Back to Journals » Infection and Drug Resistance » Volume 13
Clonal Spread of Tetracycline Resistance Among Mycoplasma hominis Clinical Strains, Tunisia
Authors Boujemaa S , Mlik B, Mardassi H , Ben Abdelmoumen Mardassi B
Received 13 February 2020
Accepted for publication 28 April 2020
Published 2 July 2020 Volume 2020:13 Pages 2093—2097
DOI https://doi.org/10.2147/IDR.S249630
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Sahil Khanna
Safa Boujemaa,1 Béhija Mlik,1 Helmi Mardassi,2 Boutheina Ben Abdelmoumen Mardassi1
1Group of Mycoplasmas, Laboratory of Molecular Microbiology, Vaccinology, and Biotechnology Development. Institut Pasteur De Tunis, Université De Tunis El Manar, Tunis, Tunisia; 2Unit of Typing & Genetics of Mycobacteria, Laboratory of Molecular Microbiology, Vaccinology, and Biotechnology Development. Institut Pasteur De Tunis, Université De Tunis El Manar, Tunis, Tunisia
Correspondence: Boutheina Ben Abdelmoumen Mardassi
Group of Mycoplasmas, Laboratory of Molecular Microbiology, Vaccinology, and Biotechnology Development. Institut Pasteur De Tunis, Université De Tunis El Manar, Tunis, Tunisia
Tel +216 71 847 609
Fax +216 71 791 833
Email [email protected]
Abstract: Antimicrobial resistance in a number of bacterial pathogens has been shown to spread clonally. To our knowledge, data about the phylodistribution of drug resistance in Mycoplasma hominis are very scarce. The aims of this study were to assess the antimicrobial susceptibility of Mycoplasma hominis clinical strains in Tunisia, to identify the molecular basis of antibiotic resistance, and to investigate the phylogenetic relationships of resistant strains. This study included 65 molecularly typed Mycoplasma hominis clinical strains recovered from Tunisian patients over 18 years (2000– 2018). The antimicrobial susceptibility was tested against nine antibacterial agents using the broth microdilution method. Minimum spanning tree was constructed to establish the phylogenetic relationships among resistant isolates. Fluoroquinolones, doxycycline, and josamycine were found to be the most effective antibacterial agents. However, 22 strains belonging to 11 expanded multilocus sequence types (eSTs) proved resistant to tetracycline. The majority of these eSTs were genetically related, indicative of clonal expansion of tetracycline resistance. The present study provides relevant information on the antibiotic susceptibility of Tunisian M. hominis clinical strains, lending support to a clonal transmission of tetracycline resistance. This is likely to have an important implication in monitoring the spread of drug resistance among M. hominis.
Keywords: Mycoplasma hominis, antibiotic resistance, tetracycline, clonal transmission, expanded multilocus sequence type
Introduction
Mycoplasma hominis has an etiological role in genitourinary tract infections (pelvic inflammatory disease, cervicitis, bacterial vaginosis) and appears to be associated with severe complications in pregnancy and serious infections in newborns and immunocompromised hosts.1 Ample evidence from recent studies conducted in Tunisia indicates that M. hominis is one of the potential causative agents that play an important role in the pathogenesis of human infertility with a prevalence of 83%.2 And only 17% were associated with gynecological infections.2 However, M. hominis is less frequent among women suffering from complications during pregnancy (2,5%), in Tunisia.3
In comparison with Ureaplasma spp. and Mycoplasma pneumoniae, Mycoplasma hominis displays high levels of genetic and antigenic heterogeneity among clinical isolates as disclosed by several molecular typing methods,2,4 which lends further support to the virulence of this species.
In recent years, due to the abuse of antibiotics, the resistance of M. hominis to antibiotics has been shown to have an increasing trend.5 The prevalence of this pathogen and the extent of its antibiotic resistance profiles vary geographically and may be related to local antibiotic use regulations.6 Thus, antimicrobial surveillance should be diligent and thorough for effective antimicrobial therapy, and to monitor the spread of resistant strains.
The present study aimed to evaluate the antibiotic susceptibility of M. hominis to various antimicrobial agents over 18 years, in Tunisia, and to identify the involved resistance markers. The association of antimicrobial susceptibility with genetic diversity was also assessed to highlight the evolution of antibiotic resistance in M. hominis.
Materials and Methods
Out of 1180 samples, sixty-five (5,5%) M. hominis clinical strains isolated from outpatients (48 non-pregnant women and 17 men) presenting to the diagnostic center of the Pasteur Institute of Tunis (Grand Tunis, Tunisia), from January 2000 to December 2018, were included in this study. As control, M. hominis strain PG21 (ATCC 23,114) has been considered. The patient group, whose ages ranged between 25 and 48 years, experienced symptoms of urogenital tract infections (10 cases) or infertility (55 cases).
