very similar to what was found in our previous study using the same experimental approach,5 the likelihood of 3p13 deletion was about twice as high in ERG-positive than in ERG-negative cancers. This frequent co-occurrence raises the question whether one of these alterations facilitates development of the other. That most cancers harboring both alterations had small tumor areas with 3p13 loss in an otherwise ERG-positive background – while the inverse situation was not seen (ie, cases with homogeneous 3p13 deletions containing small areas of ERG positivity) – strongly suggests that ERG fusion typically precedes 3p13 deletion. It seems possible that 3p13 deletion could provide a selection advantage to cancer cells which is particularly powerful in ERG-positive cells. In fact, candidate genes at 3p13, including FOXP1 and RYBP, as well as ERG are strong transcriptional regulators with a plethora of target genes.3,21–24 This offers a multitude of modes of interaction. ERG has been shown to deregulate more than 1,600 genes, while FOXP1 modifies transcription of more than 600 genes.3,21–23 Pathways affected by both genes include AKT and AR signaling.23,25–27 Both pathways are highly relevant for prostate cancer and may be subject to functional interaction with ERG and FOXP1. A hypothetical target of joint functional effects of ERG and RYBP is the transcription factor E2F3, which interacts with ERG and RYBP in cell line models.24,28
That only heterozygous 3p13 deletions were found supports the concept that biallelic inactivation (either at the same time or one allele after the other) is not a suitable mechanism for 3p tumor suppressor gene inactivation in prostate cancer. This notion is in line with data from next-generation sequencing studies not revealing any mutations of 3p13 genes occurring more frequently than the 0.9% of RYPB mutations (http://www.cbioportal.org).29,30 However, the lack of homozygous 3p13 deletions may offer novel therapeutic options. Global copy number analysis studies have shown that the 3p13 deletion typically spans two megabases including seven genes, at least one of which might be essential for cell survival.2,5 Heterozygous deletion of essential genes has been postulated to render cancer cells vulnerable to further inhibition of these genes, and 56 genes have been identified until now, suppression of which specifically inhibited the proliferation of cells harboring partial copy number loss of these genes.31 Such essential genes had been suggested as promising targets for anticancer therapies and were thus termed CYCLOPS (copy number alterations yielding cancer liabilities owing to partial loss) genes.31 However, it is currently not known whether any of the seven genes within the 3p13 deletion region may be druggable.
In this context, it is important to note that heterogeneity of a molecular alteration may limit both the applicability of diagnostic tests and the effectiveness of targeted therapies. Considering the importance of heterogeneity, the number of studies systematically analyzing targeted heterogeneity in cancer is rather small. Moreover, studies addressing heterogeneity often limit themselves to the analysis of one tumor block per cancer.32,33 However, the analysis of one tumor area cannot represent the molecular events in a large cancer. Our TMA analysis of one sample each from at least ten different tumor-containing blocks distributed across the entire tumor enables a three-dimensional assessment of molecular features in a large series of cancers. This heterogeneity TMA concept differs markedly from previous TMA studies, which used multiple tumor cores from just one tumor-containing block.34,35 Nevertheless, our study has several limitations. These include the preselection of large unifocal tumors, which may have resulted in a shift toward more advanced cancers, as well as the different amount of tissues analyzed per cancer in case that not all tissue spots contained tumor cells. In addition, fluorescence signals were not counted in each spot but rather estimated, which might have led to false deletion calling in some cases. Lastly, formalin fixation inevitably results in a certain fraction of non-interpretable tissues which may introduce a bias in our study.
Conclusion
In summary, the results of our study demonstrate a marked heterogeneity of 3p13 deletion in prostate cancer. Analyzing up to ten different regions per cancer enabled us to determine a 3p13 deletion rate of 20%–25%. Data derived from tumor mapping demonstrated that 3p13 deletions generally develop in ERG-expressing cells after TMPRSS2:ERG fusion has evolved. That all 3p13 deletions were heterozygous supports the concept that at least one gene in this region must be essential for prostate epithelial cells.
Acknowledgments
The authors are grateful to Janett Lütgens, Sünje Seekamp, Inge Brandt, Sylvia Schnöger, and Sascha Egteshadi for their excellent technical assistance and Eike Burandt, Till Krech, and Stefan Steurer for their pathological review. This work was supported by the Federal Ministry of Education and Research; grant 01ZX1302C.
