Back to Journals » Biologics: Targets and Therapy » Volume 11
Autophagy regulates the stemness of cervical cancer stem cells
Authors Yang Y, Yu L, Li J, Yuan YH, Wang XL, Yan SR, Li DS, Ding Y
Received 16 February 2017
Accepted for publication 12 April 2017
Published 21 June 2017 Volume 2017:11 Pages 71—79
DOI https://doi.org/10.2147/BTT.S134920
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Doris Benbrook
This paper has been retracted
Yi Yang, 1,2 Li Yu, 1 Jin Li, 1 Ya Hong Yuan, 1 Xiao Li Wang, 1 Shi Rong Yan, 1 Dong Sheng Li, 1 Yan Ding 1
1Hubei Key Laboratory of Embryonic Stem Cell Research, 2Reproductive Center, Taihe Hospital, Hubei University of Medicine, Shiyan, People’s Republic of China
Abstract: Cancer stem cells (CSCs) are a rare population of multipotent cells with the capacity to self-renew. It has been reported that there are CSCs in cervical cancer cells. Pluripotency-associated (PA) transcription factors such as Oct4, Sox2, Nanog and CD44 have been used to isolate CSCs subpopulations. In this study, we showed that autophagy plays an important role in the biological behavior of cervical cancer cells. The expression of the autophagy protein Beclin 1 and LC3B was higher in tumorspheres established from human cervical cancers cell lines (and CaSki) than in the parental adherent cells. It was also observed that the basal and starvation-induced autophagy flux was higher in tumorspheres than in the bulk population. Autophagy could regulate the expression level of PA proteins in cervical CSCs. In addition, CRISPR/Cas 9-mediated Beclin 1 knockout enhanced the malignancy of HeLa cells, leading to accumulation of PA proteins and promoted tumorsphere formation. Our findings suggest that autophagy modulates homeostasis of PA proteins, and Beclin 1 is critical for CSC maintenance and tumor development in nude mice. This demonstrates that a prosurvival autophagic pathway is critical for CSC maintenance.
Keywords: cervical cancer, autophagy, cancer stem cell, LC3, Oct4
Introduction
Cancer stem cells (CSCs) are a subpopulation of cells sharing some similar characteristics with embryonic stem cells (ESCs) and displaying stem cell properties, including self-renewal, tumorigenesis, tumor growth and differentiation.1 CSCs have been identified in many different types of tumors, including cervical cancer.2–4 Increasing studies have reported that CSCs could promote tumorigenesis and drug resistance.5,6 Some cell-surface markers, such as CD133, Oct4, Sox2, Nanog and CD44, have been popularly used to isolate CSC subpopulations. These CSCs are thought to demonstrate therapeutic resistance and play critical roles in the recurrence and metastasis of cancer.3
Homeostasis of intracellular proteins is essential for cell viability and proliferation. Different systems of intracellular protein quality control maintain homeostasis and cellular viability.7 These processes are largely modulated by autophagy.8 Autophagy is a conserved lysosomal degradation pathway, which involves the degradation of cytoplasmic components to maintain cellular function.9 It is generally induced by extracellular stresses such as starvation. Dysfunction of autophagy is related to a variety of diseases and cancers.10
The physiologic function of autophagy is well known to maintain homeostasis in mammalian cells. It has been reported that autophagy regulates homeostasis of pluripotency-associated (PA) proteins in human embryonic stem cells (hESCs). The pluripotency of hESCs is largely maintained by orchestrated networks of regulatory PA transcription factors such as Oct4, Sox2 and Nanog.11 Quantitative imbalances of these proteins can lead to the loss of pluripotency or differentiation of ESCs.12–14 That is to say, autophagy could maintain the pluripotency of hESCs by regulating the PA proteins. Autophagy has been highly observed in CSCs.15 However, the role of autophagy in CSC pluripotency maintenance is still poorly understood. Understanding the mechanisms underlying the maintenance of CSCs may be critical for the development of new therapeutic approaches that may be able to target this specific population and enhance the efficacy of chemotherapy in cervical cancer.
