Back to Journals » Infection and Drug Resistance » Volume 11
Bacterial profile, antibiotic resistance pattern and associated factors among cancer patients at University of Gondar Hospital, Northwest Ethiopia
Authors Fentie A, Wondimeneh Y , Balcha A, Amsalu A , Adankie BT
Received 10 August 2018
Accepted for publication 27 September 2018
Published 8 November 2018 Volume 2018:11 Pages 2169—2178
DOI https://doi.org/10.2147/IDR.S183283
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Suresh Antony
Alelign Fentie,1 Yitayih Wondimeneh,1 Abera Balcha,2 Anteneh Amsalu,1 Birhanemeskel Tegene Adankie1
1Department of Medical Microbiology, College of Medicine and Health Sciences, University of Gondar, Gondar, Ethiopia; 2School of Medicine, College of Medicine and Health Sciences, University of Gondar, Gondar, Ethiopia
Introduction: Cancer is alarmingly increased in developing countries like Ethiopia, where multidrug resistant bacterial infection is rampant. The aim of this study was to determine the bacterial profile, antimicrobial resistance pattern, and associated factors among cancer patients attending University of Gondar Hospital.
Methods: A consecutive 216 cancer patients were recruited from February to April, 2017. Socio-demographic and clinical data were collected using a structured questionnaire. Culture and antibiotic resistance were done following standard microbiological procedures.
Result: The overall prevalence of bacterial infection was 19.4%. The predominant bacterial isolates were Staphylococcus aureus (28.6%), followed by coagulase negative staphylococci (26.2%) and Escherichia coli (21.4%). Multidrug resistance was detected in 46.5% bacterial isolates. Methicillin resistance was detected in 25% of S. aureus and in 45.5% of coagulase negative staphylococci. Fluoroquinolone resistance was detected in 33.3% of E. coli isolates. Cancer patients with solid tumor, started cancer therapy, and being symptomatic had higher odds of culture positivity.
Conclusion: The overall burden of bacterial infection among cancer patients is considerably high. The findings of this study inform baseline information for policymakers and call for additional studies with large isolates in different cancer treatment centers in the region and in the country to better understand the bacterial isolate and resistance pattern.
Keywords: cancer patient, bacterial profile, antibiotic resistance, Ethiopia
Introduction
Major advances in the care of cancer patients over the past several decades have led to significant improvement in patient survival. Despite these advances, cancer patients are prone to serious infection complications with substantial morbidity and mortality. In this patient population, infection risk results from a complex interplay between the host’s underlying immunodeficiency and the nature of treatment practices they experience (like surgery, radiation therapy, chemotherapy), prophylaxis use, and application of invasive procedures (central venous catheter and urinary catheter).1 The symptoms of infection in cancer patients could be masked by the cancer treatment modalities,2 that is an indicator for considering asymptomatic infections.
Previous studies on bacterial infection and drug resistance pattern among cancer patients were mainly focused on bloodstream infection (BSI) with hematologic malignancies.2 However, cancer patients who have solid tumors might have a tendency to undergo surgery to remove the tumor or sometimes due to other medical reasons. This, thus, increases the potential of acquiring bacterial infection either by endogenously normal flora near the operative sites or exogenously from the hospital environments, such as in the air, hospital staff, inanimate objects, and medical equipment, as a result of their prolonged and frequent contact.4,5
Therefore, patients with both type of cancer are highly susceptible to almost any type of bacterial infection.6 Among Gram positive bacteria (GPB) genus Staphylococcus and from Gram negative bacteria (GNB): Escherichia coli, Klebsiella pneumoniae, and Pseudomonas aeruginosa are frequently associated.7 Moreover, frequent prescription of broad-spectrum antibiotics as prophylaxis among cancer patients may potentially alter the composition of endogenous flora and select multidrug resistant pathogens such as methicillin-resistant Staphylococcus aureus (MRSA), vancomycin-resistant enterococci (VRE), and Gram-negative bacilli (eg, fluoroquinolone-resistant Escherichia coli [FREC]).8–10 As a result, empirical antibiotic treatments of cancer patients are continually challenged by (1) the change in frequency of Gram-positive and Gram-negative bacteria and (2) the emergence of new antimicrobial resistant pathogens. The pattern and prevalence of resistance may vary with respect to geographical location and difference in infection prevention and control strategies between health care facilities.
