Back to Journals » OncoTargets and Therapy » Volume 12
Contribution of interaction between genetic variants of interleukin-11 and Helicobacter pylori infection to the susceptibility of gastric cancer
Authors Liao C, Hu S, Zheng Z, Tong H
Received 2 May 2019
Accepted for publication 24 July 2019
Published 11 September 2019 Volume 2019:12 Pages 7459—7466
DOI https://doi.org/10.2147/OTT.S214238
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Sanjeev K. Srivastava
Chuanwen Liao,1 Shuqin Hu,2 Zihan Zheng,1 Huazhang Tong3
1Department of Gastrointestinal Surgery, People’s Hospital of Jiangxi Province, Nanchang, Jiangxi Province 330006, People’s Republic of China; 2Medical Department, People’s Hospital of Jiangxi Province, Nanchang, Jiangxi Province 330006, People’s Republic of China; 3Department of Radiotherapy, People’s Hospital of Jiangxi Province, Nanchang, Jiangxi Province 330006, People’s Republic of China
Correspondence: Huazhang Tong
Department of Radiotherapy, People’s Hospital of Jiangxi Province, No 92, Aiguo Road, Nanchang, Jiangxi Province 330006, People’s Republic of China
Tel +86 7 918 689 5513
Fax +86 7 918 689 5513
Email [email protected]
Background: Gastric cancer (GC) ranks the second leading cause of cancer-related mortality worldwide. We aimed to clarify the relevance of genetic variants of IL-11, a hub of various carcinogenic pathways, as well as their interactions with Helicobacter pylori (H. pylori) infection in the development of GC.
Methods: A case–control study with 880 GC cases and 900 healthy controls was conducted in a Chinese population. Six tagSNPs were detected by Taqman Allelic Discrimination assay, while H. pylori status was detected by Typing Detection Kit for Antibody to H. pylori and serum IL-11 level was measured using ELISA method.
Results: We found that rs1126760 (C vs T: OR=1.39, 95% CIs=1.13–1.70, P=0.002) and rs1126757 (C vs T: OR=0.82, 95% CIs=0.72–0.93, P=0.002) were significantly associated with susceptibility of GC. Even adjusted for Bonferroni correction, the results were still significant (P=0.002×6=0.012). IL-11 rs1126760 was significantly associated with higher serum and expression level of IL-11, while rs1126757 was significantly associated with lower serum IL-11 level (P<0.001). Significant interaction with H. pylori infection was identified for rs1126760 (P for interaction =0.005). Higher expression of the IL-11 gene was significant with development and poor prognosis of GC.
Conclusion: Our study provides strong evidence that genetic variants of the IL-11 gene may interact with H. pylori infection and contribute to the development of GC. Further studies with larger sample size and functional experiments are needed to validate our findings.
Keywords: gastric cancer, polymorphism, IL-11, Helicobacter pylori
Introduction
Gastric cancer (GC) ranks the second leading cause of cancer-related mortality as well as the fourth most common cancer globally.1,2 Although the largest statistically significant decreases occurred for GC (decrease of 17.1–11% deaths per 100 000) worldwide, it was estimated that 8,65,000 (8,48,300–8,84,700) deaths occurred annually.3 Especially in China, according to the report of Cancer Statistics in China, 2015, 6,79,100 new GC cases and 4,98,000 deaths happened every year, ranking both the second most common cancer and the second leading cause of cancer-related mortality.4 Diet and Helicobacter pylori (H. pylori) infection have been thought to be the important risk factors for GC.5 Besides, genetic factors have also been identified to be associated with susceptibility of GC, and many loci have been identified through genetic epidemiology studies.6–11
IL, a group of cytokines expressed by leukocytes and a member of the IL-6 family of cytokines, regulates tumor-associated inflammation and tumorigenesis making them attractive clues for cancer prevention and targets for adjuvant treatment in cancers.12,13 Among them, IL-11 drives gastric tumorigenesis independent of trans-signaling and acts as a hub of various carcinogenic pathways.14–17 Many oncogenes and tumor suppressor genes function in the process of gastric carcinogenesis, development, invasion and progression through IL-11.15–22 It could promote chronic gastric inflammation and contribute to tumorigenesis mediated by excessive activation of signal transducers and activators of transcription 3 (STAT3) and signal transducers and activators of transcription 1 (STAT1).20 Genetic variants of the IL-11 gene might affect its gene expression and are associated with multiple diseases, including cancers, chronic obstructive pulmonary disease, ulcerative colitis, osteoarthritis, repeated implantation failure and pregnancy loss.23–30 However, no studies have evaluated the effect of IL-11 polymorphisms on development of GC. In this case–control study, we investigated the genetic associations and interactions between genetic variants of IL-11 and H. pylori infection in the development of GC in a Chinese population.
