Back to Journals » Journal of Inflammation Research » Volume 14
Current Status of Functional Studies on Circular RNAs in Rheumatoid Arthritis and Their Potential Role as Diagnostic Biomarkers
Received 20 January 2021
Accepted for publication 9 March 2021
Published 30 March 2021 Volume 2021:14 Pages 1185—1193
DOI https://doi.org/10.2147/JIR.S302846
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Professor Ning Quan
Yayun Xu,1– 3 Feihu Chen1– 3
1Department of Epidemiology and Biostatistics, School of Public Health, Anhui Medical University, Hefei, People’s Republic of China; 2The Key Laboratory of Major Autoimmune Diseases of Anhui Province, Anhui Institute of Innovative Drugs, School of Pharmacy, Anhui Medical University, Hefei, People’s Republic of China; 3School of Pharmacy, The Key Laboratory of Anti-Inflammatory and Immune Medicines, Ministry of Education, Hefei, People’s Republic of China
Correspondence: Feihu Chen
Department of Epidemiology and Biostatistics, School of Public Health, Anhui Medical University, 81 Meishan Road, Hefei, 230032, People’s Republic of China
Email [email protected]
Abstract: Circular RNAs (circRNAs), a new class of endogenous non-coding RNAs (ncRNAs), are highly stable and exhibit tissue-specific expression. Accumulating evidence has indicated that circRNAs play crucial roles in the development and progression of multiple diseases. Notably, circRNAs, important epigenetic modulators of gene expression in inflammation and autoimmune regulation, have a close association with the pathogenesis of rheumatoid arthritis (RA). RA, one of the most common systemic autoimmune diseases, is characterized by synovial hyperplasia and inflammation, and cartilage and bone destruction. Here, we focus on the roles of circRNAs in macrophage, synovial tissues, fibroblast-like synoviocytes (FLSs), and cartilage tissues in pathogenesis and progression of RA, highlighting the potential of circRNAs in the blood as diagnostic biomarkers, and aiming at providing new insights into the diagnosis and therapy of this disease.
Keywords: circular RNAs, rheumatoid arthritis, fibroblast-like synoviocyte, macrophage, biomarkers, cartilage, pathogenesis
Introduction
Rheumatoid arthritis (RA), characterized by joint synovitis, synovial hyperplasia, erosion of articular cartilage, and systemic immune and inflammatory manifestations, is one of the most common chronic and debilitating systemic autoimmune diseases that affects approximately 1% of the population worldwide.1,2 Although the survival rate of RA patients has improved, most patients still experience joint damage, disability, and serious illness, which seriously reduce the quality of life of patients, resulting in a large burden of care.3,4 Recent evidence has indicated that environmental factors, susceptibility genes and immune system disorders are all involved in the pathogenesis of RA.5,6 However, the specific mechanisms underlying etiology and development of this disease are not fully understood.
It is well known that early diagnosis and treatment can improve the clinical manifestations and prognosis of patients with RA to a certain extent. The current diagnosis of RA is based primarily on American College of Rheumatology (ACR)/European League Against Rheumatism (EULAR) classification criteria and traditional serological indicators, including anti-cyclic citrullinated peptide antibodies (ACPA) and rheumatoid factor (RF).7 However, several lines of evidence indicate that these diagnostic methods have various disadvantages such as subjective clinical diagnosis or limited sensitivity and specificity regarding early diagnosis, which leads to a high misdiagnosis rate and a worse clinical outcome.8 Therefore, it is of great clinical significance to explore new biomarkers aimed at improving the diagnosis and prognosis evaluation of RA.
Numerous evidence have demonstrated that non-coding RNA (ncRNA) mainly including microRNA (miRNA), circular RNA (circRNA) and long ncRNA (lncRNA) play a vital role in inflammation and immune regulation of autoimmune diseases.9,10 As a novel class of ncRNAs, circRNAs form from the covalent linkage of the 3ʹ and 5ʹ ends to produce a closed loop.11 Emerging evidence has suggested that circRNAs are not only involved in the pathogenesis and development of RA12 but also can be used as new biomarkers for the diagnosis and prognosis of this disease.13 In this review, we will summarize the available knowledge on the categories, biological characteristics, identification and functions of circRNAs, the role and possible mechanism of circRNAs in RA, and the diagnostic potential of circRNAs in RA.
