Back to Journals » International Journal of General Medicine » Volume 14
Construction of a Ferroptosis-Related Gene Signature for Predicting Survival and Immune Microenvironment in Melanoma Patients
Authors Zeng N , Ma L, Cheng Y, Xia Q, Li Y, Chen Y, Lu Z , Lu Q, Jiang F , Luo D
Received 1 July 2021
Accepted for publication 14 September 2021
Published 5 October 2021 Volume 2021:14 Pages 6423—6438
DOI https://doi.org/10.2147/IJGM.S327348
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Scott Fraser
Ni Zeng,1,2 Liwen Ma,3 Yuxin Cheng,1 Qingyue Xia,1 Yueyue Li,1 Yihe Chen,1 Zhiyu Lu,1 Qian Lu,1 Feng Jiang,4 Dan Luo1
1Department of Dermatology, The First Affiliated Hospital of Nanjing Medical University, Nanjing, 210000, People’s Republic of China; 2Department of Dermatology, Affiliated Hospital of Zunyi Medical University, Zunyi, 563000, People’s Republic of China; 3Department of Dermatology, Nanjing Drum Tower Hospital Clinical College of Nanjing Medical University, Nanjing, 210000, People’s Republic of China; 4Department of Neonatology, Obstetrics and Gynecology Hospital of Fudan University, Shanghai, 200011, People’s Republic of China
Correspondence: Dan Luo
Department of Dermatology, The First Affiliated Hospital of Nanjing Medical University, No. 300, Guangzhou Road, Nanjing, 210000, People’s Republic of China
Email [email protected]
Feng Jiang
Department of Neonatology, Obstetrics and Gynecology Hospital of Fudan University, No. 419, Fangxie Road, Shanghai, 200011, People’s Republic of China
Email [email protected]
Objective: In this research, we studied the genes associated with ferroptosis to develop a prognostic model and find out an association with tumor immune microenvironment in skin cutaneous melanoma (SKCM) patients.
Methods: To find SKCM-related ferroptosis genes, we used Cox regression and LASSO approach on 60 genes related to ferroptosis and SKCM-related RNA-seq. Following that, a ferroptosis-related gene signature was created. Time-dependent ROC curve and Kaplan–Meier analysis were calculated to determine its capability of prediction. Besides, several assessments were used to evaluate overall survival (OS), accompanied by the creation of a nomogram for the clinicopathologic factors and the ferroptosis-related gene signature we established. We also investigated the relationship between ferroptosis-related gene signature with three immune checkpoints and immune cell infiltration.
Results: Our prognostic model included two genes (ALOX5, CHAC1). In both TCGA and GEO cohorts, OS was lower in high-risk category. Using our gene signature, we can reliably predict OS. Additionally, our gene signature can predict immune cell infiltration and SKCM immunotherapy response.
Conclusion: Our gene signature has shown to be a reliable predictor of OS, reflect the immune microenvironment, and predict the effectiveness of immunotherapy for SKCM patients.
Keywords: skin cutaneous melanoma, SKCM, ferroptosis, immune microenvironment, immune checkpoint
Introduction
There are over 200,000 new cases diagnosed per year worldwide with skin cutaneous melanoma (SKCM), which is a very aggressive disease.1 Patients who have lived 5 years since the time of diagnosis of advanced melanoma have a 5-year over survival (OS) average of 10–29%, and the total success rate of their form of chemotherapy is less than 20%.1,2 Though great efforts have been devoted to the management of advanced and metastatic SKCM, the treatment and management of it is far less effective.3 Ultraviolet radiation and hereditary predetermination are the major melanoma risk factors.4 Early detection and management of melanoma contribute to improved results as well as new treatments for even more severe stages of the disease. Mortality from melanoma has actually been remaining constant, with relatively little improvement over time, which emphasizes the significance of continuing studies on the molecular mechanisms of melanoma production and clinical goals.5
In the human body, iron is a necessary micronutrient for certain biological processes, for example, cell metabolism, cellular development, and proliferation.6 Iron homeostasis is precisely controlled by iron ingestion, systemic transport, and preservation inside the body.7 In tumor cells, alterations in iron metabolism may cause two-side impact: on the heme synthesis and sequestration, leading to the accumulation of free iron and depletion of hemoglobin.8 Although the majority of tumor cells have an elevated iron requirement, an adequate level in the body can encourage tumor growth and proliferation, iron accumulation above that range may trigger cell death, or cell death may result in membrane lipid peroxidation.9 Ferroptosis may be of use in the management of some cancers. Ferroptosis has gained popularity as a potential cancer cure after the first presentation in 2012.10 A number of experiments have concluded that ferroptosis plays a crucial function in cell death and in tumor inhibition.11 Additional studies have shown the importance of ferroptosis in diagnosis and prognosis,12 but During the formation and progression of the disease, key regulators and pathways of ferroptosis are still unclear.
