Back to Journals » Infection and Drug Resistance » Volume 15
Clinical Perspective of Antimicrobial Resistance in Bacteria
Authors Zhu Y ![]() , Huang WE, Yang Q
, Huang WE, Yang Q
Received 21 October 2021
Accepted for publication 18 January 2022
Published 2 March 2022 Volume 2022:15 Pages 735—746
DOI https://doi.org/10.2147/IDR.S345574
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Suresh Antony
Ying Zhu,1,2 Wei E Huang,3 Qiwen Yang1
1Department of Clinical Laboratory, State Key Laboratory of Complex Severe and Rare Diseases, Peking Union Medical College Hospital, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing, People’s Republic of China; 2Graduate School, Peking Union Medical College, Chinese Academy of Medical Sciences, Beijing, People’s Republic of China; 3Department of Engineering Science, University of Oxford, Oxford, OX1 3PJ, UK
Correspondence: Qiwen Yang; Wei E Huang, Email [email protected]; [email protected]
Abstract: Antimicrobial resistance (AMR) has become a global clinical problem in recent years. With the discovery of antibiotics, infections were not a deadly problem for clinicians as they used to be. However, worldwide AMR comes with the overuse/misuse of antibiotics and the spread of resistance is deteriorated by a multitude of mobile genetic elements and relevant resistant genes. This review provides an overview of the current situation, mechanism, epidemiology, detection methods and clinical treatment for antimicrobial resistant genes in clinical important bacteria including methicillin-resistant Staphylococcus aureus (MRSA), vancomycin-resistant Enterococcus (VRE), penicillin-resistant Streptococcus pneumoniae (PRSP), extended-spectrum β-lactamase-producing Enterobacteriaceae, acquired AmpC β-lactamase-producing Enterobacteriaceae, carbapenemase-producing Enterobacteriaceae (CPE), multidrug-resistant (MDR) Acinetobacter baumannii and Pseudomonas aeruginosa.
Keywords: antimicrobial resistant, genes, antibiotic resistance mechanisms, epidemiology, detection methods
Introduction
The global spread of antimicrobial resistance (AMR) is a serious threat to global public health. AMR not only causes soaring economic burden on health care but also increases morbidity and mortality. When microorganisms (such as bacteria, fungi, viruses, and parasites) are exposed to antimicrobial drugs (such as antibiotics, antifungals and antivirals), they respond and develop AMR. As a result, the anti-microbial drugs become less effective gradually. Infections persistent in human body promotes the risk of spread. As O’neill et al described in 2016,2 death rates due to resistant infections were 700,000 per year and the infectious population will reach 50M people globally in 2050. On one hand, the supply of new antibiotics is insufficient to keep pace with the increase of AMR pathogens. On the other hand, unnecessary use of antibiotics globally further selectively enriches AMR pathogens, increasing health risks. In addition, AMR will compromise the effective therapies against other diseases such as cancer chemotherapy, treatment of HIV and malaria. It is important to understand the AMR mechanisms and develop rapid point-of-care diagnostic test.
The abuse of antimicrobial drugs increases the background concentration of antibiotics, and creates niches favoring AMR bacteria. It is reported that horizontal gene transfer (HGT) mechanism could play an important role to the rapid dissemination of genes for AMR via natural ecosystem and human microbiome.3–10 There are three mechanisms of HGT: gene transformation, conjugation and transduction. Gene transformation involves bacterial uptake of naked DNA from environment, conjugation is usually plasmid-mediated process between a donor and a recipient bacterium, and transduction is associated with bacterial phage infection. All three HGT mechanisms can rapidly spread genes for AMR among microbial community in natural environment and human microbiome, and be selected in the presence of antimicrobial drugs. Specific terminologies describing acquired resistance are explained in Box 1. Gene database involved in AMR have been well documented.11 To date, 263 pathogens, 4937 reference sequences have been collected by Comprehensive Antibiotic Resistance Database (CARD, https://card.mcmaster.ca/).
In 2017, the World Health Organization (WHO) published a list of bacteria for which researches and drugs were urgently needed.12 In the list, carbapenem-resistant Acinetobacter baumannii, carbapenem-resistant Pseudomonas aeruginosa, carbapenem-resistant Enterobacteriaceae and ESBL-producing Enterobacteriaceae were listed as critical priority pathogens, while vancomycin-resistant Enterococcus (VRE) faecium, methicillin-resistant Staphylococcus aureus (MRSA) were listed as high priority pathogens. What’s more, penicillin-non-susceptible Streptococcus pneumoniae was listed as medium priority pathogens. Now many reviews are reported about certain AMR rather than a broader conclusion of AMR. Our study summarizes the prevalent AMR genes, AMR mechanisms, epidemiology, detection methods and medication. Hence, taking morbidity and mortality into consideration, in this review, we focus on 3 gram-positive bacteria including MRSA, VRE, penicillin-resistant Streptococcus Pneumoniae (PRSP); and 5 gram-negative bacteria including extended-spectrum β-lactamase-producing Enterobacteriaceae, acquired AmpC β-lactamase-producing Enterobacteriaceae, carbapenemase-producing Enterobacteriaceae (CPE), multidrug-resistant (MDR) Acinetobacter baumannii and Pseudomonas aeruginosa. The antimicrobial resistant bacteria discussed are all clinically important with wide prevalence or/and high risk of death.