In Ethiopia, the cancer care services were first started in an organized manner in 1998 at Tikur Anbesa Specialized Hospital,11 and have been subsequently scaled up into eight cancer centers including the University of Gondar Hospital in 2015. The treatment of bacterial infections in patients with cancer should often rely on the use of established guidelines, along with consideration of the local epidemiology and antibiotic susceptibility patterns of the potential etiologic agents. However, in Ethiopia identification and drug susceptibility testing are not routinely performed, so patient management is based on empirical treatment. The actual burden and antimicrobial resistance pattern of bacterial isolates are not known. Thus, the present study provides baseline information on the spectrum of bacterial isolates, antimicrobial resistance patterns, and associated risk factors in cancer patients attending the University of Gondar Hospital, Northwest Ethiopia.
Materials and methods
Ethical consideration
The proposal was reviewed and approved by the School of Biomedical and Laboratory Science ethical committee. Permission was obtained from the University of Gondar Hospital administration. The purpose of the study was explained to each study participant. In addition to informed written consent, obtained from each adult participant, assent was obtained from a parent or legal guardian of patients under the age of 18 years. Patient information was anonymized and deidentified prior to analysis. The laboratory result was given to the attending physician and treated accordingly.
Study design, area, and period
A hospital-based cross-sectional study was conducted at the University of Gondar Hospital, cancer treatment center from February to April, 2017. The hospital is located in Gondar town, which is 182 km from Bahir Dar, the capital city of Amhara regional state, and 747 km from Addis Ababa, the capital city of Ethiopia. Apart from other services, the established cancer treatment center provides services for 2,300 cancer cases. The cancer treatment center has outpatient and inpatient departments with 20 beds. The most common treatment available for cancer patients in the center are surgery and chemotherapy.
Study population
All cancer confirmed patients who visited the University of Gondar Hospital cancer treatment center during the study period.
Operational definition
Multiple drug resistant (MDR) was defined as a bacterial isolate, which is resistant to one or more antibiotics in three or more classes of antimicrobials agents.31
MRSA and methicillin-resistant coagulase negative staphylococci (MRCoNS) were defined as Staphylococcus aureus and CoNS resistance for cefoxitin disk (≥21 mm).12
BSI was defined as isolation from blood of one (monomicrobial) or more (polymicrobial) clinically significant organisms during a febrile episode.
A febrile patient was defined as a cancer patient having a single auxiliary temperature of >37.5°C sustained over a 1-hour period.
Clinical and laboratory data
Clinical examination and other co-morbidity factors were diagnosed by an oncologist. Socio-demographic and clinical data were collected by using a structured questionnaire by nurses working in the center. According to the patients’ clinical status, different types of specimen were collected.
Blood sample collection and processing
Blood samples (5 mL for adults and 1 mL for children each) for cultures were obtained from each patient who developed fever at the time of diagnosis. Blood samples were drawn from two different sites of peripheral vein aseptically (disinfecting with 70% alcohol and 2% tincture of iodine) by experienced nurses prior to any antibiotic use. The collected blood samples were then transferred into culture bottles of sterile tryptic soy broth (Oxoid Ltd., Basingstoke, UK). Bottles were incubated at 37°C for 7 days and observed for signs of bacterial growth (turbidity, hemolysis, clot formation) on a daily bias for up to7 days. Bottles which showed signs of growth were gram stained and subcultured on blood agar, chocolate agar, MacConkey agar, and mannitol salt agar (Oxoid Ltd.). These plates were than aerobically incubated for 18–24 hours at 37°C. Blood sample containing broths with no bacterial growth after 7 days was subcultured before being reported as a negative result.
