Back to Journals » International Journal of Nanomedicine » Volume 16
Application of Nanotechnology in the COVID-19 Pandemic
Authors Yang D
Received 9 December 2020
Accepted for publication 8 January 2021
Published 26 January 2021 Volume 2021:16 Pages 623—649
DOI https://doi.org/10.2147/IJN.S296383
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Thomas Webster
Dongki Yang
Department of Physiology, College of Medicine, Gachon University, Incheon, 21999, South Korea
Correspondence: Dongki Yang
Department of Physiology, College of Medicine, Gachon University, Incheon 21999, South Korea
Tel +82-32-899-6072
Fax +82-32-899-6588
Email [email protected]
Abstract: COVID-19, caused by SARS-CoV-2 infection, has been prevalent worldwide for almost a year. In early 2000, there was an outbreak of SARS-CoV, and in early 2010, a similar dissemination of infection by MERS-CoV occurred. However, no clear explanation for the spread of SARS-CoV-2 and a massive increase in the number of infections has yet been proposed. The best solution to overcome this pandemic is the development of suitable and effective vaccines and therapeutics. Fortunately, for SARS-CoV-2, the genome sequence and protein structure have been published in a short period, making research and development for prevention and treatment relatively easy. In addition, intranasal drug delivery has proven to be an effective method of administration for treating viral lung diseases. In recent years, nanotechnology-based drug delivery systems have been applied to intranasal drug delivery to overcome various limitations that occur during mucosal administration, and advances have been made to the stage where effective drug delivery is possible. This review describes the accumulated knowledge of the previous SARS-CoV and MERS-CoV infections and aims to help understand the newly emerged SARS-CoV-2 infection. Furthermore, it elucidates the achievements in developing COVID-19 vaccines and therapeutics to date through existing approaches. Finally, the applicable nanotechnology approach is described in detail, and vaccines and therapeutic drugs developed based on nanomedicine, which are currently undergoing clinical trials, have presented the potential to become innovative alternatives for overcoming COVID-19.
Keywords: COVID-19, SARS-CoV-2, antiviral drug, vaccines, nanoparticles, nanotechnology
Introduction
At the end of 2019, viral infectious disease emerged in China, which spread worldwide in months. The World Health Organization (WHO) officially declared that the coronavirus outbreak is turning into a pandemic on March 11, 2020.1,2 The WHO named this novel coronavirus to SARS-CoV-2 and the disease as COVID-19.3,4 Coronaviruses (CoVs) are RNA viruses, 27–32 kb in size, and belong to the Coronaviridae family of viruses, which includes the genera Alphacoronavirus, Betacoronavirus, Gammacoronavirus, and Deltacoronavirus. It is known as “corona” virus because all CoV particles consist of crown-like peplomer spikes. The CoV particles are pleomorphic and approximately 80 to 160 nm in diameter.5,6 CoVs are known to infect humans and various types of animals. In particular, human coronaviruses (HCoVs), a notable group of CoVs, give rise to several respiratory diseases, including bronchiolitis, pneumonia, and common cold.7 HCoVs are currently the most instantly evolving viruses because of their high genomic nucleotide recombination rates.8 To date, seven known HCoVs (HCoV-229E, HCoV-NL63, HCoV-HKU1, HCoV-OC43, Middle East respiratory syndrome coronavirus (MERS-CoV), severe acute respiratory syndrome coronavirus (SARS-CoV), and SARS-CoV-2) have been classified. In addition, HCoVs are divided into two categories, the Alphacoronaviruses (including HCoV-NL63 and HCoV-229E) and Betacoronaviruses (including HCoV-HKU1, HCoV-OC43, MERS-CoV, SARS-CoV, and SARS-CoV-2).9 Virus species belonging to the Coronavirus are very diverse and reported to be the main pathogens causing upper respiratory infections, including cold, along with rhinoviruses. Studies showed that between 30% and 80% of viral colds are infections of rhinoviruses,10 and approximately 15% are infections of HCoV-229E, HCoV-NL63, HCoV-OC43, and HCoV-HKU1.11,12 In general, HCoVs have been reported to cause upper respiratory tract infections, including cold symptoms; however, the scope of HCoV infection is not limited to the upper respiratory tract alone. In severe infections, HCoVs tend to attack the lungs directly,13,14 which is one of the unique characteristics of HCoVs that cause dyspepsia.15
HCoVs are one of the top 10 known viruses fatal to human beings, with mortality rates of up to 10% for SARS-CoV and 36% for MERS-CoV.16–18 Currently, approximately 79.2 million people have already been infected, and 1,754,493 people worldwide die within 354 days of the appearance of SARS-CoV-2 (the WHO on December 27, 2020).
SARS-CoV-2 infection is characterized by serious problems, such as in an incubation period of approximately 2 weeks mild to moderate symptoms develop in infected patients, and a high infection rate. Therefore, vaccines are important, as the data show asymptomatic transmission of SARS-CoV-2.19–21 Moreover, development of vaccines and therapeutics is the most attractive option to fight SARS-CoV-2 infection and treat infected patients. Globally, scientists and doctors are working continuously to investigate and decipher the exact viral structure, mode of infection, mode of transmission, prevention, immunopathogenic mechanisms, and the most effective treatment strategies.22
Furthermore, nanotechnology tools can provide a broader overview of the new vaccine design strategies. For instance, a nano-based formulation for SARS-CoV-2 therapeutics is being developed as a delivery vehicle, along with a novel nano-vaccine metastasis platform and useful nano drugs for treating SARS-CoV-2 infections. Therefore, until now, scientists are working hard to rapidly identify and develop appropriate nano-vaccines and treatment options, including new nano-based technologies.