Categories of circRNAs
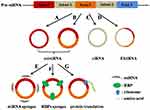
According to their components, three types of circRNAs have been identified by high-throughput sequencing: exon-intron circular RNAs (EIciRNAs),14 exonic circular RNAs (ecircRNAs),15 and circular intronic RNAs (ciRNAs),16 among which ecircRNAs account for over 80% of identified circRNAs (Figure 1). CircRNAs are generated from the precursor messenger RNAs (pre-mRNAs) through backsplicing in most cases, in which a downstream 5ʹ splice donor is linked to an upstream 3ʹ splice acceptor via a 3ʹ→5ʹ phosphodiester bond.17
Biological Characteristics of circRNAs
The biological characteristics of circRNAs can be divided into the following aspects:
(1) High stability. Owning to the covalently closed RNA circle without 5ʹ end caps or 3ʹ poly tails, circRNAs can resist ribonuclease (RNase) digestion and are usually stable with the half-life larger than 48.15,18
(2) Widely expression. CircRNAs can be found in mammals, fungi, plants, and protists. In eukaryotic cells, circRNAs, mainly located in the cytoplasm, have been reported to be expressed in peripheral blood, exosomes, and tissues.19,20
(3) Cell- and tissue-specific expression patterns. The majority of identified circRNAs possess cell- and tissue-specific expression patterns. For example, circRNAs are most abundant in neuronal tissues and accumulate with aging.21
(4) Aberrant expression in diseases. The altered expression of circRNAs between normal and pathological conditions has been observed in numerous diseases.22,23 Therefore, circRNAs (especially in PBMCs) were selected to explore the possibility of being used as biomarkers for the diagnosis of these diseases.
Biological Functions of circRNAs
Recent studies have illustrated that circRNAs function mainly through three different mechanisms (Figure 1).
(1) CircRNAs contain miRNA response elements (MRE), which promotes the combination of circRNAs and miRNAs. Specially, circRNAs work as molecular sponges for miRNAs, thereby decreasing the level of functional miRNAs and regulating the expression of miRNA target gene expression.24
(2) CircRNAs directly affect gene expression by sponging RNA binding proteins (RBPs), the key factors that involve in the post-transcriptional regulation.25
(3) Few circRNAs can serve as templates and be translated into proteins/peptides.26
Identification of circRNAs
There are various methods to identify and study the function of circRNAs.
(1) RNA-seq. RNA-seq is an increasing used methodology to study either coding and non-coding RNA expression. This method can identify new circRNA species and quantify circRNA expression.
(2) Microarray. Microarray, a robust, reliable and high throughput method used for transcriptome profiling, is a low-cost method, which depends on the hybridization probes. The microarray method can only detect and quantify the known circRNAs approximately.
(3) Northern Blot and RT-PCR. These two methods only validate known circRNAs dependent on an exoribonuclease-RNase R.
(4) Fluorescence in situ hybridization. This method can be used to analyze the subcellular localization of circRNAs.
(5) RNA immunoprecipitation and dual-luciferase reporter assay. These two methods can be used to investigate the interaction between circRNAs and miRNAs/RBPs.
Role and Possible Mechanism of circRNAs in RA
As a chronic systemic autoimmune disease, the main pathological characteristics of RA include synovial cell proliferation, multiple inflammatory cell infiltration, pannus formation, and cartilage and bone tissue destruction, which thereby eventually lead to joint deformity and loss of function.2 Owing to their stability, tissue-specific, and disease-specific expression, we will focus on the role and possible mechanism of circRNAs in macrophage, cartilage tissues, synovial tissues, and fibroblast-like synoviocytes (FLSs) in RA.
CircRNAs in Macrophage
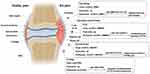
Macrophages, which promote systemic autoimmune diseases and chronic arthritis by linking innate and adaptive immunity, are essential cells in RA.27 Increased numbers of macrophages in RA can infiltrate in synovial tissues and cause joint erosion.28 Abundant activated macrophages in RA synovia have been demonstrated to be an early hallmark of RA, and after effective treatment, the number of these macrophages can be reduced.29 Recently, Yang et al proved that circRNA_09505 in macrophages aggravated inflammation and joint damage.30 Specially, the results of in vitro study showed that the proliferation and cell-cycle progression of macrophages were significantly increased and decreased when circRNA_09505 was overexpressed and knocked down, respectively, indicating that circRNA_09505 was capable of enhancing macrophage proliferation and cell-cycle progression.30 Moreover, the results of in vivo study demonstrated that knockdown of circRNA_09505 in macrophages alleviated inflammation and joint damage in collagen-induced arthritis (CIA) mice,30 the most widely used model for RA.31 Mechanistically, circRNA_09505 regulated macrophage inflammation by functioning as a ceRNA for miR-6089 through AKT1/NF-κB signaling pathway30 (Figure 2).
