Back to Journals » Infection and Drug Resistance » Volume 11
Comparative genomics of five different resistance plasmids coexisting in a clinical multi-drug resistant Citrobacter freundii isolate
Authors Ouyang J , Sun F, Zhou D, Feng J, Zhan Z, Xiong Z, Yang B, Liu Z, Li T, Tong Y , Xia P
Received 17 February 2018
Accepted for publication 2 June 2018
Published 12 September 2018 Volume 2018:11 Pages 1447—1460
DOI https://doi.org/10.2147/IDR.S165818
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Eric Nulens
Jing Ouyang,1,2,* Fengjun Sun,1,* Dongsheng Zhou,3 Jiao Feng,3 Zhe Zhan,3 Zhijian Xiong,4 Bo Yang,1 Zhirui Liu,1 Tingbo Li,1 Yigang Tong,3 Peiyuan Xia1
1Department of Pharmacy, Southwest Hospital, The Third Military Medical University, Chongqing 400038, China; 2Department of Pharmacy, Chongqing Public Health Medical Center, Chongqing 400038, China; 3State Key Laboratory of Pathogen and Biosecurity, Beijing Institute of Microbiology and Epidemiology, Beijing 10007, China; 4Outpatient Department of August First Film Studio, Beijing 100161, China
*These authors contributed equally to this work
Background: Plasmid-mediated multi-drug resistance (MDR) has been widely found in Citrobacter freundii. C. freundii P10159 was isolated from a human case of postoperative urinary tract infection in a Chinese teaching hospital.
Methods: The complete nucleotide sequences of five resistance plasmids pP10159-1, pP10159-2, pP10159-3, pP10159-4 and pP10159-5 from C. freundii P10159 were determined through high-throughput genome sequencing, and then compared with related plasmids sequences. Plasmid transfer, CarbaNP test of carbapenemase activity, and bacterial antimicrobial susceptibility test were performed to characterize resistance phenotypes mediated by these plasmids.
Results: pP10159-1 carrying blaNDM-1 and pP10159-2 harboring blaIMP-4 plus qnrS1 were almost identical to IncX3 plasmid pNDM-HN380 and IncN1 plasmid pP378-IMP, respectively. The blaKPC-2-carrying plasmids pP10159-3, pHS062105-3 and pECN49-KPC were highly similar to each other, and constituted a novel group of plasmids belonging to an unknown incomparability group. The MDR plasmids pP10159-4 and pP10159-5 had the backbones highly similar to IncHI4 plasmid pNDM-CIT and type 2 IncC plasmid pR55, respectively, but their accessory resistance regions differed from pNDM-CIT and pR55, respectively. The five plasmids from the P10159 isolate contained a total of 24 different genes or gene loci, which contributed to resistance to 13 distinct antibiotic molecules or toxic compounds.
Conclusion: This is the first report of co-occurrence of five different resistance plasmids, with determination of their complete sequences. Data presented here provide a deeper insight into co-selection and maintenance of multiple plasmids and an extremely large number of resistance genes in a single bacterial isolate.
Keywords: Citrobacter freundii, multi-drug resistance, plasmids, mobile elements
Introduction
Citrobacter freundii, a member of the family Enterobacteriaceae, is widely found in the environment as well as in the intestinal tract of humans and animals. C. freundii is generally considered a low-grade opportunistic pathogen that rarely causes infections, but it has been associated with a wide spectrum of infections of the central nervous system, the respiratory, gastrointestinal, urinary and respiratory tracts, the blood, and many other normally sterile sites in neonates and immunocompromised patients.1
Plasmid-mediated multi-drug resistance (MDR) has been widely found in C. freundii. This study disclosed the co-occurrence of five resistance plasmids pP10159-1, pP10159-2, pP10159-3, pP10159-4 and pP10159-5, containing a total of 24 different resistance markers, in a single clinical C. freundii isolate.
Materials and methods
Bacterial strain
The use of human specimens and all related experimental protocols were approved by the Committee on Human Research of Southwest Hospital, and carried out in accordance with the approved guidelines. The indicated patient signed a written informed consent.
C. freundii P10159 was isolated in 2013 from a mid-stream urine specimen from an esophageal cancer patient with hospital-acquired postoperative urinary tract infection from a teaching hospital in Chongqing City, China. Bacterial species identification was performed using Bruker MALDI Biotyper (Bruker Daltonics) and 16S rRNA gene sequencing.2 The sequence type (ST) of P10159 was determined based on the C. freundii multilocus sequence typing (MLST) scheme (https://pubmlst.org/cfreundii/). All PCR amplicons were sequenced on an ABI 3730 Sequencer.