From the 65 isolates, 59 were previously characterized using an Expanded MultiLocus Sequence Typing (eMLST) scheme, including ten genes (uvrA, gyrB, ftsY, tuf, gap, p120ʹ, vaa, lmp1, lmp3, p60).2 The remaining six isolates were genetically typed in this study by the same method. The sequences generated by eMLST were submitted to the Mycoplasma hominis MLST website (https://pubmlst.org/mhominis/) sited at the University of Oxford.7
Nine antimicrobial agents were tested for antibiotic susceptibility: tetracycline ‘TET’, doxycycline ‘DOX’, ofloxacin ‘OFX’, ciprofloxacin ‘CIP’, levofloxacin ‘LVX’, moxifloxacin ‘MXF’, erythromycin ‘ERY’, azithromycin ‘AZM’, and josamycin ‘JOS’ (Sigma-Aldrich, Germany). Antimicrobial susceptibility was determined using a broth microdilution method for M. hominis PG21 and clinical strains, as detailed previously.8 The specific breakpoints (mg/liter) indicating susceptibility (S) or resistance (R) are: tetracycline S ≤4, R ≥8; doxycycline S≤4, R≥8; ciprofloxacin S≤1, R≥2; ofloxacin S≤1, R≥4; levofloxacin S ≤1, R ≥2; moxifloxacin S ≤0.25, R≥0.5; erythromycin S ≤1, R ≥4; azithromycin S≤0.125, R≥4; and josamycin S≤2, R≥8.9,10
To confirm the occurrence of tetracycline resistance among M. hominis isolates, tet(M) and Int-Tn genes were amplified from both tetracycline-susceptible and -resistant strains, as previously detailed.11 Simultaneously, macrolide-resistant strains underwent PCR amplification of domain II and domain V of 23S rRNA, as well as the genes encoding ribosomal proteins L4 and L22.12 Amplicons were then sequenced.2 Sequences were analyzed using BioEdit and translated into amino acids. Amino acid substitutions in L4 and L22, as well as nucleotide mutations in domain II and domain V of 23S rRNA, were identified by comparison with the corresponding sequences of Escherichia coli ER1709 (GenBank accession number: CP030240.1).
A minimum spanning tree (MST) analysis based on eMLST data has been constructed, using BioNumerics software (version 7.0, Applied Maths), to illustrate the distribution of antimicrobial resistance among M. hominis clinical strains.2
Results and Discussion
Minimum inhibitory concentration (MIC) results for the 65 M. hominis clinical isolates are shown in Supplementary Table 1. All M. hominis isolates were sensitive to DOX, OFX, CIP, LVX, MXF, and JOS (MIC ≤ 1 mg/liter). The high sensitivity rate of M. hominis to fluoroquinolones is fairly consistent with that reported by Meygret et al in France,13 but distinctly different from the studies carried out in China, which reported a high resistance rates to OFX (59%) and CIP (64%).5 Interestingly, Tunisian genital mycoplasmas (Ureaplasma spp. and M. hominis) showed a difference in their resistance patterns to fluoroquinolones. Indeed, Ureaplasma spp. strains proved more resistant to CIP, OFX, and LVX (100%, 38.62%, 17.82%, respectively) than did M. hominis strains.14
Since M. hominis is known to be naturally resistant to 14- and 15-membered macrolides, all isolates were resistant to ERY (MIC ≥ 16 mg/liter) and AZM (MIC ≥ 4 mg/liter).12 No mutations were found in either L4 or L22 ribosomal proteins; however, all resistant strains harbored a G2057A transition in their 23S rRNA sequences. This is in agreement with prior reports indicating that the G2057A transition accounts for the intrinsic resistance to macrolides in M. hominis.12
Regarding tetracycline susceptibility, 43 (66.15%) strains were sensitive. This rate is distinctly different from that of other countries, which reported a higher (>80%) sensitivity rate to TET.15 We noticed through the studied period a decreased susceptibility to tetracycline, most likely as a consequence of its use as a first-line drug against M. hominis in Tunisia. Indeed, a previous study conducted in our laboratory in 2012 showed that 75% of M. hominis strains were sensitive to tetracycline.11 Screening of the tet(M) determinant, which is associated with tetracycline resistance, as well as the Int-Tn gene, were carried out, and all tetracycline-resistant isolates were positive, while no tetracycline-sensitive isolates harbored these genes. As mentioned previously, a unique sequence of the tet(M) amplicon was shared by the tetracycline-resistant strains, suggesting that they originally acquired the tet(M) determinant from a common source.11
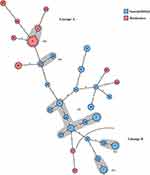
As demonstrated previously, the eMLST method segregated M. hominis strains into two distinct genetic lineages, which were differentially associated with infertility (lineage A) or gynecological infections (lineage B).2 Based on the MST, we found that tetracycline resistance was widespread in lineage A. Indeed, all M. hominis isolates from lineage B were sensitive to tetracycline, whereas 41.5% of isolates from lineage A were resistant. Interestingly, the MST showed that tetracycline resistance was linked to specific clones: eST3, eST6, eST14, eST17, eST20, eST21, eST23, eST24, eST25, and eST26 (Figure 1). The depiction of tetracycline resistance on the MST clearly showed its clonal expansion among the majority of clones (82%). However, the remaining tetracycline-resistant clones (eST14, eST17, eST21, eST24) were positioned at the edges of the MST, indicating a very distant genetic relatedness to the major resistant clones, which suggests the convergent acquisition of tetracycline resistance by these isolates.