Absolute neutrophil count
Samples (2 mL and 1 mL of blood from adults and children, respectively) were collected simultaneously using ethylene diamine tetra acetic acid tubes during blood collection for culture. Absolute neutrophil count was done using a XT-4000i hematology analyzer (Sysmex Europe GmbH, Norderstedt, Germany).
Urine sample collection and processing
Midstream urine was collected with a sterile urine container from both symptomatic and asymptomatic urinary tract infection (UTI) cases. Midstream urine specimens were inoculated on cystine lactose electrolyte deficient (CLED) by using a calibrated loop (0.001/mL). All the media were incubated at 37°C for 18–24 hours. Significant bacteriuria was defined as a colony count ≥105 CFU/mL urine.
Wound swab/ear discharge collection and processing
Before antibiotics were started, wound swab and ear discharge samples were collected aseptically based on the clinical manifestations of the cancer patients by using a sterile saline moistened cotton swab. The swabs were streaked on MacConkey agar, chocolate agar, blood agar plates, and mannitol salt agar. These plates were then aerobically incubated for 18–24 hours at 37°C.
Bacterial identification
The bacterial pathogens were identified after appearance of growth on subcultured/cultured plates of blood/wound swab/discharge samples and significant growth on CLED by standard microbiological and biochemical procedures.13 CoNS were only considered as causative pathogens if similar pathogens were isolated on two blood samples drawn on separate occasions.
Antimicrobial susceptibility testing
Antibiotic susceptibility testing of bacterial isolates was done by Kirby Bauer disc diffusion method using Muller Hinton agar (MHA) plate (Oxoid Ltd.).14 Three-to-five pure colonies from a fresh non-selective agar plate were selected and transferred to 5 mL sterile nutrient broth (Oxoid Ltd.) and thoroughly mixed to make the suspension homogenous. The mixture was transferred into sterile normal saline by sterile pipette and turbidity was adjusted visually with sterile normal saline to match with a 0.5 McFarland standard. The suspension was then inoculated uniformly over the entire surface of a MHA plate using a sterile swab. The inoculated plates were left at room temperature to dry for 3–5 minutes. Using sterile forceps, the antibiotic discs were placed on MHA by considering 24 mm distance between each disk and 15 mm from the border. The tested antibiotic discs include: ciprofloxacin (5 µg), gentamicin (10 µg), norfloxacin (10 µg), and nitrofurantoin (300 µg), for both Gram positive and Gram negative bacterial isolates; cot-trimoxazole (25 µg), penicillin (10 µg), vancomycin (30 µg), cefoxitin (30 µg), and tetracycline (30 µg) for GPB. Amikacine (30 µg), ceftriaxone (30 µg), cefepime (30 µg), cefixime (5 µg), ampicillin (10 µg), amoxicillin/clavulanic acid (20/10 µg), and nalidixic acid (30 µg) were used for GNB. After 18–24 hours incubation at 37°C, the zone of inhibition was measured and reported as susceptible (S), intermediate (I), or resistance (R), according to the clinical and laboratory standard institute (CLSI) guideline.12
Quality control
The reliability of the study findings was guaranteed by implementing quality control measures that include sterility and performance checks throughout the whole process of the laboratory work. E. coli ATCC 25922, P. aeruginosa ATCC 27853, and S. aureus ATCC 25923 control strains were used. Moreover, the validity and completeness of the questionnaire were checked daily by the principal investigator.
Data analysis
Data were entered, coded, and analyzed using SPSS software version 20. Variables that showed a significant association in a bivariate analysis were selected for further analysis using multivariable logistic regression analysis. A P-value <0.05 was considered as statistically significant. The strength of the association was interpreted using an adjusted odds ratio in a 95% confidence interval.
Ethics approval and consent to participate
This study was approved by the institutional review board (IRB) of University, College of Medicine and Health Sciences. Informed written consent and/assent was obtained from study participants. Patient information was anonymized and deidentified prior to analysis.
