Back to Journals » International Journal of General Medicine » Volume 13
Combined Expression of CD34 and FLT3-Internal Tandem Duplication Mutation Predicts Poor Response to Treatment in Acute Myeloid Leukemia
Authors Abdellateif MS , Kassem AB , EL-Meligui YM
Received 12 August 2020
Accepted for publication 14 September 2020
Published 16 October 2020 Volume 2020:13 Pages 867—879
DOI https://doi.org/10.2147/IJGM.S276138
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Scott Fraser
Mona S Abdellateif,1 Amira B Kassem,2 Yomna M EL-Meligui3
1Medical Biochemistry and Molecular Biology Unit, Cancer Biology Department, National Cancer Institute, Cairo University, Cairo, Egypt; 2Clinical Pharmacy and Pharmacy Practice Department, Faculty of Pharmacy, Damanhour University, Damanhur, Egypt; 3Clinical Pathology Department, National Cancer Institute, Cairo University, Cairo, Egypt
Correspondence: Mona S Abdellateif
Medical Biochemistry and Molecular Biology, Cancer Biology Department, National Cancer Institute, Cairo University Fax + (202)23644720
Email [email protected]
Background: Acute myeloid leukemia (AML) is a common hematological malignancy associated with different cytogenetic and genetic abnormalities.
Methods: FLT3-internal tandem duplication (FLT3/ITD) mutation and CD34 expression levels were assessed in the bone marrow (BM) aspirates of 153 de novo AML patients. Data were correlated with relevant clinic-pathological features of the patients, response to treatment, disease-free survival (DFS), and overall free survival (OS) rates.
Results: FLT3-ITD mutation was detected in 27/153 (17.6%) AML patients (P=0.001), and CD34 was expressed in 83/153 (54.2%) patients (P=0.293) compared to those with wild FLT3 and CD34− expression, respectively. Patients with FLT3-ITD mutation showed increased peripheral blood and BM blast cells, abnormal cytogenetics, poor DFS and OS compared to those with wild FLT3 (P=0.013, P< 0.001, P=0.010, P=0.008 and P=0.004, respectively), while there was no significant association with response to treatment (P=0.081). There was no significant association between CD34 expression and response to treatment, DFS, and OS (P> 0.05). FLT3-ITD mutation and FAB subtypes were independent prognostic factors for DFS. Older age ≥ 39 years, HB < 7 mg/dL PB blast ≥ 54%, and FLT3-ITD mutation were independent prognostic factors for poor OS in AML patients. The presence of both FLT3-ITD mutation and CD34 expression associated significantly with resistance to therapy (P=0.024), short DFS and OS rates (P=0.006, P=0.037, respectively).
Conclusion: Combined expression of both FLT3-ITD mutation and CD34 expression is an important prognostic and predictive factor for poor disease outcome in AML patients.
Keywords: acute myeloid leukemia, AML, FLT3-ITD, CD34
Introduction
Acute myeloid leukemia (AML) is a hematopoietic malignant disease that affects children and adults.1 It is the most common acute leukemia in adults, with a 5-year overall survival rate of about 27%.2–4 Although 60–70% of patients experienced complete remission after the induction regimen, most of them relapsed within 3 years. This relapse may occur due to the development of CD34+/CD38–AML leukemic stem cells (LSCs),5 which is characterized by self-renewal capacity that continuously produce immature blood cells. These cells harbor frequent mutations that inhibit hematopoietic differentiation pathways, which is one of the hallmarks of AML.6,7 Thus, identification of these functionally distinct CD34+ cell populations is a crucial step to evaluate the disease course and therefore relapse prediction.
CD34 is a glycosylated transmembrane protein and it is a well-established marker for human hematopoietic stem and early progenitor cells.8 The CD34 family proteins are included in enhancing proliferation, blocking differentiation, trafficking, and cell adhesion.9,10 It had been previously reported that CD34+ AML blasts are more resistant to apoptosis than their CD34- counterpart.11
AML is caused by different cytogenetics and genetic abnormalities that play a crucial role in the pathogenesis, progression, and diagnosis of AML patients.12 One of the most common mutated genes in AML is FLT3 (FMS like tyrosine kinase 3), which occurs in about 30% of AML patients with normal cytogenetics.13 FLT3 is a member of the tyrosine kinase receptor III family, and it greatly impacts the proliferation and differentiation of early hematopoietic progenitor cells. It has two types of mutations: 1) internal tandem duplication (FLT3/ITD) within or near the second juxta membrane receptor, which occurs in 15–35% of patients, and 2) the point mutations (FLT3/TKD) leading to amino acid substitutions in the loop of activating tyrosine kinase domain, which occurs in 5–10% of AML patients.13–15
The FLT3 gene is expressed only in immature hematopoietic CD34+ progenitor cells in the bone marrow. It is located on chromosome 13q12 encoding FLT3 protein. Binding of the extracellular domain of FLT3 receptor to its ligand resulted in subsequent signaling pathways activation through JAK/STAT, PI3K/AKT, and MAPK/ERK signal transduction. These pathways regulate the development and function of hematopoietic stem/progenitor cells.16
Therefore, the aim of the current study was to assess FLT3/ITD mutation and CD34 expression levels in newly-diagnosed AML patients. These markers were assessed separately and in combination, in relation to the clinical features of the patients, different chromosomal abnormalities, response to treatment, and survival rates (DFS and OS). This will help for more stratification of the patients, more accurate prediction of heir outcomes, and consequently more appropriate management.