The term “tumor microenvironment” refers to the immune cells that exist inside a tumor. TIME regulates iron synthesis and homeostasis in the body. Iron homeostasis is often maintained by Th1 cells, macrophages, etc. Further, it was discovered that immunoregulation and ferroptosis worked in concert in TIME.13 The activation of cytotoxicity in tumor cells reveals tumor antigens, enhancing the microenvironment’s immunogenicity and hence the treatment’s effectiveness and a separate research study discovered that activating CD8 T-cells can boost lipid peroxidation activity in the TIME against tumors, and the increased lipid peroxidation of the tumor cells aids the therapeutic action of immunotherapy.14,15
While there are a variety of SKCM signatures, ferroptosis studies are yet to be proven. Ferroptosis models, for the first time, are built from detailed gene expression, but representing the actual physiological status were developed for use in the SKCM population to predict the microenvironment of the patient in the patient cohort. It is possible that this approach may help with making treatment choices in the future.
Materials and Methods
Data Collection
A total of 460 SKCM RNA-seq data and accompanying clinical details were downloaded from TCGA database. RNA-seq data and SKCM clinical details for another 213 samples were extracted from Gene Expression Omnibus (GSE65904) database. TCGA and GEO data are freely accessible. Thus, the present study was excluded from ethics committees of the respective jurisdiction. The present study adheres to the TCGA and GEO guidelines.
PPI Network Construction
For differentially expressed ferroptosis-related genes, a protein-protein interaction (PPI) network was discovered using the STRING database. For exploring the molecular interactions, the Cytoscape bioinformatics tool was used.
Development of a Gene Signature Related to Ferroptosis That Can Predict Prognosis
The search for ferroptosis-related genes of prognostic significance as part of the univariate Cox analysis of OS. By minimizing the chance of overfitting, LASSO method was used to build a predictive signature. The R package “glmnet” was performed for variable chosen and shrinkage. After that, multivariate regression was used to define the model with the lowest criterion score, that is goodness of fit metric.16 Afterward, the ferroptosis gene signature risk score was divided in conjunction with linear combination of the risk factor and the expression equation (β). Risk score = β1 * gene1expression + β2 * gene2expression + β3 * gene3expression + … + βn * gene expression. Using the risk score algorithm, For each patient, a risk score was determined. To divide the patients into high-risk and low-risk categories, we used the median risk score level as a cutoff value. In this analysis, the Kaplan–Meier method was conducted to test the significance of variations in survival time between the high-risk and low-risk classes. The ROC curve (including 1-, 3-, and 5-year survival) was created to represent the ferroptosis-based signature using “survivalROC” R package to show sensitivity and specificity.
Clinical and Pathologic Factors Associated with Gene Signature Related to Ferroptosis
We used “survival” R package to complete the univariate and multivariate studies on gene signature related to ferroptosis and clinicopathologic characteristics in TCGA and GSE65904. Further, various properties were tested to determine whether the gene signature related to ferroptosis was correlated with clinicopathological factors.
Construction of Prognostic Nomogram
In order to provide a quantitative method for predicting the survival risk for SKCM patients, the nomogram was developed by R package “rms” using a ferroptosis signature as well as quantitative data. In the meanwhile, calibration curves were drawn to provide an accurate estimate of predictive and test the accuracy of nomograms.
Gene Set Enrichment Analysis
According to their calculated risk values, the SKCM samples were split into two groups (high and low risk groups). We used GSEA to differentiate between the two groups in order to discover and study the essential mechanism for KEGG pathways.17 The reference gene collection was c2.cp.kegg.v6.2.symbols.gmt, which was annotated.
Relationship of Ferroptosis-Related Gene Signature and the 22 Tumor-Infiltrating Immune Cells
In both datasets, CIBERSORT was used to measure the proportion of tumor infiltrating immune cells. Via the linear support vector principle, CIBERSORT is really effective at analyzing expression matrices of immune cell types.18 The association between 22 different types of tumor-infiltrating immune cells was investigated. An integrated study of Spearman coefficient and Wilcoxon-rank sum was performed to determine the relationship between the 22 tumor infiltrating immune cells.19 We evaluated the association between the risk score and the levels of expression levels of CTLA4, PD-1, and PD-L1, the three main immune checkpoint genes.
Statistical Analysis
Kaplan–Meier study was performed using R packages “survival” and “survminer.” The “survival” package was used to analyze the Cox study. For ROC study, the R package “survivalROC” was used. Statistical significance is shown by a p-value < 0.05.