CircRNAs in Cartilage Tissues
The progressive destruction and loss of articular cartilage is a major clinical manifestation of RA, which is considered to be an urgent clinical problem.32 It has been reported that erosion and loss of cartilage are still in progress even when joint inflammation is successfully suppressed.33,34 Therefore, the key approach in preventing the progression of RA is to elucidate the pathogenesis of the destruction of articular cartilage and find new targets that would control or be used to control the development of RA. A recent study analyzed the difference in the expression of circRNAs in RA cartilage tissue and control specimens.35 The results showed that there were significant differences in the expression of 36 circRNAs between RA and control group, among which circFADS2 showed the most obvious variation in expression.35 Moreover, further clinical studies confirmed the difference in circFADS2 expression in 26 RA patients and 8 healthy controls by reverse transcription-quantitative polymerase chain reaction amplification (RT-qPCR) analysis.35 To uncover the role of circFADS2 in chondrocytes, lipopolysaccharide (LPS)-treated chondrocyte, a well-established RA model for in vitro inflammatory investigation,36 was used. It has been showed that LPS-treated chondrocytes exhibited increased circFADS2 levels when compared to untreated chondrocytes.35 The study of mechanism demonstrated that circFADS2 protected LPS-treated chondrocytes from apoptosis and inflammation acting as an interceptor of the miR-498/mTOR cross-talking35 (Figure 2).
CircRNAs in Synovial Tissues
Synovial hyperplasia is one of the pathologic hallmarks of RA.37 During the development of RA, synovial tissue is a site where a large number of inflammatory cells infiltrate.38 Synovial fibroblasts interact with these inflammatory cells to form a pathological tissue that provokes the destruction of cartilage and bone.39 Recently, Li et al identify the expression pattern of circRNAs in synovial tissues and their inflammatory regulation mechanism.39 It has been revealed that 29 circRNAs were differentially expressed between RA and normal synovial tissues.40 Further functional loss and rescue experiments indicated that circ_0001859 plays a vital role in aggravating the inflammatory process of SW982 cells, a human synovial sarcoma cell line widely used in research examining the mechanism of RA.41,42 Mechanically, circ_0001859 can directly sponge the miR-204/211 family. Knockdown of circ_0001859 upregulated the expression of miR-204/211 and subsequently strengthen the inhibition of its target, ATF2, to suppress its function, and eventually promote inflammation in SW982 cells.40
CircRNAs in Fibroblast-Like Synoviocytes (FLSs)
As one of the main constituent cells in RA synovial tissues, FLSs contribute to joint inflammation and damage by secreting inflammatory mediators, metalloproteinases, and cathepsins, and by invading periarticular cartilage and bone.43 Thus, it will be of great significance to explore the functional and pathological significance of circRNAs in FLSs of RA. Consistent to the result that circ_0088036 is significantly elevated in PBMCs from patients with RA,44 a significant upregulation in the expression level of circ_0088036 was observed in the RA-FLSs compared with osteoarthritis (OA)-FLSs.12 Functionally, the effect of circ_0088036 on the proliferation of RA-FLSs was assessed by the 5-ethynyl-29-deoxyuridine (EdU) assay. EdU is a thymidine analogue which can substitute for thymidine to penetrate into the DNA molecule during DNA replication period. The effect of circ_0088036 on the migration of RA-FLSs was assessed by the transwell migration assay, a well established and frequently used in the field of cell migration research. The results of these two assays revealed that circ_0088036 overexpression could promote the proliferative and the migration ability of the RA-FLSs.12 Mechanistically, circ_0088036 promoted RA progression by sponging miR-140-3p and upregulating SIRT 1 expression.12
These studies shed light on the role of circRNAs in the pathogenesis and development of RA and provide novel insights into the potential role of circRNAs as treatment targets for RA.
Diagnostic Potential of circRNAs in RA
Accumulating evidence has reinforced the argument that circRNAs in peripheral blood can be used as diagnostic biomarkers for RA. CircRNAs can resist RNase digestion and exhibit high stability, which allows circRNAs to be selectively enriched during sample processing.45 Additionally, the advantage of using blood-based markers, including serum/plasma and peripheral blood mononuclear cells (PBMCs), is the ease of sample acquisition and preparation.