Methods
This prospective cohort study included 153 AML patients who were presented and diagnosed at the National Cancer Institute (NCI) during the period 2013–2015. Control samples were obtained from 20 age- and sex-matched healthy subjects who were donors for bone marrow transplantation (BMT). Patients were subjected to full history taking, clinical and radiological examination. They were confirmed for the diagnosis of AML according to the French-American-British (FAB) and World Health Organization (WHO) criteria.17
Therapeutic Regimen
Patients received their chemotherapy regimen according to the National Comprehensive Cancer Network (NCCN) guidelines.18 The treatment protocol was formed of induction therapy with a 3+7 regimen (daunorubicin 45 mg/m2 and cytosine arabinoside 100 mg/m2) followed by two cycles of a 5+2 protocol after remission (daunorubicin 45 mg/m2 for 2 days, plus cytosine arabinoside 100 mg/m2 for another 5 days). Patients who did not show complete remission (CR) morphologically after the first session of induction chemotherapy received cytosine arabinoside 500 mg/m2 by slow intravenous push twice a day for 7 days and Novantrone 12 mg/m2 daily for 3 days. Patients with acute Promyelocytic leukemia (APL, the M3 subtype of AML) were treated with induction therapy formed of three doses of doxorubicin 60 mg/m2 or idarubicin 12 mg/m2 D1–3 combined with All Trans Retinoic acid (ATRA) 45 mg/m2 in two divided doses, and Arsenic trioxide (ATO) till morphological complete remission (CR) occur, it might continue for a maximum period of 60 days. Patients who achieved CR received consolidation therapy, which was formed of two cycles of three doses of doxorubicin 60 mg/m2 or idarubicin 12 mg/m2 D1–3 in combination with ATRA 45 mg/m2 for 15 days. Then the consolidation therapy is followed by the maintenance therapy, which is formed of law dose of ATRA alone or combined with chemo-therapy (6-mercaptopurine (6-MP) and/or methotrexate). It lasts for about 1 year.
Response to Therapy
Response to therapy was evaluated according to the WHO criteria, where complete remission (CR) was achieved when the patient had a hemoglobin concentration equal to or more than 10 gm/dL, total leukocyte count equal to or more than 3000 (109/L), platelet count equal to or more than 150.000 (109/L), normocellular bone marrow (with normal differential count or presence <5% blast cells), and finally absence of all clinical and radiological manifestations of AML.
Samples Collection
Bone marrow (BM) aspirates were obtained from the participating AML patients and the control subjects. All BM samples were subjected to morphological examination, routine immunophenotyping (IPT), Conventional karyotyping, Fluorescence In Situ Hybridization (FISH), and conventional PCR for common genetic abnormalities including t(8:21), t(9,22), t(15,17), and pericentric inversion of chromosome 16 (inv16).
Flow Cytometer Detection of CD34
PB or BM samples were obtained at diagnosis and analyzed within 24 hours. Briefly, 100 µl the sample with an adjusted cell count of approximately 1x106 cells/tube were incubated with monoclonal antibodies against CD34 according to the manufacturer recommendations in the dark for 30 minutes at room temperature, lysed, and washed with phosphate buffer saline (PBS) to get rid of excess antibodies, then cells were resuspended in PBS.
Mouse anti-human CD34 monoclonal antibodies (catalog no for CD34:550,619) purchased from BD Biosciences (CA, USA) were added at diagnosis. Isotype/negative control (IgG1) was used. Samples were run on a multicolor flow cytometer (FACS Canto; BD Biosciences, San Jose, CA, USA) according to manufacturer's protocol. Analysis was performed using the FACS DIVA 6.1.3 software (BD Biosciences). Blast population was selected first on forward scatter versus side scatter and then from CD45 versus side scatter, so Gating strategy was based on CD45.19 In total, 10,000 events were required, and the percent expression of CD34 on gated myeloblasts were recorded, markers’ positivity was considered if ≥10% (Figure 1).
