Back to Journals » International Journal of General Medicine » Volume 14
Bioinformatics Analyses Reveal the Prognostic Value and Biological Roles of SEPHS2 in Various Cancers
Authors Zhang L, Zhao Q, Mao L, Li H, Zhuang M, Wang J, Liu Y, Qi M, Du X, Xia Z, Sun N, Liu Q, Chen H, Zhang R
Received 7 July 2021
Accepted for publication 2 September 2021
Published 24 September 2021 Volume 2021:14 Pages 6059—6076
DOI https://doi.org/10.2147/IJGM.S328222
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Scott Fraser
Luyu Zhang,1 Qianqian Zhao,1 Leilei Mao,2 Huanze Li,1 Miaoqing Zhuang,1 Jiayi Wang,1 Yue Liu,1 Meng Qi,3 Xiaoping Du,3 Zengrun Xia,3 Na Sun,4 Qiling Liu,4 Hongfang Chen,1,5,6 Rongqiang Zhang4
1School of Nursing, Shaanxi University of Chinese Medicine, Xianyang, Shaanxi, People’s Republic of China; 2School of Information Engineering, Chang’an University, Xi’an, Shaanxi, People’s Republic of China; 3Ankang R & D Center of Se-enriched Products, Ankang, Shaanxi, People’s Republic of China; 4School of Public Health, Shaanxi University of Chinese Medicine, Xianyang, Shaanxi, People’s Republic of China; 5Shaanxi Academy of Tradional Chinese Medicine, Xi’an, Shaanxi, People’s Republic of China; 6Shaanxi Provincial Hospital of Chinese Medicine, Xi’an, Shaanxi, People’s Republic of China
Correspondence: Hongfang Chen
Shaanxi Academy of Tradional Chinese Medicine, Shaanxi Provincial Hospital of Chinese Medicine, No. 4 Xihua Gate, Xi’an, Shaanxi, 710003; School of Nursing, Shaanxi University of Chinese Medicine, No. 1 Middle Section of Century Avenue, Xianyang, Shaanxi, 712046, People’s Republic of China
Email [email protected]
Rongqiang Zhang
School of Public Health, Shaanxi University of Chinese Medicine, No. 1 Middle Section of Century Avenue, Xianyang, Shaanxi, 712046, People’s Republic of China
Tel/Fax +86-029-38185219
Email [email protected]
Purpose: Selenophosphate synthetase 2 (SEPHS2) has been shown to regulate selenoprotein biosynthesis by catalyzing the synthesis of active selenium donor selenophosphate. SEPHS2 influences the survival of tumor cells. However, few studies have explored the expression level and prognostic of SEPHS2 in various cancers.
Methods: The expression of SEPHS2 in human tumor tissues and normal adjacent tissues was analyzed in The Cancer Genome Atlas (TCGA), Genotype-Tissue Expression (GTEx), Human Protein Atlas (HPA), and UALCAN databases. Cox regression analysis and Kaplan–Meier curve analysis were performed to analyze the association of SEPHS2 expression with the prognosis of cancer patients. The expression and prognosis of SEPHS2 in gliomas were further verified using the Chinese Glioma Genome Atlas (CGGA) dataset. The relationship between SEPHS2 and immune infiltration, tumor mutational burden (TMB), microsatellite instability (MSI), and neoantigens was comprehensively explored using a TCGA cohort. The mechanism by which SEPHS2 regulates tumor progression was explored by using the STRING database. A nomogram was constructed using the R software to predict the overall survival (OS) of patients with brain lower grade glioma (LGG).
Results: SEPHS2 was highly expressed in many cancers including LGG. Its high expression was significantly associated with poor OS, disease-free survival (DFS), and progression-free survival (PFS). Univariate and multivariate Cox analyses showed that SEPHS2 was an independent prognostic factor for LGG. Concordance index and calibration curves revealed that the nomogram had good predictive performance (concordance index: 0.791; 95% CI: 0.732– 1). A significant correlation was found between SEPHS2 and immune infiltration, TMB, MSI, and tumor neoantigens across diverse cancers. Enrichment analysis showed that SEPHS2 may regulate the PPAR signaling pathway.
Conclusion: SEPHS2 expression regulates tumor development and it is a potential treatment target and prognostic biomarker, especially for lower grade glioma.
Keywords: SEPHS2, selenoprotein, cancer, prognosis, biomarker
Introduction
Cancer is one of the leading causes of death worldwide. The latest global cancer statistics released by the International Agency for Research on Cancer (IARC) showed that there were approximately 19.3 million new cancer cases and 10 million deaths worldwide in 2020.1 The increase in global population and proportion of aging population has increased the incidence of malignant tumors worldwide.2 Malignant tumors rarely show clinical symptoms in the early stage of the disease; therefore, early diagnosis is challenging. Gliomas are the most common type of primary brain tumors. According to the histopathological characteristics, the World Health Organization (WHO) divides gliomas into four grades, among which grades II and III are brain lower grade glioma (LGG), and grades IV is glioblastoma (GBM).3 Clinically, LGG presents with aggressive growth and tends to progress to high-grade glioma in most cases. Since LGG tissues have no clear demarcation from normal brain tissues, surgical resection of the tumors is challenging. Moreover, standard treatments such as radiotherapy and chemotherapy are not effective for patients with LGG. Thus, the overall prognosis of LGG is poor. This calls for identification of new biomarkers for early diagnosis and treatment of LGG.
Selenium (Se) is one of the essential micronutrients for the human body and a component of certain proteins and enzymes.4 Se interacts with amino acids to form selenoproteins which regulate physiological functions in humans and animals.5 So far, 25 types of selenoproteins have been identified in humans, including selenophosphate synthetases (SEPHS1-2), glutathione peroxidases (GPX1-4, GPX6), iodothyronine deiodinases (DIO1-3) and thioredoxin reductases (TXNRD1-3).6 Selenocysteine (Sec) is considered the main form of Se in proteins and the active center of 25 selenoproteins. Sec insertion is a translation process mediated by UGA codons in the coding region of selenoprotein mRNA.7 Compounds that reduce Se use adenosine triphosphate (ATP) to generate selenium phosphate (Sep) under the catalysis of SEPHS2. Active Sep reacts with O-phosphoryl-tRNA [Ser]Sec produced under the action of O-phosphoseryl-tRNA-Sec kinase (PSTK) to generate Sec tRNA [Ser]Sec. Inserting selenocysteine into proteins to form selenoprotein requires a sec insertion sequence (SECIS) with a specialized stem-loop structure located in the 3ʹ untranslated region of selenoprotein mRNA and the cooperative action of several protein factors including SECIS binding protein 2 (SBP2) and sec-specific translation elongation factor (eEFSec).8–10 SEPHS2 participates in the biosynthesis of selenoproteins, and its role is to provide active Se donor Sep for the biosynthesis of selenocysteine.11 As a self-regulating factor, SEPHS2 regulates the production of selenophosphate and biosynthesis of selenoproteins.12
Se and selenoprotein have beneficial effects against tumors, oxidative stress, viral infection, cardiovascular disease, and many other aspects of human physiology. Changes in Se levels or expression of selenoproteins influence the incidence of cancer, the degree of invasion, and overall mortality. Previous prospective studies showed that a high level of Se was associated with a lower risk of colon cancer, liver cancer, prostate cancer, and esophageal cancer.13,14 Results from animal model studies have demonstrated that Se treatment reduced tumor invasiveness and improved clinical symptoms in patients with glioblastoma.15,16 It was demonstrated that selenoprotein P (SELP, SELENOP), TXNRD, and selenoprotein F (SELENOF, SEP15) promoted tumor initiation and growth, invasion and metastasis by targeting tumor-associated signaling pathways.17 Specific methylation of GPX3 has been shown to be a target for early prevention and treatment of clear cell renal cell carcinoma.18 Elsewhere, it has been reported that SEPHS2 plays a vital role in the survival of cancer cells.19 Several studies have investigated the expression of SEPHS2 in colorectal adenoma (CRA), colorectal cancer (CRC), triple-negative breast cancer (TNBC), and breast cancer.19–21 However, the expression of SEPHS2 in human pan-cancer and its potential role has not been explored.