Patients and methods
Study subjects
Totally included in this study were 880 histologically diagnosed GC patients who were recruited between July 2010 and July 2017. None of the included patients had either a previous history of tumors or a history of chemotherapy and radiotherapy. Nine hundred age- and gender matched-cancer-free controls who had no clinical history of gastroduodenal disease were randomly selected from the subjects who visited the health checkup clinics. Demographic and clinical data were collected from medical records, while 5 mL venous blood was collected from all subjects for analyzing their genetic variations and the H. pylori infection status. The study protocol was approved by the Ethics Committee of People’s Hospital of Jiangxi Province and conducted in compliance with the Declaration of Helsinki. All participants provided written informed consent.
TagSNP selection and genotyping
TagSNPs of the IL-11 gene were selected using Haploview 4.2 based on the 1000 Genomes Project database (http://www.1000genomes.org) with minor allele frequency >0.05 in the Chinese population as well as a threshold of r2>0.8. Thus, six SNPs, including rs1042505, rs1126760, rs7250912, rs4252556, rs8104023 and rs1126757, were finally selected in the current study. Genomic DNA was isolated from peripheral blood using the QIAamp DNA Blood Maxi Kit (Qiagen, Hilden, Germany) according to the manufacturer’s protocol. Genotyping for all the six SNPs was carried out by Taqman Allelic Discrimination assay using the Quantstudio 12 Kflex (Applied Biosystems, Foster City, CA). Ten percent randomly selected samples were detected in duplicates and the concordance rate was 100%. All laboratory personnel were blinded to the disease status of the study subjects.
H. pylori infection multiplex serology and serum IL-11 assay
We determined the serostatus of antibodies to four H. pylori specific antigens (CagA, VacA, UreA and UreB) using Typing Detection Kit for Antibody to H. pylori (Shenzhen Blot Biotech Co., Ltd, Shenzhen, China) according to the manufacturer’s instructions. The H. pylori seropositivity was defined as any of the positivity of the four antigens. The serum IL-11 levels of 100 randomly selected controls were measured using enzyme-linked immunosorbent assay (ELISA) method.
Bioinformatics analysis
The comparison of expression of IL-11 gene in GC tissues was analyzed using GEPIA.31 The association of expression of the IL-11 gene survival of GC was analyzed using Kaplan–Meier plotter.32 The genotype-based mRNA expression analysis of IL-11 was conducted using GTEx portal (https://www.gtexportal.org/) as described previously.33,34
Statistical analysis
The proportions of selected variables in GC cases and healthy controls were compared by the χ2 test. Hardy–Weinberg equilibrium (HWE) was evaluated by Pearson’s goodness-of-fit Chi-square (χ2) test for all tagSNPs. The OR and 95% CI were calculated to evaluate the associations between the genetic variants of the IL-11 gene and GC risk by logistic analysis adjusted for age, gender, smoking and drinking status, H. pylori infection, and education level. The false-positive report probability (FPRP) was calculated to evaluate the significant findings as previously.35 We set 0.2 as an FPRP threshold and assigned a prior probability of 0.1 to detect an OR of 0.67/1.50 (protective/risk effects). FPRP value less than 0.2 was considered a noteworthy finding. Gene–environmental interactions in GC were tested on multiplicative scales using a likelihood ratio test. All statistical analyses were performed using Stata 12.0 software (StataCorp, College Station, TX, USA). All statistical tests were two-tailed, and a threshold for significance was set at P<0.05.