AMR in Gram-Positive Bacteria
Methicillin-Resistant Staphylococcus aureus
From 1950s to 1980s, MRSA developed from a newcomer to a worldwide superbug and is still spread around not only nosocomial but also outside hospital. MRSA are generally coupled with high mortality and morbidity which arouses great concerns due to its clinical importance. mecA is the most popular relevant gene mediating methicillin-resistant in Staphylococcus aureus. It encodes PBP2a, a transpeptidase with lower affinity for the β-lactam antibiotics, which mediates the methicillin resistance in MRSA. Except mecA, other homologues (mecB, mecC, mecD) are discovered recently. These methicillin-resistant genetic components are carried on staphylococcal cassette chromosome mec (SCCmec), a mobile genetic element (MGE).13,14 mecB is often described in a transposon mec complex (Tn6045) in Macrococcus caseolyticus. Although mecC was reported in livestock-associated MRSA, mecC in MRSA can be transmitted between species.15 mecD has been recently spotted in bovine and canine M. caseolyticus isolates.16
Vancomycin-Resistant Enterococcus
From 1970s, vancomycin was approved to treat Enterococcus which came with the appearance of VRE. Until early 1990s, VRE became a serious problem in America, and great efforts have been made to prevent VRE.17 Vancomycin resistance in Enterococcus is related to various van genotypes, including vanA, vanB, vanM and another 7 types (vanC/D/E/F/G/L/N).18 It is reported that Enterococcus faecium (E. faecium) features with high recombination rates due to the lack of CRISPR-cas loci, which protect the conservation of genomic DNA in other bacteria.18 The van genes confer modifications to the d-Ala-d-Ala dipeptide at the C-terminus end of the translocated pentapeptide which is the key to formidable antimicrobial effect of glycopeptides against enterococci reduces the affinity of vancomycin binding by up to 1000 times, thus losing its efficacy.18 Genes of vanA, vanB and vanM deserve more clinical attention because they tend to cause intermediate to high levels of resistance to vancomycin in Enterococcus. The vanM, vanA, vanB, and vanD is genetically and phenotypically similar, whereas vanL and vanN are similar to vanC.19
Penicillin-Resistant Streptococcus Pneumoniae
S. pneumoniae has six types of PBPs, of which three PBPs mutation are related to penicillin-resistance: PBP1A, PBP2X and PBP2B.20 PBPs mutation is the main mechanism for PRSP to acquire penicillin-resistance. These resistant isolates of pneumococci survived after the selection of abundant treatment of bacterial infections with β-lactams and evolved gradually through accumulation of spontaneous mutations coupled with recombination of alleles from other β-lactam resistant group streptococci.21
AMR in Gram-Negative Bacteria
AMR in Enterobacteriaceae
The most popular resistant mechanism in gram-negative bacteria is hydrolytic enzymes, especially β-lactamase, a group of enzymes, which can hydrolyze the β-lactam ring to break the amide bond and inactivate the antibacterial activity of drugs. Various mechanisms of antibacterial resistance to β-lactams reported include ESBLs, AmpC, and carbapenemases.22
 |
Box 1 MDR, XDR and PDR |
Extended-Spectrum β-Lactamase-Producing (ESBL-Producing) Enterobacteriaceae
ESBLs are frequently detected in Escherichia coli (E. coli) and Klebsiella pneumoniae, which are representative MDR gram-negative bacteria.23 Most of the ESBLs belong to Ambler class A and they are generally inhibited by clavulanic acid or tazobactam.24 β-lactmases related to the AMR of ESBLs include CTX-M (blaCTX-M genes), TEM (blaTEM genes) and SHV (blaSHV genes).25 Recently, rapid worldwide spread of ST131 E. coli strains, which contain plasmids harboring CTX-M ESBL genes especially blaCTX-M-15, has attracted great attention.23 Some special OXA-type β-Lactamase, OXA‐10 and OXA‐13 to OXA‐19, also hydrolyze extended spectrum cephalosporins and they are also regarded as ESBLs.26 Currently, E.coli with CTX-Ms are the most widespread ESBLs-producing bacteria and CTX-M-15 is the most frequently detected resistant gene, followed by CTX-M-14, which is often found in South-East Asia.27 Furthermore, recent epidemiology studies have discovered CTX-M-27 in Japan and Europe.25 The population structure of ESBL-producing E. coli is dominated globally by a high-risk strain named ST131.28
Acquired AmpC β-Lactamase-Producing Enterobacteriaceae
AmpC β-lactamases, which are clinically important cephalosporinases, are mostly class C β-lactamases that also hydrolyze 3rd generation cephalosporins, but are not inhibited by clavulanic acid or tazobactam.24 However, the name of AmpC is not accurate since several so-called enzymes in the literature actually belong to class A.29 AmpC enzymes are encoded on chromosome and plasmids. The AmpC β-lactamases are at a low expression level in E. coli and the AmpC-encoded gene is absent in the chromosome of Klebsiella and Salmonella strains.30 However, plasmid-expressed AmpC β-lactamases can cause relevant resistance in these bacteria. Plasmid-mediated AmpC enzymes have been named according to the resistant drugs (CMY, FOX, MOX, LAT), according to the type of enzyme (ACC, ACT) or based on the site of discovery, such as MIR-1 or DHA.26
Carbapenemase-Producing Enterobacteriaceae
Clonal spread from chromosomes and plasmid-mediated transmission contribute to continued rise in CPE incidence.31 The blaKPC, blaNDM, blaOXA, blaIMP and blaVIM are the dominant carbapenemase gene families.32 These carbapenemase genes can be defined by Ambler classification system: Ambler class A (KPC enzymes); molecular class B (NDM, VIM, and IMP enzymes); and class D (OXA enzymes).33 As the most important antibacterial agents for multidrug-resistant infections of gram-negative Bacilli spp, carbapenem and colistin are the last resource to treat such bacterial infections. Unfortunately, bacteria have evolved carbapenemase, one of their arsenals to resist drugs.