This study explored the expression pattern of SEPHS2 in several cancer types through pan-cancer analysis. The association of SEPHS2 with the pathological characteristics of various cancer tissues was explored using RNA sequencing (RNA-seq) data retrieved from TCGA and GTEx databases. Its prognostic value and association with Tumor mutation burden (TMB), microsatellite instability (MSI), neoantigens, and immune cell infiltration was also explored. A nomogram for predicting the survival of patients with LGG was constructed. A workflow of the study showing the design and analyses is shown in Figure 1.
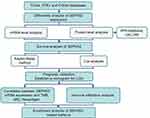 |
Figure 1 Flowchart presenting the design and analysis process of this study. |
Methods
Raw Data Collection and Preprocessing
The Cancer Genome Atlas (TCGA) database (http://cancergenome.nih.gov) provide more than 2.5 petabytes of genome, epigenome, transcriptome, and proteome data. From the Genomic Data Commons (GDC; https://portal.gdc.cancer.gov/) data portal, we downloaded RNA-seq data of 11,826 tumor samples representing 33 tumor types and for 731 adjacent non-tumor samples. To improve accuracy of the analysis results, we collected SEPHS2 data of normal tissue samples from the Genotype-Tissue Expression (GTEx) database (https://www.gtexportal.org/home/). In addition, clinical data of corresponding patients, including age, gender, race, disease stage, and tumor pathological stage were downloaded from the GDC data portal. Missing and repeated results were removed from the data. The Fragments Per Kilobase Million (FPKM) values was converted into Transcripts Per Kilobase Million (TPM) values to allow more direct comparison between samples.
SEPHS2 Expression in Human Tissues, Cell Lines
Human Protein Atlas (HPA; Https://www.proteinatlas.org/) database is a comprehensive resource website for exploring the expression and distribution of 16,975 individual proteins in all cells, tissues and organs in the human body.22 In this study, the HPA database was used to explore expression profile of SEPHS2 in normal and neoplastic tissues (n = 44). mRNA expression of SEPHS2 in normal human cell lines (n = 47) was determined using modules of Cell Atlas. In addition, immunohistochemical images of SEPHS2 in pan-cancer were obtained from the more than 13 million immunohistochemistry (IHC) images in the HPA database.
SEPHS2 Expression Profiles in Malignant Tumors
Normal tissue samples from the TCGA and GTEX databases were integrated and compared with tumor tissue samples to analyze the expression profile of SEPHS2 in pan-cancer. Statistical analysis was performed using R software v 4.0.3, and two groups of data between normal tissue samples and tumor tissue samples were compared using the rank sum test. P value < 0.05 was considered statistically significant.
The UALCAN data portal (http://ualcan.path.uab.edu/analysis-prot.html) is a website mainly used for online analytical mining of TCGA genomic data. It can be used to analyze gene expression at the protein level.23 A dataset retrieved by the CPTAC module was used to explore expression profile and clinicopathological characteristics of SEPHS2 protein in breast cancer (BRCA), ovarian cancer (OV), colon cancer (COAD), kidney renal clear cell carcinoma (KIRC), uterine corpus endometrial carcinoma (UCEC) and lung adenocarcinoma (LUAD). Results were presented as a box plot. Z-values represented standard deviations (SD) from the median (M) across samples for a given cancer type.
To explore the relationship between SEPHS2 expression and clinical characteristics of tumor patients, the expression of SEPHS2 in patients with different pathological stages was determined. “Expression DIY” module in GEPIA2 (http://gepia.cancer-pku.cn) was used to map the pathological staging plots of different tumor stages (stages I, II, III and IV).24 Log2 [TPM +1] transformation was performed on expression data and results were presented as a violin diagrams.
Survival and Prognosis Analysis
Overall survival (OS) and disease-free survival (DFS) analysis of SEPHS2 in 33 types of tumors in TCGA were analyzed using the “survival map” module of GEPIA2. The median was used as the group cut off, and samples were divided into two groups with high and low SEPHS2 expression. Kaplan–Meier curve analysis was performed using the “survival analysis” module of GEPIA2. Log rank test was used for hypothesis testing, and the hazard ratio (HR) and 95% confidence interval (95% C.I.) were calculated by Cox regression Model. The significance level was set at 0.05. To further explore the prognostic value of SEPHS2 expression in patients with different tumor types, survival analysis [OS, DFS, progression-free survival (PFS), and disease-specific survival (DSS)] were performed using “ggforest” in the R-package “survminer”.
The Chinese Glioma Genome Atlas (CGGA; http://www.cgga.org.cn/) database contains a comprehensively mapped glioma genome of Chinese population by multi-group technology and provides nearly 2000 glioma samples from Chinese cohorts.25 The CGGA database was used to explore the relationship between the expression level of SEPHS2 and survival in gliomas.
Relationship Between SEPHS2 Expression and Genetic Mutation
TMB is the total number of mutations per million bases (mut/Mb) in the genomic coding region of tumor cells.26 A high TMB is associated with higher neoantigen loads, implying that more T cells, tumor-targeted immune response are activated, and sensitivity of patients to respond to immune checkpoint blockade increases.27 MSI is a phenomenon of base pair insertion or loss in the microsatellite region due to defect in the mismatch repair system.28 R-package was used to explore the relationship between SEPHS2 expression and MSI, TMB and neoantigens in different types of cancers. The three datasets were tested by rank sum test unless otherwise stated. The results are presented by lollipop plot and Scatter density plot. P values < 0.05 were considered statistically significant.