Genomic DNA sequencing and plasmid sequence assembly
Bacterial genomic DNA was isolated using a Qiagen large construct kit and sequenced from a paired-end library with a mate-pair library with average insert size of 5 kb (ranging from 2 to 10 kb) at a mean coverage 108, using a MiSeq sequencer (Illumina, CA, USA). Reads were trimmed to remove poor quality sequences. In order to get the complete plasmid sequences, DNA contigs were assembled based on their contig coverage using Newbler 2.8.3 Gaps between contigs were filled using a combination of PCR and Sanger sequencing using an ABI 3730 Sequencer.
Sequence annotation and genome comparison
Open reading frames and pseudogenes were predicted using RAST 2.04 combined with BLASTP/BLASTN searches against the UniProtKB/Swiss-Prot5 and RefSeq6 databases. Annotation of resistance genes, mobile elements and other features was carried out using CARD,7 ResFinder,8 ISfinder9 and INTEGRALL.10 Multiple and pairwise sequence comparisons were performed using MUSCLE 3.8.3111 and BLASTN, respectively. Gene organization diagrams were drawn in Inkscape 0.48.1 (https://inkscape.org/en/).
Plasmid conjugal transfer
Plasmid conjugal transfer experiments were carried out with the rifampin-resistant Escherichia coli EC600 being used as recipient and the P10159 strain as donor. Three milliliters of overnight cultures of each of donor and recipient bacteria were mixed together, harvested and resuspended in 80 µL of Brain Heart Infusion (BHI) broth (BD Biosciences). The mixture was spotted on a 1 cm2 hydrophilic nylon membrane filter with a 0.45 µm pore size (Millipore) that was placed on a BHI agar (BD Biosciences) plate and then incubated for mating at 37°C for 12–18 h. Bacteria were washed from the filter membrane and spotted on Muller-Hinton (MH) agar (BD Biosciences) plates containing 1,000 µg/mL rifampin together with indicated additional antibiotics for selecting an E. coli transconjugant carrying one of the following resistance markers: 4 µg/mL meropenem for blaNDM-1 (pP10159-1), blaIMP-4 (pP10159-2) and blaKPC-2 (pP10159-3); 10 µg/mL chloramphenicol for catB3 (pP10159-4); and 10 µg/mL azithromycin for mph(A) (pP10159-5).
Plasmid electroporation
To prepare competent cells for plasmid electroporation, 200 mL of overnight culture of E. coli TOP10 in Super Optimal Broth (SOB) at an optical density (OD600) of 0.4 to 0.6 was washed three times with electroporation buffer (0.5 M mannitol and 10% glycerol) and concentrated into a final volume of 2 mL. One microgram of plasmid DNA, which was isolated from the P10159 strain using a Qiagen Plasmid Midi Kit, was mixed with 100 µL of competent cells for electroporation at 25 µF, 200 Ω and 2.5 Kv. The resulting cells were suspended in 500 µL of SOB and an appropriate aliquot was spotted on SOB agar plates containing the above indicated antibiotics for selecting of an E. coli electroporant carrying one of blaNDM-1, blaIMP-4, blaKPC-2, catB3 and mph(A).
Detection of carbapenemase activity
Activity of class A/B/D carbapenemases in bacterial cell extracts was determined via a modified CarbaNP test.12 Overnight bacterial cell culture in MH broth was diluted 1:100 into 3 mL of fresh MH broth and bacteria were allowed to grow at 37°C with shaking at 200 rpm to reach an OD600 of 1.0 to 1.4. If required, ampicillin was used at 200 µg/mL. Bacterial cells were harvested from 2 mL of the above culture and washed twice with 20 mM Tris-HCl (pH 7.8). Cell pellets were resuspended in 500 µL of 20 mM Tris-HCl (pH 7.8) and lysed by sonication, followed by centrifugation at 10,000 × g at 4°C for 5 minutes. Fifty microliters of the supernatant (the enzymatic bacterial suspension) were mixed with 50 µL of substrate I to V, respectively, followed by incubation at 37°C for 2 hours. Substrate I: 0.054% phenol red plus 0.1 mM ZnSO4 (pH7.8). Substrate II: 0.054% phenol red plus 0.1 mM ZnSO4 (pH7.8) and 0.6 mg/µL imipenem. Substrate III: 0.054% phenol red plus 0.1 mM ZnSO4 (pH7.8), 0.6 mg/µL mg imipenem and 0.8 mg/µL tazobactam. Substrate IV: 0.054% phenol red plus 0.1 mM ZnSO4 (pH7.8), 0.6 mg/µL mg imipenem and 3 mM EDTA (pH7.8). Substrate V: 0.054% phenol red plus 0.1 mM ZnSO4 (pH7.8), 0.6 mg/µL mg imipenem, 0.8 mg/µL tazobactam and 3 mM EDTA (pH7.8).