Results
Characteristics of the study population
A total of 880 GC cases and 900 healthy controls were enrolled in this case–control study, respectively. As shown in Table 1, we presented the distributions of selected variables in GC cases and healthy controls. Age and gender were comparable, which means the credibility of the matching effect between the two groups. Compared with the healthy controls, GC cases are more likely to be smokers, drinkers, H. pylori carriers,and have a lower education level.
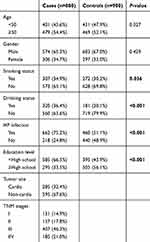 |
Table 1 Distributions of selected variables in GC cases and healthy controls |
Associations between IL-11 gene polymorphisms and susceptibility of GC
The genotype distributions of the enrolled polymorphisms of the IL-11 gene are summarized in Table 2. All tested genotypes of each polymorphism in controls did not deviate from HWE (p>0.05). Among the six tagSNPs, we found that rs1126760 (C vs T: OR =1.39, 95% CIs =1.13–1.70, P=0.002) and rs1126757 (C vs T: OR=0.82, 95% CIs =0.72–0.93, P=0.002) were significantly associated with susceptibility of GC. Even adjusted for Bonferroni correction, the results were still significant (P=0.002×6=0.012). For rs1126760, carriers of genotype TC (OR =1.33; 95%=1.05–1.69; P=0.016) and CC (OR =2.69; 95%=1.26–5.72; P=0.010) have a higher GC risk, compared with carriers of genotype TT. While for rs1126760, carriers of genotype TC (OR =0.85; 95%=0.73–0.99; P=0.038) and CC (OR =0.63; 95%=0.44–0.90; P=0.012) have a lower GC risk, compared with carriers of genotype TT. Results of the dominant and recessive model for rs1126760 and rs1126757 were also significant (P<0.05). However, we did not find any significant associations for rs1042505, rs7250912, rs4252556 and rs8104023 in any genetic models. For the positive results, FPRP was calculated (Table 3). Noteworthy findings were detected for 3 comparisons of rs1126760 (TC vs TT, C vs T and dominant model) and 2 comparisons of rs1126757 (C vs T and dominant model).
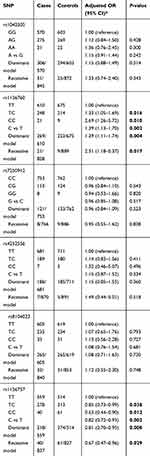 |
Table 2 Genetic variants of the IL-11 gene and susceptibility of GC |
 |
Table 3 False‐Positive Report Probability values for associations between genetic variants of the IL-11 gene and susceptibility of GC |
Interaction analyses between IL-11 gene polymorphisms and H. pylori infection
In order to evaluate the effects of the gene–environmental interaction between IL-11 polymorphisms and H. pylori infection on the susceptibility of GC, analyses of joint effects were performed for the two promising SNPs (Table 4). Significant interaction with H. pylori infection was identified for rs1126760 (P for interaction =0.005). We found 3.35-fold (95% CIs: 2.69–4.18) elevated GC risk for subjects with genotype TC+CC and with H. pylori infection. We did not find any significant interaction for SNP rs1126757.
