Back to Journals » International Journal of Chronic Obstructive Pulmonary Disease » Volume 9 » Issue 1
Bacterial microbiome of lungs in COPD
Received 29 October 2013
Accepted for publication 25 December 2013
Published 21 February 2014 Volume 2014:9(1) Pages 229—238
DOI https://doi.org/10.2147/COPD.S38932
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Marc A Sze,1 James C Hogg,2 Don D Sin1
1Department of Medicine, 2Department of Pathology and Laboratory Medicine, The James Hogg Research Centre, Providence Heart-Lung Institute, St Paul's Hospital, University of British Columbia, Vancouver, BC, Canada
Abstract: Chronic obstructive pulmonary disease (COPD) is currently the third leading cause of death in the world. Although smoking is the main risk factor for this disease, only a minority of smokers develop COPD. Why this happens is largely unknown. Recent discoveries by the human microbiome project have shed new light on the importance and richness of the bacterial microbiota at different body sites in human beings. The microbiota plays a particularly important role in the development and functional integrity of the immune system. Shifts or perturbations in the microbiota can lead to disease. COPD is in part mediated by dysregulated immune responses to cigarette smoke and other environmental insults. Although traditionally the lung has been viewed as a sterile organ, by using highly sensitive genomic techniques, recent reports have identified diverse bacterial communities in the human lung that may change in COPD. This review summarizes the current knowledge concerning the lung microbiota in COPD and its potential implications for pathogenesis of the disease.
Keywords: chronic obstructive pulmonary disease, bacterial microbiome, lungs
What is the microbiome?
The microbiome is defined as the total collection of microbiota that resides within humans or on their skin surface. Thus, the lung microbiome is the complete collection of microbiota residing in the airways and parenchymal tissues. The microbiota, in turn, is defined as all microorganisms that inhabit a particular site or place, eg, the gastrointestinal (GI) tract, skin, or airways.1–3 These microorganisms can consist of, but are not limited to, bacteria, viruses, and fungi,4,5 of which the most widely studied have been bacteria. In the human body, bacteria outnumber human cells by a factor of 10.6 Most of these bacteria are found in the GI tract.6 Other sources of frequent habitation include the mouth, nose, and skin.7 Bacteria coexist with fungi and viruses in these locations. Despite their abundance, the functional role of bacteria in most human organs is largely unknown. With the advent of new molecular techniques and animal models, the mysteries of the human microbiota are beginning to be unraveled, especially in the GI system where bacteria appear to regulate host immune responses.8–10 The gut is the home of the largest number of bacteria and, not surprisingly, the first large-scale human microbiota studies were conducted using GI tissues.11,12 The initial hypothesis was that most microorganisms residing in the GI tract caused human disease.13 However, with careful investigation, it became clear that many bacteria in the GI tract were in fact beneficial to humans.14 For example, some bacteria are required for the production of essential micronutrients (such as vitamin K), protection against pathogens (such as Clostridium difficile), regulation of the host inflammatory response,15,16 and disease expression.17–19 It is now recognized that perturbations in the gut microbiota may be responsible for a wide range of diseases, including pseudomembranous colitis,20 inflammatory bowel disease,21 and even non-GI conditions such as obesity and cardiovascular disease.22 Although less is known about the role of the lung microbiota in the pathogenesis of chronic obstructive pulmonary disease (COPD), in this paper we review some of the key new findings that have linked changes in bacterial communities in the lung with disease progression and exacerbation of COPD.
Background on COPD
COPD is currently the third leading cause of death in the world and the second leading cause of disability-adjusted life years lost in the US.23,24 Although smoking is the leading risk factor, only 15%–20% of persistent smokers go on to develop the disease.25 COPD is an umbrella term that encompasses several distinctive phenotypes,26 including chronic bronchitis, small airways disease (or bronchiolitis), mucus hypersecretion, and emphysema.25,27,28 Despite the morphologic and clinical heterogeneity of COPD, one common feature is inflammation in the small airways, which intensifies with disease progression. The inflammatory response in the lungs is characterized by infiltration of inflammatory and immune cells, including CD4+ T-cells, CD8+ T-cells, B-cells, macrophages, and neutrophils.29,30 Notable histologic findings in moderate to severe COPD (but not in mild disease) are tertiary lymphoid follicles in the walls of small airways.28 In COPD, the putative antigen for these follicles is unknown, although some have implicated elastin fragments31 and others have suggested environmental triggers such as bacteria.32
British hypothesis and airways “infection”
It was originally postulated that persistent or recurrent bacterial infections in the airways are in part responsible for chronic cough and mucus hypersecretion and play a major role in the pathogenesis of COPD.33,34 However, this theory lost favor when Fletcher et al showed that there was no significant relationship between chronic cough, sputum production, and recurrent respiratory infection or decreased lung function.35 More recently, and contrary to the original description by Fletcher et al, several studies have shown chronic mucus hypersecretion and recurrent lower respiratory tract infections to be associated with an accelerated decline in lung function.36,37 Importantly, other contemporaneous studies have demonstrated the importance of microorganisms in driving exacerbations and loss of lung function.33,38,39 These more recent data suggest that bacteria have an important role to play in the pathogenesis of the disease, although the mechanism by which it occurs is far from clear.
BAL, bronchial brushing, and endotracheal lung microbiota
Studies of the lung microbiota are now just emerging. Recently, Hilty et al showed that the lungs are not sterile and bacteria are found in the lower airways.40 Using clone libraries, they interrogated the 16S rRNA gene fragments for bacterial communities in healthy control subjects (n=8), patients with asthma (n=11), and those with moderate to severe COPD (n=5). They demonstrated that bronchoalveolar lavage (BAL) fluid and bronchial brushings contained bacterial communities that were different to those found in the nasal cavity or the oropharynx.40 More specifically, they showed that there was increased representation of Proteobacteria in COPD and asthmatic airways, which was accompanied by a reduction in Bacteriodetes in COPD samples.40 This was the first study to suggest that there is a unique bacterial community in the lungs and that this may change with disease.