Bacterial antimicrobial susceptibility test
Bacterial antimicrobial susceptibility was tested by the broth dilution method and interpreted as per Clinical and Laboratory Standards Institute (CLSI) guidelines.13
Nucleotide sequence accession numbers
The pP10159-1, pP10159-2, pP10159-3, pP10159-4 and pP10159-5 sequences were submitted to GenBank under accession numbers MF072961 to MF072965, respectively.
Results
C. freundii co-harboring five resistance plasmids C. freundii
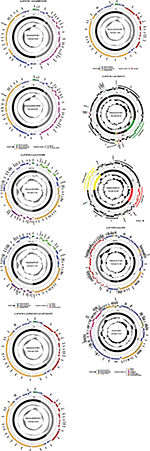
P10159 belonged to a novel ST252, with an allelic profile 108–62–71–7–1–1–57 corresponding to the seven housekeeping genes aspC, clpX, fadD, mdh, arcA, dnaG, and lysP. High-throughput sequencing with the genomic DNA of the P10159 strain revealed the circularly closed sequences of the five plasmids pP10159-1, pP10159-2, pP10159-3, pP10159-4 and pP10159-5, 42.8 kb to 228.7 kb in length with 53 to 270 predicted open reading frames (ORFs) (Figure S1 and Table 1). The modular structure of each plasmid was divided into the backbone regions and separate accessory modules, which were defined as acquired DNA regions associated with and bordered by mobile elements (Figure S1 and Table 1). Some of these accessory modules harbored drug resistance genes (Table 2).
  | Table 1 Major features of plasmids analyzed Notes: pP10159-1, pP10159-2, pP10159-3, pP10159-4 and pP10159-5 from C. freundii P10159 were sequenced in this work, while the other plasmids were derived from GenBank and used as references. Sequence comparison of each of these five sequenced plasmids with their reference plasmid(s) was interpreted in the text. #These accessory modules contained resistance genes as listed in Table 2. Abbreviation: MDR, multi-drug resistant. |
  | Table 2 Drug resistance genes in plasmids analyzed Abbreviation: MDR, multi-drug resistant. |
pP10159-1, pP10159-2 and pP10159-3 could be transferred into E. coli through conjugation or electroporation, which generated the corresponding transconjugants NDM-EC600, IMP-EC600 and KPC-EC600 plus the respective electroporants NDM-TOP10, IMP-TOP10 and KPC-TOP10 (Table 3). This assay indicated that these three plasmids were conjugative. Repeated attempts failed to transfer pP10159-4 or pP10159-5 into E. coli through conjugation and electroporation.
  | Table 3 Antimicrobial drug susceptibility profiles Abbreviations: MIC, minimum inhibitory concentration; S, sensitive; R, resistant; I, intermediately resistant. |
P10159 and all the transconjugants and transformants were resistant to ampicillin, ceftazidime, meropenem and cefoxitin (Table 3). P10159 had class A + B carbapenemase activity, while the transconjugants and transformants harboring pP10159-1 or pP10159-2 had B activity and those harboring pP10159-3 had A activity (Table S1). These resistance phenotypes were consistent with the production of one or all of NDM, IMP and KPC enzymes in the corresponding strains.
Comparison of pP10159-1 with pNDM-HN380
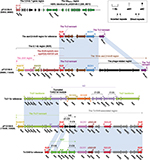
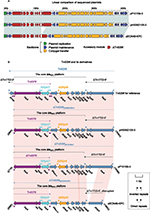
pP10159-1 showed 99% nucleotide identity (with 100% query coverage) to nine blaNDM-carrying IncX3 plasmids, including the first fully sequenced pNDM-HN38014 (Figure 1A). pP10159-1/pNDM-HN380 contained three resistance genes blaNDM-1, ble and blaSHV-12, which were all located in the blaNDM-1 regions (Figure 1B). The blaNDM-1 regions of pP10159-1/pNDM-HN380 were organized in order of a 4.1-kb truncated version of the composite transposon-like IS26-blaSHV-12-IS26 unit,15 a blaNDM-1-containing∆Tn125 element derived from the ISAba125-flanked composite transposon Tn125,16 IS3000 and a 583-bp Tn3 remnant (Figure 1B). An inversion of the truncated IS26-blaSHV-12-IS26 unit in the blaNDM-1 regions represented the only modular difference between pP10159-1 and pNDM-HN380 (Figure 1B).