Results
Patient Characteristics
A total of 460 melanoma samples were obtained from the TCGA database. There were 213 individual samples from the validation dataset. Table S1 includes all the basic demographic information. Figure 1 represents the design of the present analysis.
 |
Figure 1 Flowchart of this study’s analysis protocol. |
Characterization of Ferroptosis-Related Gene Signature
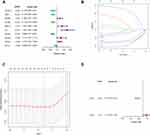
This research includes a total of 60 ferroptosis-related genes (Table S2). A PPI network was built to elucidate the interrelationships among these genes (Figure S1). We developed Kaplan–Meier curves from the TCGA database of ferroptosis-related genes to study OS. 10 genes were strongly related to patient outcome in the Log rank test (p < 0.05) among the 60 genes (Figure 2A). For the purpose of building a ferroptosis-related model, LASSO was applied to select the best optimal model (Figure 2B and C). 8 genes were found with the LASSO algorithms. Finally, a risk model was generated using a multivariate Cox regression analysis. The genes ALOX5 and CHAC1 have been found to be highly predictive (Figure 2D). After calculated the expression equation (β) The following equation is used to calculate the signature’s risk value: risk score = (−0.3258) x expression (ALOX5) + (0.1597) x expression (CHAC1). Among them, it was concluded that a protective effect was demonstrated for ALOX5 had coefficient <0 associated with long OS (Figure S2A). CHAC1 was associated with short OS and coefficient > 0 and considered as a high-risk factor (Figure S2B). Each patient in the TCGA and GEO cohorts was given a risk score, and they were divided into low and high risk categories.
Evaluation of the Risk Model
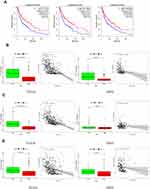
The risk scores were measured and the patients were divided into high- and low-risk categories based on the median level. (Figure 3A). In both TCGA and GSE65904, expression of ALOX5 was increased in conjunction with low risk, seen in heatmap (Figure 3B). The expression of CHAC1 was increased in conjunction with high risk in bot datasets. Patients in the TCGA population get a weaker OS as their risk level rises (Figure 3C). The mortality rate was also greater in the high-risk group, according to our findings (Figure 3D and E). Furthermore, for the prognostic classification of risk score, a ROC study was conducted. We looked at the prognosis prediction classification efficiency at 1, 3, and 5 years. For the survival rates of 1, 3, and 5 years in TCGA cohort, the prognostic signature had AUC values of 0.651, 0.638, and 0.622, respectively. At 1, 3, and 5 years, the AUC values in GEO dataset were 0.560, 0.636, and 0.557 (Figure 3F). Furthermore, for the prognostic classification of risk score, a ROC study was conducted. We looked at the prognosis prediction classification efficiency at 1, 3, and 5 years. For the survival rates of 1, 3, and 5 years in the TCGA cohort, the prognostic signature had AUC values of 0.651, 0.638, and 0.622, respectively. At 1, 3, and 5 years, the AUC values in the GEO dataset were 0.560, 0.636, and 0.557 (Figure 3F). These results showed that the developed prognostic tool has good sensitivity and specificity to estimate SKCM patients.We used immunohistochemistry findings from the Human Protein Atlas database to demonstrate that ALOX5 was significantly increased in normal skin tissue and CHAC1 was significantly increased in melanoma tissue to further establish the expression of two identified genes in the signature (Figure 4).
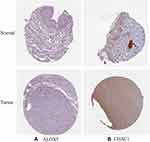 |
Figure 4 The expression of hub ferroptosis-related genes was tested using the HPA database in SKCM and normal tissue. (A) ALOX5 (B) CHAC1. |
The Ferroptosis-Related Signature Studied by Univariate and Multivariate Cox Analysis
The signature model’s independence was determined using univariate and multivariate Cox regression analyses in clinical applications in the TCGA (Figure 5A) and GEO (Figure 5B) datasets. By using univariate Cox study, risk score was positively correlated with prognosis; however, by using multivariate Cox study, it indicated that the signature was an independent prognostic risk factor. Our results showed that the two-gene signature worked effectively in clinical practice.
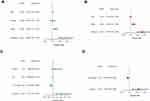 |
Figure 5 In the TCGA and GEO cohorts, Independent prognostic factors for SKCM OS were discovered using univariate and multivariate studies. (A–B) TCGA cohort (C–D) GEO cohort. |
Nomogram Development
We generated nomograms that combined both the ferroptosis-related signature and the typical clinicopathological factors centered on the TCGA cohort (Figure 6A) and GEO cohort (Figure 6E) to estimate OS risk of people with SKCM using a quantitative process. The nomograms had reasonable precision as an optimal model in both the TCGA dataset (Figure 6B–D) and the GEO dataset, according to calibration plots (Figure 6F–H).
Gene Set Enrichment Analysis
We used GSEA to determine between high and low risk groups in terms of biological pathways. In TCGA and GEO cohorts, GSEA research showed the gene sets were greatly enriched in RNA polymerase and Aminoacyl tRNA biosynthesis. Oxidative phosphorylation was also shown to be enriched in the TCGA dataset, as was base excision repair in the GEO dataset (Figure 7).