SARS-CoV
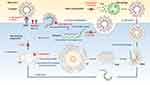
Earlier, SARS-CoV has caused a major pandemic in the new millennium.23–25 SARS-CoV has been classified as a new virus in the group II CoVs (Beta-CoVs) of the Coronaviridae family, which originated from the zoonotic pool of viruses. Owing to its epidemiological association with wild game animals and occurrence of human cases early in the 2003 pandemic,26 the SARS-CoV strain has been thought to mutate from the bat-related virus through an intermediate civet host (Figure 1A).27,28 The spike (S) protein in SARS-CoV assembles peplomers, which are located outside the lipid envelope (Figure 1B). Moreover, the S proteins of SARS-CoV, MERS-CoV, and SARS-CoV-2 have surprisingly high amino acid sequence homogeneity with each other (Figure 1C). The S protein directly interacts with the host’s cellular receptor, angiotensin-converting enzyme 2 (ACE2).29 With the binding of the S protein to ACE2, the transmembrane protease serine 2 (TMPRSS2) and furin in the host cell membrane simultaneously cleave the S protein to activate SARS-CoV (Figure 1D).30,31 The variability of the interaction between the S protein and ACE2 is particularly crucial for cross-species transmission. SARS-CoV is spread by direct contact with the mucous membrane, respiratory droplet nuclei, or fomites.32 Viral pneumonia with rapid respiratory deterioration is the most representative clinical manifestation of SARS-CoV infection. The major symptoms include chills, fever, muscle pain, discomfort, and nonproductive cough, and rhinitis and sore throat appear less frequently. Rapidly and newly occurring viral infections, such as SARS-CoV infections, are difficult to treat using vaccinology, even though no reagents are required for vaccine development. Moreover, diseases that have not been observed earlier in human beings develop and spread quickly. Novel zoonotic viruses are particularly capricious, as vaccines and remedies focused against formerly derived strains do not work against the strains related to current infectious diseases. Recently, technological advances, such as memory B cell immortalization and phage display, have achieved rapid development of human monoclonal antibodies (hu-mAbs) for SARS-CoV in just a single epidemic year.33 However, despite the ongoing efforts made by scientists and doctors, there exists no effective treatment for SARS. Antibiotics are not effective against viruses, and antiviral drugs do not show much benefit. RNA interference (RNAi) has been proposed as a new therapeutic strategy for SARS-CoV because it can degrade specific mRNAs. In addition, the inhibition of SARS-CoV by RNAi in cultured cells and animal tissues has been reported.34–43
In an experiment using Vero cells, plasmid-based siRNAs designed specifically for the viral RNA polymerase have been shown to inhibit the SARS-CoV cytopathic effect.41 In addition, a study demonstrated the suppression of the expression of S and nucleocapsid (N) proteins when RNAi is applied to SARS CoV-infected cells.36 In primate cells, siRNA duplexes targeting SARS-CoV genomic RNA have been found to block viral replication and infection.34,40
Since developing effective vaccines and achieving social immunity takes too long, the therapeutic strategy using broadly neutralized hu-mAbs targeting SARS-CoV is also a useful direction to follow at present.44 The 80R, a hu-mAb targeting SARS-CoV, has been reported to neutralize the civet (SZ3) S protein effectively in vivo.45 In the case of the hu-mAb m396, it has been reported to neutralize SARS-CoV equipped with GD03 S protein (icGD03-S) completely.46 Furthermore, animal experimental data have already been accumulated, which show that hu-mAbs can protect against SARS-CoV infection.44 Additionally, there have been some notable advances in development of small molecules targeting SARS-CoV and the design of polypeptides. However, no effective results have been published strengthening the applicability of new concepts, such as nanotechnology-based development of SARS-CoV vaccines.
The SARS-CoV epidemic is characterized by vulnerabilities in the elderly population, with a mortality rate of 25–55% among people aging 65 years and higher.47–50 Existing research suggests that the influenza virus vaccine for a young adult population is very inefficient in the elderly population at 17–53% compared with the record high level of 70–90%, which seems to be the result of an aging-related immune system dysfunction.51 Currently, in order to overcome some deficiencies pertaining to the aging immune system, co-administration of adjuvants (MF59, CpG DNA) or cytokines (IL-2) that completely activate Antigen-presenting cells (APC)/Th cells during vaccination has been attempted, and consequently, the probability of achieving a successful preventive effect has increased.52–54 Therefore, if a basic strategy aimed at developing vaccines for the elderly is continually developed, it will make a significant contribution to the research on effective SARS-CoV vaccines.
MERS-CoV
Middle East respiratory syndrome (MERS) is a novel virus-associated infectious disease identified in elderly male patients with breathing difficulties in Saudi Arabia in mid-2012.55 Most of the early MERS infections occurred in West Asia,55 and later MERS spread to Southeast Asia, North America, Europe, and North Africa.56–62
MERS-CoV has also been classified as a new virus of the group II CoVs (Beta-CoVs) in the Coronaviridae family.63 In order to enter host cells, MERS-CoV, similar to SARS-CoV, requires a receptor for the S protein. In the case of MERS-CoV, candidate proteins that act as receptors have been identified as tetopeptidases, such as dipeptidyl peptidase 4 (DPP4).64 When the S protein interacts with DPP4, the next step is the activation of the infection by TMPRSS2 and furin expressed on the host cell membrane (Figure 1D).65,66 MERS-CoV has shown little variation, except for a single mutation during the infection process between human populations. It has also been reported that this single mutation is independent of the process of binding to DPP4. Another characteristic of MERS-CoV is its ability to bind DPP4 of multiple species. Therefore, it is possible to infect other animals with MERS-CoV, except camels and humans (Figure 1A).64,65
About 60% of all MERS cases are estimated to occur via human-human transmission of MERS-CoV,67 and for the remaining cases, the cause of MERS-CoV infection has not been identified. Furthermore, the risk of virus transmission has been reported to be significantly reduced for the second case.67 Research has shown that MERS is fatal to the elderly and patients with underlying diseases, such as kidney or lung disease, chronic heart disease, high blood pressure, and diabetes.68–73 One of the most persuasive animals is the one-humped camel (Camelus dromedarius) that propagates MERS-CoV to humans, as antibodies that neutralize MERS-CoV have been detected in camel herds in the Middle East and Africa.74–83 Humans with MERS usually begin to develop symptoms after an incubation period of 2 weeks, which include respiratory infections, fever, shortness of breath, and dry or productive cough.68 To date, MERS-specific therapeutic drugs have not been developed. In vitro studies have shown that some potential drugs are effective for treating MERS; however unfortunately, most of these drugs have not been proven effective using animal models closely related to humans. Therefore, clinical treatment of MERS comprises symptomatic treatment and supportive care.
Unlike other host animals, replication of MERS-CoV is impossible in mice. This is because, mouse DPP4 (mDPP4) has two amino acid sequences different from that of human DPP4 (hDPP4), which prevents the binding of the S protein in MERS-CoV.84 Therefore, the mouse model has been developed as a strategy for replacing mDPP4 with hDPP4 or a mutant of mDPP4 so that it can bind to the viral S protein.85 Initially developed mouse models have frequently failed to reproduce the disease observed in humans with MERS-CoV infection. However, recently, several transgenic mouse models have been reported to reproduce the human disease caused by MERS-CoV infection relatively well. Therefore, it is thought that the development of such a mouse model can contribute greatly to testing the efficacy of the candidate MERS-CoV vaccine.
Since DPP4 is a specific receptor for MERS-CoV, it represents a good strategic target for designing therapeutic agents. The therapeutic agents targeting DPP4, such as DPP4 and DPP4 antagonists and specific antibodies, mainly inhibit the interaction or binding between DPP4 and MERS-CoV receptor-binding domain (RBD), thereby suppress the MERS-CoV infection. In developing therapeutics against MERS, it is essential to consider the function and structure of the S protein. Therefore, specific regions of the S1 and S2 subunits, RBD, and N-terminal domain related to S proteins are the main targets. Almost all MERS-CoV-neutralizing antibodies have been designed to target RBD. In particular, RBD-specific monoclonal antibodies have stronger neutralizing activity than that of antibodies made with other targets.
SARS-CoV-2
In Wuhan, being the most largely populated in central China, patients began to develop severe pneumonia due to an unknown cause at the end of 2019.86 Rapid research resulted in identifying the cause of the disease to be a type of coronavirus. To date, the number of infected people continues to increase rapidly.