Comparison of pP10159-2 with pP378-IMP
pP10159-2 displayed >99% nucleotide identity (with >97% query coverage) to 16 blaIMP-carrying IncN1 plasmids, including pIMP-HZ117 and pP378-IMP18 (Figure 2A). Although pIMP-HZ1 was the first fully sequenced blaIMP-carrying IncN1 plasmid, pP378-IMP18 was more appropriate as the reference for genomic comparison because it contained two relatively complete mobile elements, namely a class 1 integron In823 and a truncated Tn3-family unit transposon ΔTn6292,19 harboring blaIMP-4 and qnrS1, respectively. pP10159-2 and pP378-IMP shared the same In823 and also two resistance genes blaIMP-4 and qnrS1.
There were two major modular differences between pP10159-2 and pP378-IMP. First, the CUP (conserved upstream repeat)-controlled regulon18 was composed of seven sequentially arranged operons CUPA, CUPB, CUP5, CUP4, CUP3, CUP2 and CUP1 in the backbone of pP10159-2, while the translocation of the CUP2 operon occurred in pP378-IMP (Figure 2B). Second, the core transposition module tnpA (transposase)-res (resolution site)-tnpR (resolvase) of Tn6292 was interrupted by ISKpn19, breaking tnpA into separate ∆tnpA-3′ and ∆tnpA-5′,18 whereas distinct additional deletion events occurred within the tnpA-res-tnpR modules of the ∆Tn6292 elements from P10159-2 and pP378-IMP (Figure 2C).
Comparison of pP10159-3 with pHS062105-3 and pECN49-KPC
pP10159-3, pHS062105-3 and pECN49-KPC constituted a novel group of plasmids with almost identical backbones (>99% query coverage and >99% nucleotide identity). Their key backbone gene loci included repA for plasmid replication initiation, parA for plasmid partition, a toxin-antitoxin system relEB for post-segregational killing and a P-type IV secretion system for plasmid conjugal transfer (Figures S1C and 3A). Their RepA proteins belonging to the Rep_3 superfamily (pfam10134) and matched the RepA proteins from Klebsiella pneumoniae and Pantoea stewartii with >96% query coverage and >99% amino acid identity; all these RepA proteins could not be assigned into any known incompatibility groups.
The insertion of the core blaKPC platform (ie, the Tn6376 to ΔrepB region) into the cryptic Tn3-family transposon Tn1722, truncating and splitting it into ΔTn1722-5′ and ΔTn1722-3′, generated Tn6296 (Figure 3B) as initially observed in pKP048.20 Each of pP10159-3, pHS062105-3 and pECN49-KPC carried a single accessory region ΔTn6296, which carried the blaKPC-2 gene serving as the sole resistance determinant of these plasmids (Figure 3B). The three ΔTn6296 elements of pP10159-3, pHS062105-3 and pECN49-KPC had undergone deletions and insertions relative to the prototype Tn6296: 1) ΔTn1722-3′ was lost from all the three ΔTn6296 elements; 2) a 123-bp deletion at the 3′-terminal region of Tn6376 was found in pP10159-3; and 3) a 1,754-bp deletion within ΔTn1722-5′ as well as a 73-bp insertion within the variable number tandem repeat (VNTR) region of orf396 was identified in pECN49-KPC. Notably, the above variations within ΔTn6296 accounted for the only modular difference between pHS062105-3, pP10159-3 and pECN49-KPC.
Comparison of pP10159-4 with pNDM-CIT
The pP10159-4 backbone was highly similar (98% query coverage and 98% nucleotide identity) to the reference IncHI4 plasmid pNDM-CIT.19,21 pP10159-4 and pNDM-CIT shared the core IncHI4 backbone gene loci,19 including repHI4A and repHI4B for replication initiation, parAB and parMR for partition, and the tra1 and tra2 regions for conjugal transfer. There were two major modular differences between these two backbones (Figure S2): 1) the deletion of an 11-gene region [downstream of ISKpn26; containing the arsenic resistance (ars) locus] and a 4-gene region (downstream of ISKpn34) from pP10159-4 relative to pNDM-CIT, and 2) the deletion of a distinct 4-gene region (downstream of ISLad2) from pNDM-CIT compared with pP10159-4. These large-fragment deletions might result from the insertion of the corresponding IS elements ISKpn26, ISKpn34 and ISLad2. Of the accessory regions of pP10159-4, only the 25.3-kb MDR region and the 2.9-kb blaCTX-M-3 region contained the resistance genes (Tables 1 and 2).