Results
Sociodemographic characteristics
A total of 216 cancer patients were included in the study. Out of these, 136 (63.0%) were females and 80 (37.0%) were males, with a male-to-female ratio of a 1:1.7. The mean age±SD of study participants was 41.5±17.77 years, range 1–73 years. The majority (134, 62.0%) of study subjects had no formal education, and 83 (38.4%) had housewife as their occupation. Among the study population, 111 (51.4%) were rural in residence and 138 (63.9%) were recruited in the outpatients department (Table 1).
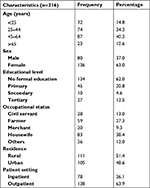  | Table 1 Socio-demographic characteristics of cancer patients at the University of Gondar Hospital, Northwest Ethiopia, 2017 |
Clinical characteristics of study populations at the time of diagnosis
Out of 216 cancer patients, 167 (77.3%) had a solid tumor. Among the total, breast cancer (41, 19.0%) was the highest, followed by lymphoma (34, 15.7%) and ovarian tumor (26, 12.0%). Thirty-six (16.7%) cancer patients were stage IV, and 155 (71.8%) had started cancer therapy. Of these, the most frequently used were chemotherapy (114, 73.5%), either alone or in combination with surgery or radiotherapy. The majority of the patients (145, 67.1%) had a history of catheterization and had at least one sign and symptoms (116, 53.7%) for different clinical features. Of these, 23 (10.6%) were febrile at the time of diagnosis (Table 2).
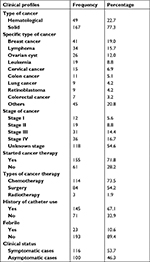  | Table 2 Clinical characteristics of cancer patients at the University of Gondar Hospital, Northwest Ethiopia, 2017 |
Bacterial profiles and site of isolation
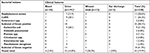
Bacterial cultures were positive in 42 patients (19.4%, 95% CI=14.4–25.0) with 43 isolates. Polymicrobial growth was recovered only in one specimen. Among the 42 culture positive cases, 6% (6/100) were from the urine of asymptomatic patients. The majority of the isolates were from urine (n=32) and wound swab (n=7), with a low frequency of the isolates from blood (n=3) and ear discharge (n=1) samples. GPB species (25, 58.1%) were higher than GNB species (18, 41.9%). GPB accounted for 41.9% (18/43) of the UTI, 9.3% (4/43) of wound infections, and 7% (3/43) of blood stream infections while GNB attributed to 32.6% (14/43) of UTI and 7% (3/43) of wound infections. The overall predominant bacterial isolates were S. aureus (12, 28.6%), followed by CoNS (11, 26.2%) and E. coli (9, 21.4%). The most common isolates identified in UTI were E. coli and CoNS (9, 28.1%) each (Table 3).
Antimicrobial resistance pattern
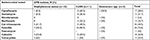
As to the antimicrobial resistant pattern for GPB, high resistance rates were observed, among total GPB, to penicillin (18, 72.0%) and cot-trimoxazole (10, 43.5%). On the other hand, a low resistance rate was detected for nitrofurantoin (1, 5.6%). Among the predominant isolates, MRSA and MRCoNS were detected in three (25%) and five (45.5%), respectively. VRE was also detected in one isolate (Table 4).
Similarly, high resistance rates to ampicillin (15, 88.2%), amoxicillin/clavulonic acid (14, 82.4%), and nalidixic acid (8, 57.1%) were observed in GNB. On the other hand, all GNB were sensitive to nitrofurantoin and had shown low resistance to amikacine (1, 5.6%). Among the most common isolated GNB, E. coli showed a higher rate of resistance to amoxicillin/clavulonic acid (8, 88.9%), ampicillin (7, 77.8%), and nalidixic acid (6, 66.7%). Moreover, fluoroquinolone (ciprofloxacin and/or norfloxacin) resistance was observed in 33.3% of E. coli (FREC) (Table 5).