Erb-Downward et al evaluated the bacterial microbiota in nonsmokers and smokers with normal lung function.17 Their study evaluated mostly BAL fluid complemented by lung tissue samples obtained from patients with very severe COPD.17 They found that there was no significant difference in overall composition of the bacterial community in the lung between nonsmokers, healthy smokers, and COPD patients.17 However, they showed that there was significant heterogeneity and diversity in the bacterial microbiota across different regions of the same lung.17
Huang et al extended these findings by using a bacterial 16S PhyloChip to determine the bacterial composition of endotracheal aspirates obtained from a small number of intubated patients with severe COPD.41 Similar to the findings of Erb-Downward et al, these investigators failed to demonstrate any significant difference in the total bacterial load between these patients and control subjects.17,41 Interestingly, however, they noted two distinct and divergent bacterial populations in the COPD samples.41 One group of COPD patients demonstrated a loss of diversity in their bacterial composition, similar to what had been reported previously by Hilty et al.40 The second group, on the other hand, showed increased diversity in bacterial community composition and in particular an increase in the number of bacteria belonging to the Firmicute phylum.41 They hypothesized that progression of COPD was associated with greater bacterial diversity and increased representation of Firmicutes in the airways.41
These observations were supported by data generated in resected lung tissue specimens from patients with very severe COPD that had increased representation of Firmicutes.42 As with other research,17,42 this study showed that there was no difference in composition of the bacterial community in the lungs between smokers and nonsmokers. Finally, although the total bacterial load was much lower in lung tissue samples than in BAL fluid, neither the tissue samples nor the BAL fluid showed any significant differences in total bacterial load between COPD patients and healthy (control) subjects.40,42
Pragman et al extended these early studies by determining the lung microbiota in a small group of control subjects and in patients with moderate to severe COPD using BAL fluid samples. They found that the bacterial communities in lungs with COPD were distinct from those of normal lungs, although there was no significant difference with regard to disease severity.18 This was the first study to use BAL fluid for evaluation of the lung microbiota according to GOLD (Global initiative for chronic Obstructive Lung Disease)43 grades of severity. These data suggest that individuals with COPD, regardless of severity, have a different bacterial microbiota in their lungs compared with those who do not have COPD and that changes in the microbial communities of the lung occur very early in the disease process. Importantly, they showed that there was segregation of bacterial communities according to use or nonuse of inhaled corticosteroids or bronchodilators.18 However, because this study was cross-sectional, causality could not be ascribed.18 Together, the studies to date17–19 suggest that changes in the lung microbiota in individuals with COPD occur early in the disease process and remain relatively stable with disease progression. Finally, although there is a lack of uniformity regarding the organisms found in the lung microbiota of people with COPD and those with normal lungs, most studies have reported an increased representation of bacteria belonging to the Firmicutes phylum in lungs with COPD.
Traditionally, it has been believed that the airways below the vocal cords are sterile,17–19,40–45 and that the lungs have several built-in protective mechanisms to limit bacterial residence and growth. One important defense system is the mucociliary escalator, which traps bacteria in the lower airways and escorts them out of the lungs.46,47 Perturbations in this system can lead to persistent lung infections.48–50 Another important defense mechanism is host immune cells. Macrophages and neutrophils, for instance, can phagocytose bacteria and clear any microorganisms that evade the mucociliary transport system.47,51 The notion of sterility of the lower airways was “validated” by traditional culture methods, which failed to grow any significant bacteria in lung tissues from normal healthy individuals. However, use of more modern molecular techniques (that do not rely on traditional culture conditions) has revealed the genetic fingerprints of bacteria in the lower airways. However, some caution is required in interpretation of these data. Most molecular studies of the lung microbiota to date have focused solely on bacterial DNA. Therefore, it is possible that some of the bacteria identified by sequencing and other molecular methods may be nonviable (ie, dead organisms). To address this limitation, future studies of the microbiota in COPD should focus on bacterial RNA, similar to those performed in the lungs of patients with cystic fibrosis.52,53 Another limitation is that almost all studies have relied on one measurement in time. Thus, it is not possible to conclude with certainty that the bacteria identified are permanently resident in lung tissues. It is possible that these bacteria are transient migrants and over time will be cleared from the airways.
Bacterial microbiota in lung tissue
The first analysis of the bacterial microbiota in COPD lung tissue was performed by Sze et al,42 and largely confirmed the previous findings of Hilty et al.41 Both these research groups showed that there was a significant difference in composition of the bacterial community between COPD and “normal” (control) lungs.42 Notably, they found that the total number of bacteria in lung tissue, which has a density of 10–100 bacterial cells per 1,000 human cells, was relatively small compared with that of the gut microbiota.42 However, even at this low concentration, using two separate techniques (terminal restriction length polymorphism analysis and pyrotag sequencing), Sze et al found that the lungs of patients with very severe COPD contained a different community of bacteria to those of controls or patients with cystic fibrosis.42 Using indicator species analysis, they noted that these differences were largely driven by bacteria belonging to either the Proteobacteria or the Firmicutes phylum,42 which is consistent with what was previously reported by Hilty et al40 and Huang et al.41 However, by using lung tissue samples rather than bronchoscopic specimens (which are prone to upper airway contamination), Sze et al42 provided the first evidence that lungs of COPD patients harbored a distinct microbiota. In a separate study done in cystic fibrosis, Goddard et al showed that the microbial diversity within the upper airways was significantly greater than that of the lower airways in explanted lung tissue.54 These data suggest that microbiota studies based on sputum and BAL may have overestimated bacterial diversity as well as bacterial density in the lower airways and parenchymal tissue.