SARS-CoV-2 belongs to the genus Betacoronavirus, and its sequence is 79% identical to the genomic sequence of SARS-CoV.87 Similar to other human betacoronaviruses, it is estimated that more than 90% of the genes in the SARS-CoV-2 genome match with those from bats, and there are several candidates for intermediate hosts existing before transmission to humans, but these are still unknown.88,89 Until now, there has been a strong hypothesis that the transmission occurred to humans by accident, such as that for SARS-CoV. Public health measures that can be used to control the transmission of SARS-CoV-2 across individuals are as much as passive approaches, such as isolation, social distancing, and refraining from small indoor gatherings. Moreover, there is a high likelihood of another major crisis occurring in the near future, as there is no vaccine or specific treatment available for SARS-CoV-2 even after an alleviation of the situation.
Like SARS-CoV, SARS-CoV-2 binds to the receptor ACE2 using the RBD of the S protein.90,91 Subsequently, the processes of fusion of cell membranes and entry of the virus into the host cell occur, similar to the mechanisms underlying other virus infections. The process by which TMPRSS2 and furin activate the S protein plays a critical role in SARS-CoV-2 infection and its spread throughout the patient’s body. Therefore, the host and host cell affinity depend on the amino acid sequence and distribution of ACE2, TMPRSS2, and furin (Figure 1D).92,93 In addition, in smokers or people with heart diseases, ACE2 levels are higher than that in healthy people, thereby increasing the susceptibility to SARS-CoV-2 infection and fastening the disease progression.94,95
SARS-CoV-2 infection is not limited to any particular class, and people of all age groups are vulnerable. The virus is mainly transmitted through droplets from symptomatic patients; however, there are many cases of infection from asymptomatic people, wherein the virus is transmitted even before the symptoms appear.96 SARS-CoV-2, present in the droplets from symptomatic patients, can usually survive on the contact surface for several days but is easily degraded by commonly available disinfectants, such as hydrogen peroxide and sodium hypochlorite.97 This droplet can cause infection via its inhalation through the respiratory tract during conversation with a SARS-CoV-2-infected individual or by touching the mucous membrane area with the hand that touched a surface contaminated by the droplet. In general, infection is caused by droplets containing SARS-CoV-2 at less than 2 m, and the risk of airborne transmission has not been reported. SARS-CoV-2 can survive for up to 3 h in droplets, and it has been known to have a survival period of about 4 h on copper compared with other metals and materials.98
Initially, after SARS-CoV-2 infection, symptoms such as dry cough, fever, and fatigue appear.99 Although not common, symptoms such as body aches, headaches, conjunctivitis, diarrhea, and sore throat may also appear.100 Currently, respiratory symptoms caused by SARS-CoV-2 vary widely, ranging from mild to severe hypoxia due to acute respiratory distress syndrome (ARDS).99 Epidemiological studies have shown that the incidence rate is significantly lower in children, and the mortality rate is very high in the elderly population.101–103 As the mortality rate in more severely ill patients increases, the disease can also be fatal to the elderly population. When infected with SARS-CoV-2, macrophages and monocytes move to the site of infection, and T and B cells together induce an immune response and begin to remove virus particles.99 In most healthy individuals, this immune response is used as a defense mechanism against viral infection; however, in patients with immunomodulatory disorders, a cytokine storm occurs, leading to severe organ failure, damaging multiple organs. This can also lead to death.104
When SARS-CoV-2 was first discovered, the most commonly searched genome was clade L, corresponding to NC_045512.2. In early 2020, the first mutant virus clades S and O appeared to have been identified. Clades V and G have appeared around the same time in mid-January. Subclades GH and GR have been reported one month after appearance of clade G. In general, clades S and GH have been observed in America, including the United States, and G and GR clades are widespread in Europe. While the appearance of clade G (including GH and GR) continues to increase gradually, that of clades L and V is gradually declining. The most common clades of the SARS-CoV-2 genome currently spread worldwide are the G clades and their derivative GH and GR clades (Figure 2). In particular, the GR clade (Nucleocapsid RG203KR mutations and the combination of the spike D614G) with high infectivity is currently the most common form of SARS-CoV-2 across the globe.105 According to a recently published study, the high mortality associated with the G clade (including GH and GR) is due to carrying the D614G mutation in the S protein, which causes SARS-CoV-2 to enter the cells at a rate more than double and be more resistant to anti-serum neutralization.106,107
Therapeutic Strategies
Antiviral Drugs
Currently, COVID-19 has increasing number of disease determinants worldwide without availability of approved treatment options, and thus, researchers are urgently developing effective vaccines and treatments. In addition, attempts of using existing medicines that have been approved for other uses may benefit COVID-19 patients to a limited extent. In vitro studies have shown that several drugs approved for other applications have some effect on SARS-CoV-2, but the results were different in small-scale non-randomization trials. These include remdesivir, which was developed as an experimental drug against Ebola virus (EBOV) during the Ebola epidemic in West Africa, chloroquine (CQ) and hydroxychloroquine (HCQ) for malaria, and lopinavir/ritonavir (LPV/r), which is used as an acquired immunodeficiency syndrome (AIDS) treatment.
First, remdesivir, a nucleotide analog prodrug that inhibits the function of the viral RNA polymerases, has been reported to reduce SARS-CoV-2 infection remarkably in Vero cells (Figure 3).108,109 Another study has found that expanded access to remdesivir in severely ill COVID-19 patients improves clinically in 36 of 53 patients.110 However, trials in severely ill COVID-19 patients in China have shown statistically insignificant clinical results.111
Second, HCQ and CQ are representative drugs for the treatment and prevention of malaria. HCQ and CQ have been shown to be effective against SARS-CoV-2 infection in an in vitro study (Figure 3).112 However, in a prospective randomized trial on COVID-19 patients in China, there was no effect of HCQ on the patients compared with those receiving conventional treatment. Rather, it has been found that a patient in the HCQ treatment group developed a serious illness.113 In clinical adjuvant therapy trials on SARS-CoV-2-infected patients, two high- and low-dose patients among patients administered 50 different CQ doses have shown a 50% lower mortality rate than that of low-dose patients.114
Finally, LPV/r is an antiretroviral drug, which is a protease inhibitor used for treating human immunodeficiency virus (HIV). In vitro studies have reported that LPV/r displays the effect of inhibiting SARS-CoV-2 replication (Figure 3).109 In addition, another clinical study has reported the administration of LPV/r and ribavirin in patients to reduce the risk of death and ARDS caused by SARS-CoV.115 However, the clinical effect of LPV/r against SAR-CoV-2 is yet to be confirmed.