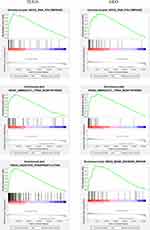 |
Figure 7 GSEA enrichment between groups of low and high risk. |
Identification of the Association Between the Ferroptosis-Related Gene Signature and Immune Cells
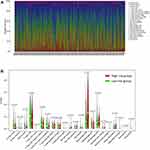
CIBERSORT was performed to better understand how the two-gene signature and the immune microenvironment interacted, and detailed comparisons with the risk score were created. Figure 8A shows the relative content distribution of 22 immune cells in TCGA cohort. Figure 8B shows in high-risk population, the concentrations of Macrophages M0 and Mast cells resting are higher than the other group. In the high-risk population, T cells CD4 memory resting, T cells CD8, T cells CD4 memory activated, and Macrophages M1 were lower than the other group. As seen in Figure 9, tumor-infiltrating immune cells are independent predictors of cancer survival. As a result, we evaluated whether ALOX5 expression is related to the amount of immune infiltration in SKCM. We examined the correlation between gene signatures (ALOX5 and CHAC1) and 24 immune cell subsets in SKCM and discovered that ALOX5 has a strong positive correlation with B cell memory, B cell naive, plasma cells, CD8 T cells, and T cells regulatory; however, ALOX5 has a robust negative correlation with macrophage M2, eosinophils, mast cells resting, and NK cells resting. Another analysis revealed that CHAC1 expression was substantially connected with the infiltration level of activated NK cells (R =0.15, p =0.007), T cell regulatory (R =0.11, p=0.022), and Eosinophils (R =0.11, p=0.022), but not with the infiltration level of T cell memory (R =−0.21, p=0.0001).
 |
Figure 9 Correlation between ALOX5, CHAC1 and infiltrating immune cells in SKCM patients. |
Potential of the Risk Score as an Immunotherapy Response Indicator
The relationship between three immune checkpoint genes and risk score was studied in the TCGA and GEO cohorts. Low levels of PD-1, PD-L1, and CTLA4 demonstrated weak survival, as seen in Figure 10A. In both the TCGA and GEO datasets, we discovered the low-risk population had higher levels of PD-L1, PD-1 and CTLA4 level than high-risk group, that risk score was significantly negatively linked with CTLA4, PD-L1, and PD-1 (Figure 10B–D), suggesting the low-risk group was far more likely to provide an immune response to immunotherapy.
Discussion
Melanoma is the most aggressive form of skin cancer, and its prevalence is on the increase across the world.20 While intense sporadic sun exposure is the most important risk factor for melanoma, other factors such as family background, genetic sensitivity, environmental factors, and immunosuppression often play a role.21 Since SKCM is a molecularly heterogeneous cancerous cancer, its molecular characteristics are linked to biological processes such as cell proliferation, microvascular infiltration, and distance metastasis, and they play a significant role in the prognosis of SKCM.22 As a consequence, It’s critical to define important molecular markers that influence the prognosis of SKCM patients, allowing for better early diagnosis and treatment to improve SKCM clinical outcomes.
The improvement of high-throughput techniques developed has opened up the possibility of discovering new genes implicated in the onset and evolution of SKCM. Ferroptosis entails iron-dependent oxidation and is a mediated autophagic cell death process.23 Excessive intracellular iron storage is caused by disturbances in iron metabolism, which may lead to ferroptosis.24 Several genes influence ferroptosis. Previous research has shown that ferroptosis is an important method for killing SKCM cells, but the exact molecular modifications and mechanism of action remain unknown.
The aim of this research was to identify ferroptosis-related genes that were correlated with SKCM prognosis by analyzing SKCM-related RNA sequences obtained from high-throughput array technologies utilizing Cox proportional hazards regression and LASSO approaches. Previous studies identified several genes, lncRNAs and miRNAs as promising therapeutic biomarkers in SKCM.25–27 However, the differentially expressed signatures were explored between the normal and tumor samples, or between the primary and metastatic tissues, and molecules associated with the progression of cancer were not taken into consideration. Our model is based on the construction of ferroptosis-related genes. We also compared the our model to other researchers such as Shou et al constructed a model based on hypoxic genes, but it did not work well in the validation set and there was no complete 1, 3 and 5 year predictive capability.28 Wu et al constructed a prediction model for SKCM, but the sample size of the validation set was too small to represent the accuracy of the model.29 Two genes (ALOX5, CHAC1) collaborated to create a prognosis model that accurately estimated the prognosis of patients with SKCM, according to our findings. Furthermore, differences in the underlying diseases of SKCM have no impact on the expression features of the two genes, meaning that the prognosis model should be used to determine prognosis in a wide range of SKCM patients. ALOX5 is a member of the arachidonic acid-derived family of proinflammatory lipid mediators. ALOX5 also plays an important role in lipid peroxidation mediation.30 ALOX5 has recently been discovered to play a key role in cell death processes such as apoptosis and ferroptosis.31 CHAC1 is a protein that belongs to the glutamylcyclotransferase family. Deglycination of the Notch receptor, which avoids receptor maturation and reduces Notch signaling, has been shown to facilitate neuronal differentiation by the encoded protein.32 This protein can also be involved in the unfolded protein reaction, glutathione control, and cellular oxidative equilibrium.33 CHAC1 was discovered to digest glutathione, converting it to 5-oxoproline and Cys-Glydipeptide, lowering intracellular GSH levels.34 Increased expression of CHAC1 in breast and ovarian cancer patients may mean a higher risk of cancer recurrence.35 Until now, the mechanism of ALOX5 and CHAC1 in SKCM has remained a mystery.