Initial location of the bacterial lung microbiota
To date, most studies of the bacterial microbiota in the lung have evaluated BAL samples. However, because the bronchoscope has to traverse through the upper airways, these data may be confounded as a result of contamination by organisms from the mouth or nose.44,55 To overcome this limitation, Charlson et al implemented several strict quality measures in their bronchoscopic techniques, including rinsing of the mouth with an antiseptic solution prior to bronchoscopy, restraints on suctioning through the bronchoscope in the upper airways, and discarding of the initial BAL fluid recovered.55 Similar to previous studies, which did not implement these stringent quality measures for bronchoscopy, they found that the overall bacterial load was higher in the BAL samples when compared with the negative controls55 and that there was good concordance between bacterial communities across the samples.55
Although the source of the lung microbiota is not fully known in COPD, based on studies in other lung conditions (eg, cystic fibrosis), the most likely source is the upper airway.56,57 This notion is supported in part by a recent study reported by Segal et al.44 They found that some healthy individuals carried organisms in BAL fluid that were commonly observed in the supraglottic region, whereas other individuals demonstrated unique organisms in BAL fluid that were not found in the upper airways.44 Interestingly, individuals in whom there was substantial overlap in the microbiota between BAL fluid and the upper airway demonstrated increased lung inflammation, characterized by increased lymphocytes and neutrophils in BAL fluid, compared with those whose BAL microbiota was distinct from that of the upper airways. These results suggest that transmigration of bacteria from the supraglottic region into the lungs may elicit a host inflammatory response and raise the possibility that bacteria from the mouth may alter the normal lung microbiota, contributing to “disease”. However, it is also possible that these data may have been confounded by upper airway contamination of the bronchoscope during the procedure.
Can the oral bacterial microbiota play a role in COPD?
Few studies have evaluated possible changes in the bacterial communities of the oral cavity of smokers compared with nonsmokers.19,58 Charlson et al found that there was indeed a difference in the oral microbiota between smokers and nonsmokers, most notably in the Firmicutes phylum.58 This was supported by Morris et al, who also showed using a much larger sample size that there were detectable differences in the oral microbiota between smokers and nonsmokers. Both studies found differences in the representation of Neisseria species,19,58 and showed that many bacteria in the oral cavity can also be found in the lungs. However, some bacteria, such as Enterobacteriaceae, Haemophilus, Methylobacterium, and Ralstonia, which are found throughout the airways, are enriched in the lungs compared with the oral cavity.19 The substantial overlap in the microbiota between the oral cavity and lungs may be related to microaspiration.45 It is well known that nearly all individuals microaspirate during sleep.45 However, aspirated bacteria are cleared by an intact mucociliary clearance system, which prevents infections in the lung.59 In individuals with COPD, there is an impairment of this mucociliary clearance system.60 Deregulation of this defense mechanism could lead to mucus hypersecretion, pooling of mucus, and mucus plugging in the airways,61 entrapping the aspirated bacteria in the lungs and causing them to acclimate and grow in this new ecosystem. This process may also elicit a local immune response, contributing to the persistent lung inflammation observed in COPD airways, even following smoking cessation. It is possible that these “aspirated” bacteria may stimulate formation of tertiary lymphoid follicles, which are prominent in the small airways of patients with severe or very severe COPD. The impaired mucociliary clearance and cilia62 in COPD lungs may also permit entry and growth of noncommensal bacterial pathogens in the lung, causing exacerbations and acute worsening of symptoms. Persistence of noncommensal organisms, coupled with the ongoing inflammatory response, may shift the lung microbiota in COPD.41,42 This hypothesis will needed to be tested in future studies.
Exacerbations of COPD
Another important feature of COPD is exacerbations, which are characterized by acute worsening of symptoms (usually cough and shortness of breath) and precipitate intensification of pharmacologic treatment and/or hospitalization.38 Most exacerbations are thought to be triggered by acute bacterial32,63 or viral38 infection. Consistent with this notion, treatment with antibiotics for 7–10 days during an acute COPD exacerbation reduces the risk of treatment failure and in-hospital mortality, providing strong indirect evidence for the importance of bacterial infection during exacerbations.64 Recently, there has been a growing interest in the potential role of bacteria during periods of clinical stability (nonexacerbation periods). Some have postulated that lungs with COPD persistently harbor “pathogenic” bacteria in the lower airways and that their gradual “overgrowth” beyond a certain threshold causes exacerbations. This hypothesis is partly supported by data from a recent randomized controlled study of azithromycin, a macrolide antibiotic that binds with the 50S subunit in bacteria and inhibits bacterial growth. This study showed that 12 months of daily prophylactic treatment with azithromycin significantly improved health status in patients with COPD and, most importantly, reduced the risk of exacerbations by 25% above and beyond standard pharmacologic treatment.65 Although azithromycin has anti-inflammatory activities (beyond antimicrobial ones), these data strongly support the importance of the microbiota in the pathogenesis of COPD.