The S Protein and ACE2 Interaction Inhibitors
The S protein is associated with the binding to host cell receptors and membrane fusion. Therefore, inhibitors that interfere with this process are used to prevent virus transmission from infected patients. In particular, interfering with the interaction of ACE2 with certain motifs in the S2 subunit of the S protein of SARS-CoV-2, which is involved in the virus fusion with the host cell, may be effective. According to recent in vitro research, the EK1 peptide, a pan-CoV fusion inhibitor, inhibits receptor-mediated infection and fusion between SARS-CoV-2 particles and host cell membrane, thereby prevents the formation of 6-helix bundles through interaction with heptad repeat 1, which is located in the S2 subunit of the S protein of SARS-CoV-2 (Figure 3).116
Neutralizing Antibodies
Unlike vaccines, monoclonal antibodies provide immediate protection; therefore, administering purified monoclonal antibodies with neutralizing capacity could be another SARS-CoV-2 treatment strategy. Currently, the development of effective neutralizing antibodies mainly focuses on the S protein immobilized on SARS-CoV-2. Two potent neutralizing camelid single-domain antibodies against SARS-CoV and MERS-CoV isolated from llama can cross-react with SARS-CoV-2, disrupting the receptor binding interface.117 Recently, it was confirmed that the 47D11 hu-mAb that binds to the S protein RBD can neutralize SARS-CoV-2 infection (Figure 3).118 The S309 antibody, also known as the SARS-CoV monoclonal antibody, also potently inactivates SARS-CoV-2 by acting on the S protein.119 Therefore, the use of various monoclonal antibody cocktails, which can target the listed non-RBD and RBD simultaneously, can be a good alternative for effective and safe COVID-19 prevention and treatment.
Immunotherapy
Excessive cytokine serum levels (cytokine storm) leading to multiple organ damage in severely ill COVID-19 patients are closely related to ARDS following exacerbation of COVID-19. Therefore, prevention and treatment of cytokine storms can be a good alternative that can interfere with COVID-19 progression. Clinical studies have shown that the main cause of inflammation is an increase in the levels of IL-6.103 The complex produced by binding of IL-6 with soluble IL-6 receptor (sIL-6R) or membrane IL-6 receptor (mIL6R) activates the inflammatory response through interaction with gp130. Tocilizumab (monoclonal antibody against IL-6) can block the signal transduction that triggers the inflammatory responses by selectively acting on sIL-6R and mIL-6R (Figure 3).120 A recent study reported that HCQ and CQ can block the development of proinflammatory cytokines, such as IL-6, which are involved in the generation of cytokine storms.121 However, the cost and safety aspects can hinder the use of tocilizumab in COVID-19 treatment. Based on a recently published study, sarilumab, another IL-6 receptor antagonist, may aid in rapid recovery in severely ill COVID-19 patients characterized by systemic hyperinflammation (Figure 3).122
Convalescent Plasma Therapy
Convalescent plasma (CP) therapy is another effective method; however, the CP should be used within at least 2 weeks post recovery to ensure high neutralizing antibody titers.123 According to a recent study, SARS-CoV-2 obtained from patients with severe respiratory disease can be neutralized by serum from several other patients (Figure 3).124 In another study, it was thought that the clinical status of five severely ill COVID-19 patients, who were administered CP containing neutralizing antibodies, would improve.125 Even today, many clinical trials are testing CP for COVID-19 treatment globally.
Preventive Vaccination Strategies
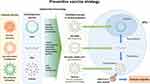
To develop effective SARS-CoV-2 vaccines, a multifaceted strategic approach to vaccine development is being attempted worldwide. Since the genomic and structural information of SARS-CoV-2 has become available much faster than that of other HCoVs, there is a possibility of rapid vaccine development.91,126–129 Moreover, data generated from vaccine development for SARS-CoV and MERS-CoV, which have been studied so far, are also helpful for developing a vaccine candidate for SARS-CoV-2.130,131
Inactivated or Live-Attenuated Vaccines
Inactivated or live-attenuated vaccines have advantages, such as stimulation of pattern recognition receptors and high immunogenicity. The viruses are alive and replicable but non-toxic. However, due to the risk of live viruses, long-term surveillance is required for assessing the safety of the vaccine. Several inactivated virus vaccines are currently being developed against SARS-CoV-2, and the first clinical trials by Sinovac Biotech, Beijing, China, have recently begun (Figure 4). More recently, recombinant SARS-CoV-2 has been synthesized from viral DNA fragments using synthetic genomics.132,133 Based on these findings, it is possible to approach a slightly more rapid generation of live-attenuated vaccines against SARS-CoV-2. Additionally, Codagenix, Farmingdale, NY, USA is exploring vaccine candidates against SARS-CoV-2 using a “codon-optimized off” strategy for virus attenuation (Figure 4).134
Recombinant Vaccines
Recombinant vaccines allow live viruses to retain some additional genes derived from pathogens through genetic manipulation, thereby translating the target protein and triggering the desired immune response.135 The advantages of recombinant vaccines are sufficient target protein expression, prolonged stability, and induction of strong immune responses.136
Vaccinia virus vector-based vaccines are currently being evaluated for use in many clinical trials based on the studies that have shown that they can induce very strong immune responses to foreign antigens.137,138 Another advantage of the vaccinia virus vector-based vaccine could be the availability of a large-scale manufacturing method, as in the case where Bavarian Nordic A/S produced and provided large amounts of its own smallpox vaccine IMVAMUNE® to the US government.139
The broad spectrum viral affinity and infectivity in dividing and non-dividing cells has made it possible to use a wide range of adenovirus (Ad) vectors to our advantage. Among the human Ad sera identified to date, human Ad serotype 5 (Ad5), which can be easily produced at high titers, is the most widely studied gene transfer vector.140,141 However, pre-existing immunity against Ad induced in many people who have already been exposed to the Ad serotype is a disadvantage of Ad vectors.