AMR in Non-Fermentative Bacteria
Multidrug-Resistant Acinetobacter Baumannii and Pseudomonas Aeruginosa
Acinetobacter baumannii and Pseudomonas aeruginosa are critical bacteria among global priority list of antibiotic-resistant bacteria that are identified by WHO12 and these troublesome MDR bacteria have potentials to cause a wide prevalence of infections and outbreaks. Acinetobacter baumannii is naturally transformable to take naked DNA and Pseudomonas aeruginosa tends to exchange AMR genes by conjugation.34 Both bacteria have various antimicrobial mechanisms, including enzymes, efflux pumps, modification of aminoglycosides, permeability defects, and alteration of target sites.35 ESBLs, metallo beta lactamases (MBLs), oxacillinases (OXAs), aminoglycoside-modifying enzymes (AMEs), 16S-rRNA methylase are all relevant to producing of enzymes in these non-fermentative bacteria.35–37 Efflux pumps are special mechanisms which can confer resistance to β-lactams, polymyxin, tigecycline and fluoroquinolones. Quinolone Resistance Determining region (QRDR) is quite important for aeruginosa and baumannii, involved in protein mutation and efflux pumps. The relevant drug resistance genes of Acinetobacter baumannii and Pseudomonas aeruginosa are listed in Table 1.
 |
Table 1 Antimicrobial Resistant Mechanisms of Acinetobacter Baumannii and Pseudomonas Aeruginosa |
Antibiotic Resistance Mechanisms
AMR is naturally occurring as a reaction of microbial organisms to environment. However, the acquired resistance in clinical settings is the result of mutations in chromosomal genes or the transmission of external genomic determinants of resistance.24 The AMR mechanisms mainly include enzymatic drug inactivation, drug target alteration, changes in outer membrane permeability and active efflux of antimicrobial compounds.
Enzymatic Drug Inactivation
Bacteria produce enzymes that can destroy antibiotics or cause them to lose their antibacterial action, causing the drug to be destroyed or fail before it acts on the bacteria cell.22 There are three main kinds of drug inactivating enzyme: hydrolase (mainly β-lactamase), passivation enzyme (aminoglycoside inactivating enzyme, chloramphenicol acetyltransferase, erythromycin esterase, etc.) and modified enzyme (aminoglycoside modifying enzyme). The enzymes encoded by both chromosomal and plasmid genes can target and cleave the vulnerable hydrolytically susceptible chemical bonds in drugs.38 The enzymatic genes of the most clinical concerns harbor amidases that cut down the β-lactam ring of the penicillin and cephalosporin classes of drugs, including blaCTX-M, blaTEM, blaSHV, blaOXA, blaCMY, blaFOX, blaMOX, blaDHA, blaKPC, blaNDM, blaVIM, blaIMP, blaOXA and so on.38
Drug Target Alteration
There are many antibiotic-binding targets in the bacteria. The target site alteration can make the antibiotics difficult to bind to the bacteria, which is an important mechanism for drug resistance. This mechanism mainly reflects in the drug-resistance of gram-positive bacteria and polymyxin-resistance. For example, the PBP of Staphylococcus aureus is converted to PBP2a (encoded by the mecA gene), and the latter is a low-affinity binding protein, resulting in resistance to all β-lactam antibiotics.24,29 The polymyxin resistance is mainly caused by the modification of the lipid A moiety of lipopolysaccharide (LPS), which is the primary target of polymyxin due to genes in PmrA/PmrB, PhoP/PhoQ, ParR/ParS, ColR/ColS or CprR/CprS two-component systems or plasmid-mediated mcr genes.39
Changes in Outer Membrane Permeability
In gram-negative bacteria, β-lactam antibiotics mainly pass through the outer membrane by hydrophilic channel proteins, and mutations leading to channel protein alteration or decreased expression will make the bacteria less sensitive to various β-lactams. For example, the loss of Opal D2 channel protein in patina causes resistance to imipenem.24,29 The low outer membrane permeability of Pseudomonas aeruginosa gives it high intrinsic resistance to antiseptics and antibiotics.40 The mutation in genes encoding outer membrane porins is expected to affect drug susceptibility.
Active Efflux of Antimicrobial Compounds
It is also known as the efflux pump system or the drug pumping system. The drug concentration in the bacteria is insufficient to exert an antibacterial effect, resulting in drug resistance. This process requires energy and acts on a variety of antibiotics. For example: Staphylococcus aureus resistance to quinolones; OprK protein of Pseudomonas aeruginosa outer membrane can transport various antibiotics to the outside of bacteria.24,29,41 The most important efflux transporters are resistance-nodulation-division (RND), major facilitator superfamily (MFS), multidrug and toxic compound extrusion (MATE), small multidrug-resistance (SMR), and ATP-binding cassette (ABC) superfamilies or families. Each efflux families possess several important genes, such as acrB, mdtF in RND, bcr, cmr in MFS, mdtK, yeeO in MATE, emrE, ydgE in SMR and macB in ABC.42
Epidemiology
With severe clinical situation of antimicrobial resistant bacteria infections, some specialists and organizations have begun to monitor the prevalence of different drug-resistant organisms. Multilocus sequence typing (MLST), with high resolution, and whole genome sequencing (WGS), with comprehensive information, are the common methods to type antimicrobial resistant bacteria in the molecular level.