Correlation Between SEPHS2 Expression and Immune Infiltration
Immuneeconv is an R-package that integrates six major immune evaluation algorithms including TIMER, CIBERSORT, QUANTISEQ, XCELL, MCPCOUNTER and EPIC, and estimates immune cell components in tumor tissues using RNA-seq data.29. Immunedeconv R package is available in github (https://github.com/icbi-lab/immunedeconv). The EPIC algorithm is used to identify non-characteristic cells and the different mRNA content in each cell type, and it is widely used in most solid tumors.30 Correlation between infiltration ratios of six immune cells (including B cell, Endothelial cell, Macrophage, NK cell, CD4+T cell, and CD8+T cell) and SEPHS2 expression were analyzed using EPIC’s algorithm and the results were presented as heatmaps. TIMER 2.0 (http://timer.cistrome.org/) is an interactive web tool that uses RNA-Seq data to determine infiltration level of different immune cells in 10,897 tumors from 32 tumor types.31 The “Immune-gene” module in TIMER was used to explore correlation between SEPHS2 expression and cancer-associated fibroblasts (CAFs) infiltration level in TCGA dataset using four deconvolution algorithms including TIDE, XCELL, MCPCOUNTER, and EPIC. P < 0.05 was considered statistically significant.
Enrichment Analysis
STRING (https://string-db.org/) is an interactive web server for study of protein–protein interactions, covering 5090 organisms and 24.6 million proteins, with comprehensive and diverse benchmark data sources.32 STRING was used to obtain the required SEPHS2-integrated proteins in human body. Active interaction sources were set to experiments with a minimum required interaction score of low confidence (0.150) and a maximum number of interactors not more than 50 in the first shell. GEPIA2 was used to identify the top 100 SEPHS2 similar genes in human tumor tissues based on Pearson’s correlation coefficient (PCC). The top five genes with the largest PCC were selected to construct scatterplots to explore the correlation between SEPHS2 expression levels and these genes. The SEPHS2-integrated proteins and SEPHS2 similar genes were combined to make up SEPHS2-related protein patterns. Enrichment analysis by Gene Ontology (GO) and Kyoto encyclopedia of genes and genes (KEGG) were conducted based on SEPHS2-related protein patterns. The false discovery rate (FDR)-corrected P value < 0.05 of each term was considered statistically significant. GO enrichment analysis included Biological Process (BP), Cellular Component (CC), and Molecular Function (MF). The top 10 terms of GO and KEGG enrichment results based on FDR values were selected and a bubble chart was generated using R package v 4.0.3.
Construction and Validation of Nomogram for LGG
Univariate and multivariate Cox regression analysis were used to identify independent prognostic parameters for LGG patients to further explore association between expression of SEPHS2 in LGG and tumor progression. A nomogram was constructed using ‘rms’ R package based on the confirmed multivariate regression analysis results. The nomogram was used to predict the OS of LGG patients. Concordance index (C-index) and calibration plots were used to estimate and verify the nomogram, respectively. The value of the C-index ranged from 0.5 to 1. A C-index closer to 1, indicates that the forecasting model is good.
Results
SEPHS2 Expression in Human Tissues and Cell Lines
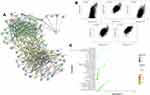
The tissue atlas and pathology atlas of HPA database were used to explore expression profiles of SEPHS2 (at mRNA and protein levels) in human tissues. Analysis showed that SEPHS2 mRNA and protein were widely expressed in various normal tissues of the human body (Figure 2A and B). At the protein level, the expression analysis of SEPHS2 in different histopathological tissues showed that SEPHS2 was moderately or lowly expressed in most cancer tissues such as thyroid cancer, colorectal cancer, melanoma, and gastric cancer (Figure 2C). IHC analysis showed that SEPHS2 protein is down-regulated in cancers, such as colorectal cancer, liver cancer, breast cancer, prostate cancer, and lung cancer (Figure 2D–H). Low expression level was also observed in normal colon tissue (Figure 2I). Detailed on IHC results are presented in Table 1. SEPHS2 mRNA expression was enriched mostly in HMC-1, THP-1 and Hep-G2 cell lines (Figure 3A). Immunofluorescent staining images showed that SEPHS2 was mainly localized in the nucleoplasm in human HEK 293, MCF7 and U-2 OS cell lines (Figure 3B–D).
 |
Table 1 Clinical Information of the Subjects and Immunohistochemistry (IHC) Results |
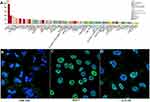 |
Figure 3 SEPHS2 expression profiles in human cell lines. (A) SEPHS2 expression levels across 69 different cell lines. (B–D) Immunofluorescence staining image of SEPHS2 in cell lines. |
SEPHS2 Expression Levels in Human Malignant Tumors
Expression of SEPHS2 mRNA in normal tissues and different types of tumor tissues was compared using the TCGA database and the GTEx database (Figure 4). Analysis showed significant differences in expression between adjacent normal tissues and tumor tissues except for 3 types of tumors (P < 0.001). These findings show that SEPHS2 is abnormally over-expressed in most malignant tumors. SEPHS2 was highly expressed in 22 tumor types including adrenocortical carcinoma (ACC), bladder urothelial carcinoma (BLCA), breast invasive carcinoma (BRCA), glioblastoma multiforme (GBM), brain lower grade glioma (LGG). On the contrary, SEPHS2 was down-regulated in KIRC and kidney renal papillary cell carcinoma (KIRP). In addition, analysis of the CGGA database, showed that mRNA expression levels of SEPHS2 increased with the gliomas grading, and its expression level in GBM was higher than that of LGG. Analysis using the CPTAC database showed significantly higher expression levels of SEPHS2 protein in the tumor tissues of UCEC, LUAD compared with normal tissues (Figure 5A, P < 0.001). Expression level of SEPHS2 in COAD and KIRC tumor tissues was significantly lower compared with that in normal tissues (Figure 5A, P < 0.001). Analysis of the relationship between SEPHS2 expression and the pathological stages in tumor patients was explored using the “Pathological Staging Chart” module in GEPIA2. Analysis showed that SEPHS2 expression was associated with five tumor pathological stages, including esophageal carcinoma (ESCA), KIRC, KIRP, testicular germ cell tumors (TGCT), and thyroid carcinoma (THCA), but not with the other tumors analyzed (Figure 5B, P < 0.05).
 |
Figure 4 Expression levels of SEPHS2 in tumor and normal tissues from TCGA and GTEx databases (**P < 0.01, ***P < 0.001). |
Relationship Between SEPHS2 Protein and Clinicopathological Characteristics
To explore the correlation between SEPHS2 protein expression levels and age, gender, ethnicity, clinical stages, clinicopathological features analysis was performed based on six different tumor datasets from UALCAN database. Analysis showed that expression of SEPHS2 protein was significantly higher in the fourth stage of OV, the first stage of UCEC, and the third stage of LUAD compared with adjacent normal tissues (Figure 6A). Further analysis showed that SEPHS2 protein expression varied in different ethnic groups, age groups and sex groups. Expression of SEPHS2 protein in OV, COAD, UCEC, KIRC, and LUAD was significantly correlated with the patient’s race. Caucasian patients showed highest SEPHS2 expression in OV, UCEC, and LUAD (Figure 6B). SEPHS2 protein was significantly expressed in all pathological stages of OV, UCEC, and LUAD (Figure 6C). In LUAD, significant differences were observed in gender, with males having significantly higher SEPHS2 protein expression levels compared with females (Figure 6D).