The MDR region (Figure 4) of pP10159-4 was organized as IS1R, a 14.1-kb 11-gene region encoding unknown functions, ∆IS1R, ISKpn19, ΔISPa38, orf402 (hypothetical protein), orf291 (DNA polymerase), ΔTn6308 and ISKpn19. The unit transposon Tn6308 belonged to the Tn21 subgroup of the Tn3 family and was initially identified in plasmid pP10164-3.22 Tn6308 (Figure 4) had a hybrid backbone, which was composed of the core transposition module tnpAR-res of Tn1696 and the mercury resistance (mer) region of Tn21 and bordered by an intact 38-bp IRL and an IS5075-disrupted IRR (inverted repeat right); the class 1 integron In37 was inserted into the res site of Tn6308, truncating it from an original 120-bp fragment into an 83-bp remnant. Compared with the prototype class 1 integron, In37 in Tn6308 had a very complex mosaic structure and had undergone two major events: 1) an IS26-flanked composite transposon Tn6309, carrying the class C tetracycline resistance module tetA(C)-tetR(C), was inserted into intI1; and 2) an 8.1-kb region (8KR) was inserted at a site between 3′-conserved segment (3′-CS), and the Tn402 tni module, leading to truncation of 3′-CS and tni. 8KR included the chromate-resistance unit IRLchrA-chrA-orf98 and the macrolide-resistance unit IS26-mph(A)-mrx-mphR(A)-IS6100;23 IRLchrA was further disrupted by IS5075 in 8KR. Tn6309, a 217-bp region containing attI1, and a 15.5-kb region (8KR-∆tniA-mer:IS5075-IRRTn6308) were not found in ΔTn6308 of pP10159-4 relative to Tn6308.
ISEcp1 was able to capture and arrange blaCTX-M-3-Δorf477 at its downstream end while it moved,24 while the blaCTX-M-3 region of pP10159-4 was a close derivative of this ISEcp1-base unit with truncation at its 3′-end region.
Comparison of pP10159-5 with pR55
pP10159-5 shared 94% of its backbone with the reference type 2 IncC plasmid pR55,25 with 99% nucleotide identity. All the type 2 IncC signature sequences, including two small insertions i1 and i2 as well as two genes orf1847 and rhs2,26 were found in pR55 and pP10159-5, with the only exception that a 246-bp deletion occurred at the 3′-terminus of rhs2 in pP10159-5 (Figure S1E). pP10159-5 and pR55 possessed the core IncC backbone gene loci including repA for replication initiation, parAB and parM for partition, and the tra1 and tra2 regions for conjugal transfer (Figure S1E).
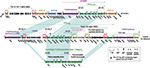
Linear sequence comparison of pP10159-5 and pR55 revealed five different regions (DIERs), namely DIER-1 to DIER-5 (Figure S3). DIER-1 was located between orf564 and orf312, and manifested as the traE and orf225 region composed of a 3′-terminal Tn629218 remnant (designated ΔTn6292; harboring no resistance genes) and eight conjugal transfer genes [found in another type 2 IncC plasmid pSRC119-A/C27 but not in pR55] in pP10159-5; however, DIER-1 was manifested as the orf546 to sul2 region carrying the floR (florfenicol/chloramphenicol resistance)-ISCR2 region and five putative plasmid maintenance genes in pR55. DIER-2 existed as the orf861 to orf852 region (consisting of 34 putative plasmid maintenance genes) in pR55 and was deleted due to the insertion of ISEc52 at a site between traN and orf147 in pP10159-5; the ISEc52 insertion also resulted in the truncation of traN in pP10159-5. The loss of the orf861 to orf852 region, a part of the plasmid maintenance region as observed in pR55, did not destroy the plasmid maintenance of pP10159-5. A 58.2-kb MDR region (DIER-3) was inserted into rhs2 in pP10159-5 relative to pR55, leading to the truncation of rhs2 as well as the deletion of the downstream orf273-orf243 region. The Tn3-family unit transposon Tn6187 (DIER-4; 32.6 kb in length) was inserted into orf492 in pR55 relative to pP10159-5, splitting orf492 into two separate parts and meanwhile leaving 5-bp direct repeats (DRs; target site duplication signals of transposition) at both ends of Tn6187. The traF to orf186 region (DIER-5; containing not only conjugal transfer but plasmid maintenance genes) was located between uvrD and mobI in pR55, but replaced by a second copy of ISEc52 in pP10159-5; the ISEc52 insertion further led to the truncation of uvrD in pP10159-5. These five DIERs were associated with not only accessory modules but backbone regions, and the insertion and deletion events occurred within the two conjugal transfer regions tra1 and tra2 of pP10159-5 would cause this plasmid to be non-conjugative.