Several studies have been conducted to determine the aberrant expressions of circRNAs in PBMCs from RA patients (Table 1). For example, it has been identified and verified that five circRNAs including circ_092516, circ_003524, circ_103047, circ_104871, and circ_101873 were significantly increased in PBMCs from RA patients.44 Based on microarray chip technology and RT-qPCR, it has been found that circ_0092285 and circ_0058794 were significantly higher in PBMC of RA patients, while circ_0088088 and circ_0038644 were significantly lower.46 Recently, Luo et al showed that the expression level of circ_0044235 in peripheral blood from 77 RA patients was decreased significantly compared to 50 healthy controls.47 As one of the most widely studied circRNAs, the expression of ciRS-7 in patients with RA was investigated. It has been shown that the relative expression of ciRS-7 was significantly higher in PBMCs from RA patients.48 More recently, 59 new-onset RA patients were selected to detect the differentially expressed circRNAs between RA patients and healthy controls. The results showed that the levels of circ_0002715, circ_0001947, circ_0000367, and circ_0035197 were significantly increased in the peripheral blood of new-onset RA patients than in the peripheral blood of healthy controls.49 Due to the intrinsic defects of microarrays, such as their inability to detect unidentified circRNAs,50 RNA sequencing was used to investigate the expression profiles of circRNAs in PBMCs from RA patients and healthy controls. The results showed 71 markedly dysregulated circRNAs, including 41 upregulated and 30 downregulated circRNAs. These selected dysregulated circRNAs were verified by RT-qPCR in the PBMCs of 32 RA patients and 20 healthy controls, and the results indicated that circ_0000396 and circ_0130438 were downregulated in the RA group versus the healthy group.51 Similarly, RNA sequencing and RT-qPCR validation were used to identify the differentially expressed circRNAs in RA in another study. RT-qPCR validation demonstrated that the aberrant expression levels of circ_0001200, circ_0001566, circ_0003972 and circ_0008360 were consistent with the results from the sequencing analysis.13
 |
Table 1 Aberrant Expressed circRNAs in PBMCs in Patients with RA |
To evaluate the diagnostic value of PBMC circRNAs in RA, several studies conducted receiver operating characteristic (ROC) analysis to estimate the sensitivity and specificity (Table 2). It has been shown that the area under the curve of circ_104871 in PBMCs was 0.833 with the sensitivity of 83.30% and specificity of 68.00% in distinguishing RA patients from healthy controls.44 Moreover, the area under the curve of circ_0044235 in PBMCs from ROC analysis was 0.757 with a sensitivity of 61.04% and specificity of 82.72% in distinguishing RA patients from healthy controls.46 The combination of multiple circRNAs can improve the sensitivity and specificity of diagnosis, and has been proven to be a more comprehensive and effective disease detection indicator than a single circRNA. Accordingly, Luo et al investigated the predictive power of two circRNAs (circ_0000175 and circ_0008410) in PBMCs as a holistic panel for RA diagnosis.52 The results showed the combined area under the ROC curve was 0.971, with 93.10% sensitivity and 93.33% specificity. Similarly, combined AUC for both circ_104871 and circ_003524 was 0.855, slightly higher than that of circ_10487 (AUC = 0.833).
 |
Table 2 AUC for circRNAs in PBMCs in Discriminating RA Patients from Healthy Controls |
To determine whether the significantly and differentially expressed circRNAs in the RA patients were relevant biomarkers for the disease activity of RA, the correlation between RA clinical features and circRNA levels was assessed (Figure 3). RA is characterized by excessive production of autoantibodies, such as RF and ACPA. RF and ACPA were included in the 2010 American College of Rheumatology/European League Against Rheumatism (ACR/EULAR) classification criteria for RA.53 RFs are autoantibodies against the fragment-crystallizable (Fc) region of IgG that are detected in RA.54 Recently, circRNA_09505 in PBMCs was found to be positively associated with RF in RA.30 ACPA is a clinically useful diagnostic and prognostic marker in RA.55 It has been shown that ACPA can be detected in serum years before the onset of arthritis.56 Circ_0001200,13 circ_0001566,13 and ciRS-748 in PBMCs of RA patients were positively correlated with the levels of ACPA, while circ_00017552 and circ_000836013 were negatively correlated with the levels of ACPA. The 28-joint disease activity scores (DAS-28) are mainly used to assess joint inflammation and structural damage in patients with RA.57 A positive relationship between DAS-28 and circ_0003972 has been reported.13 DAS28 can be characterized using erythrocyte sedimentation rate (ESR) or C-reactive protein (CRP).58 Moreover, ESR and CRP levels are also part of the 2010 ACR/EULAR criteria.53 It has been reported that the expression of circRNA_09505 was positively associated with ESR and CRP levels in serum from RA patients.30 Additionally, circ_0001200 levels13 and circ_000841052 were positively associated with silk time, while circ_0008360 level13 was negatively associated with the silk time. The results of correlation analysis indicated that these circRNAs may play critical roles in the pathogenesis mechanism of RA and be used as a biomarker of disease severity.