For Gram-positive bacteria, the main antimicrobial resistant bacteria include MRSA, VRE and PRSP. MRSAs are widely distributed all over the world, especially in South America, North America and Japan with high prevalence rate of over 40%, while PRSP is a challenge for Europe.43–45 As for VRE, Enterococcus faecium is more ubiquitous than Enterococcus faecalis.43–50
As for gram-negative bacteria, the main antimicrobial resistant bacteria include cephalosporin- and/or carbapenem-resistant Escherichia coli and Klebsiella pneumoniae, and carbapenem-resistant Acinetobacter baumannii and Pseudomonas aeruginosa. The global spread of ESBL-producing bacteria in the community since the 2000s has threatened the public health. In recent years, the clinical medication and intensity of carbapenem antibiotics have increased year by year, and the carbapenem resistance rate of clinical strains (including Acinetobacter baumannii, Pseudomonas aeruginosa and Klebsiella pneumoniae) is of an increasing trend. MDR Acinetobacter baumannii and Pseudomonas aeruginosa may gradually turn into a major clinical challenge.23,50
To be more specific, information about global AMR is plotted in Table 2 to present the difference situation in 9 countries.23,43–52 This table indicates the percentage of prevalent antimicrobial resistant bacteria in different countries of six continents relatively. The data were collected from references and several surveillance systems and monitoring reports. Australia is on behalf of Oceania, Argentina on behalf of South America, USA and Canada on behalf of South America, UK on behalf of Europe, China, India, and Japan on behalf of Asia and South Africa on behalf of Africa. The data of Australia were collected from Australian Group on Antimicrobial Resistance (AGAR),46–48 Argentina, South Africa and Japan from The Center for Disease Dynamics, Economics & Policies,43 USA from reports or articles published by Chong et al.23 Monaco et al51 and Sader et al.52 Canada from Canadian Antimicrobial Resistance Alliance (CARA),44 UK from article published by Chong Yong et al23 and European Centre for Disease Prevention and Control (ECDC),45 India from India Council of Medical Research49 and China from article published by Chong et al23 and China Antimicrobial Resistance Surveillance System (CARSS) monitoring report.50 MRSA, VRE (faecalis), VRE (faecium), PRSP, ESBL, CRkp, MDR-PA and MDR-AB are included in the study.
 |
Table 2 The Percentage of Prevalent Antimicrobial Resistant Bacteria from 9 Countries on 6 Continents |
According to obtained data, VRE (faecium), accounting for 50%, 68% and 69% ranks the first in Australia, USA and Argentina; MDR-PA (21%) in Canada; VRE (22.2%) in UK; MDR-AB (56.1%) in China; CRkp (59%) in India; MRSA (41% and 27%, respectively) in Japan and South Africa.23,43–52
Detection Methods
Antimicrobial Susceptibility Testing
Antimicrobial susceptibility tests (ASTs) include K-B test, dilution test, Epsilometer test (E-test) and commercialized automatic drug susceptibility detection and analysis systems. K-B test uses antibiotic-containing wafers or disks, sometimes added with enzyme inhibitors, to test whether specific bacteria strains are sensitive to specific antibiotics. Broth microdilution test, the gold standard of AST, is used to quantitatively determine the minimal inhibitory concentration (MIC) of antimicrobial agent to inhibit or kill the bacteria.53 MIC is standardized by globally recognized institutions such as CLSI and EUCAST. To be more convenient, E-test comes into the market as a plastic strip with a gradient concentration of antimicrobial agents impregnated in it. With advanced technology, commercial systems with consistent and rapid results, including bioMérieux VITEK® 2 Automated instrument for ID/AST testing, Sensititre™ Complete Automated AST System and BD Phoenix™ automated identification and susceptibility testing system, are recommended for clinical microbiology laboratories.
Matrix-Assisted Laser Desorption/Ionization Time of Flight Mass Spectrometry (MALDI-TOF MS)
MALDI-TOF MS is widely used in the rapid identification of bacteria based on the detection of proteins. Furthermore, MALDI-TOF MS can be utilized in AST with the combination of other techniques, such as surface-enhanced Raman scattering (SERS) and isotope incorporation.54
Molecular Biology Tests
With the rapid development of molecular biology, it’s widely used in microbiology, especially in the tests of AMR genes of MRSA, tuberculosis and superbugs. Methods that are commonly used in laboratory include polymerase chain reaction (PCR), quantitative PCR (qPCR), Real-Time Fluorescent PCR Assay (RT-PCR), DNA Microarrays/Genechip and WGS, which are all in need of large volumes of blood and specific for antibiotic resistance gene identification.54
Raman Micro-Spectroscopy
Raman micro-spectroscopy is a non-invasive and label-free technology which provides an intrinsic biochemical “fingerprint” of a single bacterial cell.55 Raman micro-spectroscopy coupled with the stable-isotope technique using heavy water (D2O rather than H2O) has been proven as a universal method to evaluate in vivo metabolic activity of a cell.56–60 In the presence of heavy water (D2O), metabolically active cells will form a C-D band by incorporating D (deuterium) from D2O via NADPH electron transport chain. It has been confirmed that C-D Raman biomarker in Single-Cell Raman Spectra quantitatively indicates general metabolic activity of a cell. The higher the metabolic activity, the higher the C-D intensity. When cells are exposed to D2O and antibiotics in AST test, the C-D band in cells becomes detectable in as short a time as 20 minutes and reaches the highest value after 1–3 hours.56–60 Raman micro-spectroscopy has also been applied to bacterial identification.55,61
Microfluidic Chip
Microfluidic Chip has been tested in identification and AST of multiple uropathogens for research, which combines the spatial resolution of the cell culture arrays and the color resolution from the chromogenic reaction. The AST is determined by MIC which is reflected from the degree of chromogenic reaction.62,63
Clinical Treatment for AMR Bacteria
Due to various AMR bacteria, clinical medication is in a dilemma where clinicians worried that no effective drugs to fight against superbugs in the future. Table 3 listed recommendations of medications for AMR bacteria according to update guidelines and references.