Materials and methods
Cell culture and chemical treatments
HeLa (ATCC, BH0999) and CaSki cells (ATCC, ATCC-1398) were cultured in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% fetal bovine serum, as in our previous study.16
Tumorsphere formation
Cervical cancer cells were plated in low attachment culture dishes at a density of 500 cells/ml in DMEM/F12 basic medium, supplemented with 1% B27 (1:50; Invitrogen), 20 ng/ml basic fibroblast growth factor (BD Biosciences), 20 ng/ml epidermal growth factor (BD Biosciences) and 0.4% bovine serum albumin (Sigma-Aldrich), as per the study by Cho et al.17 To propagate tumorspheres in vitro, primary tumorspheres were enzymatically dissociated with 0.05% trypsin for 3 min at 37°C to obtain a single-cell suspension, and then cultured in suspension to produce the next generation of tumorspheres.
Chemical treatments of cervical cancer cells
Cervical cancer adherent and stem cells were stabilized in corresponding media. For inhibition of autophagy, cervical cancer cells were cultured in media supplemented with 100 nM rapamycin (Sigma-Aldrich) for 48 h. For starvation, cells were incubated in HBSS buffer for different times. For inhibition of mTOR signaling, cervical cancer cells were cultured in media supplemented with 100 nM rapamycin (A.G. Scientific, R-1018).
CRISPR vector and single guide RNA (sgRNA) cloning
The pGK1.2 vector (Genloci, Beijing, People’s Republic of China) expressing Cas 9 and containing a cloning site for the sgRNA sequence was digested with BbsI (NEB). Beclin 1 sgRNAs were constructed online (http://crispr.mit.edu/) and then cloned into the pGK1.2 vector according to the manufacture’s instruction. The complementary oligos with two overhanging sequences CACC or AAAC at 5′ for each sgRNA were annealed to form double sequence, and finally ligated with the linearized pGK1.2. Competent cells were transformed with 2 µl of the ligated plasmid, plated with selection and single colonies were expanded prior to plasmid extraction using Maxiprep kit (Qiagen). The correct insertion of the sgRNA sequences was confirmed using Sanger sequencing.
Selection of Beclin 1 mutation-corrected HeLa clones
HeLa cells were transfected using the Amaxa cell line Nucleofector Kit V (Lonza) according to manufacturer’s guidelines. A total of 1×106 cells were transfected with 10 µg of pGK1.2 plasmid and then transfected for the second time after 24 h. In all, 3 μg/ml puromycin was used to select the positive cells. After 3 days of selection, cells were plated as single in 96-well plates and incubated at 37°C for 2 weeks before being harvested for DNA extraction. The percentages of Beclin 1 targeting were analyzed with T7 endonuclease 1 as in our previous study.16
Cell growth assay
The proliferation capability of HeLa cells was detected with RTCA DP Analyzer (ACEA). HeLa cells with or without TET1 disruption were digested with 0.05% trypsin-EDTA and plated into E-Plate 16 at a density of 1×103/well and incubated at 37°C for 96 h. The experiment was performed in triplicates.
Invasion assay
The invasion ability of HeLa and CaSki cells were analyzed with Transwell filters, as in our previous study.16 The upper surface of Transwell filters (diameter 12 mm, pore size 8 mm; Millipore, Billerica, MA, USA) was coated with Matrigel, and then 1×105 cells suspended in serum-free medium were plated into the top side of the polycarbonate. Medium with serum was used as a chemoattractant in the lower chamber. The cells were incubated at 37°C for 48 h before removing the medium from the top chamber. The nonmigratory or non-invasive cells were scraped off by a cotton swab. Cells on the lower surface were fixed in 70% ethanol for at least 10 min, then washed and stained with 0.1% crystal violet for 5–10 min. Images were collected under an inverted microscope (Olympus) and the number of migrated cells were counted. Each assay was repeated three times.