Adeno-associated virus (AAV) is a non-pathogenic, low immunogenic, vector-enclosed, single-stranded DNA virus. AAV has both the characteristics and advantages of Ads. AAV vectors require a very efficient large-scale production method, such as the baculovirus system, which has been developed because of their low titer production efficiency.142,143 AAV is better than Ad when continuous transgene expression is required for treatment.144,145 The capsid modification vector is an alternative to overcome the low immunogenicity of AAV vectors. Mixed capsids generated from different serotypes provide effective gene transfer and tropism to host cells.146 Since AAV vectors often require integration with the host genome for viral gene expression, genotoxicity risk assessment must be considered when using AAV vectors.147
ChAdOx1 nCoV-19 (AZD1222), an Ad-based recombinant vaccine developed at the University of Oxford, Oxford, UK, was found to be resistant in the Phase I/II COV001 trial, and a strong immune response to SARS-CoV-2 was generated in all participants (Figure 4).148 Almost all participants receiving AZD1222 showed a four-fold increase in the antibody neutralizing activity against the SARS-CoV-2 S protein.148 In addition, no serious side effects were reported with the use of AZD1222.148
Application of Nanotechnology in COVID-19 Therapeutics
Scientists in the field of nanomedicine have steadily conducted research on linking the gene delivery ability of various nanosystems and viral vectors to high infectivity. Nanomedical researchers have studied the molecular mechanisms of vectors to develop delivery systems that can be used in a variety of fields.149,150 Nanoparticles (NPs) and viruses act at the same scale, which makes the nanotechnology approach very powerful in vaccine development and immunoengineering.151 NPs are tools that can reproduce the structural and functional properties of viruses, and nanomedicine can be the best alternative to innovative vaccine development technologies.151–153 From the perspective of vaccine technology development, the present time, wherein SARS-CoV-2 is a major threat worldwide, is most important, and nanotechnology and nanomedicine are presented as new therapeutic technologies and approaches that can have a clinical impact.154–157
Theranostic Nanoparticles
Recently, the application of NPs has emerged as groundbreaking in the medical field and allows accurate diagnosis and specific treatment of several diseases at once. The small size, low toxicity, electrical charge, and chemical plasticity make it possible to overcome several barriers encountered in various routes of administration of a generic drug. Treatment with NPs can target the SARS-CoV-2 entry and life cycle. The S protein is the most important factor in preventing the entry of SARS-CoV-2 via the first process of membrane fusion. Thus, therapeutic NPs can be designed to pre-block SARS-CoV-2 entry by inhibiting the S protein from binding to host cells. Since the introduction of nanotechnology in the treatment of general viral infections, nanomedicines that can effectively treat viruses, including Influenza A and B viruses (IAV and IBV),158–160 EBOV,161 HIV1 and HIV2,162–165 Herpes simplex virus type 1 and 2 (HSV1 and HSV2),166–169 hepatitis B virus (HBV),170–176 hepatitis C virus (HCV),177 and human norovirus (HuNoV),178 have been developed and commercialized in various ways (Table 1). In particular, a few months ago, the first SARS-CoV-2 therapeutic drug, dexamethasone has been developed using nanotechnology. It has been reported of treating infections caused by SARS-CoV-2 using an anti-edema and anti-fibrotic mechanism, and effective delivery and treatment can be expected using various nano-formulating dexamethasones.179
 |
Table 1 Conventional and Nanomedical Treatments for Major Viruses |
Intranasal Delivery Therapy
Currently, many studies are being conducted on developing a method for delivering nanoparticles into the nasal cavity as a safe and more effective countermeasure against viral infection and treatment.180 Since SARS-CoV-2 initiates infection on the mucosal surface of the eye or nasal cavity, mucosal therapy is the most important strategy for treating such infectious diseases. Delivery through the nasal cavity is not only simple and inexpensive but also non-invasive, and the NP is rapidly absorbed due to the cavity’s abundant capillary plexus and large surface area.181 The properties of the NPs, such as the surface charge, size, and shape, are important factors to be considered while optimizing the method of delivery to the nasal cavity and play a critical role in effective and safe treatment.182 Studies have been conducted using small animals to evaluate the system that is delivered to the lungs by administering NPs to the nasal cavity. Therefore, findings of these animal studies cannot be easily generalized to humans. To date, three types of NPs (organic, inorganic, and virus-like NPs) have been designed with delivery capabilities that are suitable for therapeutic purposes, which can also be administered intranasally for effective delivery.
Lipid nanoparticles (LNs) are biocompatible due to their lipid properties; hence, they can be selectively applied in fields such as biomedical science. Among the various LNs, liposomes in the form of spherical capsules, which are hydrophilic on the inside and consist of a phospholipid bilayer on the outside, are most suitable for intranasal delivery.183 The advantages184–188 and disadvantages189–192 obtained by using liposomes have been summarized (Table 2). Using lipid-coated mesoporous silica nanoparticles, a form of LNs, an antiviral molecule ML336, which is unstable and highly hydrophilic against Venezuelan equine encephalitis virus (VEEV), has been delivered into VEEV-infected mice. The suitability, cycle time, and viral titer have been shown to improve.193 Drug candidates in the form of nucleic acids, such as siRNA, have a significant limitation of being unstable during systemic circulation.194–196 However, transporting siRNA using LNs can target specific organs and has the great advantage of preventing degradation during systemic circulation.197
 |
Table 2 Advantages and Disadvantages of Nanomedicine on Therapeutic Strategies for COVID-19 |
Polymer nanoparticles (PNs) are an effective choice of delivery systems because their properties and functions can be adjusted according to their specific application. Conjugation of a therapeutic compound to chitosan-made PNs can improve penetration of the mucosal tissue and the persistence of PNs in the mucosal environment.198,199 Antibody-drug conjugates using auristatin are used for relatively safe treatment of blood cancer by eliminating the risk of high toxicity but have a critical disadvantage of the drug payload being too low. To overcome this limitation, nanoparticle-drug conjugates of monomethyl auristatin E, developed using PN technology, enable the availability of a large amount of auristatin payload, and have high safety.200 In addition, in the case of accurin PNs encapsulating the Aurora B kinase inhibitor AZD2811, the toxicity has been observed to reduce significantly and the efficacy has been found to increase compared with that before introducing PNs, which has been shown to cause decisive side effects in a Phase 2 clinical trial.201
Dendrimer nanoparticles (DNs) have strong interactions with viruses. The resulting system improves antiviral activity and has a powerful effect in preventing infection in the host. In addition, effective cases have been reported, wherein DNs are used as a treatment for viral infectious diseases, such as influenza virus and HIV.202 The advantages203–212 and disadvantages209,210,213–215 obtained by applying PNs and DNs are summarized (Table 2).
The ability of gold nanoparticles (GNs) to induce an immune response by antigen-presenting cells easily is attractive for use in vaccine development. GNs have the advantage of being easily transformed for delivery through the nasal cavity.216 It also has the advantage of activating the immune response associated with CD8+ (cytotoxic) T cells by spreading to the lymph nodes.217 The advantages218 and disadvantages219–222 of applying GNs to nanomedicine have been separately summarized (Table 2).
Virus-like nanoparticles (VLPs) are capsids, comprising virus-derived structural proteins and adjuvants. VLPs can generate a potential immunogenic epitope, resulting in higher immunogenicity. Furthermore, since VLPs are small, they can act as adjuvants, and changing adjuvants can induce a much more effective immune response than viruses.223–225 As a result of intranasal delivery of VLPs using influenza virus, it has been found that VLP functions as a vaccine by producing a very large number of T cells and antibodies that can induce various types of immune reactions to improve immunity and prevent further infection.226 The advantages227–233 and disadvantages227,234,235 of applying VLPs in terms of drug delivery or treatment have been summarized separately (Table 2).
Cell membrane nanovesicles and exosomes have been reported to have the ability to bind and neutralize bacterial toxins by previous studies.236–238 In addition, recently, the development of cell membrane nanovesicles containing proteins having the same structure and activity as native cells is being made by biomimetic synthesis technology that includes the synthesis and display of proteins on the cell surface.239–241 Cell membrane nanovesicles, built to display high levels of ACE2 and abundant cytokine receptors, are nanodecoys that can compete with host cells for viral and cytokine binding. Studies have shown that nanodecoy significantly inhibited the replication and infection of SARS-CoV-2 and efficiently binds and neutralizes inflammatory cytokines such as IL-6 and GM-CSF.242,243 Therefore, a treatment method using cell membrane nanovesicles can be an effective alternative to SARS-CoV-2 and cytokine storms.
Exosomes are tiny nanovesicles with a size of 30nm to 150nm, secreted for all types of cell-to-cell communication, and are emerging nanomaterials in recent cell regeneration, treatment, and diagnostic research.244 It has already been reported that exosomes containing the S protein of SARS-CoV induced an accelerated neutralizing antibody titer by priming with a vaccine of the S protein of SARS-CoV and then increasing with an adenovirus vector vaccine.245 Therefore, this strategy using exosomes has the potential to be sufficiently applied to treatment for SARS-CoV-2. The advantages246–248 and disadvantages249–254 of applying cell-derived vesicles to nanomedicine have been separately summarized (Table 2).