Microbiota and exacerbations of COPD
Exacerbations are a major source of morbidity and mortality in patients with COPD.38 To date, most studies that have evaluated the role of the bacterial microbiota in exacerbations have been performed in patients with cystic fibrosis,66–68 with few studies having investigated the role of the lung microbiota during exacerbations of COPD.41,69 An early report by Fodor et al indicated that there was no change in the lung microbiota during exacerbations of COPD compared with periods of stability;67 this was supported by Tunney et al who obtained sputum samples from 40 individuals at baseline and further samples in the 24–48 hours following the first dose of an antibiotic during exacerbations.69 They found that more anaerobic bacteria could be cultivated at the end of the antibiotic treatment period than prior to antibiotic initiation.69 When 16S (small ribosomal subunit) bacterial sequencing analysis was performed, they found no significant difference in bacterial composition between the stable and exacerbation groups.69 However, using sequencing technology, they detected small changes in bacterial community structure following use of antibiotics compared with prior samples from the same patients.69 These data suggest that the bacterial community in the airways does not change by a large amount during exacerbations compared with periods of stability. This notion is consistent with the findings of Sethi et al,32 who demonstrated that exacerbations are caused by introduction of new strains of bacteria rather than by infection with new families or genera of bacteria.63 Using traditional culture-based methods, Sethi et al showed that exacerbations could be attributed to new strains of Haemophilus influenzae.32
Microbiota and inflammation in COPD
Very little research has been done on the role played by the microbiota in inflammation of the lung in COPD. One of the first relevant studies was reported by Segal et al,44 and was discussed earlier in this paper. A more detailed study was performed by Larsen et al,70 who determined the cytokine profile of dendritic cells in response to exposure to various types of bacteria that had been shown previously to be associated with asthma or COPD.40 They found that bacteria identified as likely commensal organisms induced muted interleukin (IL)-23, IL-12p70, and IL-10 expression, whereas bacteria identified as potential pathogens (eg, Haemophilus spp. and Moraxella spp.) induced a more robust cytokine response.70 They also went on to show that coexposure of Prevotella spp., identified as commensal organisms, reduced the dendritic IL-12p70 response related to H. influenzae.70 Others have evaluated Lactobacillus and have found that its presence reduces airway inflammation in mouse models of asthma.71,72 Together, these data suggest that commensal organisms may regulate the inflammatory response of the lungs induced by possible pathogenic organisms and protect the lungs against inflammatory and oxidative stress. This hypothesis will require validation in in vivo and in vitro studies.
Our knowledge so far about the lung microbiota in COPD
Overall, our understanding of the bacterial lung microbiota in COPD is still in its infancy. Despite the excitement about the lung microbiota, the data are inconsistent and poorly reproducible across studies. For example, although the first few studies to investigate COPD (mostly using BAL or sputum samples) reported that bacterial diversity decreased with increasing severity of disease,17,40 subsequent studies using lung tissue samples have failed to show significant differences in bacterial diversity.42 One possible explanation for this discrepancy is that tissue samples contain mostly parenchyma (mixed with airways and blood vessels) while BAL and sputum mostly sample the airways (Table 1). This suggests that the bacterial microbiota in the airways is different from that in the alveolar tissue, giving rise to differential microniches across different compartments of the lung. It should also be noted that the concept of reduced diversity with disease progression has not been consistently replicated, even in studies using BAL samples.18 In contrast with earlier reports, one recent study showed that diversity paradoxically increased in severe COPD compared with controls.18
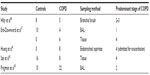  | Table 1 Breakdown of different bacterial microbiota studies in COPD |
One common finding so far from the COPD studies is that there is increased expression of bacteria in the Firmicute phylum in moderate, severe, and even very severe disease,18,40–42 with one notable dissenting study.17 Erb-Downward et al found that the predominant bacteria in lung tissue from patients with very severe disease were those belonging to the Proteobacteria phylum.17 This finding is similar to that reported by Hilty et al.40 Huang et al provided a plausible explanation to reconcile these differences. They speculated that there are two types of bacterial microbiota related to COPD, one dominated by Proteobacteria and another dominated by Firmicutes, and that the latter may be associated with increased diversity.41 Further studies will be needed to investigate and resolve this controversy and determine the role of the lung microbiota in progression of COPD.
Although cystic fibrosis and bronchiectasis are both chronic airway diseases, they are likely to harbor a different lung microbiota than that in the COPD lung. For example, in COPD, some (but not all) studies have shown reduced bacterial diversity with disease progression; however, no study has shown that this leads to an outgrowth of a single organism, unlike in cystic fibrosis where loss of bacterial diversity leads to bacterial outgrowth (usually Pseudomonas). Moreover, the bacterial load in COPD is far less than that observed in cystic fibrosis or idiopathic bronchiectasis, which makes cross-comparisons difficult.
Important findings in the GI microbiota
An interesting emerging concept is that the gut microbiota modulates the body’s entire immune response and that disturbances in the gut microbiota can lead to disease in extraintestinal organs. A large body of literature has been published on the role of the gut microbiota in health and disease.73–77 These investigations have provided new insights regarding a variety of bowel diseases and yielded novel therapeutic targets. Good examples are the use of stool transplantation to treat refractory cases of pseudomembranous colitis secondary to C. difficile infection76 and antibiotic treatment for severe malnutrition.76 A seminal study in this field was published by Turnbaugh et al in 2006.73 In that study, they closely examined the gut flora of genetically engineered obese and lean mice and found that their bacterial microbiotas were different.73 They noted that obese mice had an increased capacity to harvest and store dietary energy. Importantly, when the gut microbiota of these obese mice was transplanted into germ-free mice, the recipients gained a significant amount of weight in the form of fat. In contrast, when these mice were transplanted with gut flora from lean mice, no significant weight gain was observed.73
Cho et al investigated whether exposure to subtherapeutic doses of antibiotics early in life could modify body weight.75 They found that subtherapeutic doses of antibiotics modified the bacterial microbiota in the gut, which in turn influenced short-chain fatty acid synthesis by modulating prokaryotic genes for butyryl coenzyme A transferase.75 They also found that this antibiotic regimen modified the processing of fatty acids and lipids in mice by upregulating genes involved in both lipogenesis and triglyceride synthesis.75 Trehan et al extended this observation by showing a beneficial effect of antibiotic therapy on body weight.78 In their study, they treated individuals suffering from severe malnutrition with antibiotics to modify their gut microbiota77,78 and found that the group treated with antibiotics gained the most weight.78
Together, these studies indicate that modulating the gut microbiota not only impacts the GI tract but also influences extraintestinal organs (eg, adipose tissue), indicating the importance of the gut microbiota in the overall health of individuals. To date, there are no similar data for the lung microbiota. Study of the lung microbiota is difficult because of its low total bacterial load, possible contamination of oral or nasal origin, and barriers to access.