Autophagosome detection
Cells were fixed with 4% paraformaldehyde for 30 min, then washed and stained with 0.1% Acridine orange for 5–10 min. Images were collected under a fluorescence microscope (Leica).
Western blot analysis
Western blot analysis was carried out, as in our previous study.16 All experiments were performed at least three times. Representative autoradiograms are shown. The antibody dilutions were as follows: anti-LC3 rabbit mAb (Beyotime, AL221, 1:500); anti-Beclin 1 rabbit mAb (CS, 3495, 1:250); anti-phospho-mTOR rabbit mAb (CS, 5536, 1:250); anti-phospho-4EBP1 rabbit mAb (CS, 2855P, 1:250); Oct-4A rabbit mAb (CS, 2840P, 1:250); Sox2 rabbit mAb (CS, 3579P, 1:250); CD133 rabbit mAb (Sigma-Aldrich, SAB4300882, 1:250) and GAPDH mouse monoclonal antibody (Beyotime, AF0006, 1:200). Goat anti-mouse and goat anti-rabbit antibodies were obtained from Bio-Rad (Beyotime, A0258 and A0239, 1:500).
Statistical analysis
Statistical analysis of the differences between the groups was performed using Student’s t-test. P<0.05 was considered statistically significant.
Results
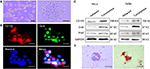
To elucidate the basal and starvation-stimulated autophagic capacities of cervical CSCs, we used an in vitro system in which primitive mammary CSCs can be propagated in culture as floating spherical colonies termed “tumorspheres”. A single-cell suspension was obtained from HeLa and CaSki cells and cultured in tumorsphere media in low adhesion plates for 7–10 days and the ability of CSCs to form tumor spheroids was monitored. A size of 60 µm was selected as cutoff for counting the number of spheroids as described previously16 (Figure 1A). Further, CSCs were characterized by the expression of stemness markers Oct-4, Sox2 and CD133 (Figure 1B and C). CSCs are known to have differentiation ability as that of stem cells. Therefore, we also measured the adipogenic differentiation ability and found lipid droplet-like cells when cultured in adipogenic induction media for about 2 weeks (Figure 1C).
Robust autophagic flux in tumorspheres is correlated with CSC/progenitor phenotype
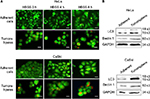
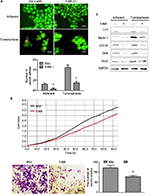
To find out whether the level of autophagy was different between general cervical cancer cells and CSC tumorspheres, we firstly tested the autophagic flux of them. Acridine orange staining indicated that the autophagic flux in CSCs in both basal and HBSS-induced conditions was higher than in the general adherent cervical cancer cells (Figure 2A). Western blot analysis of LC3 and Beclin 1 showed that the amount of LC3 II was markedly increased in cervical CSCs, and the level of Beclin 1 was also higher in the CSC group (Figure 2B). Taken together, these findings indicate that autophagosome synthesis was greater in CSCs than in the general cancer cells, both at the basal level and during HBSS treatment.
Autophagy regulates the proliferation and invasion of cervical CSCs
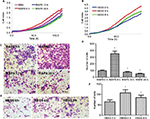
HeLa CSCs treated with rapamycin or HBSS with different times were used to detect the proliferation and invasion ability. xCELLigence RTCA DP Instrument showed that cell viability was enhanced when treated with rapamycin for 6 h or HBSS for 4 h, while decreased at 12 and 24 h when treated with rapamycin or 8 h with HBSS (Figure 3A and B).
Transwell assay indicated that HeLa cell invasion ability was enhanced when treated with rapamycin for 6 h or HBSS for 4 h, while decreased at 12 and 24 h treated with rapamycin or 8 h with HBSS (Figure 3C–F). These results indicated that autophagy could regulate the proliferation and invasion of cervical CSCs.