All the resistance genes of pP10159-5 were harbored in the MDR region (Figure 5), which was sequentially organized as IS1R, a 3.6-kb 7-gene region of unknown functions, a 9.6-kb blaSFO-1 (β-lactam resistance) region as observed in pHD0149-2,28 the chrA- and mph(A)-carrying 8KR fragment as found in In37 of Tn6308,22 a 15-kb aacC2 (aminoglycoside resistance)-tmrB (tunicamycin resistance) region that was generated from the insertion of a 7-kb phage-related region at a site between ISCfr1 and blaTEM-1 (β-lactamase resistance) of the original aacC2-tmrB region as identified in pEl1573,29 a 7.4-kb Tn21 remnant that was the 3′-terminal IRR to ΔtniA region (containing the mer locus) of Tn2130 with the disruption of IRR by IS4321R and a Tn1548-associated region.
Tn1548 was a IS26-flanked composite transposon lacking DRs at its ends, and had an IS26-In27-ISCR1-ΔISEc28-armA (aminoglycoside resistance)-ISEc29-msr(E)-mph(E) (macrolide resistance)-orf543-repAciN-IS26 structure. There were a number of Tn1548-associated elements with the replacement of In27 by different class 1 integrons,31 eg, In0 carrying no gene cassette array in pP10159-5 (Figure 5).
Remarkably, pP10159-5/pR55 belonged to the same incompatibility group and were genetically very closed related with respect to their plasmid backbones, but they carried totally different profiles of accessory modules that were inserted at different sites of the plasmid backbones; a similar observation was also found for pP10159-4/pNDM-CIT.
Discussion
C. freundii isolates are resistant to cephalosporins due to inducible expression of chromosomally encoded AmpC β-lactamase,32 but in general still remain susceptible to carbapenems. C. freundii can persist in the hosts for long periods, which likely facilitates the acquisition and accumulation of various resistance determinants under high selective pressure within the hospital environments. Carbapenem-resistant C. freundii isolates expressing plasmid-encoding carbapenemases, such as KPC,33 NDM,34 and VIM,35 have emerged during the past decade and, moreover, the coproduction of two or more carbapenemases, eg, KPC-2 + NDM-136 and NDM-1 + VIM-4 + OXA-181,37 have also been identified in C. freundii, making the corresponding isolates highly resistant to β-lactams including carbapenems.
Coexistence of two to four different resistance plasmids has been observed in C. freundii.36,38,39 We recently determined the complete nucleotide sequences of four plasmids p112298-KPC (belonging to an unknown incompatibility group; accession number KP987215),36 p112298-NDM (IncX3 type; KP987216),36 p112298-catA (IncHI2; KY270851) and p112298-tetA (type 1 IncC)40 coexisting in a clinical C. freundii isolate 112,298. In this follow-up study, high-throughput genomic sequencing disclosed the co-occurrence of five resistance plasmids pP10159-1, pP10159-2, pP10159-3, pP10159-4 and pP10159-5 in a clinical C. freundii isolate P10159. This is the first report of co-occurrence of five resistance plasmids of different incomparability groups, with determination of their complete sequences, coexisting in a bacterial isolate.
There is the increasing prevalence of MDR C. freundii strains, which resulted from co-selection of genes encoding resistance to multiple antimicrobial classes, thereby leaving few or no options of antimicrobial treatment.36,38,39 In this study, the five plasmids from C. freundii P10159 contain a total of 24 different genes or gene loci involved in the resistance to β-lactams including carbapenems, aminoglycosides, quinolones, macrolides, phenicols, rifampicin, sulfonamides, tunicamycin, bleomycin, quaternary ammonium compounds, chromate, mercury and tellurium, which are associated with several mobile elements including insertion sequences, integrons and transposons. Notably, the existence of redundant resistance genes made the P10159 strain extremely highly resistant to the corresponding classes of antibiotics, including carbapenems (blaNDM-1, blaIMP-4 and blaKPC-2), aminoglycosides (armA and aacC2), quinolones (qnrS1 and aacA4cr), macrolides [mph(A) and mph(E)] and sulfonamides (two copies of sul1).
The co-selection and maintenance of multiple plasmids and an extremely large number of resistance genes in a single bacterial isolate reflect active and complex horizontal genetic transfer events that have taken place under selective pressures associated with many kinds of antibiotic molecules or toxic compounds.
Acknowledgment
This work was supported by the Foundation of Southwest Hospital (SWH2017JCZD-04) and the National Natural Science Foundation of China (81403164).
Disclosure
The authors report no conflicts of interest in this work.