Since immune cell penetration is essential in tumors, In each SKCM specimen, CIBERSORT was also performed to measure proportional proportion of 22 different types of immune cells.36 According to some data, the interaction between the tumor and the microenvironment is important in the development of SKCM and the likelihood of responding to immunotherapies.37 As a result, we investigated whether a ferroptosis-related gene signature may be used to detect immune cell infiltration. The proportion of T cells CD4 memory resting, T cells CD8, and Macrophages M1 and T cells CD4 memory activated in low-risk group contributed more to immune response than the other group, according to our findings.
Immunotherapy has shone new light on the management of SKCM, with immune checkpoint inhibitors (ICIs) emerging as a theoretically successful treatment option.38 Anti-tumor immunity could be boosted by targeting immune checkpoint molecules.39,40 The association between ferroptosis-related gene signature and ICI reactivity was used to forecast ICI reactivity. The low-risk population had higher levels of PD-L1, PD-1, and CTLA4 expression than the high-risk category. In SKCM patients, low PD-L1, PD-1, and CTLA4 expression are linked to a weak prognosis, indicating that a ferroptosis-related gene signature has the ability to identify immunogenic and ICI-responsive SKCM patients. The therapeutic selection of ICIs in clinical practice is theoretically based on the predictive ability of ferroptosis-related gene signature. Hopefully, this predictive approach can help to speed up the development of personalized cancer immunotherapy.
Additionally, in order to evaluate the prognostic significance of the new risk model, we performed the Log rank test and the ROC curve analysis to investigate the association between the model and clinical parameters.To improve the precision of prognostic prediction, we created and validated a nomogram by combining risk score, era, and level, which could help predict clinical outcomes in SKCM patients. By the use of AUC curves, we next interrogated whether the ferroptosis-related gene patterns could serve as an early predictor for the incidence of SKCM. Our model demonstrated an AUC of 0.651, 0.638, and 0.622 in the TCGA at 1, 3, and 5 years respectively. More specifically, these modern prognostic methods have the potential to not only increase prognostic prediction precision but also to estimate the real mortality risk of particular patients, which is critical in clinical practice. Combining our prognostic model with clinicopathological indications improved prediction sensitivity and specificity for 1-, 3-, and 3-year OS, resulting in better medical therapy. To sum up, our research results indicate that a two-gene prognostic model is a reliable tool for predicting the overall survival of SKCM; it may be useful for guiding therapeutic strategies to improve the clinical outcome of melanoma patients This research has a number of advantages. First, this signature has been thoroughly tested and analyzed through a variety of databases, demonstrating its robustness and durability. Second, an extensive and in-depth study was conducted on a variety of topics, including discussions on the relationship between ferroptosis-related gene signatures and immune cells, as well as immune checkpoints. Third, a nomogram for quantitative measurement was created, which is helpful for clinical promotion and implementation. Nonetheless, there are a few flaws in our study. As a result, further SKCM patients and validations are required to validate this signature in prospective studies. However, there were several limitations to this study. Firstly, it was based solely on the TCGA and GEO databases; so, the finding must be validated using large clinical samples. Furthermore, because this study is based on a retrospective analysis, a prospective study should be conducted to confirm the model. Thirdly, more research into the processes of ALOX5, CHAC1 in SKCM is needed.
Finally, we developed a ferroptosis-based gene signature that is strongly related to the immune microenvironment and can better predict survival and represent immunotherapy efficacy in SKCM patients. The ferroptosis-related gene signature may potentially offer an important method to fulfill the therapeutic criteria of SKCM therapy to some extent in the era of precision medicine.Ferroptosis is a type of cell death that varies from apoptosis in that the formation of iron-dependent lipid peroxides causes it.41 Much or insufficient ferroptosis is associated with a rising number of physiological and pathological processes, as well as dysregulated immune responses.42 Despite being mechanistically revealed in vitro,43,44 accumulating data suggest that ferroptosis may be involved in many pathogenic scenarios.45 Ferroptosis’ role in T cell immunity and cancer immunotherapy, however, is uncertain.