Author contributions
GS, TS, and RS designed the study. MK, HV, MH, BT, and SF performed the FISH analysis. MCT and SM performed the immunohistochemical analysis. FB, FJ, AML, AH, TC, and WW contributed to the pathological validation of the tumors. MG, HH, and HHu prepared the tumor sections. CMK, MCT, and SM constructed the tissue microarray. MK, HV, and RS carried out the data analysis. MK, HV, and RS wrote the manuscript. All authors contributed toward data analysis, drafting and critically revising the paper, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Disclosure
The authors report no conflicts of interest in this work.
References
Sun J, Liu W, Adams TS, et al. DNA copy number alterations in prostate cancers: a combined analysis of published CGH studies. Prostate. 2007;67(7):692–700. | ||
Taylor BS, Schultz N, Hieronymus H, et al. Integrative genomic profiling of human prostate cancer. Cancer Cell. 2010;18(1):11–22. | ||
Weischenfeldt J, Simon R, Feuerbach L, et al. Integrative genomic analyses reveal an androgen-driven somatic alteration landscape in early-onset prostate cancer. Cancer Cell. 2013;23(2):159–170. | ||
Huang S, Gulzar ZG, Salari K, Lapointe J, Brooks JD, Pollack JR. Recurrent deletion of CHD1 in prostate cancer with relevance to cell invasiveness. Oncogene. 2012;31(37):4164–4170. | ||
Krohn A, Seidel A, Burkhardt L, et al. Recurrent deletion of 3p13 targets multiple tumour suppressor genes and defines a distinct subgroup of aggressive ERG fusion-positive prostate cancers. J Pathol. 2013;231(1):130–141. | ||
Mao X, Boyd LK, Yáñez-Muñoz RJ, et al. Chromosome rearrangement associated inactivation of tumour suppressor genes in prostate cancer. Am J Cancer Res. 2011;1(5):604–617. | ||
Wise HM, Hermida MA, Leslie NR. Prostate cancer, PI3K, PTEN and prognosis. Clin Sci. 2017;131(3):197–210. | ||
Minner S, Gärtner M, Freudenthaler F, et al. Marked heterogeneity of ERG expression in large primary prostate cancers. Mod Pathol. 2013;26(1):106–116. | ||
Kluth M, Jung S, Habib O, et al. Deletion lengthening at chromosomes 6q and 16q targets multiple tumor suppressor genes and is associated with an increasingly poor prognosis in prostate cancer. Oncotarget. 2017;8(65):108923–108935. | ||
Krohn A, Freudenthaler F, Harasimowicz S, et al. Heterogeneity and chronology of PTEN deletion and ERG fusion in prostate cancer. Mod Pathol. 2014;27(12):1612–1620. | ||
Kluth M, Meyer D, Krohn A, et al. Heterogeneity and chronology of 6q15 deletion and ERG-fusion in prostate cancer. Oncotarget. 2016;7(4):3897–3904. | ||
Mcneal JE, Redwine EA, Freiha FS, Stamey TA. Zonal distribution of prostatic adenocarcinoma. Correlation with histologic pattern and direction of spread. Am J Surg Pathol. 1988;12(12):897–906. | ||
Erbersdobler A, Hammerer P, Huland H, Henke RP. Numerical chromosomal aberrations in transition-zone carcinomas of the prostate. J Urol. 1997;158(4):1594–1598. | ||
Wise AM, Stamey TA, Mcneal JE, Clayton JL. Morphologic and clinical significance of multifocal prostate cancers in radical prostatectomy specimens. Urology. 2002;60(2):264–269. | ||
Minner S, Enodien M, Sirma H, et al. ERG status is unrelated to PSA recurrence in radically operated prostate cancer in the absence of antihormonal therapy. Clin Cancer Res. 2011;17(18):5878–5888. | ||
Krohn A, Diedler T, Burkhardt L, et al. Genomic deletion of PTEN is associated with tumor progression and early PSA recurrence in ERG fusion-positive and fusion-negative prostate cancer. Am J Pathol. 2012;181(2):401–412. | ||