The gut microbiota has been shown to have effects not just in obesity but also in diabetes79–81 and atherosclerosis.82,83 Some bacteria in the GI tract may be modulated by serum female sex hormone levels,81,84 which may have an impact on the brain and behavior.85,86 Therefore, a reasonable question to ask would be: can changes in the microbiota of the gut also affect the response of the lung to different stimuli?
Gut-lung axis
One of the first studies to show that the response of the lung could be affected by the gut bacterial microbiota was reported by Ichinohe et al, who investigated how changes in the GI microbiota could affect lung inflammation during acute influenza infection.15 The first observation they made was that treatment with antibiotics reduced production of influenza virus-specific antibodies and attenuated CD4 T-cell responses (T-cell isolation followed by restimulation with virus).15 They also found that animals treated with oral antibiotics had smaller mediastinal lymph nodes, less cellularity, and fewer dendritic cells than those not treated with antibiotics. Dendritic cells in the antibiotic-treated group demonstrated deficiencies in the regulatory T-cell response.15 Interestingly, Ichinohe et al found that rectal stimulation of Toll-like receptors restored the immune response to viral infection.15
Russell et al performed a different set of experiments involving very young mouse pups and found that low-dose oral antibiotics altered the GI flora, which in turn upregulated the mouse’s immune response to an allergen challenge with ovalbumin.87 After the animals were treated with antibiotics, they mounted greater antigen-specific immunoglobulin E responses, showed increased eosinophilia in BAL fluid, and demonstrated increased airway hyperresponsiveness when challenged with ovalbumin.87 This study also showed that treatment with low-dose vancomycin in young mice severely reduced the number of regulatory T-cells in the gut.87 These data suggest that alterations in the GI microbiota early in life can modify an individual’s risk of asthma. Some have suggested that it is the antibiotics (and not the alterations in the gut microbiota per se) that are responsible for modulating the risk of asthma. However, the data would argue against this hypothesis. A recent study showed that babies delivered vaginally harbored a microbiota (in the skin, oral cavity, nasopharynx, and GI tract) that was predominated by Lactobacillus and very similar to that in the mother’s vaginal tract. In contrast, babies born via cesarean section harbored a microbiota that was very similar to that found on the mother’s skin, where Staphylococcus and Acinetobacter were much more common colonizers.88 Epidemiologic studies have shown that babies born by cesarean section have a significantly higher risk of asthma than babies delivered vaginally.89 Russell et al showed that antibiotic therapy enhanced the risk of asthma in young but not adult mice.87 Together, these data suggest that the microbiota is responsible for modulating the risk of asthma.
There is growing evidence that smoking and COPD can have an effect on bacteria in the GI tract.90–92 Further, smoking is an important risk factor for certain inflammatory bowel diseases, such as ulcerative colitis, which has been associated with perturbations of the GI microbiota.93,94 Ekbom et al showed that there was an increased risk of COPD in patients with ulcerative colitis or Crohn’s disease.90 More recent studies, such as those by Biedermann et al, have clearly shown that smoking alters the gut bacterial microbiota and that smoking cessation increases the diversity of this microbiota.92 These observations suggest that cigarette smoke can directly impact the immune response in the lung and indirectly modulate the gut microbiota. One possibility for the latter is reducing “immune tolerance” conferred by the gut microbiota. Cigarette smoke may downregulate certain commensal organisms in the gut, which in turn may amplify immune responses elsewhere in the body. This could then enhance the immune response in the lungs. Future studies are needed to address the issue concerning how this assault by cigarette smoke on the bacterial microbiota affects the host immune system in individuals with COPD (Figure 1).
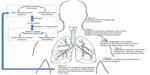  | Figure 1 Potential role of the bacterial microbiota in the pathogenesis of COPD. |
Future research directions
There are presently no longitudinal data on the microbiota in COPD. Further, human studies have been hampered by their inability to explain the large variation in bacterial communities between individual samples and across subjects. Although molecular techniques have identified a novel lung microbiota in COPD, there is little information on whether alterations in the lung microbiota induce changes in the host inflammatory response. Future studies will need to address how the COPD lung microbiota affects innate and adaptive immunity and in turn drives the formation of tertiary lymphoid follicles. Although there is strong evidence that chronic prophylactic treatment with azithromycin reduces the risk of exacerbation in COPD, it is not certain whether this effect is due to the drug’s antimicrobial or anti-inflammatory properties. Although studies of the bacterial microbiota in mice have yielded some fascinating and interesting data, their transferability to humans is not certain. At the moment, one potential role for the bacterial microbiota in the lung may be as a biomarker for worsening disease. The one generally consistent observation across the studies of the COPD microbiome is a loss of bacterial diversity with increasing severity of disease. However, the clinical relevance of this observation is not known. Despite these and other limitations, the success of studies of the gut microbiota in effecting new therapies for GI conditions such as pseudomembranous colitis provide hope that well designed and executed lung microbiota studies in humans and animals can generate new knowledge that can be translated into novel therapies and thereby reduce the growing worldwide burden of COPD.
Disclosure
The authors report no conflicts of interest in this work.