Pulmonary Delivery Using NP Inhalation Aerosols
If the advantage of drug delivery through the nasal cavity is to act on the mucous membrane area where the infection occurs, then the lungs are an important organ for drug delivery because they are another target for treating SARS-CoV-2 infection, which infects primarily through the respiratory tract (the upper airways and lung).255–257 Therefore, the use of inhaled aerosols is suggested as an effective non-invasive mode of administration. Additionally, the delivery of inhalable NPs to the lungs overcomes disadvantages, such as side effects caused by high drug concentrations in the serum with conventional oral or intravenous drug administration methods. Various nanotechnologies have been applied to develop NPs that can function as lung inhalation aerosols. These respirable NPs can be encapsulated by microparticles manufactured down to five microns to fit the aerodynamic size range or agglomerate into an aerodynamic size range. Most NPs are delivered directly to the lungs either by spraying colloidal dispersions or via dry powder inhalers and pressurized metered dose inhalers in solid form.255
The LNs mentioned previously are also one of the most widely studied NPs for effective delivery of drugs into the lungs.258 Due to the unique advantages of LNs, which include their production from substances present in the lungs, such as components of lung surfactants, they are the highest priority candidates for delivering therapeutics to the lungs.259 Liposomes are generally liquid, and the application of aerosol through a nebulizer has mainly been attempted to deliver them to the lungs in the early days;260 however, drug stability and nebulizer leakage have been pointed out as the disadvantages.261 Therefore, to compensate for these shortcomings, various studies have been conducted on the development of liposome formulations in the form of dry powder.262–264 Moreover, cationic liposomes, which have the advantage of self-assembly with nucleic acids, are attracting the most attention as systems that deliver genes to the lungs and are known to be suitable for transporting peptides and substances having high molecular weight.265
PNs play important roles in drug delivery to the lungs by achieving efficient delivery of drugs, maintaining the stability of drugs, and controlling the release of drugs.266 Currently, cationic LNs have many clinical advantages over PNs; however, cationic PNs are one of the important carriers for pulmonary delivery of genes.267,268
Although studies on delivery through the nasal cavity using DNs have already been mentioned above, many studies on pulmonary delivery using DN are also being conducted for delivering DNA drugs to the cell nucleus, with properties similar to that of liposomes.269 Research has already focused on drug delivery to the lungs as one of the applications of the method of delivering high molecular weight substances into the body using DNs.270,271 In order to use DNs effectively while delivering drugs to the lungs, additional studies are needed that consider aspects such as the biocompatibility and cytotoxicity.
Nanotechnology-Based Diagnosis
Nano biosensors have the advantage of selectively detecting all types of analytes by combining the excellent electrical and optical properties of nanomaterials with biological or synthetic molecules used as receptors.272 Using these advantages, various methods of detecting SARS-CoV-2 are being studied.273
Currently, using a Silicon-on-insulator nanowire sensor made using complementary metal-oxide-semiconductor compatible technology, the SARS-CoV-2 antibody can be detected in 5–15 minutes with an expected sensitivity of 10−12–10−15 M.274
In the case of applying Graphene, the detection of SARS-CoV-2 in clinical samples was attempted with a sensor produced by coating the graphene sheet of the field-effect transistor with a specific antibody against the SARS-CoV-2 spike protein.275 As a result of the study, it was possible to detect SARS-CoV-2 spike protein at a concentration of 1 fg/mL in phosphate-buffered saline and 100 fg/mL clinical transport medium.275
The SARS-CoV-2 biosensor using thiol-modified antisense oligonucleotides-capped GNs can diagnose positive COVID-19 cases with the naked eye through color change within 10 minutes from total RNA isolated from infected biosamples.276 As another application method for GNs, the glycan bond between the polymer-stabilized multivalent GNs bearing sialic acid derivative and the S protein of SARS-CoV-2 was identified using a glyconanoparticle platform. Applying these characteristics has the advantage of building a low-cost detection platform that can be detected in less than 30 minutes with a lateral flow diagnostic device.277
Nanotechnology-Based Vaccine Development
Subunit Vaccines
Subunit vaccine candidates are required to enhance immunogenicity effectively by eliciting an immune response when co-administered with molecular adjuvants using specific parts of the structural components of SARS-CoV-2. Therefore, developing a vaccine that targets the subunit of the SARS-CoV-2 S protein is a top priority. This is because membrane fusion and receptor-binding sites are present on the S protein.278 Vaccines based on the S protein inhibit viral infection by activating antibodies that prevent viral binding and subsequent membrane fusion.279 The SARS-CoV-2 S protein, which interacts with ACE2, is a notable candidate sufficient for both vaccine and therapeutic development.87,124,280 In addition, NPs similar to immunogenic viruses have been developed and produced with the Novavax® proprietary recombinant nanoparticle vaccine technology with the S protein (Figure 4).281 The University of Queensland, Brisbane, Australia is also developing a new SARS-CoV-2 subunit vaccine using a “molecular clamp” technology that pre-blocks the binding of viral proteins.282 As an alternative, the development of subunit vaccines using NPs, such as VLPs and protein NPs, is also actively underway. A higher binding affinity of RBD in SARS-CoV-2 for ACE2 than that of RBD in SARS-CoV has been found.283 Therefore, the RBD-based SARS-CoV vaccine can help prevent SARS-CoV-2 infection and be important for SARS-CoV-2 vaccine development. Moreover, RBD-based vaccines are effective in preventive and therapeutic strategies and are currently being developed by many research institutes and multinational pharmaceutical companies.134 RBD-based vaccines also have the advantage of minimizing host immunity enhancement.279
Nucleic Acid Vaccines