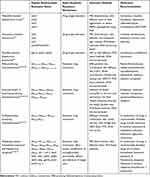 |
Table 3 Major Antibiotic Resistance Mechanisms, Detection Methods and Medical Recommendations for Popular Antimicrobial Resistance Genes of Antimicrobial Resistant Bacteria |
Conclusion
The study of drug-resistant bacterial genotypes, epidemiology, detection methods and clinical treatment is meaningful for monitoring, diagnosing and treating of drug-resistant colonies in nosocomial or community infections outbreaks. Quick and accurate detection is the primary assurance to handle worsened situation. Traditional AST still needs more research to maximize the revealed information and shorten the sample turn-around time. Under the guidance of the concept of “One Health”, focus on the AMR genes should not only be restricted among people but also in food animals and even plants.
Acknowledgments
This study was supported by National Natural Science Foundation of China (82072318), National Key Research and Development Program of China (2021YFC2301002), Beijing Key Clinical Specialty for Laboratory Medicine - Excellent Project (No. ZK201000). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author Contributions
All authors made a significant contribution to the work reported, whether that is in the conception, study design, execution, acquisition of data, analysis and interpretation, or in all these areas; took part in drafting, revising or critically reviewing the article; gave final approval of the version to be published; have agreed on the journal to which the article has been submitted; and agree to be accountable for all aspects of the work.
Disclosure
We have no conflict of interest.
References
1. Magiorakos AP, Srinivasan A, Carey RB, et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect. 2012;18(3):268–281. doi:10.1111/j.1469-0691.2011.03570.x
2. O’neill J. Tackling drug-resistant infections globally; 2016.
3. Partridge SR, Kwong SM, Firth N, Jensen SO. Mobile genetic elements associated with antimicrobial resistance. Clin Microbiol Rev. 2018;31(4):UNSPe00088–17. doi:10.1128/cmr.00088-17
4. Heuer H, Schmitt H, Smalla K. Antibiotic resistance gene spread due to manure application on agricultural fields. Curr Opin Microbiol. 2011;14(3):236–243. doi:10.1016/j.mib.2011.04.009
5. von Wintersdorff CJH, Penders J, van Niekerk JM, et al. Dissemination of antimicrobial resistance in microbial ecosystems through horizontal gene transfer. Front Microbiol. 2016;7:173. doi:10.3389/fmicb.2016.00173.
6. Carattoli A. Plasmids and the spread of resistance. Int J Med Microbiol. 2013;303(6–7):298–304. doi:10.1016/j.ijmm.2013.02.001
7. Modi SR, Collins JJ, Relman DA. Antibiotics and the gut microbiota. J Clin Invest. 2014;124(10):4212–4218. doi:10.1172/jci72333
8. Liu YY, Wang Y, Walsh TR, et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect Dis. 2016;16(2):161–168. doi:10.1016/s1473-3099(15)00424-7
9. Zhu YG, Johnson TA, Su JQ, et al. Diverse and abundant antibiotic resistance genes in Chinese swine farms. Proc Natl Acad Sci USA. 2013;110(9):3435–3440. doi:10.1073/pnas.1222743110
10. Karkman A, Do TT, Walsh F, Virta MPJ. Antibiotic-resistance genes in waste water. Trends Microbiol. 2018;26(3):220–228. doi:10.1016/j.tim.2017.09.005
11. Jia B, Raphenya AR, Alcock B, et al. CARD 2017: expansion and model-centric curation of the comprehensive antibiotic resistance database. Nucleic Acids Res. 2017;45(D1):D566–D573. doi:10.1093/nar/gkw1004
12. World Health Organization. WHO publishes list of bacteria for which now antibiotics are urgently needed. Available from: https://www.who.int/news/item/27-02-2017-who-publishes-list-of-bacteria-for-which-new-antibiotics-are-urgently-needed.