References
Torsvik V, Ovreas L. Microbial diversity and function in soil: from genes to ecosystems. Curr Opin Microbiol. 2002;5(3):240–245. | |
Relman DA. New technologies, human-microbe interactions, and the search for previously unrecognized pathogens. J Infect Dis. 2002; 186 Suppl 2:S254–S258. | |
Shanahan F. The host-microbe interface within the gut. Best Pract Res Clin Gastroenterol. 2002;16(6):915–931. | |
Rodriguez-Brito B, Li L, Wegley L, et al. Viral and microbial community dynamics in four aquatic environments. ISME J. 2010;4(6):739–751. | |
Dollive S, Chen YY, Grunberg S, et al. Fungi of the murine gut: episodic variation and proliferation during antibiotic treatment. PLoS One. 2013;8(8):e71806. | |
Turnbaugh PJ, Ley RE, Hamady M, Fraser-Liggett CM, Knight R, Gordon JI. The human microbiome project. Nature. 2007;449(7164):804–810. | |
Li K, Bihan M, Yooseph S, Methè BA. Analyses of the microbial diversity across the human microbiome. PLoS One. 2012;7(6):e32118. | |
Hoffmann C, Dollive S, Grunberg S, et al. Archaea and fungi of the human gut microbiome: correlations with diet and bacterial residents. PLoS One. 2013;8(6):e66019. | |
Scupham AJ, Presley LL, Wei B, et al. Abundant and diverse fungal microbiota in the murine intestine. Appl Environ Microbiol. 2006;72(1):793–801. | |
Reyes A, Haynes M, Hanson N, et al. Viruses in the faecal microbiota of monozygotic twins and their mothers. Nature. 2010;466(7304):334–338. | |
de Dombal FT, Burch PR, Watkinson G. Aetiology of ulcerative colitis. Gut. 1969;10(4):270–277. | |
Macfarlane GT, Macfarlane S. Human colonic microbiota: ecology, physiology and metabolic potential of intestinal bacteria. Scand J Gastroenterol Suppl. 1997;222:3–9. | |
Nesbitt B. On the presence of cholin and neurin in the intestinal canal during its complete obstruction : a research on autointoxication. J Exp Med. 1899;4(1):1–18. | |
Khoury KA, Floch MH, Herskovic T. Effects of neomycin and penicillin administration on mucosal proliferation of the mouse small intestine. With morphological and functional correlations. J Exp Med. 1969;129(5):1063–1078. | |
Ichinohe T, Pang IK, Kumamoto Y, et al. Microbiota regulates immune defense against respiratory tract influenza A virus infection. Proc Natl Acad Sci U S A. 2011;108(13):5354–5359. | |
Ferreira RB, Gill N, Willing BP, et al. The intestinal microbiota plays a role in Salmonella-induced colitis independent of pathogen colonization. PLoS One. 2011;6(5):e20338. | |
Erb-Downward JR, Thompson DL, Han MK, et al. Analysis of the lung microbiome in the “healthy” smoker and in COPD. PLoS One. 2011;6(2):e16384. | |
Pragman AA, Kim HB, Reilly CS, Wendt C, Isaacson RE. The lung microbiome in moderate and severe chronic obstructive pulmonary disease. PLoS One. 2012;7(10):1–10. | |
Morris A, Beck JM, Schloss PD, et al. Comparison of the respiratory microbiome in healthy non-smokers and smokers. Am J Respir Crit Care Med. 2013;187(10):1067–1075. | |
Bartlett JG, Chang TW, Gurwith M, Gorbach SL, Onderdonk AB. Antibiotic-associated pseudomembranous colitis due to toxin-producing clostridia. N Engl J Med. 1978;298(10):531–534. | |
Knights D, Lassen KG, Xavier RJ. Advances in inflammatory bowel disease pathogenesis: linking host genetics and the microbiome. Gut. 2013;62(10):1505–1510. | |
Le Chatelier E, Nielsen T, Qin J, et al. Richness of human gut microbiome correlates with metabolic markers. Nature. 2013;500(7464):541–546. | |
Mathers CD, Loncar D. Projections of global mortality and burden of disease from 2002 to 2030. PLoS Med. 2006;3(11):e442. | |
World Health Organization. The global burden of disease: 2004 update. Geneva, Switzerland: World Health Organization Press; 2008. Available from: http://www.who.int/healthinfo/global_burden_disease/2004_report_update/en/. Accessed December 28, 2013. | |
Fletcher C, Peto P. The natural history of chronic airflow obstruction. BMJ. 1977;1(6077):1645–1648. | |
Han MK, Agusti A, Calverley PM, et al. Chronic obstructive pulmonary disease phenotypes: the future of COPD. Am J Respir Crit Care Med. 2010;182(5):598–604. | |
Franklin W, Lowell FC, Michelson AL, Schiller IW. Chronic obstructive pulmonary emphysema; a disease of smokers. Ann Intern Med. 1956;45(2):268–274. | |
Hogg JC, Chu F, Utokaparch S, et al. The nature of small-airway obstruction in chronic obstructive pulmonary disease. N Engl J Med. 2004;350(26):2645–2653. | |
Di Stefano A, Turato G, Maestrelli P, et al. Airflow limitation in chronic bronchitis is associated with T-lymphocyte and macrophage infiltration of the bronchial mucosa. Am J Respir Crit Care Med. 1996;153(2):629–632. | |
Saetta M, Di Stefano A, Maestrelli P, et al. Activated T-lymphocytes and macrophages in bronchial mucosa of subjects with chronic bronchitis. Am Rev Respir Dis. 1993;147(2):301–306. | |
Low TB, Greene CM, O’Neill SJ, McElvaney NG. Quantification and evaluation of the role of antielastin autoantibodies in the emphysematous lung. Pulm Med. 2011;2011:826160. | |
Sethi S, Evans N, Grant BJ, Murphy TF. New strains of bacteria and exacerbations of chronic obstructive pulmonary disease. N Engl J Med. 2002;347(7):465–471. | |
Anthonisen NR. The British hypothesis revisited. Eur Respir J. 2004;23(5):657–658. | |
Fletcher CM. Chronic bronchitis – its prevalence, nature, and pathogenesis. Am Rev Respir Dis. 1959;80(4):483–494. | |