Prognostic Potential of SEPHS2 in Human Malignant Tumors
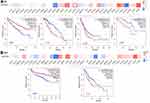
The relationship between SEPHS2 and the prognosis of different types of tumors was explored using TCGA database (Figure 7). Kaplan–Meier plots showed that expression of SEPHS2 is associated with prognosis in several types of tumors. Increased expression of SEPHS2 was correlated with a poor clinical outcome (OS) in LGG (P = 3.00×10−6), LUAD (P = 0.039) and UVM (P = 0.025) (Figure 7A). Furthermore, DFS analysis showed that high expression of SEPHS2 was associated with a poor prognosis for LGG (P = 2.50×10−4), whereas low expression of SEPHS2 was correlated with poor OS and DFS prognosis for KIRC (OS, P = 3.50×10−7; DFS, P = 3.80×10−4, Figure 7A and B). These findings indicated that SEPHS2 overexpression was a risk factor for poor prognosis of OS and DFS in LGG patients, and a favorable factor for a better prognosis in KIRC patients.
Cox regression model was used to further explore the effect of SEPHS2 on survival outcomes (OS, PFS, DFS, DSS) of patients with different types of tumors. Analysis showed that up-regulated expression of SEPHS2 was associated with poor OS in acute myeloid leukemia (LAML), LGG and uveal melanoma (UVM) (Figure 8A). On the contrary, increased SEPHS2 expression predicted good OS in KIRC and lung squamous cell carcinoma (LUSC) (Figure 8A). Further analysis showed that SEPHS2 over-expression was predicted poor DFS of patients with LGG, pheochromocytoma and paraganglioma (PCPG), prostate adenocarcinoma (PRAD), thymoma (THYM), uterine carcinosarcoma (UCS), and UVM, and was correlated with poor PFS of LGG, READ, UVM patients (Figure 8B and C). Over-expression of SEPHS2 predicted good DFS and PFS in KIRC. Analysis of the relationship between SEPHS2 and DSS in cancer patients showed that high expression levels of SEPHS2 were correlated with poor prognosis in COAD, KIRP, PRAD, and UCS, and better prognosis in READ (Figure 8D). Integration the results of Kaplan–Meier survival curves and Cox regression forest plots showed that SEPHS2 was significantly correlated with OS, DFS, and PFS in KIRC, LGG, and UVM patients. However, the sample size of KIRC and UVM was small, therefore, further studies should explore the prognostic value of SEPHS2 in LGG. To further confirm these results, glioma dataset was downloaded from CGGA database. The cohort included 144 LGG cases and 84 GBM cases. In general, consistent with the findings obtained in the TCGA database, high expression of SEPHS2 predicted worse outcomes in gliomas patients.
Relationships Between SEPHS2 and Neoantigens, TMB, MSI
TMB and MSI are implicated in occurrence and development of tumors. Association of SEPHS2 expression with TMB and MSI across different types of tumors is presented in Figure 9A and B. Analysis showed that SEPHS2 expression was positively correlated with TMB in 19 tumor types including cholangiocarcinoma (CHOL), lymphoid Neoplasm Diffuse Large B-cell Lymphoma (DLBC), pancreatic adenocarcinoma (PAAD), LGG, and KIRC. On the other hand, increased SEPHS2 expression was negatively correlated with TMB in 13 types of tumors, including THYM, COAD, and UVM (Figure 9A). SEPHS2 expression was associated with MSI in 33 tumor types (Figure 9B). High expression of SEPHS2 was positively correlated with MSI in 22 types of tumors, including CHOL, ACC, KIRC, UVM, and KIRP. On the other hand, SEPHS2 expression was negatively correlated with MSI in 11 tumor types, including OV, stomach adenocarcinoma (STAD), mesothelioma (MESO) and LGG. Notably, the CHOL cohort had the highest correlation coefficients with either TMB or MSI, followed by LGG and KIRC cohorts compared with other cancer types.
Neoantigens are only expressed on the tumor cell surface which has a high degree of immunogenicity. Previous studies report that neoantigens play important roles in antitumor immune response and the effectiveness of checkpoint blockade immunotherapies.33 The association between SEPHS2 expression and neoantigens in different types of tumor was explored. Analysis showed significant correlation between SEPHS2 expression and neoantigens in COAD, KIRC, and skin cutaneous melanoma (SKCM). In COAD and SKCM, SEPHS2 expression was significantly and negatively correlated with neoantigens. On the contrary, the neoantigens in KIRC were positively correlated with the expression level of SEPHS2 (Figure 9C).
Relationship Between SEPHS2 Expression and Immune Infiltration
The tumor microenvironment affects initiation and progression of malignant tumors and plays a pivotal role in clinical therapy. Therefore, the correlation between SEPHS2 and immune components was explored in this study. In this study, the EPIC algorithm was used to explore the correlation between SEPHS2 and immune cell infiltration. Association between SEPHS2 expression and abundance of immunocyte infiltration varied significantly among cancer types. Expression of SEPHS2 was positively correlated with infiltration levels of CD4+T cell in 11 tumor types, CD8+T cell in 10 tumor types, endothelial cell in 5 tumor types, macrophages in 3 tumor types, and B cell only in UVM. In addition, the infiltration level of NK cell was negatively correlated with the expression of SEPHS2 in BRCA, COAD, KIRC, KIRP, SKCM, THCA, and THCA. Notably, immune infiltrating levels in KICH, LGG, and UVM were showed highest positive correlation with SEPHS2 expression, whereas immune infiltrating levels in BRCA, LUAD, THCA, and THYM were the most significantly negatively correlated with SEPHS2 expression (Figure 10A).
The TIMER 2.0 interactive web tool, was used to explore the density of CAFS infiltration in diverse cancers based on EPIC, MCPCOUNTER, XCELL, and TIDE algorithms. The relationship between CAFS infiltration and expression of SEPHS2 was also explored (Figure 10B). Analyses indicated that SEPHS2 expression was negatively correlated with the abundance of CAFs infiltration across most tumor types, with highest correlation shown in KIRC and THCA. However, expression of SEPHS2 in head and neck squamous cell carcinoma (HNSC), and HNSC-HPV- showed a significant positive correlation with CAFs. These findings indicated that SEPHS2 plays an important role in immuno-oncological effects.
Enrichment Analysis
PPI network based on 130 SEPHS2-related partners were constructed (Figure 11A). Analysis showed that CLIC3, UBA2, LAGE3, UMPS and PLA2G12b interacted with SEPHS2. UMPS and PLA2G12B were also connected with several other proteins and played important roles in the PPI, implying UMPS and PLA2G12B may have key potential interaction associations with SEPHS2 proteins in tumor occurrence. The top five genes related to SEPHS2 in human cancers were identified. Analysis showed a positive correlation of all the five genes with SEPHS2 expression levels, namely ABCC6 (PCC = 0.56), CERS2 (PCC = 0.5), BPHL (PCC = 0.49), XYLB (PCC = 0.49), and MTHFS (PCC = 0.49) (Figure 11B). GO enrichment analysis showed that SEPHS2-related patterns mainly participated in metabolic and catabolic biological processes (Figure 11C). KEGG pathways analysis showed that most of the terms were mainly involved in signaling pathways related to metabolic activities, including metabolic pathways, Fatty acid degradation pathway, valine, leucine and isoleucine degradation pathway, and PPAR signaling pathway (Figure 11C).