Immune checkpoint blockade medications are novel immunotherapies that selectively activate T cells’ innate ability to attack tumors.46 The important function of iron in tumor development is linked to its potential to modulate both innate and acquired immune responses, particularly in T cells and macrophages. Our findings revealed a strong positive correlation between gene signatures (ALOX5 and CHAC1) and 24 immune cell subsets in SKCM, with ALOX5 having a strong positive correlation with B cell memory, B cell naive, plasma cells, CD8 T cells, and T cells regulatory; however, ALOX5 has a robust negative correlation with macrophage M2, eosinophils, mast cells resting, and NK cells resting. CHAC1 expression was found to be significantly linked to the infiltration levels of activated NK cells, T cell regulatory cells, and Eosinophils, but not to the infiltration level of T cell memory cells, according to another study.
According to a recent study showed the specific makeup of the lymphatic environment may inhibit melanoma cells from undergoing ferroptosis, therefore boosting metastasis.47 The immune system’s interaction with ferroptosis is still unknown. Macrophages have a critical function in iron metabolism regulation.48 ALOX5 was found to be involved in forming leukotriene B4 (LTB4), a pro-inflammatory lipid mediator that acts as a phagocyte chemoattractant in previous investigations.49,50 Researchers also suggested that melanoma’s ferroptosis cells release lipid mediators such LTB4 via ALOX5 to recruit macrophages to the ferroptosis cell location. Previous research has shown that immunotherapy-activated CD8 + T cells make tumors more susceptible to ferroptosis and, as a result, improve immunotherapy efficacy in melanoma patients.51 The era of immunity and iron has dawned in cancer treatment. A potential cancer treatment is ferroptosis-driven nanotherapeutics integrated with immunomodulation.52 Immunotherapy combined with radiotherapy has been shown to trigger ferroptosis and T-cell immunity in tumors. Thus, T cell-promoted tumor ferroptosis is a novel anti-tumor mechanism. Targeting tumor ferroptosis pathway constitutes a therapeutic approach in combination with checkpoint blockade.
Conclusion
In conclusion, we discovered two ferroptosis-related genes in the OS of SKCM with strong predictive capacity, and the prognostic model based on the two genes worked well. The ferroptosis-related gene signature may also reflect the immune microenvironment and the efficacy of immunotherapy in SKCM patients.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Jenkins RW, Fisher DE. Treatment of advanced melanoma in 2020 and beyond. J Invest Dermatol. 2021;141:23–31. doi:10.1016/j.jid.2020.03.943
2. Scolyer RA, Rawson RV, Gershenwald JE, Ferguson PM, Prieto VG. Melanoma pathology reporting and staging. Mod Pathol. 2020;33:15–24. doi:10.1038/s41379-019-0402-x
3. Pham D, Guhan S, Tsao H. KIT and melanoma: biological insights and clinical implications. Yonsei Med J. 2020;61:562–571. doi:10.3349/ymj.2020.61.7.562
4. Namikawa K, Yamazaki N. Targeted Therapy and Immunotherapy for Melanoma in Japan. Curr Treat Options Oncol. 2019;20:7. doi:10.1007/s11864-019-0607-8
5. Davis LE, Shalin SC, Tackett AJ. Current state of melanoma diagnosis and treatment. Cancer Biol Ther. 2019;20:1366–1379. doi:10.1080/15384047.2019.1640032
6. Mou Y, Wang J, Wu J, et al. Ferroptosis, a new form of cell death: opportunities and challenges in cancer. J Hematol Oncol. 2019;12:34. doi:10.1186/s13045-019-0720-y
7. Xu T, Ding W, Ji X, et al. Molecular mechanisms of ferroptosis and its role in cancer therapy. J Cell Mol Med. 2019;23:4900–4912. doi:10.1111/jcmm.14511
8. Zhou B, Liu J, Kang R, Klionsky DJ, Kroemer G, Tang D. Ferroptosis is a type of autophagy-dependent cell death. Semin Cancer Biol. 2020;66:89–100. doi:10.1016/j.semcancer.2019.03.002
9. Conrad M, Pratt DA. The chemical basis of ferroptosis. Nat Chem Biol. 2019;15(12):1137–1147. doi:10.1038/s41589-019-0408-1