When viruses enter the host cell by infection, the antigen encoded by the nucleic acid is expressed, which induces a cell-mediated reaction with the antibody. Based on this principle, nucleic acid vaccination is another effective immunization method that uses artificially synthesized nucleic acids to elicit an immune response, such as that induced by live-attenuated vaccines. The improved immunogenic properties that mimic the infectious process are the potential advantages of mRNA vaccines. To maximize the effect, several mRNAs are mixed into a single vaccine.134,284 An RNA vaccine candidate against SARS-CoV-2 is now known as mRNA-1273 (Moderna, Cambridge, MA, USA) (Figure 4). This vaccine comprises a synthetic mRNA strand such that the binding site for ACE2 can be translated to the previously modified SARS-CoV-2 S protein.285 After inoculation with intramuscular injection, a specific antiviral response to the SARS-CoV-2 S protein is induced. Moreover, the synthesis of nucleic acid vaccines does not require viruses, unlike conventional vaccines made of small subunits of inactivated or live pathogens.285 Therefore, as the safety is guaranteed, only the passing of the Phase I trial for mRNA-1273 will help a continuous evaluation of efficacy to progress quickly.285 mRNA-1273 is designed based on the LN platform; however, new nanotechnology is being introduced for the effective delivery of nucleic acid vaccines. In the case of mRNA-based vaccines, not only LNs but also DNs and PNs are being used for effective delivery and high stability.286,287 BNT162b1, under development by Pfizer, New York, NY, USA, is a codon-optimized mRNA vaccine encoding the SARS-CoV-2 RBD (Figure 4). This vaccine uses the RBD antigen to which the trimerization domain of T4 fibritin has been added to increase immunogenicity.288 Coalition for Epidemic Preparedness Innovation had begun developing vaccines as soon as the first gene sequence was released through partnership with a group developing vaccines using a novel platform. As a result, the mRNA-based SARS-CoV-2 candidate progressed to the human clinical trial stage.289 In addition, INO-4800, developed by Inovio Pharmaceuticals, Inc., Plymouth Meeting, PA, USA, is a candidate DNA vaccine among nucleic acid vaccines (Figure 4). Similar to RNA vaccines, INO-4800 is a nucleic acid vaccine that can induce an immune response by being translated into proteins within human cells. Compared with conventional vaccines, nucleic acid vaccines have great advantages in terms of production cost and purification methods. Furthermore, the nucleic acid-only structure also prevents the production of misfolded proteins that can occur in recombinant vaccines.290,291 However, the immunogenicity of nucleic acid vaccines is greatly influenced by the amount of plasmid injected into the cell and the appropriate administration interval and route. Through nanotechnology, NPs, including cationic liposomes, DNs, or PNs, have been applied to the development of nucleic acid-based vaccines to enhance the delivery efficacy and stability.286,287
NP-Based Vaccines
Unlike SARS-CoV, MERS-CoV has been utilized multiple times to introduce nanotechnology into vaccines or therapeutic research. Importantly, it has been recently reported that VLPs are suitable for the development of vaccines or treatments for symptoms of MERS-CoV infection. Nano-sized VLPs, which have the characteristic function of the virus, have the advantage of being better delivered through the lymph and capillaries than other small vaccines.292–294 In addition, it has the effect of reducing the systemic inflammatory response, and similar to viruses, has the advantage of being able to very easily enter cells.293 Furthermore, the delivery of many antigens makes the antigen-presenting cell functioning more effective. Therefore, the synthesized complex recognized by the T cell receptor increases the vaccine’s immunogenicity and efficacy, thereby ensuring patient safety.293 Nano-sized VLPs entering the host cell are directly involved in B cell activation and boosting the immune system.292,295 Indeed, the characteristics of these synthetic nano-sized VLPs are principle to developing vaccine platforms.296–298 Nano-sized VLPs have also been reported to overcome viruses by increasing the immune response effectively in animal experiments.281,299,300
Recently, the MERS-CoV S protein has been synthesized using silkworm larvae. This has then been applied to the nano-sized VLPs,223 which exhibit native conformational epitopes produced via incubation with surfactant and cell membrane vesicles.239 In another study, the development of nano-sized VLPs capable of acting as a nanocarrier in red blood cells has been achieved by single compression of red blood cells through a 1-μm filter.301 MERS-CoV nano-sized VLPs have been synthesized using the recombinant S, membrane, and envelope proteins, tested in animal models, and linked to having increased immunogenicity.223 Nano-sized VLPs have a wide range of applications, can enhance vaccine safety and effectiveness, and have tremendous advantages that can be utilized for specific purposes. Since these findings have been derived for the S protein commonly present in MERS-CoV and SARS-CoV, they can be effectively applied for treating SARS-CoV-2 infection.
Inactivation of SARS-CoV-2 in the External Environment Using Nanotechnology
SARS-CoV-2 is activated at temperatures ranging from 1 to 35°C and is easily inactivated under UV, highly alkaline, or acidic conditions.302 In addition, the degree of stability of SARS-CoV-2 varies greatly depending on the components that make up the surface of the infectious particle, and SARS-CoV-2 can be easily inactivated with commonly available disinfectants.303 The activation of SARS-CoV-2 in aerosols and on surfaces is similar to that of SARS-CoV; therefore, surface treatment using NPs that have been proven effective against SARS-CoV will be sufficiently applicable to SARS-CoV-2.98 The use of nanotechnology can provide alternatives more effective than conventional disinfection protocols for viruses used in general or medical settings that typically rely on chemical, physical, and biological strategies. Moreover, by using NPs, one can freely control the release rate of metal ions, which have proven to be antibacterial, on the surface of substances requiring antibacterial action. Because NPs can accumulate in cells owing to their nature, they can overcome the disadvantages of antimicrobial substances or metal ions that easily leak out of cells. Silver, which has been used as an antibacterial agent since ancient times, is now applied to paints and food trays.304–306 Silver nanoparticles (Ag-NPs) have already been proven to display antiviral effects against various viruses.307–310 Ag-NPs exert antiviral activity by dissolving and releasing Ag+ ions with microbial toxicity. Ag+ ions can interact with proteins present on the surface of the virus or infiltrate and accumulate in host cells, disrupting the function of proteins that play an important role in virus replication, such as enzymes involving thiols.308,311 Another antiviral function of Ag-NPs has been hypothesized, wherein they competitively interfere with virus binding to host cells because of their physical interactions with the viral surface, depending on their size.307 As a result, it has been found that Ag-NPs with a size of about 10 nm show the strongest physical interaction and antiviral effect compared with that of particles in other sizes.308 In addition, Ag-NPs have an antiviral effect of damaging the virus structure using reactive oxygen species that are released after binding to the virus surface. Ag-NPs have already been applied to and used in medical equipment. When applied to face masks and air filters, they can be used to inactivate SARS-CoV-2 via the antiviral effect of Ag+ ions. Currently, it has been reported that bacteriophage MS2 from dust can effectively be blocked by applying Ag-NPs to filters.312
Copper, which has recently been proven to exhibit antiviral effect against HuCoV-229E, may be a suitable candidate for the inactivation of SARS-CoV-2.313 When a virus incubates on a surface coated with Cu, the virus genome is degraded and inactivated.313 This antiviral mechanism involves the inactivation of virions by disrupting the function of certain viral proteins using hydroxyl radicals produced by Cu2+ ions present on the surface of the material and inactivation by direct contact with the surface.314 Similarly, studies have reported that SARS-CoV-2 is easily deactivated on the surface of Cu-loaded materials.98 Furthermore, Cu is far more advantageous in terms of economy than Ag, and it can easily be used to produce PNs and has excellent stability. Therefore, the development and application of NPs with Cu or copper oxide (CuO) is the most suitable strategy to inactivate SARS-CoV-2 in the external environment. For example, in an experiment using a mask containing CuO-NPs, the influenza virus has been shown to be inactivated remarkably.315