Future Directions
(1) Although specific expression profiles of circRNAs in RA have been well documented in many published studies, the sample size is relative small and should be enlarged in a homogeneous population in future multicentral clinical study.
(2) Extracellular vesicles (EVs) and exosomes in synovial fluid have been proposed to contribute to joint inflammation in RA by stimulating the release of pro-inflammatory cytokines.59,60 Given that EVs and exosomes contain a cargo of circRNAs, it is rational to assume that circRNAs in EVs and exosomes in synovial fluid may function as crucial regulators of RA, which needs to be confirmed in the future study.
(3) Currently studies on the aberrant expressions of circRNAs in PBMCs in RA are all cross-sectional, the dynamic changes of circRNA types and levels during the progression of the disease have not been reported.
(4) The regulatory effects of non-steroidal anti-inflammatory drugs (NSAIDs) and disease-modifying anti-rheumatic drugs (DMARDs) on the expression levels of circRNAs for the treatment of RA have not been confirmed. This direction will assist in identifying objective markers that can effectively predict the efficacy of treatment.
(5) Considering the heterogeneous features of RA immunopathogenesis and the critical role of T cell subset, cytokines, and autoantibodies in its pathogenesis, the role of circRNAs in T cell, B cell, and cytokine responses in the context of RA pathogenesis should be investigated.
Conclusions
RA has emerged as an important public health problem, often leading to disability. Current studies have demonstrated that circRNAs are closely involved in the progression of RA and might be promising biomarkers for RA. These findings may provide a new avenue for the prevention, diagnosis and therapeutic intervention of RA in the future.
Funding
This work was supported by the National Natural Science Foundation of China (grant number 81873986).
Disclosure
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
1. Van, der Woude D, van der Helm-van Mil A. Update on the epidemiology, risk factors, and disease outcomes of rheumatoid arthritis. Best Pract Res Clin Rheumatol. 2018;32:174–187. doi:10.1016/j.berh.2018.10.005
2. Aletaha D, Smolen J. Diagnosis and Management of Rheumatoid Arthritis: a Review. JAMA. 2018;320(13):1360–1372. doi:10.1001/jama.2018.13103
3. Conigliaro P, Triggianese P, De Martino E, et al. Challenges in the treatment of Rheumatoid Arthritis. Autoimmun Rev. 2019;18(7):706–713. doi:10.1016/j.autrev.2019.05.007
4. Burmester G, Pope J. Novel treatment strategies in rheumatoid arthritis. Lancet. 2017;389(10086):2338–2348. doi:10.1016/S0140-6736(17)31491-5
5. Veale D, Orr C, Fearon U. Cellular and molecular perspectives in rheumatoid arthritis. Semin Immunopathol. 2017;39:343–354. doi:10.1007/s00281-017-0633-1
6. Alpízar-Rodríguez D, Finckh A. Environmental factors and hormones in the development of rheumatoid arthritis. Semin Immunopathol. 2017;39:461–468. doi:10.1007/s00281-017-0624-2
7. Aletaha D, Neogi T, Silman AJ, et al. 2010 rheumatoid arthritis classification criteria: an American College of Rheumatology/European League Against Rheumatism collaborative initiative. Ann Rheum Dis. 2010;69(9):1580–1588. doi:10.1136/ard.2010.138461