10. Friedmann AJ, Krysko DV, Conrad M. Ferroptosis at the crossroads of cancer-acquired drug resistance and immune evasion. Nat Rev Cancer. 2019;19:405–414. doi:10.1038/s41568-019-0149-1
11. Hassannia B, Vandenabeele P, Vanden BT. Targeting Ferroptosis to Iron Out Cancer. Cancer Cell. 2019;35:830–849. doi:10.1016/j.ccell.2019.04.002
12. Tang S, Xiao X. Ferroptosis and kidney diseases. Int Urol Nephrol. 2020;52:497–503. doi:10.1007/s11255-019-02335-7
13. Vitale I, Manic G, Coussens LM, Kroemer G, Galluzzi L. Macrophages and Metabolism in the Tumor Microenvironment. Cell Metab. 2019;30:36–50. doi:10.1016/j.cmet.2019.06.001
14. Roma-Rodrigues C, Mendes R, Baptista PV, Fernandes AR. Targeting Tumor Microenvironment for Cancer Therapy. Int J Mol Sci. 2019;20. doi:10.3390/ijms20040840
15. Lei X, Lei Y, Li JK, et al. Immune cells within the tumor microenvironment: biological functions and roles in cancer immunotherapy. Cancer Lett. 2020;470:126–133. doi:10.1016/j.canlet.2019.11.009
16. Aho K, Derryberry D, Peterson T. Model selection for ecologists: the worldviews of AIC and BIC. Ecology. 2014;95:631–636. doi:10.1890/13-1452.1
17. Thomas MA, Yang L, Carter BJ, Klaper RD. Gene set enrichment analysis of microarray data from Pimephales promelas (Rafinesque), a non-mammalian model organism. Bmc Genomics. 2011;12:66. doi:10.1186/1471-2164-12-66
18. Ali HR, Chlon L, Pharoah PD, Markowetz F, Caldas C. Patterns of Immune Infiltration in Breast Cancer and Their Clinical Implications: a Gene-Expression-Based Retrospective Study. PLoS Med. 2016;13:e1002194. doi:10.1371/journal.pmed.1002194
19. Zhao E, Xie H, Zhang Y. Predicting Diagnostic Gene Biomarkers Associated With Immune Infiltration in Patients With Acute Myocardial Infarction. Front Cardiovasc Med. 2020;7:586871. doi:10.3389/fcvm.2020.586871
20. Ramalingam K, Allamaneni SS. Staging Melanoma: what’s Old and New. Surg Clin North Am. 2020;100:29–41. doi:10.1016/j.suc.2019.09.007
21. Twitty CG, Huppert LA, Daud AI. Prognostic Biomarkers for Melanoma Immunotherapy. Curr Oncol Rep. 2020;22:25. doi:10.1007/s11912-020-0886-z
22. Davis LE, Shalin SC, Tackett AJ. Current state of melanoma diagnosis and treatment. Cancer Biol Ther. 2019;20:1366. doi:10.1080/15384047.2019.1640032
23. Zhang X, Du L, Qiao Y, et al. Ferroptosis is governed by differential regulation of transcription in liver cancer. Redox Biol. 2019;24:101211. doi:10.1016/j.redox.2019.101211
24. Ashrafizadeh M, Mohammadinejad R, Tavakol S, Ahmadi Z, Roomiani S, Katebi M. Autophagy, anoikis, ferroptosis, necroptosis, and endoplasmic reticulum stress: potential applications in melanoma therapy. J Cell Physiol. 2019;234:19471–19479. doi:10.1002/jcp.28740
25. Zhang Q, Wang Y, Liang J, Tian Y, Zhang Y, Tao K. Bioinformatics analysis to identify the critical genes, microRNAs and long noncoding RNAs in melanoma. Medicine. 2017;96:e7497. doi:10.1097/MD.0000000000007497
26. Wei CY, Zhu MX, Lu NH, et al. Bioinformatics-based analysis reveals elevated MFSD12 as a key promoter of cell proliferation and a potential therapeutic target in melanoma. Oncogene. 2019;38:1876–1891. doi:10.1038/s41388-018-0531-6
27. Xu Y, Han W, Xu WH, et al. Identification of differentially expressed genes and functional annotations associated with metastases of the uveal melanoma. J Cell Biochem. 2019;120:19202–19214. doi:10.1002/jcb.29250
28. Shou Y, Yang L, Yang Y, Zhu X, Li F, Xu J. Identification of Signatures of Prognosis Prediction for Melanoma Using a Hypoxia Score. Front Genet. 2020;11:570530. doi:10.3389/fgene.2020.570530
29. Wu XR, Chen Z, Liu Y, et al. Prognostic signature and immune efficacy of m(1) A-, m(5) C- and m(6) A-related regulators in cutaneous melanoma. J Cell Mol Med. 2021;25:8405–8418. doi:10.1111/jcmm.16800
30. Nejatian N, Hafner AK, Shoghi F, Badenhoop K, Penna-Martinez M. 5-Lipoxygenase (ALOX5): genetic susceptibility to type 2 diabetes and vitamin D effects on monocytes. J Steroid Biochem Mol Biol. 2019;187:52–57. doi:10.1016/j.jsbmb.2018.10.022