MDR
In this study, the prevalence of MDR was 20 (46.5%), while five (11.6%) bacterial isolates were sensitive to all antibiotics tested, and none of the isolated pathogens were resistant to all antibiotic classes tested. On the other hand, the result of drug resistance patterns compared within species specific showed that two (100%) of the K. pneumoniae and six (66.7%) E. coli were MDR isolates (Table 6).
Factors associated with culture positivity rate
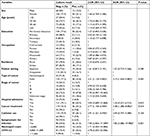
In a bivariate logistic regression analysis, cancer patients with solid tumor were 3.3 times more likely to be culture positive (COR=3.31; 95% CI=1.120–9.805; P=0.03) as compared to patients with hematological cancer. Moreover, the culture positivity rate among cancer patients started cancer therapy (COR=2.77; 95% CI=1.104–6.969; P=0.015), being catheterized (COR=2.91; 95% CI=1.221–6.930; P=0.016) and being symptomatic during diagnosis (COR=7.85; 95% CI=2.826–17.589; P=0.00) were significantly higher as compared to their counterparts. None of the socio-demographic factors, patient setting, specific type of cancer, stage of cancer, or hospital admission had statistically significant association with culture positivity.
In multivariate logistic analysis, cancer patients who started cancer therapy were 3.38 times more likely culture positive (aOR=3.38; 95% CI=1.286–8.882: P=0.014) as compared to patients who had not yet started therapy, and cancer patients with solid tumor had 3.9 times higher odds of culture positivity (aOR=3.91; 95% CI=1.026–9.882: P=0.045) as compared to hematological malignancy. Moreover, being symptomatic during diagnosis had 7.9 times higher odds of culture positivity (aOR=7.85; 95% CI=3.086–19.980: P=0.001) as compared to asymptomatic cases. However, in the multivariate regression model, we observed that being catheterized is no more an independent predictor (Table 7).
Discussion
The overall prevalence of bacterial infections among cancer patients in this study was 19.4%. This finding was comparable with the studies conducted in Romania (14.92%)15 and Iran (24.6%) among cancer patients with BSI.16 However, this finding was lower as compared to other studies conducted in Sudan (48.1%)5 and Iraq (mean of 44.2%).17 The variation in prevalence might be explained by the fact that differences in geographical location and the use of a sensitive blood culture system (BD Phoenix automated microbiology system, Waldorf, Germany) that might increase culture positivity rate.16,17 Besides the difference in the study population, in Iraq the study population were children with febrile neutropenia, while in Sudan the study was on cancer patients with nosocomial infection, that might have a high culture positivity rate as compared to our study populations, in which almost half of the study population were asymptomatic cancer patients.
The epidemiology of bacterial infections among cancer patients showing a shift in the prevalence from Gram-positive to Gram-negative bacteria and vice versa remains controversial. In this study, GPB were the predominant cause of bacterial infection, as compared to GNB. This finding is in line with studies conducted elsewhere in cancer patients with BSI, febrile neutropenia, or hematological neoplasia.16,18,19 The possible reasons for the preponderance of GPB in this study might be the increased use of indwelling catheters. Moreover, chemotherapy-induced mucositis and use of both prophylactic and empiric antibiotic regimens targeting GNB diminishes recovery of Gram-negative pathogens, while selecting for GPB were reported.9,10 In contrary, a study from other African countries reported that GNB were significantly more predominant isolates from cancer patients,5,7 and current data from other studies indicates the re-emergence of GNB among febrile neutropenic cancer patients.8,20 This might be due to minimal use or the discontinuation of fluoroquinolones prophylaxis.21,22 These findings strengthen the need for frequent surveillance for understanding the local epidemiology of bacterial infection among cancer patients.