Fletcher C, Peto R, Tinker C, Speizer F. The Natural History of Chronic Bronchitis and Emphysema. New York, NY, USA: Oxford University Press; 1976. | |
Vestbo J, Prescott E, Lange P. Association of chronic mucus hypersecretion with FEV1 decline and chronic obstructive pulmonary disease morbidity. Copenhagen City Heart Study Group. Am J Respir Crit Care Med. 1996;153(5):1530–1535. | |
Kanner RE, Anthonisen NR, Connett JE; Lung Health Study Research Group. Lower respiratory illnesses promote FEV(1) decline in current smokers but not ex-smokers with mild chronic obstructive pulmonary disease: results from the Lung Health Study. Am J Respir Crit Care Med. 2001;164(3):358–364. | |
Hurst JR, Vestbo J, Anzueto A, et al. Susceptibility to exacerbation in chronic obstructive pulmonary disease. N Engl J Med. 2010;363(12):1128–1138. | |
Donaldson GC, Seemungal TA, Bhowmik A, Wedzicha JA. Relationship between exacerbation frequency and lung function decline in chronic obstructive pulmonary disease. Thorax. 2002;57(10):847–852. | |
Hilty M, Burke C, Pedro H, et al. Disordered microbial communities in asthmatic airways. PLoS One. 2010;5(1):e8578. | |
Huang YJ, Kim E, Cox MJ, et al. A persistent and diverse airway microbiota present during chronic obstructive pulmonary disease exacerbations. OMICS. 2010;14(1):9–59. | |
Sze MA, Dimitriu PA, Hayashi S, et al. The lung tissue microbiome in chronic obstructive pulmonary disease. Am J Respir Crit Care Med. 2012;185(10):1073–1080. | |
Rabe KF, Hurd S, Anzueto A, et al; Global Initiative for Chronic Obstructive Lung Disease. Global strategy for the diagnosis, management, and prevention of chronic obstructive pulmonary disease: GOLD executive summary. Am J Respir Crit Care Med. 2007;176(6):532–555 | |
Segal LN, Alekseyenko AV, Clemente JC, et al. Enrichment of lung microbiome with supraglottic taxa is associated with increased pulmonary inflammation. Microbiome. 2013;1:1–12. | |
Gleeson K, Eggli DF, Maxwell SL. Quantitative aspiration during sleep in normal subjects. Chest. 1997;111(5):1266–1272. | |
Satir P, Sleigh MA. The physiology of cilia and mucociliary interactions. Annu Rev Physiol. 1990;52:137–155. | |
Sibille Y, Reynolds HY. Macrophages and polymorphonuclear neutrophils in lung defense and injury. Am Rev Respir Dis. 1990;141(2):471–501. | |
Smith CM, Kulkarni H, Radhakrishnan P, et al. Ciliary dyskinesia is an early feature of respiratory syncytial virus infection. Eur Respir J. March 21, 2013. [Epub ahead of print.] | |
Sagel SD, Davis SD, Campisi P, Dell SD. Update of respiratory tract disease in children with primary ciliary dyskinesia. Proc Am Thorac Soc. 2011;8(5):438–443. | |
Tambascio J, Lisboa RM, Passarelli Rde C, Martinez JA, Gastaldi AC. Adhesiveness and purulence of respiratory secretions: implications for mucociliary transport in patients with bronchiectasis. J Bras Pneumol. 2010;36(5):545–553. | |
Hogg JC, van Eeden S. Pulmonary and systemic response to atmospheric pollution. Respirology. 2009;14(3):336–346. | |
Rogers GB, Carroll MP, Serisier DJ, et al. Bacterial activity in cystic fibrosis lung infections. Respir Res. 2005;6:49. | |
Rogers GB, van der Gast CJ, Cuthbertson L, et al. Clinical measures of disease in adult non-CF bronchiectasis correlate with airway microbiota composition. Thorax. 2013;68(8):731–737. | |
Goddard AF, Staudinger BJ, Dowd SE, et al. Direct sampling of cystic fibrosis lungs indicates that DNA-based analyses of upper-airway specimens can misrepresent lung microbiota. Proc Natl Acad Sci U S A. 2012;109(34):13769–13774. | |
Charlson ES, Bittinger K, Haas AR, et al. Topographical continuity of bacterial populations in the healthy human respiratory tract. Am J Respir Crit Care Med. 2011;184(8):957–963. | |
Ramakrishnan VR, Ferril GR, Suh JD, Woodson T, Green TJ, Kingdom TT. Upper and lower airways associations in patients with chronic rhinosinusitis and bronchiectasis. Int Forum Allergy Rhinol. 2013;3(11):921–927. | |
Berkhout MC, Rijntjes E, El Bouazzaoui LH, Fokkens WJ, Brimicombe RW, Heijerman HG. Importance of bacteriology in upper airways of patients with cystic fibrosis. J Cyst Fibros. 2013;12(5):525–529. | |
Charlson ES, Chen J, Custers-Allen R, et al. Disordered microbial communities in the upper respiratory tract of cigarette smokers. PLoS One. 2010;5(12):e15216. | |
Lourenço RV, Klimek MF, Borowski CJ. Deposition and clearance of 2 micron particles in the tracheobronchial tree of normal subjects – smokers and nonsmokers. J Clin Invest. 1971;50(7):1411–1420. | |
Smaldone GC, Foster WM, O’Riordan TG, Messina MS, Perry RJ, Langenback EG. Regional impairment of mucociliary clearance in chronic obstructive pulmonary disease. Chest. 1993;103(5):1390–1396. | |
Fanta CH. Clinical aspects of mucus and mucous plugging in asthma. J Asthma. 1985;22(6):295–301. | |
Leopold PL, O’Mahony MJ, Lian XJ, Tilley AE, Harvey BG, Crystal RG. Smoking is associated with shortened airway cilia. PLoS One. 2009;4(12):e8157. | |
Sethi S. Bacterial infection and the pathogenesis of COPD. Chest. 2000;117(5 Suppl 1):286S–291S. | |
Quon BS, Gan WQ, Sin DD. Contemporary management of acute exacerbations of COPD: a systematic review and metaanalysis. Chest. 2008;133(3):756–766. | |
Albert RK, Connett J, Bailey WC, et al. Azithromycin for prevention of exacerbations of COPD. N Engl J Med. 2011;365(8):689–698. | |