 |
Table 4 Effects of interactions between HP infection and genetic variants of the IL-11 gene and susceptibility of GC |
Associations between IL-11 gene polymorphisms and serum IL-11 level, and mRNA expression correlation analysis of IL-11
As shown in Table 5, we analyzed the associations between IL-11 gene polymorphisms and serum IL-11 level in control. IL-11 rs1126760 was significantly associated with higher serum IL-11 level, while rs1126757 was significantly associated with lower serum IL-11 level (P<0.001). In the mRNA expression correlation analysis of IL-11, we also found that minor allele of rs1126760 was associated with a higher expression level of IL-11 in testis tissues (Figure 2).
 |
Table 5 Serum IL-11 levels in healthy controls |
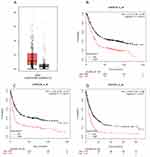
Expression of IL-11 gene with development and prognosis of GC
Figure 1 presents the association of expression of IL-11 gene with the development and prognosis of GC. The expression of the IL-11 gene in GC tissues was significantly higher than that in adjacent normal tissues (Figure 1A, P<0.001). We also found that the expression of the IL-11 gene was significantly associated with overall survival, first progression and post-progression survival of GC (Figure 1B–D, P<0.001).
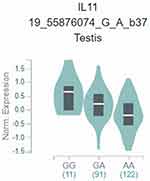 |
Figure 2 mRNA expression correlation analysis of IL-11. |
Discussion
IL-11 functions as a hub of various carcinogenic pathways and plays an essential role in the carcinogenesis of GC. In the current study, we first explored the genetic associations of the IL-11 gene as well as its interaction with H. pylori infection in the development of GC in a Chinese population. We found that IL-11 rs1126760 and rs1126757 were significantly associated with susceptibility of GC and higher serum and expression level of IL-11. Significant interaction with H. pylori infection was identified for rs1126760. Bioinformatics analyses revealed that the expression of IL-11 gene was significantly associated with the development and prognosis of GC.
IL-11 functions in many carcinogenic pathways of the cancers and acts as a function hub of many oncogenes. Since first described for its function and molecular structure by Kawashima et al36 in 1992, many investigators have focused on its biological functions in carcinogenesis.13,20,21,37–40 IL-11 was considered as a potent anti-melanoma factor by Dams-Kozlowska.20,37 It could up-regulate the invasive and proliferative activity of human colorectal carcinoma cells.39 It was also a crucial cytokine promoting chronic gastric inflammation and associated tumorigenesis mediated by excessive activation of STAT3 and STAT1.20 The IL-6 family cytokine IL-11, more than a sidekick, has linked inflammation to cancer and may represent novel, therapeutic targets.13 IL-11 was involved in a variety of gastrointestinal malignancies and laid down a framework for its potential inhibition in many human cancers.40 Results in the current study also revealed that expression of IL-11 gene in tissues was significantly associated with development and prognosis of GC and provided strong evidence for the crucial role of IL-11 gene in the carcinogenesis process of GC.
In this study, IL-11 rs1126760 and rs1126757 were significantly associated with serum IL-11 level and susceptibility of GC. SNP rs1126760 (T/C) was located in the 3ʹ UTR region of the IL-11 gene, which would result in a target loss for hsa-miR-371a-5p.41 Kim et al26 found that rs1126760 was significantly associated with increased risk of Hirschsprung disease. Medrano et al42 reported its association with response to infliximab in Crohn’s disease. SNP rs1126757 (A82A) was located in the exon 3 of the IL-11 gene. Different from our results, Kim et al26 found that rs1126757 was significantly associated with increased risk of Hirschsprung disease. Differences in IL-11 after treatment were found to be related to rs1126757.43 A CpG unit by rs1126757 interaction predictor of antidepressant response was also identified by Powell et al44.
Strength for this study included the following items. First, the large sample size ensured the enough statistical power for the finding of rs1126760 (93.8%). Second, interactions between the IL-11 gene polymorphisms and H. pylori infection were evaluated on multiplicative scales, resulting in one significantly positive interaction for rs1126760. Several limitations should be addressed in this study, including the potential selection bias for case–control study and moderate sample size for interaction analyses.