8. Boeters D, Gaujoux-Viala C, Constantin A, et al. The 2010 ACR/EULAR criteria are not sufficiently accurate in the early identification of autoantibody-negative rheumatoid arthritis: results from the Leiden-EAC and ESPOIR cohorts. Semin Arthritis Rheum. 2017;47(2):170–174. doi:10.1016/j.semarthrit.2017.04.009
9. Lodde V, Murgia G, Simula ER, et al. Long noncoding RNAs and Circular RNAs in autoimmune diseases. Biomolecules. 2020;10. doi:10.3390/biom10071044
10. Garo L, Murugaiyan G. Contribution of MicroRNAs to autoimmune diseases. Cell Mol Life Sci. 2016;73:2041–2051. doi:10.1007/s00018-016-2167-4
11. Li I, Chen Y. Emerging roles of circular RNAs in innate immunity. Curr Opin Immunol. 2020;68:107–115. doi:10.1016/j.coi.2020.10.010
12. Zhong S, Ouyang Q, Zhu D, et al. Hsa_circ_0088036 promotes the proliferation and migration of fibroblast-like synoviocytes by sponging miR-140-3p and upregulating SIRT 1 expression in rheumatoid arthritis. Mol Immunol. 2020;125:131–139. doi:10.1016/j.molimm.2020.07.004
13. Wen J, Liu J, Zhang P, et al. RNA-seq reveals the circular RNA and miRNA expression profile of peripheral blood mononuclear cells in patients with rheumatoid arthritis. Biosci Rep. 2020;40(4). doi:10.1042/BSR20193160
14. Li Z, Huang C, Bao C, et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol. 2015;22:256–264. doi:10.1038/nsmb.2959
15. Jeck W, Sorrentino JA, Wang K, et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA. 2013;19(2):141–157. doi:10.1261/rna.035667.112
16. Zhang Y, Zhang X-O, Chen T, et al. Circular intronic long noncoding RNAs. Mol Cell. 2013;51(6):792–806. doi:10.1016/j.molcel.2013.08.017
17. Ebbesen K, Hansen TB, Kjems J, et al. Insights into circular RNA biology. RNA Biol. 2017;14(8):1035–1045. doi:10.1080/15476286.2016.1271524
18. Enuka Y, Lauriola M, Feldman ME, et al. Circular RNAs are long-lived and display only minimal early alterations in response to a growth factor. Nucleic Acids Res. 2016;44(3):1370–1383. doi:10.1093/nar/gkv1367
19. Quan G, Circular LJ. RNAs: biogenesis, expression and their potential roles in reproduction. J Ovarian Res. 2018;11:9. doi:10.1186/s13048-018-0381-4
20. Barrett S, Salzman J. Circular RNAs: analysis, expression and potential functions. Development. 2016;143(11):1838–1847. doi:10.1242/dev.128074
21. Guerra B, Lima J, Araujo BH, et al. Biogenesis of circular RNAs and their role in cellular and molecular phenotypes of neurological disorders. Semin Cell Dev Biol. 2020. doi:10.1016/j.semcdb.2020.08.003
22. Zhang L, Zhang Y, Wang Y, et al. Circular RNAs: functions and clinical significance in cardiovascular disease. Front Cell Dev Biol. 2020;8:584051. doi:10.3389/fcell.2020.584051
23. Limb C, Liu DSK, Veno MT, et al. The Role of Circular RNAs in Pancreatic Ductal Adenocarcinoma and Biliary-Tract Cancers. Cancers. 2020;12:3250. doi:10.3390/cancers12113250
24. Hansen T, Jensen TI, Clausen BH, et al. Natural RNA circles function as efficient microRNA sponges. Nature. 2013;495(7441):384–388. doi:10.1038/nature11993
25. Abdelmohsen K, Panda AC, Munk R, et al. Identification of HuR target circular RNAs uncovers suppression of PABPN1 translation by CircPABPN1. RNA Bio. 2017;14(3):361–369. doi:10.1080/15476286.2017.1279788
26. Wesselhoeft R, Kowalski PS, Anderson DG, et al. Engineering circular RNA for potent and stable translation in eukaryotic cells. Nat. Commun. 2018;9(1):2629. doi:10.1038/s41467-018-05096-6