Reid DW, Latham R, Lamont IL, Camara M, Roddam LF. Molecular analysis of changes in Pseudomonas aeruginosa load during treatment of a pulmonary exacerbation in cystic fibrosis. J Cyst Fibros. 2013;12(6):688–699. | |
Fodor AA, Klem ER, Gilpin DF, et al. The adult cystic fibrosis airway microbiota is stable over time and infection type, and highly resilient to antibiotic treatment of exacerbations. PLoS One. 2012;7(9):e45001. | |
Sibley CD, Parkins MD, Rabin HR, Surette MG. The relevance of the polymicrobial nature of airway infection in the acute and chronic management of patients with cystic fibrosis. Curr Opin Investig Drugs. 2009;10(8):787–794. | |
Tunney MM, Einarsson GG, Wei L, et al. Lung microbiota and bacterial abundance in patients with bronchiectasis when clinically stable and during exacerbation. Am J Respir Crit Care Med. 2013;187(10):1118–1126. | |
Larsen JM, Steen-Jensen DB, Laursen JM, et al. Divergent pro-inflammatory profile of human dendritic cells in response to commensal and pathogenic bacteria associated with the airway microbiota. PLoS One. 2012;7(2):e31976. | |
Forsythe P, Inman MD, Bienenstock J. Oral treatment with live Lactobacillus reuteri inhibits the allergic airway response in mice. Am J Respir Crit Care Med. 2007;175(6):561–569. | |
Karimi K, Inman MD, Bienenstock J, Forsythe P. Lactobacillus reuteri-induced regulatory T cells protect against an allergic airway response in mice. Am J Respir Crit Care Med. 2009;179(3):186–193. | |
Turnbaugh PJ, Ley RE, Mahowald MA, Magrini V, Mardis ER, Gordon JI. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature. 2006;444(7122):1027–1031. | |
Turnbaugh PJ, Hamady M, Yatsunenko T, et al. A core gut microbiome in obese and lean twins. Nature. 2009;457(7228):480–484. | |
Cho I, Yamanishi S, Cox L, et al. Antibiotics in early life alter the murine colonic microbiome and adiposity. Nature. 2012;488(7413):621–626. | |
Rohlke F, Stollman N. Fecal microbiota transplantation in relapsing Clostridium difficile infection. Therap Adv Gastroenterol. 2012;5(6):403–420. | |
Smith MI, Yatsunenko T, Manary MJ, et al. Gut microbiomes of Malawian twin pairs discordant for kwashiorkor. Science. 2013; 339(6119):548–554. | |
Trehan I, Goldbach HS, LaGrone LN, et al. Antibiotics as part of the management of severe acute malnutrition. N Engl J Med. 2013;368(5):425–435. | |
Graessler J, Qin Y, Zhong H, et al. Metagenomic sequencing of the human gut microbiome before and after bariatric surgery in obese patients with type 2 diabetes: correlation with inflammatory and metabolic parameters. Pharmacogenomics J. 2013;13(6):514–522. | |
Qin J, Li Y, Cai Z, et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature. 2012;490(7418):55–60. | |
Markle JG, Frank DN, Mortin-Toth S, et al. Sex differences in the gut microbiome drive hormone-dependent regulation of autoimmunity. Science. 2013;339(6123):1084–1088. | |
Karlsson FH, Fåk F, Nookaew I, et al. Symptomatic atherosclerosis is associated with an altered gut metagenome. Nat Commun. 2012;3:1245. | |
Koren O, Spor A, Felin J, et al. Human oral, gut, and plaque microbiota in patients with atherosclerosis. Proc Natl Acad Sci U S A. 2011; 108 Suppl 1:4592–4598. | |
Flores R, Shi J, Fuhrman B, et al. Fecal microbial determinants of fecal and systemic estrogens and estrogen metabolites: a cross-sectional study. J Transl Med. 2012;10:253. | |
El Aidy S, Kunze W, Bienenstock J, Kleerebezem M. The microbiota and the gut-brain axis: insights from the temporal and spatial mucosal alterations during colonisation of the germfree mouse intestine. Benef Microbes. 2012;3(4):251–259. | |
Ohland CL, Kish L, Bell H, et al. Effects of Lactobacillus helveticus on murine behavior are dependent on diet and genotype and correlate with alterations in the gut microbiome. Psychoneuroendocrinology. 2013;38(9):1738–1747. | |
Russell SL, Gold MJ, Hartmann M, et al. Early life antibiotic-driven changes in microbiota enhance susceptibility to allergic asthma. EMBO Rep. 2012;13(5):440–447. | |
Dominguez-Bello MG, Costello EK, Contreras M, et al. Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc Natl Acad Sci U S A. 2010;107(26):11971–11975. | |
Renz-Polster H, David MR, Buist AS, et al. Caesarean section delivery and the risk of allergic disorders in childhood. Clin Exp Allergy. 2005;35(11):1466–1472. | |
Ekbom A, Brandt L, Granath F, Löfdahl CG, Egesten A. Increased risk of both ulcerative colitis and Crohn’s disease in a population suffering from COPD. Lung. 2008;186(3):167–172. | |
Jess T, Loftus EV Jr, Harmsen WS, et al. Survival and cause specific mortality in patients with inflammatory bowel disease: a long term outcome study in Olmsted County, Minnesota, 1940–2004. Gut. 2006;55(9):1248–1254. | |
Biedermann L, Zeitz J, Mwinyi J, et al. Smoking cessation induces profound changes in the composition of the intestinal microbiota in humans. PLoS One. 2013;8(3):e59260. | |
Nos P, Domènech E. Management of Crohn’s disease in smokers: is an alternative approach necessary? World J Gastroenterol. 2011;17(31):3567–3574. | |
Mahid SS, Minor KS, Soto RE, Hornung CA, Galandiuk S. Smoking and inflammatory bowel disease: a meta-analysis. Mayo Clin Proc. 2006;81(11):1462–1471. |
 © 2014 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2014 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