Although horizontal transfer mediated by mobile elements was considered as the major process underlying the worldwide dissemination of tetracycline resistance, our study indicates that clonal spread of tetracycline resistance might account significantly in particular settings. Moreover, the observation that tetracycline-resistant M. hominis strains shared a common tet(M) sequence type further support the clonal expansion of tetracycline resistance. However, it has been previously shown that this tet(M) sequence type was also harbored by Ureaplasma parvum tetracycline-resistant strains, indicating that it might be particularly competent to spread between these species.11
The clonal dissemination of tetracycline resistance observed here is in sharp disagreement with previous studies. Férandon et al have reported that 68 M. hominis isolates resistant to tetracycline were not related and grouped into 25 MLVA types (Multi Locus VNTR Analysis).4 Similarly, a previous study from our group, based on p120ʹ gene fragment polymorphisms, found no genetic relationships among tetracycline-resistant M. hominis strains.11 Hence, aside from segregating M. hominis strains into two genetically differentiable urogenital pathotypes, the eMLST scheme used herein proved to be a useful epidemiological tool to track the clonal transmission of tetracycline resistance in M. hominis strains. Similar results have been reported for other bacteria. For example, a combination of multilocus sequence typing scheme with molecular evolutionary analysis, such as the minimum spanning tree, provided reliable insight into the evolutionary pathways and the transmission of antibiotic resistance among Neisseria gonorrhoeae strains.16
Conclusion
In conclusion, the upward trend in the incidence of tetracycline-resistant M. hominis strains in Tunisia appears to be mainly due to clonal spread. The identification of resistant clones associated with tetracycline resistance could contribute to investigate novel genetic biomarkers for predicting drug resistance in M. hominis strains, which would have an impact on controlling resistance and disease monitoring.
However, this finding should prompt additional validation studies using larger and geographically diverse, strain collections. Hence, we recommend the use at first intention of fluoroquinolones, doxycycline and josamycin for treating M. hominis infections in Tunisian patients and break down the transmission of tetracycline resistance. Furthermore, the appropriate treatment of M. hominis may prove to be important in controlling infertility in Tunisia.
Data Sharing Statement
All data generated or analyzed during this study are included in this article and its additional file.
Ethical Approval
Neither human/animal subjects nor human cell lines/tissues were involved in this study. Only fully anonymized data were processed, and hence no ethical approval was required.
Author Contributions
All authors made substantial contributions to conception and design, acquisition of data, or analysis and interpretation of data; took part in drafting the article or revising it critically for important intellectual content; gave final approval of the version to be published; and agree to be accountable for all aspects of the work.
Disclosure
The authors declare that they have no conflicts of interest in this work.