Autophagy regulates the stemness of cervical CSCs
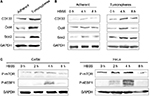
The mTOR signaling pathway associated with autophagy activity is known to be a key regulator in the pluripotency of hESCs.17 To verify whether this phenomenon exists in cancer, cervical CSCs were starved to inhibit mTOR signaling for 2, 4 and 8 h. mTOR-downstream markers (p-mTOR and p-4EBP) and PA proteins (Oct4 and Sox2) were tested. It was found that autophagy markers indicate a time-dependent induction of autophagy, LC3 II and Beclin 1 were enhanced at 2 and 4 h, but repressed at 8 h. We also observed that the expressions of Oct4, Sox2 as well as p-mTOR and p-4EBP were changed in the same way as in autophagy (Figure 4A–C). Both the PA proteins (Figure 4A and B) and the mTOR-downstream markers (Figure 4C) were increased at 2 and 4 h, and reached the peak at 4 h, but then declined at 8 h. From these results, we hypothesized a possibility that autophagy activity might be associated with the regulation of PA proteins in cervical CSCs.
The effect of autophagy inhibitor 3-MA on cervical cancer cells
To further confirm whether autophagy could regulate the stemness of cervical cancer cell, an autophagy inhibitor 3-methyladenine (3-MA) was used to inhibit the autophagy level of HeLa cells. It was found that after been treated with 3-MA, the number of autophagosome was decreased significantly (Figure 5A), and the production and invasion abilities were also inhibited (Figure 5B). Furthermore, we also observed that the expression levels of PA proteins were decreased in both in 3-MA treated group adherent and tumorsphere stem cells (Figure 5C).
Beclin 1 is critical for the malignance of cervical cancer cells
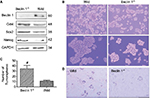
Beclin 1 gene of HeLa cell was targeted, as in our previous study (data not shown).16 To test whether PA proteins are really degraded via autophagy pathway, the protein levels of Oct4, Sox2 and CD133 in Beclin 1-/- cells were detected, and it was found that they are all inhibited compared to the wild-type groups (Figure 6A). This showed more clearly a reverse correlation in the protein regulation between autophagy and PA proteins. These results indicate that degradation of PA proteins is directly correlated with autophagy regulatory machineries.
The number of tumorspheres formed after serial passages at clonal density reflects the self-renewal of primitive CSCs, whereas the size of the tumorspheres reflects progenitor cell proliferation. To investigate the function of autophagy in the self-renewal potential of cervical CSCs and progenitor cell proliferation, we investigated HeLa tumorsphere formation following CRISPR/Cas 9-mediated Beclin 1 knockout. We found that the size and number of tumorspheres (35 versus 11) were lower (Figure 6B and C) following Beclin 1 knockout. Furthermore, when cultured in adipogenic induction media for about 2 weeks, there were more lipid droplet-like cells in Beclin 1-/- group (Figure 6D). It suggested that Beclin 1 is critical for maintaining the proliferation of cervical CSCs and progenitor cells.
Discussion
Autophagy is a process of cytoplasm and cellular organelle degradation in which a large number of small proteins are involved. It contributes to the maintenance of a proper cellular homeostasis and is emerging as an attractive therapeutic target in human cancers, including cervical cancer.15 A precise understanding of the complex autophagy machinery is essential to understand the underlying cellular and molecular mechanisms in carcinogenesis, including cervical cancer.
CSCs, also known as tumor-initiating cells, share some characteristics with adult stem cells like unlimited capacity for proliferation, self-renewal and differentiation.2 An appropriate CSC model is critical for analyzing autophagy and its involvement in CSC biology. In this study, we characterized tumorspheres from human cervical cancer cell lines. We observed that HeLa and CaSki cells could form tumorspheres when cultured in single cells. Furthermore, these tumorspheres express CD133 and Oct4 at the same time. When cultured in adipogenic induction media, the cervical CSCs could differentiate into lipid droplet-like cells. These data indicate that tumorsphere cultures of cervical cancer cells have higher stemness marker expression and the tumorspheres contain more CSCs/progenitor cells than the parental adherent cultures, which is consistent with other studies on growth of cervical CSCs as tumorspheres in three-dimensional cultures.18 Although the “whole” mouse mammary and human cervical tumors probably arise from a combination of “adherent” and “tumorsphere”-like cells, CSCs play a critical role in “total” tumor progression.