27. Siouti E, Andreakos E. The many facets of macrophages in rheumatoid arthritis. Biochem Pharmacol. 2019;165:152–169.
28. Udalova I, Mantovani A, Feldmann M, et al. Macrophage heterogeneity in the context of rheumatoid arthritis. Nat Rev Rheumatol. 2016;12(8):472–485. doi:10.1038/nrrheum.2016.91
29. Yang X, Chang Y, Wei W, et al. Emerging role of targeting macrophages in rheumatoid arthritis: focus on polarization, metabolism and apoptosis. Cell Prolif. 2020;53(7):e12854. doi:10.1111/cpr.12854
30. Yang J, Cheng M, Gu B, et al. CircRNA_09505 aggravates inflammation and joint damage in collagen-induced arthritis mice via miR-6089/AKT1/NF-κB axis. Cell Death Dis. 2020;11(10):833. doi:10.1038/s41419-020-03038-z
31. Holmdahl R, Bockermann R, Bäcklund J, et al. The molecular pathogenesis of collagen-induced arthritis in mice–a model for rheumatoid arthritis. Ageing Res Rev. 2002;1:135–147. doi:10.1016/S0047-6374(01)00371-2
32. Hillen J, Geyer C, Heitzmann M, et al. Structural cartilage damage attracts circulating rheumatoid arthritis synovial fibroblasts into affected joints. Arthritis Res Ther. 2017;19(1):40. doi:10.1186/s13075-017-1245-9
33. Rezaei H, Saevarsdottir S, Forslind K, et al. In early rheumatoid arthritis, patients with a good initial response to methotrexate have excellent 2-year clinical outcomes, but radiological progression is not fully prevented: data from the methotrexate responders population in the SWEFOT trial. Ann Rheum Dis. 2012;71(2):186–191. doi:10.1136/annrheumdis-2011-200038
34. Zhou R, Dai -B-B, Xie -Y-Y, et al. Interleukin-1β and tumor necrosis factor-α augment acidosis-induced rat articular chondrocyte apoptosis via nuclear factor-kappaB-dependent upregulation of ASIC1a channel. Biochim Biophys Acta Mol Basis Dis. 2018;1864(1):162–177. doi:10.1016/j.bbadis.2017.10.004
35. Li G, Tan W, Fang Y, et al. circFADS2 protects LPS-treated chondrocytes from apoptosis acting as an interceptor of miR-498/mTOR cross-talking. Aging. 2019;11(10):3348–3361. doi:10.18632/aging.101986
36. Zhang H, Wei Q-F, Wang S-J, et al. LncRNA HOTAIR alleviates rheumatoid arthritis by targeting miR-138 and inactivating NF-κB pathway. Int Immunopharmacol. 2017;50:283–290. doi:10.1016/j.intimp.2017.06.021
37. Edilova MI, Akram A, Abdul-Sater AA. Innate immunity drives pathogenesis of rheumatoid arthritis. Biomed J. 2020.
38. Liu H, Zhu Y, Gao Y, et al. NR1D1 modulates synovial inflammation and bone destruction in rheumatoid arthritis. Cell Death Dis. 2020;11:129. doi:10.1038/s41419-020-2314-6
39. Kok S, Lin L-D, Hou K-L, et al. Simvastatin inhibits cysteine-rich protein 61 expression in rheumatoid arthritis synovial fibroblasts through the regulation of sirtuin-1/FoxO3a signaling. Arthritis Rheum. 2013;65(3):639–649. doi:10.1002/art.37807
40. Li B, et al. Hsa_circ_0001859 Regulates ATF2 expression by functioning as an MiR-204/211 Sponge in human rheumatoid arthritis. J Immunol Res. 2018;2018:9412387. doi:10.1155/2018/9412387
41. Yamazaki T, Yokoyama T, Akatsu H, et al. Phenotypic characterization of a human synovial sarcoma cell line, SW982, and its response to dexamethasone. Vitro Cell Dev Biol Anim. 2003;39:337–339. doi:10.1290/1543-706X(2003)039<0337:PCOAHS>2.0.CO;2
42. Chang J, Lee K, Kim S, et al. Validity of SW982 synovial cell line for studying the drugs against rheumatoid arthritis in fluvastatin-induced apoptosis signaling model. Indian J Med Res. 2014;139:117–124.