 |
Figure 1 Flow histogram shows (A) negative expression of CD34 on myeloblasts, and (B) positive expression of CD34 on myeloblasts. |
Mutation Analysis of FLT3-ITD Gene
Total DNA was extracted from BM samples obtained from 153 AML patients compared to 20 healthy control subjects using a QIAamp DNA blood Mini Kit (QIAGEN) according to manufacturer’s instructions. PCR amplification was performed using the primer sequence for FLT3-ITD gene (Thermo Fisher Scientific, USA). F: 5ʹ GCAATTTAGGTATGAAAGCCAGC−3ʹ and R: 5ʹ- CTTTCAGCATTTTGACGGCAACC−3ʹ. In brief, 1 μl DNA was amplified in a volume of 25 μL containing 50 mM KCl, 1.5 Mm MgCl2, 10 mM Tris-HCl, PH 8.3, 200 mM dNTPs, 0.5 μM of each primer and 1U Taq DNA polymerase (QIAGEN). The thermal reaction conditions included a denaturation step for 150 seconds at 94°C followed by 35 amplification cycles at 94°C for 30 seconds, 57°C for 60 seconds, and 72°C for 120 seconds, and a final elongation step for 10 minutes at 72°C. The PCR product was analyzed on standard 3% agarose gel. A wild-type (WT) allele was produced at a fragment of 328 base pair (bp). Patients with an additional higher molecular weight band were considered to be FLT3/ITD+.
Statistical Methods
Statistical analysis was performed using IBM© SPSS© Statistics version 22 (IBM© Corp., Armonk, NY, USA). Numerical data were expressed as median and range according to the performed normality tests. Qualitative data were expressed as frequency and percentage. The relation between qualitative variables was assessed using Chi-square or Fisher’s exact test as appropriate. Comparison between groups was done using Mann–Whitney test. The area under the receiver operating curve (ROC) was calculated to investigate the best cut-off value, sensitivity, and specificity for the diagnosis of AML. Survival analysis was done using Kaplan-Meier test and comparison between survival curves was done using Log rank test. All tests were two-tailed, and a P-value<0.05 was considered significant.
Results
Clinico-Pathological Features of the Patients
The current study included 153 AML patients, with a median age of 39 (range=18–68) years, and a mean of 39±13.01 years. Males represented 49.7% (76/153), and females were 50.3% (77/153) of the assessed patients. Bone marrow (BM) examination showed that 131 patients (85.6%) presented with hypercellular BM, and 22 patients (14.4%) with normocellular BM. Molecular and genetic data were available for 116 patients, of them 40 (34.5%) patients showed normal karyotype, while 36 (31%) patients had t(8:21), 14 (12.1%) had t(9,22), 16 (13.8%) had t(15,17), and 10 (8.6%) patients had (inv16). Patients were classified according to the 2017 risk stratification by the European Leukemia Net (ELN)20 into 58 (42%) patients who had favorable risk, 28 (20.3%) patients with poor risk, and 52 (37.7%) patients had intermediate risk. There were 77 (50.3%) patients who had hepato and/or splenomegaly, and 42 (27.5%) patients presented with lymphadenopathy. At the end of the study, there were 83 (76.9%) patients who showed complete remission (CR), 11 (10.2%) patients had delayed CR after 28 days of treatment, while 14 (12.9%) patients were resistant to therapy (Table 1).
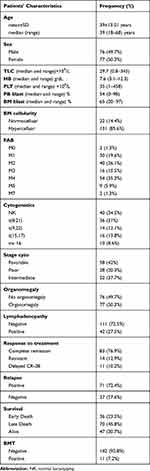 |
Table 1 Clinic-Pathological Features of the Assessed AML Patients |
Assessment of CD34 Expression and FLT3-ITD Gene Mutation
The CD34 was found to be positive in 83/153 (54.2%) patients, compared to 70/153 (45.8%) with negative CD34 expression (P=0.293). In addition, 27/153 (17.6%) patients had mutant FLT3-ITD, compared to 126/153 (82.4%) with wild FLT3 (P=0.001, Figure 2).
 |
Figure 2 Expression of CD34 and FLT3-ITD gene mutation in AML patients. |
Association Between FLT3-ITD Gene Mutation, CD34 Expression, and Relevant Clinic-Pathological Features of the Patients
Patients with FLT3-ITD mutation had a significant increase in the peripheral blood blast cells compared to those with wild FLT3 [median=64.5% (range=20–98%) vs median: 51.5% (range=0–97%); respectively, P=0.013]. Also, FLT3-ITD mutation associated significantly with increased bone marrow blast cells [median: 76% (range=46–92%)] in comparison to those with wild FLT3 [median=60.5% (range= 20–97%), P<0.001]. There was a significant association between the presence of wild FLT3 and normal karyotyping [36/40 (90%) with wild FLT3 vs 4/40 (10%) with mutant FLT3-ITD]. In addition, patients with wild FLT3 had favorable cytogenetics compared to those with FLT3-ITD mutations; as all patients who had inv16 expressed wild type FLT3. On the other hand, the most common detected abnormal cytogenetics in patients with FLT3-ITD Mutation was t(15,17) in 8/14 (57.1%), followed by t(8:21) in 2/14 (14.3%) patients (P=0.010).