S. aureus, CoNS, and E. coli were the most common bacterial pathogens isolated in cancer patients in this study. More or less similar patterns have been reported in cancer patients in different countries, although the proportion of the bacterial agents varied.7,17 The GPB S. aureus and CoNS are ubiquitous in nature, which are frequently found on the skin and are the main cause of various infections, mainly in patients with solid tumors following indwelling devices, invasive surgical procedures, and contamination from hospital environments.5,23 Likewise, E. coli is a normal member of gastro-intestinal flora and a common cause of both community and hospital acquired UTI. Particularly immunocompromised cancer patients are easily colonized by the bacteria, due to the fact that infection of cancer patients by this bacterium is inevitable.
Several studies reported very high rates of methicillin resistance (MR) among S. aureus as well as CoNS from cancer patients.7,16 In this study, 25% of MRSA and 45.5% of MRCoNS were detected. Similarly, consistent findings were also claimed in the same hospital (26.5%),24 in Ethiopia (pooled prevalence) (32.5%),25 and in other countries in Saudi Arabia (28.62%)26 and Libya (35.3%).7 Although emergence of VRE in cancer patients has been reported, one isolate of VRE was recovered from a urine specimen in this study. This suggests that the utilization of empirical vancomycin in our hospital must be thoroughly scrutinized.
GNB revealed a high resistance rate to ampicillin (88.2%) and for amoxicillin/clavulonic acid (82.4%), similar to the study conducted in Libya.7,17 Among the predominant GNB isolates, E. coli was highly resistant to amoxicillin/clavulonic acid, and ampicillin, which is in agreement with a study conducted in Sudan.5 In this study, 33.3% of E. coli was resistant to fluoroquinolone, which is also comparable with the results reported from Libya.7 However, it is lower than the report from developed countries where fluoroquinolone is most commonly prescribed as a prophylaxis in cancer patients receiving chemotherapy.3 When the prevalence of fluoroquinolone resistance in GNB exceeds 20%, it indicates there is widespread use of fluoroquinolone that would encourage multiclass drug resistance.10
In this study, nitrofurantoin showed a low level of resistance, which makes this drug a reasonable alternative in the management of UTI in cancer patients. This may have been due to its multiple mechanisms of action and under-usage of this drug in the study area. However, it cannot be used to treat UTI in complicated and febrile infants, because it is excreted in the urine and does not achieve therapeutic concentrations in the bloodstream.27
Although the development of MDR is a natural phenomenon, an interestingly extensive raise in the number of immunocompromised conditions, like cancer, increases MDR, thereby contributing to a further spread of MDR isolates, since these patients had frequent follow-up within the hospital.28 In this study, the overall prevalence of MDR was 46.5% lower than a recent report in the same place from neonatal septicemia (65%).24 Among the isolated bacteria, K. pneumoniae and E. coli were the principal MDR strains concordant with a previous study in the same place.24
Most studies have shown that cancer patients with hematological malignancy had a higher risk of developing bacterial infection as compared to solid tumor patients.1,26,29 In this study, cancer patients with a solid tumor were 3.9 times more likely to be culture positive than hematological malignancy, because the majority of patients with solid tumors undergo surgical treatment and catheterization as a result of the development of postoperative or catheter-associated infections is common. Besides, patients who started cancer therapy had higher odds of culture positivity as compared to their counterparts, concordant with the studies conducted in Qatar29 and Spain.30 Moreover, cancer patients who were symptomatic during sample collection had ~7.8 times higher odds of culture positivity as compared to asymptomatic cases, indicating that the use of culture and sensitivity test for symptomatic cases are so substantial to underpin effective treatment and prevention strategies. However, 6% asymptomatic bacteriuria reported in this study should not be undermined in the era of MDR.
Limitations
Despite a number of strengths, our prospective cross-sectional study has some limitations that should be acknowledged. First, it was performed in a single center, which may not reflect the epidemiology of different centers and/or different geographical areas. Second, due to the lack of budget, identification of extended-spectrum β-lactamase and carbapenemase-producing isolates was not done. Third, the duration, type of treatment modality, prior infection histories, and prophylaxis used were not collected.