Determination of Prognostic Factors for LGG and Development of Nomogram Model
Survival prognosis analysis, immune infiltration, and genetic mutation analysis, showed a correlation between SEPHS2 and different cancer types. LGG and SEPHS2 showed a significant relationship. A Nomogram model was developed and validated to further determine the value of SEPHS2 mRNA expression levels on the clinical prognosis of LGG patients. Univariate and multivariate Cox regression analysis were performed to determine the independent prognostic factors of LGG patients, using the TCGA cohort. Univariate Cox regression analysis showed that age, grade, SEPHS2, and radiation therapy were correlated with OS (P < 0.01, Figure 12A). Age, SEPHS2, and grade were identified as independent prognostic factors for LGG using multivariate Cox regression analysis (P < 0.01, Figure 12B). The identified independent prognostic risk parameters were used to construct a nomogram for predicting 1-year, 2-year, 3-year, and 5-year OS in LGG patients to further verify the correlation between SEPHS2 mRNA expression and cancer occurrence (Figure 12C). The C-index of the nomogram was 0.791, indicating that the model had a high prediction accuracy. Integrated calibration curves showed that the predicted survival outcome of LGG nomogram had a good match to the observed actual survival outcome, especially in the 3-year and 5-year survival (Figure 12D). These findings showed that the nomogram model performed well in predicting overall survival of LGG patients.
Discussion
SEPHS2 plays a crucial role in selenoprotein biosynthetic pathway and its main function is synthesis of active Se donor Sep. Recent in vivo studies and bioinformatics studies explored differences in the expression levels of SEPHS2 between normal tissues and specific tumor types such as gastric cancer34 and lung adenocarcinoma.35 However, SEPHS2 expression in other cancers and its regulatory mechanism has not been fully explored, and analysis from pan-cancer perspective has not been reported. Therefore, in the present study, multiple online public databases were used to comprehensively explore expression pattern of SEPHS2 in pan-cancer, and its prognostic and the therapeutic significance.
SEPHS2 participates in the biosynthesis of selenoproteins. The selenoprotein synthesis involves decoding, recoding, and binding of se to polypeptide chains through the UGA codon. Studies have shown that in addition to SEPHS2, other proteins such as, SBP2, eEFSec, eukaryotic initiation factor 4a3 (eIF4a3), Ribosomal protein L30 (L30), 43-kDa RNA-binding protein (SECp43), nucleolin, and soluble liver antigen protein (SLA) are involved in selenoprotein biosynthesis.8,36 SBP2, nucleolin, and eIF4a3 are regulators of selenoprotein synthesis. UGA usually exists as a stop codon, and in selenoprotein synthesis, the SECIS element interprets the stop codon and converts it into selenocysteine. SBP2 is a key factor in the translation of selenoproteins. It can recognize and bind to SECIS elements, interact with elongation factors eEFSec, and bind stably to ribosomes.37 Aside from interacting with SBP2, the main function of EFSec is regulating trafficking of the SEC tRNA [Ser]Sec to the ribosomal “A” site.38 The exact role of L30, Secp43, and SLA in the synthesis of selenoprotein is still unclear, and further research is awaited.39
Expression of SEPHS2 does not exhibit tissue specificity in normal tissues, thus providing a pathologic basis for the study of the effect of this gene on various tumors. Several studies have explored association between SEPHS2 expression and tumors.34,35 Expression of SEPHS2 mRNA and protein across different tumor tissues was investigated using data available in different databases. Analysis showed that SEPHS2 expression was up-regulated in 22 tumors including ACC, BRCA, LGG, and LUAD. Carlisle et al reported that SEPHS2 protein was up-regulated in human breast-cancer tissues compared with normal breast tissue.19 In addition, SEPHS2 is over-expressed in triple-negative breast cancer (TNBC) tissues, one of the subtypes of breast cancer. Notably, gene expression level in TNBC tissues was positively correlated with the malignant grade of the tumor, compared with normal breast tissue.21 A previous study reported that SEPHS2 is up-regulated in LUAD.35 The findings from these studies were consistent with the findings of the current study that SEPHS2 is over-expressed in breast cancer tissue and lung adenocarcinoma tissue. Analysis using the CPTAC database showed that SEPHS2 protein was highly expressed in UCEC, and LUAD, but was down-regulated in COAD and KIRC. Moreover, analysis using ULCAN database showed that age, gender, and race of cancer patients were correlated with the expression levels of SEPHS2. In summary, these findings indicated that SEPHS2 expression across different tumor types was tissue-specific due to various clinicopathological features.
Furthermore, analysis showed that SEPHS2 expression was correlated with individual cancer stages. Cancer staging helps to determine the severity of cancer. Analysis using ULCAN database showed that expression of SEPHS2 was up-regulated in OV, KIRC, LUAD, and UCEC in each cancer stage compared with normal tissues. These findings indicated that SEPHS2 expression is significantly correlated with the severity of cancer. Furthermore, Kaplan–Meier curves and Cox regression forest plots showed that high expression level of SEPHS2 in LGG patients was correlated with poor prognosis, as shown by OS, DFS, and PFS. A nomogram model was developed to further explore the role of SEPHS2 in prognosis of LGG patients. Cox regression analysis showed that SEPHS2 was an independent prognostic parameter for LGG patients. Analysis of nomogram and calibration plots showed that SEPHS2 accurately predicts the overall survival of LGG patients. This nomogram can help clinicians and LGG patients to make more accurate survival prognostic judgments, and it is valuable for choosing better clinical treatment options. These results indicate that SEPHS2 is a potential prognosis factor for the LGG. In addition, LGG progresses to GBM following inactivation of tumor suppressor genes, mutations (such as PTEN), oncogene amplification (such as EGFR), and abnormal regulation of the cell cycle.40 Nevertheless, we did not find any statistical significance of SEPHS2 expression on the survival of patients with GBM in the TCGA cohort. We speculate that the reason may be related to the pathological features of GBM and the complicated molecular mechanism. Tumor necrosis and microvascular proliferation are prominent features of GBM.41 GBM exhibits a high degree of intra-tumor heterogeneity and plasticity at the cytopathological characteristics, as well as a high degree of invasion and diffusion of tumor cells, which together make it difficult to treat.42 The prognosis of GBM patients is extremely poor, and the median survival time of GBM patients receiving standard treatment is only 12–15 months.40,41 Recent evidence from clinical trials shows that targeted therapy does not improve prognosis of patients with GBM.43
These findings indicated that the expression of SEPHS2 is related to the occurrence and development of many kinds of tumors, especially LGG. Evidence shows that SEPHS2 affects the development of drug-resistance in glioma stem cells and is closely related to the survival of glioma stem cells.16 A bioinformatics study showed that expression level of GPX1 is related to the prognosis of patients with glioma.44 In LGG, we found that SEPHS2 expression was negatively correlated with five selenoprotein genes (GPX3, SELENOK, SELENOO, TXNRD2, and TXNRD3) expression, which suggested that SEPHS2 might have antagonistic effects with these genes. However, SEPHS2 expression was positively correlated with nine genes (DIO2, GPX2, SELENOF, SELENOH, SELENOI, SELENOM, SELENON, SELENOT, and TXNRD1) expression, which indicated that SEPHS2 have potential synergistic effects with these genes.