Likewise, the absence of CD34 expression was significantly associated with favorable cytogenetics, as all patients (n=16) who had t(15,17) were negative for CD34 expression. Meanwhile, out of all patients (n=14) who had t(9,22), 10 (71.4%) patients were CD34 positive, compared to four (28.6%) patients who were negative for CD34 (P=0.012). CD34 was significantly expressed in patients with FAB M0 (2.4%), M1 (25.3%), M2 (34.9%), and M4 (33.7%) due to increased monocytic and granulocytic lineage (P=0.004, Table 2).
 |
Table 2 Association Between FLT3-ITD Gene Mutation, CD34 Expression and Relevant Clinic-Pathological Features of the Patients |
Consequently, patients were categorized according to FLT3-ITD mutational status and CD34 expression into four groups; G1 (n=9): FLT3-ITD mutant and CD34+, G2 (n=18): FLT3-ITD mutant and CD34-, G3 (n=74): Wild FLT3 and CD34+, and G4 (n=52): Wild FLT3 and CD34-. Patients with wild FLT3 and CD34- (G4) significantly had FAB M5 and M7 types. While patients with Wild FLT3 and CD34+ (G3) significantly had FAB M1 (P=0.036). Normal karyotyping was present significantly in G4 (wild FLT3, CD34-), and t(8,21) was present significantly in G3 (Wild FLT3, CD34+). On the other hand, patients in G1 (FLT3-ITD mutant, CD34+) significantly did not have favorable cytogenetics like t(15,17): 0/9 (0.0%), inv16: 0/9 (0.0%), and t(8,21): 2/9 (33.3%, P<0.001).
Patients groups who had mutant FLT3-ITD (G1 and G2) showed a significant increase in the peripheral blood and bone marrow blast cells (P=0.033 and P=0001, respectively) compared to those with wild FLT3 (G3 and G4, Table 3).
 |
Table 3 Associations Between Combined Expression of CD34 and FLT3-ITD Mutation with Patients’ Clinico-Pathological Features |
Patients’ Response to Treatment
There was no significant association between response to treatment and FLT3-ITD mutation (P=0.081) or CD34 expression (P=0.597, Table 2);, however, the incidence of relapse associated significantly with FLT3-ITD mutation (60%) compared to (40%) in patients with wild FLT3 (P=0.025, Table 1).
Resistance to treatment was significantly observed in G1 (FLT3-ITD mutant, CD34+) where it represented 57.1%, while complete remission (CR) was detected in 28.6% of the patients, and delayed CR was detected only in 14.3%. On the other side, data in G3 (wild FLT3, CD34+) and G4 (wild FLT3, CD34-) were comparable, as CR was significantly observed in 46/74 (80.7%) and 30/52 (78.9%), respectively. Delayed CR (after 28 days) was detected in 5/74 (8.8%) and 4/52 (10.5%), respectively, while resistance to treatment was detected in 6/74 (10.5%) and 4/52 (10.5%), respectively (P=0.024, Table 3 and Figure 3).
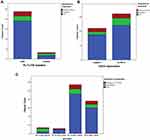 |
Figure 3 Association between response to treatment and (A) FLT3-ITD gene mutation, (B) CD34 expression, and (C) different combined expression of both markers in patients’ groups. |
Disease Free Survival (DFS) and Overall Survival (OS) Rates
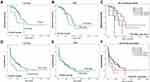
The median DFS time for all patients was 16.76 months, there was a significant association between DFS rate and FLT3-ITD mutational status (16.77ms in wild FLT3 compared to 8.93ms in mutant FLT3-ITD, P=0.008, Figure 4A). There was no significant association between DFS rate of the patients and CD34 expression (18ms in CD34- and 13.6 ms in CD34+, P=0.419, Figure 4B).
Combined expression of CD34 and FLT3-ITD mutational status showed that the median DFS time for G1 (FLT3-ITD mutant, CD34+) was 8.13 ms, G2 (FLT3-ITD mutant, CD34-) was 8.93 ms, G3 (wild FLT3, CD34+) was 13.57 ms, and G4 (wild FLT3, CD34-) was 19.83 ms (P=0.006, Figure 4C).
The median OS time for all patients was 5.47 months, there was a significant association between OS rate and FLT3-ITD mutational status (7.07 ms in wild FLT3 compared to 1.17 ms in mutant FLT3-ITD, P=0.004, Figure 4D). There was no significant association between DFS rate of the patients and CD34 expression (6.5 ms in CD34- and 4.17 ms in CD34+, P=0.652, Figure 4E).
Combined expression of CD34 and FLT3-ITD mutational status showed that the median OS time for G1 (FLT3-ITD mutant, CD34+) was 2.07 ms, G2 (FLT3-ITD mutant, CD34-) was 0.9 ms, G3 (wild FLT3, CD34+) was 6.73 ms, and G4 (wild FLT3, CD34-) was 7.07 ms (P=0.037, Figure 4F).