Perner S, Mosquera JM, Demichelis F, et al. TMPRSS2-ERG fusion prostate cancer: an early molecular event associated with invasion. Am J Surg Pathol. 2007;31(6):882–888. | ||
Schaefer G, Mosquera JM, Ramoner R, et al. Distinct ERG rearrangement prevalence in prostate cancer: higher frequency in young age and in low PSA prostate cancer. Prostate Cancer Prostatic Dis. 2013;16(2):132–138. | ||
Tsourlakis MC, Stender A, Quaas A, et al. Heterogeneity of ERG expression in prostate cancer: a large section mapping study of entire prostatectomy specimens from 125 patients. BMC Cancer. 2016;16:641. | ||
Mertz KD, Horcic M, Hailemariam S, et al. Heterogeneity of ERG expression in core needle biopsies of patients with early prostate cancer. Hum Pathol. 2013;44(12):2727–2735. | ||
Gabut M, Samavarchi-Tehrani P, Wang X, et al. An alternative splicing switch regulates embryonic stem cell pluripotency and reprogramming. Cell. 2011;147(1):132–146. | ||
Brase JC, Johannes M, Mannsperger H, et al. TMPRSS2-ERG -specific transcriptional modulation is associated with prostate cancer biomarkers and TGF-β signaling. BMC Cancer. 2011;11:507. | ||
Tomlins SA, Rhodes DR, Yu J, et al. The role of SPINK1 in ETS rearrangement-negative prostate cancers. Cancer Cell. 2008;13(6):519–528. | ||
Schlisio S, Halperin T, Vidal M, Nevins JR. Interaction of YY1 with E2Fs, mediated by RYBP, provides a mechanism for specificity of E2F function. Embo J. 2002;21(21):5775–5786. | ||
Bock J, Mochmann LH, Schlee C, et al. ERG transcriptional networks in primary acute leukemia cells implicate a role for ERG in deregulated kinase signaling. PLoS One. 2013;8(1):e52872. | ||
Halacli SO, Dogan AL. FOXP1 regulation via the PI3K/Akt/p70S6K signaling pathway in breast cancer cells. Oncol Lett. 2015;9(3):1482–1488. | ||
Takayama K, Suzuki T, Tsutsumi S, et al. Integrative analysis of FOXP1 function reveals a tumor-suppressive effect in prostate cancer. Mol Endocrinol. 2014;28(12):2012–2024. | ||
Bilke S, Schwentner R, Yang F, et al. Oncogenic ETS fusions deregulate E2F3 target genes in Ewing sarcoma and prostate cancer. Genome Res. 2013;23(11):1797–1809. | ||
Cerami E, Gao J, Dogrusoz U, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012;2(5):401–404. | ||
Gao J, Aksoy BA, Dogrusoz U, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6(269):pl1. | ||
Nijhawan D, Zack TI, Ren Y, et al. Cancer vulnerabilities unveiled by genomic loss. Cell. 2012;150(4):842–854. | ||
Hanna W, Nofech-Mozes S, Kahn HJ. Intratumoral heterogeneity of HER2/neu in breast cancer--a rare event. Breast J. 2007;13(2):122–129. | ||
de Winter JA, Trapman J, Brinkmann AO, et al. Androgen receptor heterogeneity in human prostatic carcinomas visualized by immunohistochemistry. J Pathol. 1990;160(4):329–332. | ||
Rubin MA, Dunn R, Strawderman M, Pienta KJ. Tissue microarray sampling strategy for prostate cancer biomarker analysis. Am J Surg Pathol. 2002;26(3):312–319. | ||
Kristiansen G, Fritzsche FR, Wassermann K, et al. GOLPH2 protein expression as a novel tissue biomarker for prostate cancer: implications for tissue-based diagnostics. Br J Cancer. 2008;99(6):939–948. |
Supplementary material
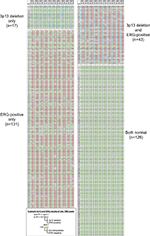  | Figure S1 3p13 deletion and ERG expression status of all 317 arrayed prostate cancers. Shown are the results of all ten tissue spots of all cancers. Abbreviations: TMA, tissue microarray. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.