Previous studies report that TMB levels affect the response rate of immunotherapy.45 Tumor mutation burden-high (TMB-H) is associated with improved OS in patients receiving immunocheckpoint inhibitors (ICI) for various types of cancer.46 Moreover, TMB is significantly correlated with tumor neoantigen loads, and high non-synonymous mutant expression promotes production of neoantigens, activates more tumor-targeted immune responses, and increases patient response rates to immunosuppressive therapy.27 Neoantigens are potential targets for tumor immunotherapy and can be used to predict efficacy of immunotherapy approaches such as PD-1 inhibitors.47,48 MSI is a predictor of efficacy of immunotherapy for advanced solid tumors.49 MSI phenotype determines the efficacy of immunocheckpoint inhibitors, and not the specific solid tumor type. The PD-1 inhibitor Pembrolizumab was approved by the Food and Drug Administration (FDA) in 2020 as a first-line treatment for advanced MSH-H metastatic colorectal cancer.50 TMB, MSI and neoantigens play important roles in the salvage therapy of cancer patients, therefore, in this study association between TMB, MSI, and neoantigens and the expression of SEPHS2 in cancer patients was explored. Spearman correlation analysis showed that the TMB level in 19 tumors types and MSI level in 22 tumors types were positively correlated with SEPHS2 expression. In addition, SEPHS2 expression was significantly correlated with the neoantigens of the three tumor types. These findings imply that tumor patients with up-regulated SEPHS2 expression and elevated TMB or MSI may exhibit better therapeutic effects after treatment with immune checkpoint blockade (ICB) thus prolonging their survival. In summary, these findings showed the correlation between SEPHS2 expression and tumor immunotherapy, and that TMB, MSI and neoantigens are potential predictive biomarkers for determining efficacy of immunotherapy.
Selenoprotein enhance immune system functions by regulating, the migration and proliferation of immune cells, and the secretion of inflammation-related cytokines. Previous studies have shown that the expression of selenoprotein regulates M1 macrophages, affects the cytotoxic activities of NK cells, and affects the proliferation of T cells.51 SEPHS2 is one of the enzymatic selenoproteins that regulate immune cell function.52 Cancer-associated fibroblasts, which are highly heterogeneous in the tumor microenvironment, inhibit immune cells and regulate proliferation and metastasis of tumor cells indirectly or directly.53 In this study, we found that SEPHS2 expression was positively correlated with immune infiltration level of CD8 + T cells, CD4 + T cells, NK cells, macrophages, endothelial cells and B cells in LGG. In addition, SEPHS2 was significantly negatively correlated with CAFS in KIRC. These findings indicate that abnormal expression of SEPHS2 may affect immune responses and tumor immune microenvironment.
Previous studies have reported that peroxisome proliferator-activated receptors (PPAR) subtypes are involved in lipid metabolism and affect the occurrence and development of tumors.54,55 Consistent with previous studies, enrichment analysis of SEPHS2-related patterns showed that PPAR signaling pathway was enriched in different cancer types in the present study. Recent studies have reported that selenoproteins modify the mode of action of PPAR-γ.56,57 Se deficiency inhibits PPAR-γ activity, promotes nuclear factor-κB (NF-κB) activation in tumor cells, and increases tumor invasiveness and immunosuppression.58 As mentioned earlier, SEPHS2 is a selenoprotein that is required for the synthesis of selenoproteins. It is reasonable to speculate that SEPHS2 modify tumor progression by regulating PPAR signaling pathway.
This study had a few limitations. First, the sample size of mRNA expression data of some tumor types was too small, which makes it difficult to determine the role of SEPHS2 in these tumors. For example, the sample size of CHOL normal tissue was 9, and the sample size of tumor tissue was 36. Second, the results of this study were based on analyses of public databases which not been verified using in vivo or in vitro experiments. Therefore, there is need to verify these findings through experimental studies or clinical studies using large samples to further explore the mechanism of SEPHS2 in tumor development and its clinical value.
Conclusion
High expression of SEPHS2 is associated with poor prognosis of LGG patients and is an independent predictor of LGG. In addition, expression of SEPHS2 is related to immune infiltration and immunotherapy markers (TMB, MSI, neoantigens). SEPHS2 may influence occurrence and development of tumors through the PPAR signaling pathway. These findings demonstrate that SEPHS2 may be a potential prognostic biomarker and treatment target for LGG.
Abbreviations
Se, Selenium; Sep, Selenium phosphate; SEPHS2, selenophosphate synthetase 2; TCGA, The Cancer Genome Atlas; GTEx, Genotype-Tissue Expression; CGGA, Chinese Glioma Genome Atlas; HPA, Human Protein Atlas; EPIC, Estimating the Proportions of Immune and Cancer cells; MSI, microsatellite instability; TMB, tumor mutational burden; OS, overall survival; DFS, disease-free survival; PFS, progression-free survival; DSS, disease-specific survival; RNA-seq, RNA sequencing; CAFs, cancer-associated fibroblasts; PCC, Pearson’s correlation coefficient; GO, Gene Ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes; ACC, adrenocortical carcinoma; BLCA, bladder urothelial carcinoma; BRCA, breast invasive carcinoma; CESC, cervical squamous cell carcinoma and endocervical adenocarcinoma; CHOL, cholangiocarcinoma; COAD, colon adenocarcinoma; DLBC, lymphoid neoplasm diffuse large B-cell lymphoma; ESCA, esophageal carcinoma; GBM, glioblastoma multiforme; HNSC, head and neck squamous cell carcinoma; KICH, kidney chromophobe; KIRC, kidney renal clear cell carcinoma; KIRP, kidney renal papillary cell carcinoma; LAML, acute myeloid leukemia; LGG, brain lower grade glioma; LIHC, liver hepatocellular carcinoma; LUAD, lung adenocarcinoma; LUSC, lung squamous cell carcinoma; MESO, mesothelioma; OV, ovarian serous cystadenocarcinoma; PAAD, pancreatic adenocarcinoma; PCPG, pheochromocytoma and paraganglioma; PRAD, prostate adenocarcinoma; READ, rectum adenocarcinoma; SARC, sarcoma; SKCM, skin cutaneous melanoma; STAD, stomach adenocarcinoma; TGCT, testicular germ cell tumors; THCA, thyroid carcinoma; THYM, thymoma; UCEC, uterine corpus endometrial carcinoma; UCS, uterine carcinosarcoma; UVM, uveal melanoma.