Univariate and Multivariate Analysis for Survival Rates
Univariate Cox Regression analysis demonstrated that FLT3-ITD mutation (HR=0.310, CI=0.123–0.779: P=0.013) and FAB subtypes (HR=3.047, CI=1.313–7.075, P=0.010) associated with shorter DFS rates. Age ≥39 years (HR=0.427, CI=0.287–0.634, P<0.001), HB concentration <7 mg/dL (HR=1.678, CI=1.134–2.483, P=0.010), PB blast ≥54% (HR=0.627, CI=0.415–0.947, P=0.026), and FLT3-ITD mutation (HR=0.508, CI=0.319–0.809: P=0.004) associated significantly with poorer OS rates of the AML patients.
However, multivariate Cox Regression analysis demonstrated that FAB subtypes (HR=2.933, CI=1.237–6.951, P=0.015) and FLT3-ITD mutation (HR=0.347, CI=0.137–0.880, P=0.026) were independent prognostic factors for DFS, while older age (HR=0.405, CI=0.263–0.623: P=0.001), HB <7 mg/dL (HR=1.591, CI=1.042–2.429, P=0.031), PB blast ≥54% (HR=0.653, CI=0.427–0.999, P=0.049), and FLT3-ITD mutation (HR=0.537, CI=0.318–0.904, P=0.019) were independent prognostic factors for OS rates of the AML patients (Table 4).
 |
Table 4 Univariate and Multivariate Analysis for Survival Rates |
Discussion
Acute myeloid leukemia (AML) is the most common form of acute leukemia in adults which is characterized by poor survival rate. Although cytogenetics is the most valuable prognostic marker in AML, however in patients with normal karyotype, molecular tasting like FLT3-ITD mutation has an important role as a prognostic risk stratification for those patients, and therefore optimizing better therapeutic approaches.21 The current study demonstrated that FLT3-ITD mutation represented 17.6% of the assessed Egyptian AML patients. This percentage was comparable to other studies which reported the prevalence of FLT3-ITD mutation which were 17.4%, 15.9%, and14.6% in Latin American, Chinese, and Indian patients, respectively.21–23 However, it was higher in South Korea (22.4%) and Australia (30.8%), while it was lower in Poland (8%) and Saudi Arabia (11.6%).24–27
In line with many previously published studies,28,30 it was reported that t(15,17) and chromosome 8 trisomy were the most significant recurrent alterations associated with FLT3-ITD mutation. Our data showed that most of all patients with FLT3-ITD mutation had t(15,17) followed by t(8:21). In addition, AML patients who did not have FLT3-ITD mutation showed significantly normal karyotyping.
The current study demonstrated that patients with FLT3-ITD mutation had a significant increased bone marrow and peripheral blood blast cell counts compared to those with wild FLT3. Also, there was an increased total leukocyte count (TLC) in patients with FLT3-ITD mutation, however it did not reach a significant level. But there was no significant association with other clinic-pathological features including hemoglobin concentration, BM cellularity, FAB subtypes, or the presence of organomegaly. These data are in agreement with those of Fröhling et al,31 who reported a significant increase in the BM blast cells count, peripheral blood blast cell count, and TLC in patients with FLT3-ITD mutation. Also, Wang et al32 concluded a significant increase in the BM blast cell count and TLC in AML patients with FLT3-ITD mutation. On the other hand, Bhattacharyya et al21 detected a significant increase in the TLC in AML patients with FLT3-ITD mutation, however no significant association with PB or BM blast cell count.
The present data also showed an increased incidence of disease relapse, inferior overall survival, and disease-free survival rates in patients with FLT3-ITD mutation compared to patients with wild FLT3. These data are consistent with our previous study,33 and many other recent studies reported a significant association between FLT3-ITD mutation and increased risk of relapse, poor OS or DFS rates in AML patients.21,–31–35 Similarly, Garcia and Stone36 concluded that patients with FLT-ITD AML had a higher rate of relapse after transplantation compared to patients with wild FLT3. Additionally, Hu and Chen37 reported in their reviews that AML patients with mutant FLT3-ITD showed increased recurrence after chemotherapy and hematopoietic stem cell transplantation, as well as an creased mortality rate and short survival time in comparison to AML patients with wild-type FLT3. On the other hand, Karabacak et al38 found that the presence of FLT3/ITD mutation associated significantly with shorter OS rate in 40 AML patients, but did not associate with DFS. Melo et al28 observed that FLT3-ITD mutation had no effect on overall survival, remission status, or relapse rate in AML patients. This discrepancy in the results could be explained by the small number of patients assessed in their study (34 patients), and the short median follow-up period (2 years) of the patients.