Conclusion
The overall burden of bacterial infections among cancer patients is considerably higher, particularly in patients with solid tumor, who had started cancer therapy, and those being symptomatic during diagnosis. GPB isolates were higher than GNB. The most common bacterial isolates were S. aureus, CoNS, and E. coli. Multidrug resistance was detected in 46.5% bacterial isolates. MR was detected in 25% of S. aureus and in 45.5% of CoNS isolates. Fluoroquinolone resistance was detected in 33.3% of E. coli isolates. The findings of the current study call for a more robust study with a larger isolate number to better understand the resistance pattern of the common bacterial isolates and advise policymakers to develop guideline.
Availability of data and material
All the raw data supporting the findings can be requested from the principal investigator Alelign Fentie.
Acknowledgments
We express our gratitude to our study participants who took part in the study. We also thank the staff members of the University of Gondar Hospital Cancer Treatment Center for their professional assistance during the data collection and processing. Finally, our special acknowledgment goes to Dr Gizachew Yismaw and Dr Eueal Negash for their professional advice during data collection. Lastly, we acknowledge the University of Gondar for financial support. This research was partially funded by the University of Gondar for data collection.
Author contributions
All authors contributed to data analysis, drafting and revising the article, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Disclosure
The authors report no conflicts of interest in this work.
References
Zembower TR. Epidemiology of infections in cancer patients. Cancer Treat Res. 2014;161:43–89. | ||
Greene JN. Cheson BD. Infections in Cancer Patients (Basic and Clinical Oncology). 5th ed. New York: Marcel Dekker, Inc; 2004. | ||
Freifeld AG, Bow EJ, Sepkowitz KA, et al. Clinical practice guideline for the use of antimicrobial agents in neutropenic patients with cancer: 2010 update by the infectious diseases society of America. Clin Infect Dis. 2011;52(4):e56–e93. | ||
Homsi J, Walsh D, Panta R, Lagman R, Nelson KA, Longworth DL. Infectious complications of advanced cancer. Support Care Cancer. 2000;8(6):487–492. | ||
Nurain AM, Bilal NE, Ibrahim ME. The frequency and antimicrobial resistance patterns of nosocomial pathogens recovered from cancer patients and hospital environments. Asian Pac J Trop Biomed. 2015;5(12):1055–1059. | ||
Thom KA, Kleinberg M, Roghmann M. Infection Prevention in the Cancer Center. Clin Infect Dis. 2013:57(4):579–585. | ||
Zorgani AA, Belgasim Z, Ziglam H, Ghenghesh KS. Antimicrobial susceptibility profiles of gram- negative bacilli and gram-positive cocci isolated from cancer patients in Libya. Clin Microbiol. 2012;3(3):1–8. | ||
Markman M. Principles and Practice of Cancer Infectious Diseases. Lubbock, TX: Humana press; 2011. | ||
Maschmeyer G, Haas A. The epidemiology and treatment of infections in cancer patients. Int J Antimicrob Agents. 2008;31(3):193–197. | ||
Bow EJ. Fluoroquinolones, antimicrobial resistance and neutropenic cancer patients. Curr Opin Infect Dis. 2011;24(6):545–553. | ||
Tigeneh W, Molla A, Abreha A, Assefa M. Pattern of cancer in Tikur Anbessa specialized hospital oncology center in Ethiopia from 1998 to 2010. Int J Cancer Res Mol Mech. 2015;1:1–5. | ||
CLSI. Performance Standards for Antimicrobial Susceptibility Testing M100. 27th ed. Wayne, PA: CLSI; 2017. | ||
Cheesbrough M. District Laboratory Practice in Tropical Countries: Part 2. 2nd ed. New York: Cambridge University; 2006:1–43. | ||