Conclusively, our study provides strong evidence that genetic variants of the IL-11 gene may interact with H. pylori infection and contributes to the development of GC. Our results increased the understanding of the possible mechanism of IL-11 gene in the carcinogenesis and development of GC. Further studies with larger population and laboratory-based functional experiments are needed to validate our findings.
Acknowledgment
This work was supported by a grant from Natural Science Foundation of Jiangxi Province (No. 20132BAB205108).
Disclosure
The authors report no conflicts of interest in this work.
References
1. Ang TL, Fock KM. Clinical epidemiology of gastric cancer. Singapore Med J. 2014;55(12):621–628.
2. Sitarz R, Skierucha M, Mielko J, Offerhaus GJA, Maciejewski R, Polkowski WP. Gastric cancer: epidemiology, prevention, classification, and treatment. Cancer Manag Res. 2018;10:239–248. doi:10.2147/CMAR.S149619
3. Collaborators GBDCoD. Global, regional, and national age-sex-specific mortality for 282 causes of death in 195 countries and territories, 1980-2017: a systematic analysis for the Global Burden of Disease Study 2017. Lancet. 2018;392(10159):1736–1788. doi:10.1016/S0140-6736(18)32203-7
4. Chen W, Zheng R, Baade PD, et al. Cancer statistics in China, 2015. CA Cancer J Clin. 2016;66(2):115–132. doi:10.3322/caac.21338
5. Rawla P, Barsouk A. Epidemiology of gastric cancer: global trends, risk factors and prevention. Prz Gastroenterol. 2019;14(1):26–38. doi:10.5114/pg.2018.80001
6. Berthenet E, Yahara K, Thorell K, et al. A GWAS on Helicobacter pylori strains points to genetic variants associated with gastric cancer risk. BMC Biol. 2018;16(1):84. doi:10.1186/s12915-018-0550-3
7. Gu D, Zheng R, Xin J, et al. Evaluation of GWAS-identified genetic variants for gastric cancer survival. EBioMedicine. 2018;33:82–87. doi:10.1016/j.ebiom.2018.06.028
8. Yu F, Tian T, Deng B, et al. Multi-marker analysis of genomic annotation on gastric cancer GWAS data from Chinese populations. Gastric Cancer. 2019;22(1):60–68. doi:10.1007/s10120-018-0841-y
9. Cai M, Dai S, Chen W, et al. Environmental factors, seven GWAS-identified susceptibility loci, and risk of gastric cancer and its precursors in a Chinese population. Cancer Med. 2017;6(3):708–720. doi:10.1002/cam4.1038
10. He J, Zhuo ZJ, Zhang A, et al. Genetic variants in the nucleotide excision repair pathway genes and gastric cancer susceptibility in a southern Chinese population. Cancer Manag Res. 2018;10:765–774. doi:10.2147/CMAR.S160080
11. He J, Qiu LX, Wang MY, et al. Polymorphisms in the XPG gene and risk of gastric cancer in Chinese populations. Hum Genet. 2012;131(7):1235–1244. doi:10.1007/s00439-012-1152-8
12. Grivennikov SI, Karin M. Inflammatory cytokines in cancer: tumour necrosis factor and interleukin 6 take the stage. Ann Rheum Dis. 2011;70 Suppl 1:i104–i108. doi:10.1136/ard.2010.140145
13. Putoczki T, Ernst M. More than a sidekick: the IL-6 family cytokine IL-11 links inflammation to cancer. J Leukoc Biol. 2010;88(6):1109–1117. doi:10.1189/jlb.0410226
14. Balic JJ, Garbers C, Rose-John S, Yu L, Jenkins BJ. Interleukin-11-driven gastric tumourigenesis is independent of trans-signalling. Cytokine. 2017;92:118–123. doi:10.1016/j.cyto.2017.01.015
15. Buzzelli JN, O’Connor L, Scurr M, et al. Overexpression of IL-11 promotes premalignant gastric epithelial hyperplasia in isolation from germline gp130-JAK-STAT driver mutations. Am J Physiol Gastrointest Liver Physiol. 2019;316(2):G251–G262. doi:10.1152/ajpgi.00304.2018
16. Wang DQ, Ding XP, Yin S, Mao YD. Role of the IL-11/STAT3 signaling pathway in human chronic atrophic gastritis and gastric cancer. Genet Mol Res. 2016; 5(2): gmr7358.