43. Rodriguez-Trillo A, Mosquera N, Pena C, et al. Non-Canonical WNT5A signaling through ryk contributes to aggressive phenotype of the rheumatoid fibroblast-like synoviocytes. Front Immunol. 2020;11:555245. doi:10.3389/fimmu.2020.555245
44. Ouyang Q, Wu J, Jiang Z, et al. Microarray Expression Profile of Circular RNAs in Peripheral Blood Mononuclear Cells from Rheumatoid Arthritis Patients. Cell Physiol Biochem. 2017;42(2):651–659. doi:10.1159/000477883
45. Rybak-Wolf A, Stottmeister C, Glažar P, et al. Circular RNAs in the mammalian brain are highly abundant, conserved, and dynamically expressed. Mol Cell. 2015;58(5):870–885. doi:10.1016/j.molcel.2015.03.027
46. Zheng F, Yu X, Huang J, et al. Circular RNA expression profiles of peripheral blood mononuclear cells in rheumatoid arthritis patients, based on microarray chip technology. Mol Med Rep. 2017;16:8029–8036. doi:10.3892/mmr.2017.7638
47. Luo Q, Zhang L, Li X, et al. Identification of circular RNAs hsa_circ_0044235 in peripheral blood as novel biomarkers for rheumatoid arthritis. Clin Exp Immunol. 2018;194:118–124. doi:10.1111/cei.13181
48. Tang X, Wang J, Xia X, et al. Elevated expression of ciRS-7 in peripheral blood mononuclear cells from rheumatoid arthritis patients. Diagn Pathol. 2019;14(1):11. doi:10.1186/s13000-019-0783-7
49. Luo Q, Liu J, Fu B, et al. Circular RNAs Hsa_circ_0002715 and Hsa_circ_0035197 in peripheral blood are novel potential biomarkers for new-onset rheumatoid arthritis. Dis Markers. 2019;2019:2073139. doi:10.1155/2019/2073139
50. Li J, Hou R, Niu X, et al. Comparison of microarray and RNA-Seq analysis of mRNA expression in dermal mesenchymal stem cells. Biotechnol Lett. 2016;38:33–41. doi:10.1007/s10529-015-1963-5
51. Yang X, Li J, Wu Y, et al. Aberrant dysregulated circular RNAs in the peripheral blood mononuclear cells of patients with rheumatoid arthritis revealed by RNA sequencing: novel diagnostic markers for RA. Scand J Clin Lab Invest Suppl. 2019;79(8):551–559. doi:10.1080/00365513.2019.1674004
52. Luo Q, Zeng L, Zeng L, et al. Expression and clinical significance of circular RNAs hsa_circ_0000175 and hsa_circ_0008410 in peripheral blood mononuclear cells from patients with rheumatoid arthritis. Int J Mol Med. 2020;45(4):1203–1212. doi:10.3892/ijmm.2020.4498
53. Van Hoovels L, Jacobs J, Vander Cruyssen B, et al. Performance characteristics of rheumatoid factor and anti-cyclic citrullinated peptide antibody assays may impact ACR/EULAR classification of rheumatoid arthritis. Ann Rheum Dis. 2018;77:667–677. doi:10.1136/annrheumdis-2017-212365
54. Tan E, Smolen J. Historical observations contributing insights on etiopathogenesis of rheumatoid arthritis and role of rheumatoid factor. J Exp Med. 2016;213:1937–1950. doi:10.1084/jem.20160792
55. Murata K, Ito H, Hashimoto M, et al. Fluctuation in anti-cyclic citrullinated protein antibody level predicts relapse from remission in rheumatoid arthritis: kurama cohort. Arthritis Res Ther. 2020;22:268. doi:10.1186/s13075-020-02366-x
56. Kokkonen H, et al. Antibodies of IgG, IgA and IgM isotypes against cyclic citrullinated peptide precede the development of rheumatoid arthritis. Arthritis Res Ther. 2011;13:R13. doi:10.1186/ar3237
57. Hensor E, Emery P, Bingham SJ, et al. Discrepancies in categorizing rheumatoid arthritis patients by DAS-28(ESR) and DAS-28(CRP): can they be reduced? Rheumatology. 2010;49(8):1521–1529. doi:10.1093/rheumatology/keq117
58. Greenmyer J, Stacy J, Sahmoun A, et al. DAS28-CRP cutoffs for high disease activity and remission are lower than DAS28-ESR in rheumatoid arthritis. ACR Open Rheumatol. 2020;2:507–511. doi:10.1002/acr2.11171
59. Foers A, Cheng L, Hill AF, et al. Review: extracellular vesicles in joint inflammation. Arthritis Rheumatol. 2017;69(7):1350–1362. doi:10.1002/art.40076
60. Tavasolian F, Moghaddam AS, Rohani F, et al. Exosomes: effectual players in rheumatoid arthritis. Autoimmun Rev. 2020;19(6):102511. doi:10.1016/j.autrev.2020.102511
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.