31. Ivanov I, Golovanov AB, Ferretti C, et al. Mutations of Triad Determinants Changes the Substrate Alignment at the Catalytic Center of Human ALOX5. Acs Chem Biol. 2019;14:2768–2782. doi:10.1021/acschembio.9b00674
32. Chen MS, Wang SF, Hsu CY, et al. CHAC1 degradation of glutathione enhances cystine-starvation-induced necroptosis and ferroptosis in human triple negative breast cancer cells via the GCN2-eIF2alpha-ATF4 pathway. Oncotarget. 2017;8:114588–114602. doi:10.18632/oncotarget.23055
33. Wang N, Zeng GZ, Yin JL, Bian ZX. Artesunate activates the ATF4-CHOP-CHAC1 pathway and affects ferroptosis in Burkitt’s Lymphoma. Biochem Biophys Res Commun. 2019;519:533–539. doi:10.1016/j.bbrc.2019.09.023
34. Ogawa T, Wada Y, Takemura K, et al. CHAC1 overexpression in human gastric parietal cells with Helicobacter pylori infection in the secretory canaliculi. Helicobacter. 2019;24:e12598. doi:10.1111/hel.12598
35. Goebel G, Berger R, Strasak AM, et al. Elevated mRNA expression of CHAC1 splicing variants is associated with poor outcome for breast and ovarian cancer patients. Br J Cancer. 2012;106:189–198. doi:10.1038/bjc.2011.510
36. Ge P, Wang W, Li L, et al. Profiles of immune cell infiltration and immune-related genes in the tumor microenvironment of colorectal cancer. Biomed Pharmacother. 2019;118:109228. doi:10.1016/j.biopha.2019.109228
37. Zhang C, Zheng JH, Lin ZH, et al. Profiles of immune cell infiltration and immune-related genes in the tumor microenvironment of osteosarcoma. Aging. 2020;12:3486–3501. doi:10.18632/aging.102824
38. Darvin P, Toor SM, Sasidharan NV, Elkord E. Immune checkpoint inhibitors: recent progress and potential biomarkers. Exp Mol Med. 2018;50:1–11. doi:10.1038/s12276-018-0191-1
39. Khan M, Arooj S, Wang H, Cell-Based Immune NK. Checkpoint Inhibition. Front Immunol. 2020;11:167. doi:10.3389/fimmu.2020.00167
40. Madden K, Kasler MK. Immune Checkpoint Inhibitors in Lung Cancer and Melanoma. Semin Oncol Nurs. 2019;35:150932. doi:10.1016/j.soncn.2019.08.011
41. Dixon SJ, Lemberg KM, Lamprecht MR, et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell. 2012;149:1060–1072. doi:10.1016/j.cell.2012.03.042
42. Chen X, Kang R, Kroemer G, Tang D. Ferroptosis in infection, inflammation, and immunity. J Exp Med. 2021;218. doi:10.1084/jem.20210518
43. Kagan VE, Mao G, Qu F, et al. Oxidized arachidonic and adrenic PEs navigate cells to ferroptosis. Nat Chem Biol. 2017;13:81–90. doi:10.1038/nchembio.2238
44. Doll S, Proneth B, Tyurina YY, et al. ACSL4 dictates ferroptosis sensitivity by shaping cellular lipid composition. Nat Chem Biol. 2017;13:91–98. doi:10.1038/nchembio.2239
45. Conrad M, Angeli JP, Vandenabeele P, Stockwell BR. Regulated necrosis: disease relevance and therapeutic opportunities. Nat Rev Drug Discov. 2016;15:348–366. doi:10.1038/nrd.2015.6
46. Sanmamed MF, Chen L, Paradigm A. Shift in Cancer Immunotherapy: from Enhancement to Normalization. Cell. 2018;175:313–326. doi:10.1016/j.cell.2018.09.035
47. Ubellacker JM, Tasdogan A, Ramesh V, et al. Lymph protects metastasizing melanoma cells from ferroptosis. Nature. 2020;585:113–118. doi:10.1038/s41586-020-2623-z
48. Shen L, Zhou Y, He H, et al. Crosstalk between Macrophages, T Cells, and Iron Metabolism in Tumor Microenvironment. Oxid Med Cell Longev. 2021;2021:8865791. doi:10.1155/2021/8865791
49. Serezani CH, Lewis C, Jancar S, Peters-Golden M. Leukotriene B4 amplifies NF-kappaB activation in mouse macrophages by reducing SOCS1 inhibition of MyD88 expression. J Clin Invest. 2011;121:671–682. doi:10.1172/JCI43302
50. Afonso PV, Janka-Junttila M, Lee YJ, et al. LTB4 is a signal-relay molecule during neutrophil chemotaxis. Dev Cell. 2012;22:1079–1091. doi:10.1016/j.devcel.2012.02.003
51. Wang W, Green M, Choi JE, et al. CD8(+) T cells regulate tumour ferroptosis during cancer immunotherapy. Nature. 2019;569:270–274. doi:10.1038/s41586-019-1170-y
52. Shan X, Li S, Sun B, et al. Ferroptosis-driven nanotherapeutics for cancer treatment. J Control Release. 2020;319:322–332. doi:10.1016/j.jconrel.2020.01.008
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.