For more confirmation, multivariate Cox Regression analysis demonstrated that FLT3-ITD mutation and FAB subtypes were independent prognostic factors for DFS rate. Older age ≥39 years, HB <7 mg/dL, PB blast ≥54%, and FLT3-ITD mutation were negative prognostic factors for poor OS rates of the AML patients. Our results in this context are consistent with Fröhling et al,31 who stated that FLT3-ITD mutation was an independent negative marker affecting remission duration and OS rate of the patients. Similarly, Colovic et al39 concluded that FLT3/ITD mutation was the most significant prognostic factor for overall survival in a cohort of 113 Serbian adult AML patients. Hence, assessment of FLT3 expression level in AML patients is an indicator to evaluate the prognosis of the disease course and to monitor the small residual focus.31,40 Moreover, Arellano et al41 concluded that peripheral blood blast was found to be able to predict complete remission, relapse-free survival, and overall survival in AML patients.
Regarding assessment of CD34 expression in the bone marrow of adult AML patients, the present study demonstrated that CD34 expression was expressed in 54.2% of the assessed AML patients and most of these patients had a significant t(9,22) karyotyping. Meanwhile those with CD34 negative expression had a favorable cytogenetics like t(15,17). However, there was no significant association between CD34 expression and disease relapse, overall survival, or disease-free survival rates of the patients. Our results in this concern are contradictory to previous studies which reported increased CD34 percentages at relapse in AML patients.42,43 This variability in the results may be due to the time of assessment of CD34+ cells, whether at diagnosis or at relapse. Baer et al42 observed that CD34+ cells were absent at diagnosis in 47 patients, and gained at relapse in 17 (36%) AML cases.
Another important finding in the current is that we tried to assess the combined expression of both FLT3-ITD and CD34 in AML patients, and whether this combination will add a value to the prognosis or the outcome of the patients. Accordingly, we found that patients who were positive for both FLT3-ITD and CD34 expression showed the worst prognosis and outcome in the form of absence of favorable cytogenetics like t(15,17) and inv-16, as well as the shortest disease-free survival and overall survival time among patients’ groups. Meanwhile, patients who had both wild type FLT3 and CD34 negative expression showed significantly normal cytogenetics, as well as the most favorable overall and disease free survival rates in the assessed AML patients.
Many recent studies proposed the important role of CD34 and FLT3-ITD mutation in predicting patients’ response to therapy. For example, Yiau et al,44 who stated that CD34 was significantly higher in chemo-resistant cells compared to chemo-sensitive samples, and this chemo-resistance was mediated through a pBAD signaling pathway. Similarly, Perl et al45 and Hu and Chen37 reported through their works that FLT3 is a good promising target for AML treatment. However, our results in this context are controversial, as we were not able to detect a significant association between patients’ response to treatment and neither FLT3-ITD mutation nor CD34 expression. Patients who expressed both markers together showed significant resistance to chemotherapy compared to those who had only CD34 expression or FLT3-ITD mutation. Therefore, the present study provides evidence that both FLT3-ITD and CD34 are necessary for prediction of patients’ response to therapy, and we cannot rely only on one of them.
Thus, we can conclude that both FLT3-ITD mutation and CD34 expression could be considered as an important predictor and prognostic factor for poor outcome of AML patients, as patients with combined expression of both factors showed the shortest disease-free survival and overall survival rates, as well as the poorest response to therapy. This will help for better understanding and prediction of disease course, and therefore proper management of those patients.
One of the main drawbacks in the current study was that the number of patients’ groups were not equal, that meant some groups had a small number of patients. This is because all patients were recruited randomly during the assigned period of research, and then they were classified according to FLT3-ITD mutation and CD34 expression. Hence, these preliminary results required a further study on a larger number of patients to extensively assess this issue.
Ethical Consideration
The study protocol was approved by the ethical committee of the National Cancer Institute, Cairo University, which was in accordance with the 2011 Declaration of Helsinki. Signed informed consent was obtained from each patient before enrollment in the study.
Disclosure
All authors declare that there are no conflicts of interest.
References
1. Chaudhury S, O’Connor C, Canete A, et al. Age-specific biological and molecular profiling distinguishes paediatric from adult acute myeloid leukaemias. Nat Commun. 2018;9(1):5280. doi:10.1038/s41467-018-07584-1
2. De Kouchkovsky I, Abdul-Hay M. Acute myeloid leukemia: a comprehensive review and 2016 update. Blood Cancer J. 2016;6(7):e441. doi:10.1038/bcj.2016.50
3. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2017. CA Cancer J Clin. 2017;67(1):7–30. doi:10.3322/caac.21387
4. Miller KD, Siegel RL, Lin CC, et al. Cancer treatment and survivorship statistics, 2016. CA Cancer J Clin. 2016;66:271–289.
5. Shlush LI, Zandi S, Mitchell A, et al. Identification of pre-leukaemic haematopoietic stem cells in acute leukaemia. Nature. 2014;506(7488):328–333. doi:10.1038/nature13038
6. Reinisch A, Chan SM, Thomas D, Majeti R. Biology and clinical relevance of acute myeloid Leukemia stem cells. Semin Hematol. 2015;52:150–164.