13. Lakhundi S, Zhang K. Methicillin-resistant Staphylococcus aureus: molecular characterization, evolution, and epidemiology. Clin Microbiol Rev. 2018;31(4). doi:10.1128/CMR.00020-18
14. Liu J, Chen D, Peters BM, et al. Staphylococcal chromosomal cassettes mec (SCCmec): a mobile genetic element in methicillin-resistant Staphylococcus aureus. Microb Pathog. 2016;101:56–67. doi:10.1016/j.micpath.2016.10.028
15. Paterson GK, Harrison EM, Holmes MA. The emergence of mecC methicillin-resistant Staphylococcus aureus. Trends Microbiol. 2014;22(1):42–47. doi:10.1016/j.tim.2013.11.003
16. Becker K, van Alen S, Idelevich EA, et al. Plasmid-encoded transferable mecb-mediated methicillin resistance in Staphylococcus aureus. Emerg Infect Dis. 2018;24(2):242–248. doi:10.3201/eid2402.171074
17. Frieden TR, Munsiff SS, Low DE, et al. Emergence of vancomycin-resistant enterococci in New York City. Lancet. 1993;342(8863):76–79. doi:10.1016/0140-6736(93)91285-t
18. Lee T, Pang S, Abraham S, Coombs GW. Antimicrobial-resistant CC17 Enterococcus faecium: the past, the present and the future. J Glob Antimicrob Resist. 2019;16:36–47. doi:10.1016/j.jgar.2018.08.016
19. Ahmed MO, Baptiste KE. Vancomycin-resistant enterococci: a review of antimicrobial resistance mechanisms and perspectives of human and animal health. Microb Drug Resist. 2018;24(5):590–606. doi:10.1089/mdr.2017.0147
20. El Moujaber G, Osman M, Rafei R, Dabboussi F, Hamze M. Molecular mechanisms and epidemiology of resistance in Streptococcus pneumoniae in the Middle East region. J Med Microbiol. 2017;66(7):847–858. doi:10.1099/jmm.0.000503
21. Straume D, Stamsas GA, Havarstein LS. Natural transformation and genome evolution in Streptococcus pneumoniae. Infect Genet Evol. 2015;33:371–380. doi:10.1016/j.meegid.2014.10.020
22. Dever LA, Dermody TS. Mechanisms of bacterial resistance to antibiotics. Arch Intern Med. 1991;151(5):886–895. doi:10.1001/archinte.1991.00400050040010
23. Chong Y, Shimoda S, Shimono N. Current epidemiology, genetic evolution and clinical impact of extended-spectrum beta-lactamase-producing Escherichia coli and Klebsiella pneumoniae. Infect Genet Evol. 2018;61:185–188. doi:10.1016/j.meegid.2018.04.005
24. Munita JM, Arias CA. Mechanisms of antibiotic resistance. Microbiol Spectr. 2016;4(2). doi:10.1128/microbiolspec.VMBF-0016-2015
25. Peirano G, Pitout JDD. Extended-spectrum beta-lactamase-producing Enterobacteriaceae: update on molecular epidemiology and treatment options. Drugs. 2019;79(14):1529–1541. doi:10.1007/s40265-019-01180-3
26. Smet A, Martel A, Persoons D, et al. Broad-spectrum beta-lactamases among Enterobacteriaceae of animal origin: molecular aspects, mobility and impact on public health. FEMS Microbiol Rev. 2010;34(3):295–316. doi:10.1111/j.1574-6976.2009.00198.x
27. Bevan ER, Jones AM, Hawkey PM. Global epidemiology of CTX-M beta-lactamases: temporal and geographical shifts in genotype. J Antimicrob Chemother. 2017;72(8):2145–2155. doi:10.1093/jac/dkx146
28. Ghosh H, Doijad S, Falgenhauer L, Fritzenwanker M, Imirzalioglu C, Chakraborty T. blaCTX-M-27-encoding Escherichia coli sequence type 131 lineage C1-M27 clone in clinical isolates, Germany. Emerg Infect Dis. 2017;23(10):1754–1756. doi:10.3201/eid2310.170938
29. Jacoby GA. AmpC beta-lactamases. Clin Microbiol Rev. 2009;22(1):161–82, Table of Contents. doi:10.1128/CMR.00036-08
30. Xia J, Gao J, Tang W. Nosocomial infection and its molecular mechanisms of antibiotic resistance. Biosci Trends. 2016;10(1):14–21. doi:10.5582/bst.2016.01020
31. van Duin D, Doi Y. The global epidemiology of carbapenemase-producing Enterobacteriaceae. Virulence. 2017;8(4):460–469. doi:10.1080/21505594.2016.1222343
32. Samuelsen O, Overballe-Petersen S, Bjornholt JV, et al. Molecular and epidemiological characterization of carbapenemase-producing Enterobacteriaceae in Norway, 2007 to 2014. PLoS One. 2017;12(11):e0187832. doi:10.1371/journal.pone.0187832
33. Jesús Rodríguez-Baño B-G-G, Machuca I, Pascuala A. Treatment of infections caused by extended-spectrum-betalactamase-, ampc-, and carbapenemase-producing Enterobacteriaceae. Clin Microbiol Rev. 2017. doi:10.1128/CMR.00079-17
34. Dijkshoorn L, Nemec A, Seifert H. An increasing threat in hospitals: multidrug-resistant Acinetobacter baumannii. Nat Rev Microbiol. 2007;5(12):939–951. doi:10.1038/nrmicro1789
35. Lee CR, Lee JH, Park M, et al. Biology of Acinetobacter baumannii: pathogenesis, antibiotic resistance mechanisms, and prospective treatment options. Front Cell Infect Microbiol. 2017;7:55. doi:10.3389/fcimb.2017.00055
36. Antunes LC, Visca P, Towner KJ. Acinetobacter baumannii: evolution of a global pathogen. Pathog Dis. 2014;71(3):292–301. doi:10.1111/2049-632X.12125
37. Subedi D, Vijay AK, Willcox M. Overview of mechanisms of antibiotic resistance in Pseudomonas aeruginosa: an ocular perspective. Clin Exp Optom. 2018;101(2):162–171. doi:10.1111/cxo.12621
38. Wright GD. Bacterial resistance to antibiotics: enzymatic degradation and modification. Adv Drug Deliv Rev. 2005;57(10):1451–1470. doi:10.1016/j.addr.2005.04.002
39. Olaitan AO, Morand S, Rolain JM. Mechanisms of polymyxin resistance: acquired and intrinsic resistance in bacteria. Front Microbiol. 2014;5:643. doi:10.3389/fmicb.2014.00643
40. Chevalier S, Bouffartigues E, Bodilis J, et al. Structure, function and regulation of Pseudomonas aeruginosa porins. FEMS Microbiol Rev. 2017;41(5):698–722. doi:10.1093/femsre/fux020
41. Correia S, Poeta P, Hébraud M, Capelo JL, Igrejas G. Mechanisms of quinolone action and resistance: where do we stand? J Med Microbiol. 2017;66(5):551–559. doi:10.1099/jmm.0.000475
42. Li XZ, Plesiat P, Nikaido H. The challenge of efflux-mediated antibiotic resistance in gram-negative bacteria. Clin Microbiol Rev. 2015;28(2):337–418. doi:10.1128/CMR.00117-14
43. The center for disease dynamics Economics & Policy. ResistanceMap: Antibiotic Resistance. Available from: https://resistancemap.cddep.org/AntibioticResistance.php.