17. Necula LG, Chivu-Economescu M, Stanciulescu EL, et al. IL-6 and IL-11 as markers for tumor aggressiveness and prognosis in gastric adenocarcinoma patients without mutations in Gp130 subunits. J Gastrointestin Liver Dis. 2012;21(1):23–29.
18. Ma J, Song X, Xu X, Mou Y. Cancer-associated fibroblasts promote the chemo-resistance in gastric cancer through secreting IL-11 targeting JAK/STAT3/Bcl2 pathway. Cancer Res Treat. 2019;51(1):194–210. doi:10.4143/crt.2018.031
19. Zhou C, Ji J, Cai Q, et al. MTA2 enhances colony formation and tumor growth of gastric cancer cells through IL-11. BMC Cancer. 2015;15:343. doi:10.1186/s12885-015-1584-3
20. Ernst M, Najdovska M, Grail D, et al. STAT3 and STAT1 mediate IL-11-dependent and inflammation-associated gastric tumorigenesis in gp130 receptor mutant mice. J Clin Invest. 2008;118(5):1727–1738. doi:10.1172/JCI34944
21. Merchant JL. What lurks beneath: IL-11, via Stat3, promotes inflammation-associated gastric tumorigenesis. J Clin Invest. 2008;118(5):1628–1631. doi:10.1172/JCI35344
22. Nakayama T, Yoshizaki A, Izumida S, et al. Expression of interleukin-11 (IL-11) and IL-11 receptor alpha in human gastric carcinoma and IL-11 upregulates the invasive activity of human gastric carcinoma cells. Int J Oncol. 2007;30(4):825–833.
23. Lokau J, Gottert S, Arnold P, et al. The SNP rs4252548 (R112H) which is associated with reduced human height compromises the stability of IL-11. Biochim Biophys Acta Mol Cell Res. 2018;1865(3):496–506. doi:10.1016/j.bbamcr.2017.12.003
24. Klein W, Rohde G, Arinir U, et al. A promotor polymorphism in the Interleukin 11 gene is associated with chronic obstructive pulmonary disease. Electrophoresis. 2004;25(6):804–808. doi:10.1002/elps.200305773
25. Klein W, Tromm A, Griga T, et al. A polymorphism in the IL11 gene is associated with ulcerative colitis. Genes Immun. 2002;3(8):494–496. doi:10.1038/sj.gene.6363897
26. Kim LH, Cheong HS, Shin JG, et al. Genetic variants of IL-11 associated with risk of Hirschsprung disease. Neurogastroenterol Motil. 2015;27(10):1371–1377. doi:10.1111/nmo.12629
27. Haase MG, Schulze A, Grover S, Kemnitz I, Konig IR, Fitze G. GT-repeat extension in the IL11 promoter is associated with Hirschsprung’s disease (HSCR). Gene. 2018;677:163–168. doi:10.1016/j.gene.2018.07.054
28. Tachmazidou I, Hatzikotoulas K, Southam L, et al. Identification of new therapeutic targets for osteoarthritis through genome-wide analyses of UK Biobank data. Nat Genet. 2019;51(2):230–236. doi:10.1038/s41588-018-0327-1
29. Styrkarsdottir U, Lund SH, Thorleifsson G, et al. Meta-analysis of Icelandic and UK data sets identifies missense variants in SMO, IL11, COL11A1 and 13 more new loci associated with osteoarthritis. Nat Genet. 2018;50(12):1681–1687. doi:10.1038/s41588-018-0247-0