7. Schepers K, Campbell TB, Passegué E. Normal and leukemic stem cell niches: insights and therapeutic opportunities. Cell Stem Cell. 2015;16(3):254–267. doi:10.1016/j.stem.2015.02.014
8. Lei H, Yang L, Zhou L, Tong Y, Wu Y. Targeting acute myeloid leukemia CD34+ stem/progenitor cells with small molecule inhibitor MK-8776. Leuk Res. 2018;72:71. doi:10.1016/j.leukres.2018.08.003
9. Nielsen JS, McNagny KM. Novel functions of the CD34 family. J Cell Sci. 2008;121(24):36833–36892. doi:10.1242/jcs.03504
10. AbuSamra DB, Aleisa FA, Al-Amoodi AS, et al. Not just a marker: CD34 on human hematopoietic stem/progenitor cells dominates vascular selectin binding along with CD44. Blood Adv. 2017;1(27):2799. doi:10.1182/bloodadvances.2017004317
11. van Stijn A, van der Pol MA, Kok A, et al. The differential role of the CD34+ and CD34- blast compartments in apoptosis resistance in acute myeloid leukemia. Haematologica. 2003;88:497–508.
12. Rollins-Raval M, Pillai R, Warita K, et al. CD123 immunohistochemical expression in acute myeloid leukemia is associated with underlying FLT3-ITD and NPM1 mutations. Appl Immunohistochem Mol Morphol. 2013;21(3):212–217.
13. Levis M. FLT3 mutations in acute myeloid leukemia: what is the best approach in 2013? Hematol Am Soc Hematol Educ Program. 2013;2013:220–226.
14. Volpe G, Clarke M, Garcia P, et al. Regulation of the Flt3 gene in haematopoietic stem and early progenitor cells. PLoS One. 2015;10(9):e0138257. doi:10.1371/journal.pone.0138257
15. Cheng J, Qu L, Wang J, Cheng L, Wang Y. High expression of FLT3 is a risk factor in leukemia. Mol Med Rep. 2018;17(2):2885–2892.
16. Takahashi S. Downstream molecular pathways of FLT3 in the pathogenesis of acute myeloid leukemia: biology and therapeutic implications. J Hematol Oncol. 2011;4(13). doi:10.1186/1756-8722-4-13
17. Daniel AA, Attilio O, Robert H, et al. Revision to the World Health Organization classification of myeloid neoplasms and acute Leukemia. Blood. 2016;127(20):2391–2405.
18. Tallman MS, Wang ES, Altman JK, et al. Acute myeloid leukemia, version 3.2019, NCCN clinical practice guidelines in oncology. J Natl Compr Cancer Network. 2019;17(6):721–749. doi:10.6004/jnccn.2019.0028
19. Ge F, Li B, Gao X, et al. Immuno-phenotypes and prognosis of acute leukemia in elderly patients. Int J Clin Exp Med. 2014;7(10):3714.
20. Döhner H, Estey E, Grimwade D, et al. Diagnosis and management of AML in adults:2017 ELN recommendations from an international expert panel. Blood. 2017;129(4):424–447.
21. Bhattacharyya J, Nath S, Saikia KK, et al. Prevalence and clinical significance of FLT3 and NPM1 mutations in acute myeloid leukaemia patients of Assam, India. Indian J Hematol Blood Transfusion. 2018;34(1):32–42. doi:10.1007/s12288-017-0821-0
22. Cuervo-Sierra J, Go´mez-Almaguer D, Jaime-Pe´rez JC, et al. Prevalence of FLT3 mutations in acute myeloid leukemia: a multicenter Latin America study. Blood. 2013;122(21):4979. doi:10.1182/blood.V122.21.4979.4979
23. Wang W, Wang XQ, Xu XP, Lin GW. Prevalence and prognostic significance of FLT3 gene mutations in patients with acute leukaemia: analysis of patients from the Shanghai Leukaemia Co-operative Group. J Int Med Res. 2010;38(2):432–442.