References
1. Waites KB, Schelonka RL, Xiao L, Grigsby PL, Novy MJ. Congenital and opportunistic infections: ureaplasma species and Mycoplasma hominis. Semin Fetal Neonatal Med. 2009;14(4):190–199. doi:doi:10.1016/j.siny.2008.11.009
2. Boujemaa S, Ben Allaya A, Mlik B, Mardassi H, Ben Abdelmoumen Mardassi B. Phylogenetics of Mycoplasma hominis clinical strains associated with gynecological infections or infertility as disclosed by an expanded multilocus sequence typing scheme. Sci Rep. 2018;8(1):14854. doi:doi:10.1038/s41598-018-33260-x
3. Mardassi BBA, Ayari H, Béjaoui-Khiari A, Mlik B, Moalla I, Amouna F. Genetic variability of the P120ʹ surface protein gene of Mycoplasma hominis isolates recovered from Tunisian patients with urogenital and infertility disorders. BMC Infect Dis. 2007;7:1. doi:doi:10.1186/1471-2334-7-142
4. Férandon C, Peuchant O, Renaudin H, Bébéar C. Diversity of Mycoplasma hominis clinical isolates from Bordeaux, France, as assessed by multiple-locus variable-number tandem repeat analysis. BMC Microbiol. 2013;13(1):120. doi:doi:10.1186/1471-2180-13-120
5. Song T, Ye A, Xie X, et al. Epidemiological investigation and antimicrobial susceptibility analysis of ureaplasma species and Mycoplasma hominis in outpatients with genital manifestations. J Clin Pathol. 2014;67(9):817–820. doi:doi:10.1136/jclinpath-2014-202248
6. Domingues D, Távora Tavira L, Duarte A, Sanca A, Prieto E, Exposto F. Genital mycoplasmas in women attending a family planning clinic in Guiné-Bissau and their susceptibility to antimicrobial agents. Acta Trop. 2003;86(1):19–24. doi:10.1016/S0001-706X(03)00002-0
7. Jolley KA, Bray JE, Maiden MCJ. Open-access bacterial population genomics: bIGSdb software, the PubMLST.org website and their applications. Wellcome Open Res. 2018;3:124. doi:doi:10.12688/wellcomeopenres.14826.1
8. Waites KB, Duffy LB, Bébéar CM, et al. Standardized methods and quality control limits for agar and broth microdilution susceptibility testing of Mycoplasma pneumoniae, Mycoplasma hominis, and Ureaplasma urealyticum. J Clin Microbiol. 2012;50(11):3542–3547. doi:doi:10.1128/JCM.01439-12
9. Kasprzykowska U, Sobieszczańska B, Duda-Madej A, Secewicz A, Nowicka J, Gościniak G. A twelve–year retrospective analysis of prevalence and antimicrobial susceptibility patterns of Ureaplasma spp. and Mycoplasma hominis in the province of Lower Silesia in Poland. Eur J Obstet Gynecol Rep Biol. 2018;220:44–49. doi:doi:10.1016/j.ejogrb.2017.11.010
10. Clinical and Laboratory Standards Institute (CLSI), ed. Methods for Antimicrobial Susceptibility Testing for Human Mycoplasmas; Approved Guideline. 31. CLSI document M43-A. Wayne, PA:Clinical and Laboratory Standards Institute; 2011
11. Mardassi BBA, Aissani N, Moalla I, Dhahri D, Dridi A, Mlik B. Evidence for the predominance of a single tet(M) gene sequence type in tetracycline-resistant Ureaplasma parvum and Mycoplasma hominis isolates from Tunisian patients. J Med Microbiol. 2012;61(Pt 9):1254–1261. doi:doi:10.1099/jmm.0.044016-0
12. Pereyre S, Gonzalez P, De Barbeyrac B, et al. Mutations in 23S rRNA account for intrinsic resistance to macrolides in Mycoplasma hominis and Mycoplasma fermentans and for acquired resistance to macrolides in M. hominis. Antimicrob Agents Chemother. 2002;46(10):3142–3150. doi:10.1128/AAC.46.10.3142-3150.2002
13. Meygret A, Le Roy C, Renaudin H, Bébéar C, Pereyre S. Tetracycline and fluoroquinolone resistance in clinical Ureaplasma spp. and Mycoplasma hominis isolates in France between 2010 and 2015. J Antimicrobial Chemother. 2018;73(10):2696–2703. doi:doi:10.1093/jac/dky238
14. Boujemaa S, Mlik B, Ben Allaya A, Mardassi H, Ben Abdelmoumen Mardassi B. Spread of multidrug resistance among Ureaplasma serovars, Tunisia. Antimicrob Resist Infect Control. 2020;9(1):19. doi:doi:10.1186/s13756-020-0681-5
15. Dégrange S, Renaudin H, Charron A, Bébéar C, Bébéar CM. Tetracycline resistance in Ureaplasma spp. and Mycoplasma hominis: prevalence in Bordeaux, France, from 1999 to 2002 and description of two tet(M)-positive isolates of M. hominis susceptible to tetracyclines. Antimicrob Agents Chemother. 2008;52(2):742–744. doi:doi:10.1128/AAC.00960-07
16. Vidovic S, Thakur SD, Horsman GB, Levett PN, Anvari V, Dillon J-AR. Longitudinal analysis of the evolution and dissemination of neisseria gonorrhoeae strains (Saskatchewan, Canada, 2005 to 2008) reveals three major circulating strains and convergent evolution of ciprofloxacin and azithromycin resistance. J Clin Microbiol. 2012;50(12):3823–3830. doi:doi:10.1128/JCM.01402-12
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.