44. CARA. Antibiogram. Available from: http://www.can-r.com/study.php?study=antb2017&year=2017.
45. ECED. Surveillance Atlas of Infectious Diseases. Available from: https://atlas.ecdc.europa.eu/public/index.aspx?Dataset=27&HealthTopic=4.
46. Coombs GW, Daley DA, Lee YT, Pang S. Australian Group on Antimicrobial Resistance (AGAR) Australian Enterococcal Sepsis Outcome Programme (AESOP) annual report 2017. Commun Dis Intell (2018). 2019;43. doi:10.33321/cdi.2019.43.42.
47. Bell JM, Gottlieb T, Daley DA, Coombs GW. Australian Group on Antimicrobial Resistance (AGAR) Australian Gram-negative Sepsis Outcome Programme (GNSOP) annual report 2017. Commun Dis Intell (2018). 2019;43. doi:10.33321/cdi.2019.43.37.
48. Coombs GW, Daley DA, Lee YT, Pang S. Australian Group on Antimicrobial Resistance (AGAR) Australian Staphylococcus aureus Sepsis Outcome Programme (ASSOP) Annual Report 2017. Commun Dis Intell (2018). Sep 16 2019;43. doi:10.33321/cdi.2019.43.43.
49. Research ICoM. AMRSN_Annual_Report_2018; 2018.
50. CARSS. 2019 national bacterial resistance monitoring report; 2019.
51. Monaco M, Pimentel de Araujo F, Cruciani M, Coccia EM, Pantosti A. Worldwide epidemiology and antibiotic resistance of Staphylococcus aureus. Curr Top Microbiol Immunol. 2017;409:21–56. doi:10.1007/82_2016_3
52. Sader HS, Castanheira M, Arends SJR, Goossens H, Flamm RK. Geographical and temporal variation in the frequency and antimicrobial susceptibility of bacteria isolated from patients hospitalized with bacterial pneumonia: results from 20 years of the SENTRY Antimicrobial Surveillance Program (1997–2016). J Antimicrob Chemother. 2019;74(6):1595–1606. doi:10.1093/jac/dkz074
53. Institute CaLS. Performance Standards for Antimicrobial Susceptibility Testing. M100-E302020. Wayne, PA: Clinical and Laboratory Standards Institute; 2020.
54. Dekter HE, Orelio CC, Morsink MC, et al. Antimicrobial susceptibility testing of Gram-positive and -negative bacterial isolates directly from spiked blood culture media with Raman spectroscopy. Eur J Clin Microbiol Infect Dis. 2017;36(1):81–89. doi:10.1007/s10096-016-2773-y
55. Huang WE, Griffiths RI, Thompson IP, Bailey MJ, Whiteley AS. Raman microscopic analysis of single microbial cells. Anal Chem. 2004;76(15):4452–4458. doi:10.1021/ac049753k
56. Song YZ, Cui L, Lopez JAS, et al. Raman-deuterium isotope probing for in-situ identification of antimicrobial resistant bacteria in Thames River. Sci Rep. 2017;7:16648. doi:10.1038/s41598-017-16898-x.
57. Tao Y, Wang Y, Huang S, et al. Metabolic-activity-based assessment of antimicrobial effects by D2O-labeled single-cell raman microspectroscopy. Anal Chem. 2017;89(7):4108–4115. doi:10.1021/acs.analchem.6b05051
58. Wang Y, Xu JB, Kong LC, et al. Raman-deuterium isotope probing to study metabolic activities of single bacterial cells in human intestinal microbiota. Microb Biotechnol. 2020;13(2):572–583. doi:10.1111/1751-7915.13519
59. Zhang SH, Guo LZ, Yang K, et al. Induction of Escherichia coli into a VBNC state by continuous-flow UVC and subsequent changes in metabolic activity at the single-cell level. Front Microbiol. 2018;9:2243. doi:10.3389/fmicb.2018.02243.