24. Park BG, Chi HS, Park SJ, et al. Clinical implications of non-A-type NPM1 and FLT3 mutations in patients with normal karyotype acute myeloid leukemia. Acta Haematol. 2012;127(2):63–71. doi:10.1159/000331509
25. Al-Mawali A, Gillis D, Lewis I. Characteristics and prognosis of adult acute myeloid leukemia with internal tandem duplication in the FLT3 gene. Oman Med J. 2013;28(6):432. doi:10.5001/omj.2013.121
26. Koczkodaj D, Zmorzyński S, Michalak-Wojnowska M, Wąsik-Szczepanek E, Filip AA. Examination of the FLT3 and NPM1 mutational status in patients with acute myeloid leukemia from southeastern Poland. Arch Med Sci AMS. 2016;12(1):120. doi:10.5114/aoms.2015.49811
27. Gari M, Abuzenadah A, Chaudhary A, et al. Detection of FLT3 oncogene mutations in acute myeloid leukemia using conformation sensitive gel electrophoresis. Int J Mol Sci. 2008;9(11):2194–2204. doi:10.3390/ijms9112194
28. Melo CP, Campos CB, Dutra ÁP, et al. Correlation between FLT3–ITD status and clinical, cellular and molecular profiles in promyelocytic acute leukemias. Leuk Res. 2015;39(2):131–137. doi:10.1016/j.leukres.2014.11.010
29. Schnittger S, Bacher U, Haferlach C, Kern W, Alpermann T, Haferlach T. Clinical impact of FLT3 mutation load in acute promyelocytic leukemia with t(15;17)/PML–RARA. Haematologica. 2011;96(12):1799–1807. doi:10.3324/haematol.2011.049007
30. De Lourdes Chauffaille M, Borri D, Proto-Siqueira R, Moreira ES, Alberto FL. Acute promyelocytic leukemia with t (15; 17): frequency of additional clonal chromosome abnormalities and FLT3 mutations. Leuk Lymphoma. 2008;49(12):2387–2389. doi:10.1080/10428190802511248
31. Fröhling S, Schlenk RF, Breitruck J, et al. Prognostic significance of activating FLT3 mutations in younger adults (16 to 60 years) with acute myeloid leukemia and normal cytogenetics: a study of the AML Study Group Ulm. Blood J Am Soc Hematol. 2002;100(13):4372–4380.
32. Wang L, Xu WL, Meng HT, et al. FLT3 and NPM1 mutations in Chinese patients with acute myeloid leukemia and normal cytogenetics. J Zhejiang Univ Sci B. 2010;11(10):762–770. doi:10.1631/jzus.B1000052
33. Ebrahim EK, Assem MM, Amin AI, Kamel MM, El Meligui YM, Metwally AM. FLT3 internal tandem duplication mutation, cMPL and CD34 expressions predict low survival in acute myeloid leukemia patients. Ann Clin Lab Sci. 2016;46(6):592–600.
34. Fernandez HF, Sun Z, Yao X, et al. Anthracycline dose intensification in acute myeloid leukemia. N Engl J Med. 2009;361(13):1249–1259. doi:10.1056/NEJMoa0904544
35. Adnan-Awad S, Gaber O, Eltokhy SA, et al. FLT3-ITD mutations in Egyptian patients of acute myeloid leukemia: correlation with cytogenetic, FAB subgroups and prognosis. Clin Lab. 2017;63(5):1027–1034. doi:10.7754/Clin.Lab.2017.170121
36. Garcia JS, Stone RM. The development of FLT3 inhibitors in acute myeloid leukemia. Hematol Oncol Clinics. 2017;31(4):663–680. doi:10.1016/j.hoc.2017.03.002
37. Hu X, Chen F. Targeting on glycosylation of mutant FLT3 in acute myeloid leukemia. Hematology. 2019;24(1):651–660.
38. Karabacak BH, Erbey F, Bayram I, et al. Fms-like tyrosine kinase 3 mutations in childhood acute leukemias and their association with prognosis. Asian Pac J Cancer Prev. 2010;11(4):923–927.
39. Colovic N, Tosic N, Aveic S, et al. Importance of early detection and follow-up of FLT3 mutations in patients with acute myeloid leukemia. Ann Hematol. 2007;86(10):741–747. doi:10.1007/s00277-007-0325-3
40. Reiter K, Polzer H, Krupka C, et al. Tyrosine kinase inhibition increases the cell surface localization of FLT3-ITD and enhances FLT3-directed immunotherapy of acute myeloid leukemia. Leukemia. 2018;32(2):313–322. doi:10.1038/leu.2017.257
41. Arellano M, Pakkala S, Langston A, et al. Early clearance of peripheral blood blasts predicts response to induction chemotherapy in acute myeloid leukemia. Cancer. 2012;118(21):5278. doi:10.1002/cncr.27494
42. Baer MR, Stewart CC, Dodge RK, et al. High frequency of immunophenotype changes in acute myeloid leukemia at relapse: implications for residual disease detection (Cancer and Leukemia Group B Study 8361). Blood. 2001;97(11):3574–3580. doi:10.1182/blood.V97.11.3574
43. Thomas X, Campos L, Archimbaud E, et al. Surface marker expression in acute myeloid leukaemia at first relapse. Br J Haematol. 1992;81(1):40–44. doi:10.1111/j.1365-2141.1992.tb08168.x
44. Yiau SK, Lee C, Mohd. Tohit ER, Chang KM, Abdullah M. Potential CD34 signaling through phosphorylated-BAD in chemotherapy-resistant acute myeloid leukemia. J Recept Signal Transduction. 2019;39(3):276–282. doi:10.1080/10799893.2019.1660899
45. Perl AE, Altman JK, Cortes J, et al. Selective inhibition of FLT3 by gilteritinib in relapsed or refractory acute myeloid leukaemia: a multicentre, first-in-human, open-label, Phase 12 study. Lancet Oncol. 2017;18(8):1061–1075. doi:10.1016/S1470-2045(17)30416-3
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.