Recent reports suggest a potential role for autophagy in the origin, maintenance and distribution of CSCs.19 In the present study, we showed that at both basal level and under starvation conditions, autophagic flux was significantly higher in the tumorspheres than in the adherent cells. Tumorspheres are formed by growing cervical cancer cells under low attachment conditions; extracellular matrix detachment can induce robust autophagy in all cells during tumorsphere formation. These findings suggest that CSC/progenitor phenotype displays higher autophagic flux than the non-CSC/progenitor phenotype.
Although the existence of the CSCs in various tumor types has been confirmed, the mechanism that regulates their generation and maintenance remains elusive. In the present study, we observed that induction of autophagy by HBSS or rapamycin could regulate the population of CSCs. Firstly, HBSS- or rapamycin-treated cervical cancer cells showed increased ability of proliferation and invasion. It confirms the crucial role of autophagy in cervical CSC stemness maintaining. Secondly, we observed an increase in the expression of Oct-4 and Sox2, two potential stem cell markers, when autophagy was induced. At the same time, autophagy also helps to maintain the CSC pool, which may be crucial for developing cancer relapse. However, autophagy-mediated regulation stemness and drug-resistant genes in CSCs remain to be explored further. Moreover, in the present study, for the first time we are showing that CSCs of cervical cancer may rely highly on autophagy to sustain their proliferation and survival.
The pluripotency of ESCs is maintained by intracellular networks of many PA proteins such as Oct4, Sox2 and Nanog. However, the mechanisms underlying the regulation of protein homeostasis for pluripotency remain elusive. Here, we firstly demonstrated that autophagy could modulate the levels of PA proteins in human cervical cancer cells. A previous study had shown that rapamycin inhibits mTORC1 activity in cell cycle-arrested senescent REF cells, thereby preventing senescence and activating transcription of pluripotent genes.20 Our results were consistent with this study. But we found that autophagy and PA proteins began to decline as treatment time was prolonged. This may be because the cells tend toward apoptosis when the HBSS handles beyond the limitation.
It has reported that the PTEN/mTOR/STAT3 pathway is required for the maintenance of cervical CSCs. Activation of mTOR downregulates autophagy and leads to the phosphorylation of eukaryotic initiation factor 4E binding protein 1 (4EBP1) and 70 kDa polypeptide 1 ribosomal protein S6 kinase-1 (p70S6K).21 We monitored the activity of mTOR by detecting the levels of phosphorylation of its substrate, p4EBP1, and found that it was negatively correlated with the level of autophagy, which means the inhibition of mTOR signaling.
Beclin 1 depletion in monolayer cultures was found to increase tumorigenesis.22 In this study, in order to further confirm the effect of autophagy inhibition, we used CRISPR/Cas 9-mediated gene targeting to knock out Beclin 1, which is implicated in the initiation of autophagic pathway. We found that the knockdown of Beclin 1 in the adherent cervical cancer cells led to an increase in the expression level of PA proteins. These findings suggest that autophagy may be protumorigenic in cervical CSCs/progenitor cells.
Acknowledgments
This study was funded by the National Natural Science foundation of China (81541147 and 81602297) and Science and Technology Department of Hubei Province (2016CFB11).
Disclosure
The authors report no conflicts of interest in this work.