References
Ranjan KP, Ranjan N. Citrobacter: An emerging health care associated urinary pathogen. Urol Ann. 2013;5(4):313–314. | ||
Frank JA, Reich CI, Sharma S, Weisbaum JS, Wilson BA, Olsen GJ. Critical evaluation of two primers commonly used for amplification of bacterial 16S rRNA genes. Appl Environ Microbiol. 2008;74(8):2461–2470. | ||
Nederbragt AJ. On the middle ground between open source and commercial software - the case of the Newbler program. Genome Biol. 2014;15(4):113. | ||
Brettin T, Davis JJ, Disz T, et al. RASTtk: a modular and extensible implementation of the RAST algorithm for building custom annotation pipelines and annotating batches of genomes. Sci Rep. 2015;5:8365. | ||
Boutet E, Lieberherr D, Tognolli M, et al. UniProtKB/Swiss-Prot, the manually annotated section of the uniprot knowledge base: how to use the entry view. Methods Mol Biol. 2016;1374:23–54. | ||
O’Leary NA, Wright MW, Brister JR, et al. Reference sequence (RefSeq) database at NCBI: current status, taxonomic expansion, and functional annotation. Nucleic Acids Res. 2016;44(D1):D733–D745. | ||
Jia B, Raphenya AR, Alcock B, et al. CARD 2017: expansion and model-centric curation of the comprehensive antibiotic resistance database. Nucleic Acids Res. 2017;45(D1):D566–D573. | ||
Zankari E, Hasman H, Cosentino S, et al. Identification of acquired antimicrobial resistance genes. J Antimicrob Chemother. 2012;67(11):2640–2644. | ||
Siguier P, Perochon J, Lestrade L, Mahillon J, Chandler M. ISfinder: the reference centre for bacterial insertion sequences. Nucleic Acids Res. 2006;34(Database issue):D32–D36. | ||
Moura A, Soares M, Pereira C, Leitão N, Henriques I, Correia A. INTEGRALL: a database and search engine for integrons, integrases and gene cassettes. Bioinformatics. 2009;25(8):1096–1098. | ||
Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32(5):1792–1797. | ||
Chen Z, Li H, Feng J, et al. NDM-1 encoded by a pNDM-BJ01-like plasmid p3SP-NDM in clinical Enterobacter aerogenes. Front Microbiol. 2015;6:294. | ||
CLSI. Performance standards for antimicrobial susceptibility testing: twenty-fifth informational supplement M100-S25. Wayne, PA, USA: CLSI; 2015. | ||
Ho P, Li Z, Lo W, et al. Identification and characterization of a novel incompatibility group X3 plasmid carrying blaNDMin Enterobacteriaceae isolates with epidemiological links to multiple geographical areas in China. Emerg Microbes Infect. 2012;11(11):e39. | ||
Ford PJ, Avison MB. Evolutionary mapping of the SHV beta-lactamase and evidence for two separate IS26-dependent blaSHV mobilization events from the Klebsiella pneumoniae chromosome. J Antimicrob Chemother. 2004;54(1):69–75. | ||
Poirel L, Bonnin RA, Boulanger A, Schrenzel J, Kaase M, Nordmann P. Tn125-related acquisition of blaNDM-like genes in Acinetobacter baumannii. Antimicrob Agents Chemother. 2012;56(2):1087–1089. | ||
Lo WU, Cheung YY, Lai E, Lung D, Que TL, Ho PL, Wu L, Pl H. Complete sequence of an IncN plasmid, pIMP-HZ1, carrying blaIMP-4 in a Klebsiella pneumoniae strain associated with medical travel to China. Antimicrob Agents Chemother. 2013;57(3):1561–1562. | ||
Feng W, Zhou D, Wang Q, et al. Dissemination of IMP-4-encoding pIMP-HZ1-related plasmids among Klebsiella pneumoniae and Pseudomonas aeruginosa in a Chinese teaching hospital. Sci Rep. 2016;6:33419. | ||
Liang Q, Yin Z, Zhao Y, et al. Sequencing and comparative genomics analysis of the IncHI2 plasmids pT5282-mphA and p112298-catA and the IncHI5 plasmid pYNKP001-dfrA. Int J Antimicrob Agents. 2017;49(6):709–718. | ||
Shen P, Wei Z, Jiang Y, et al. Novel genetic environment of the carbapenem-hydrolyzing beta-lactamase KPC-2 among Enterobacteriaceae in China. Antimicrob Agents Chemother. 2009;53(10):4333–4338. | ||
Dolejska M, Villa L, Poirel L, Nordmann P, Carattoli A. Complete sequencing of an IncHI1 plasmid encoding the carbapenemase NDM-1, the ArmA 16S RNA methylase and a resistance-nodulation-cell division/multidrug efflux pump. J Antimicrob Chemother. 2013;68(1):34–39. | ||