30. Turienzo A, Lledo B, Ortiz JA, et al. Prevalence of candidate single nucleotide polymorphisms on p53, IL-11, IL-10, VEGF and APOE in patients with repeated implantation failure (RIF) and pregnancy loss (RPL). Hum Fertil (Camb);2018. 1–6. doi:10.1080/14647273.2018.1524935
31. Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017;45(W1):W98–W102. doi:10.1093/nar/gkx247
32. Szasz AM, Lanczky A, Nagy A, et al. Cross-validation of survival associated biomarkers in gastric cancer using transcriptomic data of 1,065 patients. Oncotarget. 2016;7(31):49322–49333. doi:10.18632/oncotarget.10337
33. Zhuo ZJ, Liu W, Zhang J, et al. Functional polymorphisms at ERCC1/XPF genes confer neuroblastoma risk in Chinese children. EBioMedicine. 2018;30:113–119. doi:10.1016/j.ebiom.2018.03.003
34. Zhu J, Fu W, Jia W, Xia H, Liu GC, He J. Association between NER pathway gene polymorphisms and Wilms tumor risk. Mol Ther Nucleic Acids. 2018;12:854–860. doi:10.1016/j.omtn.2018.08.002
35. He J, Wang MY, Qiu LX, et al. Genetic variations of mTORC1 genes and risk of gastric cancer in an Eastern Chinese population. Mol Carcinog. 2013;52 Suppl 1:E70–79. doi:10.1002/mc.22013
36. Kawashima I, Ohsumi J, Miyadai K, Takiguchi Y. [Function, molecular structure and gene expression of interleukin-11 (IL-11/AGIF)]. Nihon Rinsho. 1992;50(8):1833–1839.
37. Dams-Kozlowska H, Izycki D, Mackiewicz A. IL-11 is a potent anti-melanoma factor. Adv Exp Med Biol. 2001;495:373–377. doi:10.1007/978-1-4615-0685-0_54
38. Singh B, Berry JA, Shoher A, Lucci A. COX-2 induces IL-11 production in human breast cancer cells. J Surg Res. 2006;131(2):267–275. doi:10.1016/j.jss.2005.11.582
39. Yoshizaki A, Nakayama T, Yamazumi K, Yakata Y, Taba M, Sekine I. Expression of interleukin (IL)-11 and IL-11 receptor in human colorectal adenocarcinoma: IL-11 up-regulation of the invasive and proliferative activity of human colorectal carcinoma cells. Int J Oncol. 2006;29(4):869–876.
40. Grivennikov SI. IL-11: a prominent pro-tumorigenic member of the IL-6 family. Cancer Cell. 2013;24(2):145–147. doi:10.1016/j.ccr.2013.07.018
41. Gong J, Tong Y, Zhang HM, et al. Genome-wide identification of SNPs in microRNA genes and the SNP effects on microRNA target binding and biogenesis. Hum Mutat. 2012;33(1):254–263. doi:10.1002/humu.21641
42. Medrano LM, Taxonera C, Gonzalez-Artacho C, et al. Response to infliximab in Crohn’s disease: genetic analysis supporting expression profile. Mediators Inflamm. 2015;2015:318207. doi:10.1155/2015/125380
43. Powell TR, Schalkwyk LC, Heffernan AL, et al. Tumor necrosis factor and its targets in the inflammatory cytokine pathway are identified as putative transcriptomic biomarkers for escitalopram response. Eur Neuropsychopharmacol. 2013;23(9):1105–1114. doi:10.1016/j.euroneuro.2012.09.009
44. Powell TR, Smith RG, Hackinger S, et al. DNA methylation in interleukin-11 predicts clinical response to antidepressants in GENDEP. Transl Psychiatry. 2013;3:e300. doi:10.1038/tp.2013.73
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.