60. Berry D, Mader E, Lee TK, et al. Tracking heavy water (D2O) incorporation for identifying and sorting active microbial cells. Proc Natl Acad Sci U S A. 2015;112(2):E194–E203. doi:10.1073/pnas.1420406112/-/DCSupplemental
61. Wang Y, Ji YT, Wharfe ES, et al. Raman activated cell ejection for isolation of single cells. Anal Chem. 2013;85(22):10697–10701. doi:10.1021/ac403107p
62. Xu B, Du Y, Lin J, et al. Simultaneous identification and antimicrobial susceptibility testing of multiple uropathogens on a microfluidic chip with paper-supported cell culture arrays. Anal Chem. 2016;88(23):11593–11600. doi:10.1021/acs.analchem.6b03052
63. Sun H, Chan CW, Wang Y, et al. Reliable and reusable whole polypropylene plastic microfluidic devices for a rapid, low-cost antimicrobial susceptibility test. Lab Chip. 2019;19(17):2915–2924. doi:10.1039/c9lc00502a
64. Amin M, Navidifar T, Saleh Shooshtari F, Goodarzi H. Association of the genes encoding metallo-beta-lactamase with the presence of integrons among multidrug-resistant clinical isolates of Acinetobacter baumannii. Infect Drug Resist. 2019;12:1171–1180. doi:10.2147/IDR.S196575
65. Goic-Barisic I, Hrenovic J, Kovacic A, Music MS. Emergence of oxacillinases in environmental carbapenem-resistant Acinetobacter baumannii associated with clinical isolates. Microb Drug Resist. 2016;22(7):559–563. doi:10.1089/mdr.2015.0275
66. Murugan N, Malathi J, Umashankar V, Madhavan HN. Resistome and pathogenomics of multidrug resistant (MDR) Pseudomonas aeruginosa VRFPA03, VRFPA05 recovered from alkaline chemical keratitis and post-operative endophthalmitis patient. Gene. 2016;578(1):105–111. doi:10.1016/j.gene.2015.12.022
67. Blackwell GA, Holt KE, Bentley SD, Hsu LY, Hall RM. Variants of AbGRI3 carrying the armA gene in extensively antibiotic-resistant Acinetobacter baumannii from Singapore. J Antimicrob Chemother. 2017;72(4):1031–1039. doi:10.1093/jac/dkw542
68. Forsberg KJ, Patel S, Wencewicz TA, Dantas G. The tetracycline destructases: a novel family of tetracycline-inactivating enzymes. Chem Biol. 2015;22(7):888–897. doi:10.1016/j.chembiol.2015.05.017
69. Deng M, Zhu MH, Li JJ, et al. Molecular epidemiology and mechanisms of tigecycline resistance in clinical isolates of Acinetobacter baumannii from a Chinese university hospital. Antimicrob Agents Chemother. 2014;58(1):297–303. doi:10.1128/AAC.01727-13
70. Ostrer L, Khodursky RF, Johnson JR, Hiasa H, Khodursky A. Analysis of mutational patterns in quinolone resistance-determining regions of GyrA and ParC of clinical isolates. Int J Antimicrob Agents. 2019;53(3):318–324. doi:10.1016/j.ijantimicag.2018.12.004
71. Nang SC, Han ML, Yu HH, et al. Polymyxin resistance in Klebsiella pneumoniae: multifaceted mechanisms utilized in the presence and absence of the plasmid-encoded phosphoethanolamine transferase gene mcr-1. J Antimicrob Chemother. 2019;74(11):3190–3198. doi:10.1093/jac/dkz314
72. Mlynarcik P, Kolar M. Molecular mechanisms of polymyxin resistance and detection of mcr genes. Biomed Pap Med Fac Univ Palacky Olomouc Czech Repub. 2019;163(1):28–38. doi:10.5507/bp.2018.070
73. Moffatt JH, Harper M, Boyce JD. Mechanisms of polymyxin resistance. Adv Exp Med Biol. 2019;1145:55–71. doi:10.1007/978-3-030-16373-0_5
74. Li X, Quan J, Yang Y, et al. Abrp, a new gene, confers reduced susceptibility to tetracycline, glycylcine, chloramphenicol and fosfomycin classes in Acinetobacter baumannii. Eur J Clin Microbiol Infect Dis. 2016;35(8):1371–1375. doi:10.1007/s10096-016-2674-0
75. Alcalde-Rico M, Hernando-Amado S, Blanco P, Martinez JL. Multidrug efflux pumps at the crossroad between antibiotic resistance and bacterial virulence. Front Microbiol. 2016;7:1483. doi:10.3389/fmicb.2016.01483
76. Chuanchuen R, Narasaki CT, Schweizer HP. The MexJK efflux pump of Pseudomonas aeruginosa requires OprM for antibiotic efflux but not for efflux of triclosan. J Bacteriol. 2002;184(18):5036–5044. doi:10.1128/jb.184.18.5036-5044.2002
77. Kale P, Dhawan B. The changing face of community-acquired methicillin-resistant Staphylococcus aureus. Indian J Med Microbiol. 2016;34(3):275–285. doi:10.4103/0255-0857.188313
78. Thaden JT, Pogue JM, Kaye KS. Role of newer and re-emerging older agents in the treatment of infections caused by carbapenem-resistant Enterobacteriaceae. Virulence. 2017;8(4):403–416. doi:10.1080/21505594.2016.1207834
 © 2022 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms
and incorporate the Creative Commons Attribution
- Non Commercial (unported, 3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2022 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms
and incorporate the Creative Commons Attribution
- Non Commercial (unported, 3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