References
Clevers H. The cancer stem cell: premises, promises and challenges. Nat Med. 2011;17(3):313–319. | ||
Al-Hajj M, Wicha MS, Benito-Hernandez A, Morrison SJ, Clarke MF. Prospective identification of tumorigenic breast cancer cells. Proc Natl Acad Sci U S A. 2003;100(7):3983–3988. | ||
Singh SK, Clarke ID, Terasaki M, et al. Identification of a cancer stem cell in human brain tumors. Cancer Res. 2003;63(18):5821–5828. | ||
Kim CF, Jackson EL, Woolfenden AE, et al. Identification of bronchioalveolar stem cells in normal lung and lung cancer. Cell. 2005;121(6):823–835. | ||
Clement V, Sanchez P, de Tribolet N, Radovanovic I, Ruiz i Altaba A. HEDGEHOGGLI1 signaling regulates human glioma growth, cancer stem cell self-renewal, and tumorigenicity. Curr Biol. 2007;17(2):165–172. | ||
Gangemi R, Paleari L, Orengo AM, et al. Cancer stem cells: a new paradigm for understanding tumor growth and progression and drug resistance. Curr Med Chem. 2009;16(14):1688–1703. | ||
Chen B, Retzlaff M, Roos T, Frydman J. Cellular strategies of protein quality control. Cold Spring Harb Perspect Biol. 2011;3(8):a004374. | ||
Rubinsztein DC. The roles of intracellular protein-degradation pathways in neurodegeneration. Nature. 2006;443(7113):780–786. | ||
Ciechanover A. Proteolysis: from the lysosome to ubiquitin and the proteasome. Nat Rev Mol Cell Biol. 2005;6(1):79–87. | ||
Levine B, Kroemer G. Autophagy in the pathogenesis of disease. Cell. 2008;132(1):27–42. | ||
Boyer LA, Lee TI, Cole MF, et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell. 2005;122(6):947–956. | ||
Niwa H, Miyazaki J, Smith AG. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nat Genet. 2000;24(4):372–376. | ||
Mitsui K, Tokuzawa Y, Itoh H, et al. The homeoprotein Nanog is required for maintenance of pluripotency in mouse epiblast and ES cells. Cell. 2003;113(5):631–642. | ||
Wang Z, Oron E, Nelson B, Razis S, Ivanova N. Distinct lineage specification roles for NANOG, OCT4, and SOX2 in human embryonic stem cells. Cell Stem Cell. 2012;10(4):440–454. | ||
Gong C, Bauvy C, Tonelli G, et al. Beclin 1 and autophagy are required for the tumorigenicity of breast cancer stem-like/progenitor cells. Oncogene. 2013;32(18):2261–2272, 2272e. | ||
Ding Y, Yu AQ, Li CL, Fang J, Zeng Y, Li DS. TALEN-mediated Nanog disruption results in less invasiveness, more chemosensitivity and reversal of EMT in Hela cells. Oncotarget. 2014;5(18):8393–8401. | ||
Cho YH, Han KM, Kim D, et al. Autophagy regulates homeostasis of pluripotency-associated proteins in hESCs. Stem Cells. 2014;32(2):424–435. | ||
Ponti D, Costa A, Zaffaroni N, et al. Isolation and in vitro propagation of tumorigenic breast cancer cells with stem/progenitor cell properties. Cancer Res. 2005;65(13):5506–5511. | ||
Hippert MM, O’Toole PS, Thorburn A. Autophagy in cancer: good, bad, or both? Cancer Res. 2006;66(19):9349–9351. | ||
Pospelova TV, Bykova TV, Zubova SG, Katolikova NV, Yartzeva NM, Pospelov VA. Rapamycin induces pluripotent genes associated with avoidance of replicative senescence. Cell Cycle. 2013;12(24):3841–3851. | ||
Zhou J, Wulfkuhle J, Zhang H, et al. Activation of the PTEN/mTOR/ STAT3 pathway in breast cancer stem-like cells is required for viability and maintenance. Proc Natl Acad Sci U S A. 2007;104(41):16158–16163. | ||
Yue Z, Jin S, Yang C, Levine AJ, Heintz N. Beclin 1, an autophagy gene essential for early embryonic development, is a haploinsufficient tumor suppressor. Proc Natl Acad Sci U S A. 2003;100(25):15077–15082. |
 © 2017 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms.php
and incorporate the Creative Commons Attribution
- Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2017 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms.php
and incorporate the Creative Commons Attribution
- Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.