Sun F, Zhou D, Sun Q, et al. Genetic characterization of two fully sequenced multi-drug resistant plasmids pP10164-2 and pP10164-3 from Leclercia adecarboxylata. Sci Rep. 2016;6:33982. | ||
Partridge SR. Analysis of antibiotic resistance regions in Gram-negative bacteria. FEMS Microbiol Rev. 2011;35(5):820–855. | ||
Lartigue MF, Poirel L, Aubert D, Nordmann P. In vitro analysis of ISEcp1B-mediated mobilization of naturally occurring beta-lactamase gene blaCTX-M of Kluyvera ascorbata. Antimicrob Agents Chemother. 2006;50(4):1282–1286. | ||
Doublet B, Boyd D, Douard G, Praud K, Cloeckaert A, Mulvey MR. Complete nucleotide sequence of the multidrug resistance IncA/C plasmid pR55 from Klebsiella pneumoniae isolated in 1969. J Antimicrob Chemother. 2012;67(10):2354–2360. | ||
Harmer CJ, Hall RM. The A to Z of A/C plasmids. Plasmid. 2015;80:63–82. | ||
Harmer CJ, Holt KE, Hall RM. A type 2 A/C2 plasmid carrying the aacC4 apramycin resistance gene and the erm(42) erythromycin resistance gene recovered from two Salmonella enterica serovars. J Antimicrob Chemother. 2015;70(4):1021–1025. | ||
Zhao J-Yi, Zhu Y-Qi, Li Y-Nian, et al. Coexistence of SFO-1 and NDM-1 β-lactamase genes and fosfomycin resistance gene fosA3 in an Escherichia coli clinical isolate. FEMS Microbiol Lett. 2015;362(1):1–7. | ||
Partridge SR, Ginn AN, Paulsen IT, Iredell JR. pEl1573 Carrying blaIMP-4, from Sydney, Australia, is closely related to other IncL/M plasmids. Antimicrob Agents Chemother. 2012;56(11):6029–6032. | ||
Liebert CA, Hall RM, Summers AO. Transposon Tn21, flagship of the floating genome. Microbiol Mol Biol Rev. 1999;63(3):507–522. | ||
Du XD, Li DX, Hu GZ, et al. Tn1548-associated armA is co-located with qnrB2, aac(6′)-Ib-cr and blaCTX-M-3 on an IncFII plasmid in a Salmonella enterica subsp. enterica serovar Paratyphi B strain isolated from chickens in China. J Antimicrob Chemother. 2012;67(1):246–248. | ||
Lindberg F, Normark S. Contribution of chromosomal beta-lactamases to beta-lactam resistance in enterobacteria. Rev Infect Dis. 1986;8 Suppl 3(Suppl 3):S292–S304. | ||
Deshpande LM, Rhomberg PR, Sader HS, Jones RN. Emergence of serine carbapenemases (KPC and SME) among clinical strains of Enterobacteriaceae isolated in the United States Medical Centers: report from the MYSTIC Program (1999-2005. Diagn Microbiol Infect Dis. 2006;56(4):367–372. | ||
Rimrang B, Chanawong A, Lulitanond A, et al. Emergence of NDM-1- and IMP-14a-producing Enterobacteriaceae in Thailand. J Antimicrob Chemother. 2012;67(11):2626–2630. | ||
Yan JJ, Ko WC, Chuang CL, Wu JJ, Wc K, Jj W. Metallo-beta-lactamase-producing Enterobacteriaceae isolates in a university hospital in Taiwan: prevalence of IMP-8 in Enterobacter cloacae and first identification of VIM-2 in Citrobacter freundii. J Antimicrob Chemother. 2002;50(4):503–511. | ||
Feng J, Qiu Y, Yin Z, et al. Coexistence of a novel KPC-2-encoding MDR plasmid and an NDM-1-encoding pNDM-HN380-like plasmid in a clinical isolate of Citrobacter freundii. J Antimicrob Chemother. 2015;70(11):2987–2991. | ||
Poirel L, Ros A, Carricajo A, et al. Extremely drug-resistant Citrobacter freundii isolate producing NDM-1 and other carbapenemases identified in a patient returning from India. Antimicrob Agents Chemother. 2011;55(1):447–448. | ||
Wu W, Espedido B, Feng Y, Zong Z. Citrobacter freundii carrying blaKPC-2 and blaNDM-1: characterization by whole genome sequencing. Sci Rep. 2016;6:30670. | ||
Xiong J, Déraspe M, Iqbal N, et al. Genome and plasmid analysis of blaIMP-4-carrying Citrobacter freundii B38. Antimicrob Agents Chemother. 2016;60(11):6719–6725. | ||
Ma L, Yin Z, Zhang D, et al. Comparative genomics of type 1 IncC plasmids from China. Future Microbiol. 2017;12:1511–1522. |
Supplementary material
  | Table S1 Results of modified CARBA-NP test |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.