Data Sharing Statement
The datasets generated and/or analyzed during the current study are available in the TCGA (https://portal.gdc.cancer.gov/) and GTEx (http://commonfund.nih.gov/GTEx/) repository.
Ethics Approval and Informed Consent
All data for this study are derived from publicly available databases, and no ethical approval has been applied.
Acknowledgments
We thank the School of Nursing, Shaanxi University of Chinese Medicine, and Ankang R & D Center of Se-enriched Products for assistance.
Author Contributions
All authors contributed to study design, acquisition, and analysis of data, and writing and revising manuscripts and reached a consensus on the journal to which the article was submitted, approved the final version to be published and agreed to be responsible for all content of the article.
Funding
This study was supported by Special R & D Program Project of Chinese Academy of Se-enriched Industry (2020FXZX05-01), Key Research and Development Program in Shaanxi Province (2020SF-076), Research Project from Health Commission of Shaanxi Provincial Government (2018A017), Education Department of Shaanxi Provincial Government (19JS015).
Disclosure
The authors report no conflicts of interest in this work.
References
1. International Agency for Research on Cancer [homepage on the Internet]. Latest global cancer data: cancer burden rises to 19.3 million new cases and 10.0 million cancer deaths in 2020; 2020. Available from: https://www.iarc.who.int/news-events/latest-global-cancer-data-cancer-burden-rises-to-19-3-million-new-cases-and-10-0-million-cancer-deaths-in-2020/.
2. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. doi:10.3322/caac.21492
3. Louis DN, Perry A, Wesseling P, et al. The 2021 WHO classification of tumors of the central nervous system: a summary. Neuro Oncol. 2021;23(8):1231–1251. doi:10.1093/neuonc/noab106
4. Rayman MP. Selenium and human health. Lancet. 2012;379(9822):1256–1268. doi:10.1016/S0140-6736(11)61452-9
5. Hossain A, Skalicky M, Brestic M, et al. Selenium biofortification: roles, mechanisms, responses and prospects. Molecules. 2021;26(4):881. doi:10.3390/molecules26040881
6. Gladyshev VN, Arner ES, Berry MJ, et al. Selenoprotein gene nomenclature. J Biol Chem. 2016;291(46):24036–24040. doi:10.1074/jbc.M116.756155
7. Xu XM, Carlson BA, Mix H, et al. Biosynthesis of selenocysteine on its tRNA in eukaryotes. PLoS Biol. 2007;5(1):e4. doi:10.1371/journal.pbio.0050004
8. Labunskyy VM, Hatfield DL, Gladyshev VN. Selenoproteins: molecular pathways and physiological roles. Physiol Rev. 2014;94(3):739–777. doi:10.1152/physrev.00039.2013
9. Turanov AA, Xu XM, Carlson BA, Yoo MH, Gladyshev VN, Hatfield DL. Biosynthesis of selenocysteine, the 21st amino acid in the genetic code, and a novel pathway for cysteine biosynthesis. Adv Nutr. 2011;2(2):122–128. doi:10.3945/an.110.000265
10. Tujebajeva RM, Copeland PR, Xu XM, et al. Decoding apparatus for eukaryotic selenocysteine insertion. EMBO Rep. 2000;1(2):158–163. doi:10.1093/embo-reports/kvd033
11. Xu XM, Carlson BA, Irons R, et al. Selenophosphate synthetase 2 is essential for selenoprotein biosynthesis. Biochem J. 2007;404(1):115–120. doi:10.1042/BJ20070165
12. Kim IY, Guimaraes MJ, Zlotnik A, Bazan JF, Stadtman TC. Fetal mouse selenophosphate synthetase 2 (SPS2): characterization of the cysteine mutant form overproduced in a baculovirus-insect cell system. Proc Natl Acad Sci U S A. 1997;94(2):418–421. doi:10.1073/pnas.94.2.418
13. Hughes DJ, Duarte-Salles T, Hybsier S, et al. Prediagnostic selenium status and hepatobiliary cancer risk in the European Prospective Investigation into Cancer and Nutrition cohort. Am J Clin Nutr. 2016;104(2):406–414. doi:10.3945/ajcn.116.131672
14. Hughes DJ, Fedirko V, Jenab M, et al. Selenium status is associated with colorectal cancer risk in the European prospective investigation of cancer and nutrition cohort. Int J Cancer. 2015;136(5):1149–1161. doi:10.1002/ijc.29071
15. Hervouet E, Staehlin O, Pouliquen D, et al. Antioxidants delay clinical signs and systemic effects of ENU induced brain tumors in rats. Nutr Cancer. 2013;65(5):686–694. doi:10.1080/01635581.2013.789541
16. Yakubov E, Eibl T, Hammer A, Holtmannspotter M, Savaskan N, Steiner HH. Therapeutic Potential of selenium in glioblastoma. Front Neurosci. 2021;15:666679. doi:10.3389/fnins.2021.666679
17. Short SP, Williams CS. Selenoproteins in tumorigenesis and cancer progression. Adv Cancer Res. 2017;136:49–83.
18. Liu Q, Jin J, Ying J, et al. Frequent epigenetic suppression of tumor suppressor gene glutathione peroxidase 3 by promoter hypermethylation and its clinical implication in clear cell renal cell carcinoma. Int J Mol Sci. 2015;16(5):10636–10649. doi:10.3390/ijms160510636
19. Carlisle AE, Lee N, Matthew-Onabanjo AN, et al. Selenium detoxification is required for cancer-cell survival. Nat Metab. 2020;2(7):603–611. doi:10.1038/s42255-020-0224-7
20. Fedirko V, Jenab M, Meplan C, et al. Association of Selenoprotein and selenium pathway genotypes with risk of colorectal cancer and interaction with selenium status. Nutrients. 2019;11:4. doi:10.3390/nu11040935
21. Nunziata C, Polo A, Sorice A, et al. Structural analysis of human SEPHS2 protein, a selenocysteine machinery component, over-expressed in triple negative breast cancer. Sci Rep. 2019;9(1):16131. doi:10.1038/s41598-019-52718-0
22. Uhlen M, Fagerberg L, Hallstrom BM, et al. Proteomics. Tissue-based map of the human proteome. Science. 2015;347(6220):1260419. doi:10.1126/science.1260419
23. Chandrashekar DS, Bashel B, Balasubramanya S, et al. UALCAN: a portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia. 2017;19(8):649–658. doi:10.1016/j.neo.2017.05.002
24. Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017;45(W1):W98–W102. doi:10.1093/nar/gkx247
25. Zhao Z, Zhang KN, Wang Q, et al. Chinese Glioma Genome Atlas (CGGA): a comprehensive resource with functional genomic data from Chinese glioma patients. Genomics Proteomics Bioinformatics. 2021. doi:10.1016/j.gpb.2020.10.005
26. Chalmers ZR, Connelly CF, Fabrizio D, et al. Analysis of 100,000 human cancer genomes reveals the landscape of tumor mutational burden. Genome Med. 2017;9(1):34. doi:10.1186/s13073-017-0424-2
27. Buttner R, Longshore JW, Lopez-Rios F, et al. Implementing TMB measurement in clinical practice: considerations on assay requirements. ESMO Open. 2019;4(1):e442. doi:10.1136/esmoopen-2018-000442
28. Bonneville R, Krook MA, Kautto EA, et al. Landscape of microsatellite instability across 39 cancer types. Jco Precision Oncol. 2017;3(1):1–15. doi:10.1200/PO.17.00073
29. Sturm G, Finotello F, List M. Immunedeconv: an R package for unified access to computational methods for estimating immune cell fractions from Bulk RNA-Sequencing Data. Methods Mol Biol. 2020;2120:223–232.