Bauer AW, Kirby WM, Sherris JC, Turck M. Antibiotic susceptibility testing by a standardized single disk method. Am J Clin Pathol. 1966;45(4):493–496. | ||
Manafu EG, Filimon RM, Jari I, Petrariu FD, Manole A. Clinical epidemiological study on the incidence of Escherichia coli infections in the cancer patients admitted to Surgery Department II of the IASI Regional Oncology Institute in 2013. Rev Med Chir Soc Med Nat Iasi. 2014;118(3):796–800. | ||
Nejad ZE, Ghafouri E, Farahmandi-Nia Z, Kalantari B, Saffari F. Isolation, Identification, and Profile of Antibiotic Resistance of Bacteria in Patients with Cancer. IJMS. 2010;35(2):109–115. | ||
Almaziny MA. Isolation, identification, and profile of antibiotic resistance of bacteria in childhood febrile neutropenia patients. Eur J Exp Biol. 2014;4(2):1–6. | ||
Meidani M, Bagheri A, Khorvash F. A Population-Based Study of Bacterial Spectrum in Febrile Neutropenic Patients. Jundishapur J Microbiol. 2013;6(2):150–156. | ||
Gaytan-Martinez J, Avila-Moran M, Mata-Marín J, et al. Prevalence of bacterial infections and patterns of bacterial susceptibility among patients with fever, neutropenia, and hematological neoplasias. Gaceta medica de Mexico. 2010;147(4):325–332. | ||
Walwyn M, Nicholson A, Lee MG, Wharfe G, Frankson MA. Febrile neutropaenia in cancer patients. West Indian Med J. 2010;59(2):209–214. | ||
Montassier E, Batard E, Gastinne T, Potel G, de La Cochetière MF. Recent changes in bacteremia in patients with cancer: a systematic review of epidemiology and antibiotic resistance. Eur J Clin Microbiol Infect Dis. 2013;32(7):841–850. | ||
Chong Y, Yakushiji H, Ito Y, Kamimura T. Clinical impact of fluoroquinolone prophylaxis in neutropenic patients with hematological malignancies. Int J Infect Dis. 2011;15(4):e277–e281. | ||
Safdar A. Principles and Practice of Cancer Infectious Diseases. Curr Clin Oncol. 2011:3–15. | ||
G/eyesus T, Moges F, Eshetie S, Yeshitela B, Abate E, Ebba A. Bacterial etiologic agents causing neonatal sepsis and associated risk factors in Gondar, Northwest Ethiopia. BMC Pediatr. 2017;17(1):137. | ||
Eshetie S, Tarekegn F, Moges F, Amsalu A, Birhan W, Huruy K. Methicillin resistant Staphylococcus aureus in Ethiopia: a meta-analysis. BMC Infect Dis. 2016;16(1):689. | ||
Sirkhazi M, Sarriff A, Aziz NA, Almana F, Arafat O, Shorman M. Bacterial Spectrum, Isolation Sites and Susceptibility Patterns of Pathogens in Adult Febrile Neutropenic Cancer Patients at a Specialist Hospital in Saudi Arabia. World J Oncol. 2014;5(5–6):196–203. | ||
Vélez Echeverri C, Serna-Higuita LM, Serrano AK, et al. Resistance profile for pathogens causing urinary tract infection in a pediatric population, and antibiotic treatment response at a university hospital, 2010–2011. Colomb Med. 2014;45(1):39–44. | ||
Tanwar J, Das S, Fatima Z, Hameed S. Multidrug Resistance: An Emerging Crisis. Interdiscip Perspect Infect Dis. 2014;2014(12):1–7. | ||
Al-Mulla NA, Taj-Aldeen SJ, El Shafie S, Janahi M, Al-Nasser AA, Chandra P. Bacterial bloodstream infections and antimicrobial susceptibility pattern in pediatric hematology/oncology patients after anticancer chemotherapy. Infect Drug Resist. 2014;7:289–299. | ||
Gudiol C, Bodro M, Simonetti A, et al. Changing aetiology, clinical features, antimicrobial resistance, and outcomes of bloodstream infection in neutropenic cancer patients. Clin Microbiol Infect. 2013;19(5):474–479. | ||
Magiorakos AP, Srinivasan A, Carey RB, et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect. 2012;18(3):268–281. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.