30. Racle J, Gfeller D. EPIC: a tool to estimate the proportions of different cell types from bulk gene expression data. Methods Mol Biol. 2020;2120:233–248.
31. Li T, Fu J, Zeng Z, et al. TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res. 2020;48(W1):W509–W514. doi:10.1093/nar/gkaa407
32. Szklarczyk D, Gable AL, Lyon D, et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019;47(D1):D607–D613. doi:10.1093/nar/gky1131
33. Jiang T, Shi T, Zhang H, et al. Tumor neoantigens: from basic research to clinical applications. J Hematol Oncol. 2019;12(1):93. doi:10.1186/s13045-019-0787-5
34. Lan X, Xing J, Gao H, et al. Decreased expression of selenoproteins as a poor prognosticator of gastric cancer in humans. Biol Trace Elem Res. 2017;178(1):22–28. doi:10.1007/s12011-016-0908-8
35. Jia Y, Dai J, Zeng Z. Potential relationship between the selenoproteome and cancer. Mol Clin Oncol. 2020;13(6):83. doi:10.3892/mco.2020.2153
36. Papp LV, Lu J, Holmgren A, Khanna KK. From selenium to selenoproteins: synthesis, identity, and their role in human health. Antioxid Redox Signal. 2007;9(7):775–806. doi:10.1089/ars.2007.1528
37. Squires JE, Stoytchev I, Forry EP, Berry MJ. SBP2 binding affinity is a major determinant in differential selenoprotein mRNA translation and sensitivity to nonsense-mediated decay. Mol Cell Biol. 2007;27(22):7848–7855. doi:10.1128/MCB.00793-07
38. Donovan J, Caban K, Ranaweera R, Gonzalez-Flores JN, Copeland PR. A novel protein domain induces high affinity selenocysteine insertion sequence binding and elongation factor recruitment. J Biol Chem. 2008;283(50):35129–35139. doi:10.1074/jbc.M806008200
39. Howard MT, Copeland PR. New directions for understanding the codon redefinition required for selenocysteine incorporation. Biol Trace Elem Res. 2019;192(1):18–25. doi:10.1007/s12011-019-01827-y
40. Louis DN. Molecular pathology of malignant gliomas. Annu Rev Pathol. 2006;1:97–117. doi:10.1146/annurev.pathol.1.110304.100043
41. Furnari FB, Fenton T, Bachoo RM, et al. Malignant astrocytic glioma: genetics, biology, and paths to treatment. Genes Dev. 2007;21(21):2683–2710. doi:10.1101/gad.1596707
42. Gimple RC, Bhargava S, Dixit D, Rich JN. Glioblastoma stem cells: lessons from the tumor hierarchy in a lethal cancer. Genes Dev. 2019;33(11–12):591–609. doi:10.1101/gad.324301.119
43. Le Rhun E, Preusser M, Roth P, et al. Molecular targeted therapy of glioblastoma. Cancer Treat Rev. 2019;80:101896. doi:10.1016/j.ctrv.2019.101896
44. Lv S, Luo H, Huang K, Zhu X. The prognostic role of glutathione peroxidase 1 and immune infiltrates in glioma investigated using public datasets. Med Sci Monit. 2020;26:e926440. doi:10.12659/MSM.926440
45. Chan TA, Yarchoan M, Jaffee E, et al. Development of tumor mutation burden as an immunotherapy biomarker: utility for the oncology clinic. Ann Oncol. 2019;30(1):44–56. doi:10.1093/annonc/mdy495
46. Samstein RM, Lee CH, Shoushtari AN, et al. Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat Genet. 2019;51(2):202–206. doi:10.1038/s41588-018-0312-8
47. Ott PA, Hu Z, Keskin DB, et al. An immunogenic personal neoantigen vaccine for patients with melanoma. Nature. 2017;547(7662):217–221. doi:10.1038/nature22991
48. Schumacher TN, Schreiber RD. Neoantigens in cancer immunotherapy. Science. 2015;348(6230):69–74. doi:10.1126/science.aaa4971
49. Chang L, Chang M, Chang HM, Chang F. Microsatellite Instability: a predictive biomarker for cancer immunotherapy. Appl Immunohistochem Mol Morphol. 2018;26(2):e15–e21. doi:10.1097/PAI.0000000000000575
50. Andre T, Shiu KK, Kim TW, et al. Pembrolizumab in Microsatellite-instability-high advanced colorectal cancer. N Engl J Med. 2020;383(23):2207–2218.
51. Avery JC, Hoffmann PR. Selenium, Selenoproteins, and Immunity. Nutrients. 2018;10(9):1203. doi:10.3390/nu10091203
52. Razaghi A, Poorebrahim M, Sarhan D, Bjornstedt M. Selenium stimulates the antitumour immunity: insights to future research. Eur J Cancer. 2021;155:256–267. doi:10.1016/j.ejca.2021.07.013
53. Santi A, Kugeratski FG, Zanivan S. Cancer associated fibroblasts: the architects of stroma remodeling. Proteomics. 2018;18(5–6):e1700167. doi:10.1002/pmic.201700167
54. Michalik L, Desvergne B, Wahli W. Peroxisome-proliferator-activated receptors and cancers: complex stories. Nat Rev Cancer. 2004;4(1):61–70. doi:10.1038/nrc1254
55. Sertznig P, Seifert M, Tilgen W, Reichrath J. Present concepts and future outlook: function of peroxisome proliferator-activated receptors (PPARs) for pathogenesis, progression, and therapy of cancer. J Cell Physiol. 2007;212(1):1–12. doi:10.1002/jcp.20998
56. Gandhi UH, Kaushal N, Ravindra KC, et al. Selenoprotein-dependent up-regulation of hematopoietic prostaglandin D2 synthase in macrophages is mediated through the activation of peroxisome proliferator-activated receptor (PPAR) gamma. J Biol Chem. 2011;286(31):27471–27482. doi:10.1074/jbc.M111.260547
57. Shi Y, Zou Y, Shen Z, et al. Trace Elements, PPARs, and metabolic syndrome. Int J Mol Sci. 2020;21(7):2612. doi:10.3390/ijms21072612
58. Diwakar BT, Korwar AM, Paulson RF, Prabhu KS. The regulation of pathways of inflammation and resolution in immune cells and cancer stem cells by selenium. Adv Cancer Res. 2017;136:153–172.
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.