Graphene derivatives (GDs), together with metal NPs, can effectively inactivate viruses.316 The antiviral mechanism of GDs involves electrostatic interactions, wherein the negative charge on the coated surface of the GDs promotes its binding to the positively charged viral particles.317 When GDs are applied to antibodies against viruses using nanotechnology, they show excellent effects on rotavirus and influenza virus infections.318–320 In addition, this characteristic of GDs can also be applied to the prevention, diagnosis, and treatment of SARS-CoV-2, according to recent studies.321
Iron oxide nanoparticles (IONPs) have already proven antibacterial activity through many studies.322,323 It has also been approved by the US Food and Drug Administration (FDA) for the treatment of anemia because of the excellent biocompatibility of IONPs.324 The interaction between IONPs and the S protein of SARS-CoV-2 has been identified in recent studies and the potential antiviral activity of IONPs has been reported.325 In addition, the ability of IONPs to produce ROS can be applied to inactivate SARS-CoV-2 in the external environment.326,327
Conclusion
In the past, treatment and vaccine candidates for SARS and MERS have not been fully researched and developed, as they have not been recognized for adequate investment and effectiveness due to the significantly lower infection rates than that for COVID-19. However, unlike the case of SARS or MERS, COVID-19 has been a worldwide threat for almost a year. Research and development using innovative methods, such as nanotechnology, is essential to end this pandemic effectively in a short time. Various treatments using nanotechnology have been developed and commercialized for common viral infections, such as IAV and IBV,158–160 EBOV,161 HIV1 and 2,162–165 HSV1 and 2,166–169 HBV and HCV,170–177 and HuNoV.178 The accumulated advancements in these virus-fighting nanotechnologies can play an important role in taking SARS-CoV-2 treatment and vaccine development to the next level. The tedious COVID-19 pandemic, which has not yet been put to end, is now moving in the direction of overcoming the virus in a step-by-step fashion with the help of nanomedicine. Currently, several companies are moving away from traditional SARS-CoV-2 treatment and prevention strategies and using nanotechnology to develop various types of vaccines and therapeutics and conduct clinical evaluations. For example, dexamethasones, a COVID-19 therapeutic agent that has introduced via various nano-formulations, has led to a big turn in the treatment of COVID-19.179 In addition, the clearance of Phase 3 clinical trials of the liposomal mRNA vaccine (BNT162b) developed by Pfizer can be considered a great achievement of nanomedicine.328 Moreover, the technology that deactivates SARS-CoV-2 in the external environment using nanomaterials, such as Ag-NPs,307–310 NPs with Cu or CuO,314 and GDs,316 and diagnostic technology that can quickly detect SARS-CoV-2 without the use of expensive equipment by applying GNs,329 are also contributing towards the prevention and control of COVID-19. Nonetheless, owing to the complex situation caused by COVID-19, it is believed that the existing platform needs to be modified in order for the research in various fields globally to be more efficient. Therefore, nanotechnology and nanomedicine can be suitable alternatives to this change in the research and development paradigm.
Acknowledgment
This work was supported by the Basic Science Research Program, through the National Research Foundation of Korea (NRF), funded by the Korea government (MSIT) (NRF-2018R1D1A1B07041381).
Disclosure
The author declares that they have no conflicts of interest for this work.
References
1. Wang C, Horby PW, Hayden FG, Gao GF. A novel coronavirus outbreak of global health concern. Lancet. 2020;395(10223):470–473. doi:10.1016/S0140-6736(20)30185-9
2. Perlman S. Another decade, another coronavirus. New Engl J Med. 2020;382(8):760–762. doi:10.1056/NEJMe2001126
3. Gorbalenya AE, Baker SC, Baric RS, et al. The species severe acute respiratory syndrome-related coronavirus: classifying 2019-nCoV and naming it SARS-CoV-2. Nat Microbiol. 2020;5(4):536–544.
4. Wu JT, Leung K, Leung GM. Nowcasting and forecasting the potential domestic and international spread of the 2019-nCoV outbreak originating in Wuhan, China: a modelling study (vol 395, pg 689, 2020). Lancet. 2020;395(10225):E41. doi:10.1016/S0140-6736(20)30260-9
5. Woo PCY, Huang Y, Lau SKP, Yuen KY. Coronavirus genomics and bioinformatics analysis. Viruses Basel. 2010;2(8):1804–1820. doi:10.3390/v2081803
6. Guy JS, Breslin JJ, Breuhaus B, Vivrette S, Smith LG. Characterization of a coronavirus isolated from a diarrheic foal. J Clin Microbiol. 2000;38(12):4523–4526. doi:10.1128/JCM.38.12.4523-4526.2000
7. Pene F, Merlat A, Vabret A, et al. Coronavirus 229E-related pneumonia in immunocompromised patients. Clin Infect Dis. 2003;37(7):929–932. doi:10.1086/377612
8. Vijgen L, Keyaerts E, Moes E, Maes P, Duson G, Van Ranst M. Development of one-step, real-time, quantitative reverse transcriptase PCR assays for absolute quantitation of human coronaviruses OC43 and 229E. J Clin Microbiol. 2005;43(11):5452–5456. doi:10.1128/JCM.43.11.5452-5456.2005
9. Chan JF, Lau SK, To KK, Cheng VC, Woo PC, Yuen KY. Middle east respiratory syndrome coronavirus: another zoonotic betacoronavirus causing SARS-like disease. Clin Microbiol Rev. 2015;28(2):465–522.
10. Palmenberg AC, Spiro D, Kuzmickas R, et al. Sequencing and analyses of all known human rhinovirus genomes reveal structure and evolution. Science. 2009;324(5923):55–59. doi:10.1126/science.1165557
11. Cecil RL, Goldman L, Schafer AI. Goldman’s Cecil Medicine.
12. van der Hoek L. Human coronaviruses: what do they cause? Antivir Ther. 2007;12(4 Pt B):651–658.
13. Ding Y, Wang H, Shen H, et al. The clinical pathology of severe acute respiratory syndrome (SARS): a report from China. J Pathol. 2003;200(3):282–289. doi:10.1002/path.1440
14. Rabi FA, Al Zoubi MS, Kasasbeh GA, Salameh DM, Al-Nasser AD. SARS-CoV-2 and Coronavirus disease 2019: what we know so far. Pathogens. 2020;9(3):231. doi:10.3390/pathogens9030231
15. Lotfi M, Hamblin MR, Rezaei N. COVID-19: transmission, prevention, and potential therapeutic opportunities. Clin Chim Acta. 2020;508:254–266. doi:10.1016/j.cca.2020.05.044
16. Shi H, Han X, Zheng C. Evolution of CT manifestations in a patient recovered from 2019 novel Coronavirus (2019-nCoV) pneumonia in Wuhan, China. Radiology. 2020;295(1):20. doi:10.1148/radiol.2020200269
17. Su S, Wong G, Shi W, et al. Epidemiology, genetic recombination, and pathogenesis of coronaviruses. Trends Microbiol. 2016;24(6):490–502. doi:10.1016/j.tim.2016.03.003
18. Wolfe ND, Dunavan CP, Diamond J. Origins of major human infectious diseases. Nature. 2007;447(7142):279–283. doi:10.1038/nature05775
19. Day M. Covid-19: four fifths of cases are asymptomatic, China figures indicate. BMJ. 2020;369:m1375. doi:10.1136/bmj.m1375
20. Sutton D, Fuchs K, D’Alton M, Goffman D. Universal screening for SARS-CoV-2 in women admitted for delivery. N Engl J Med. 2020;382(22):2163–2164. doi:10.1056/NEJMc2009316
21. Mizumoto K, Kagaya K, Zarebski A, Chowell G. Estimating the asymptomatic proportion of coronavirus disease 2019 (COVID-19) cases on board the Diamond Princess cruise ship, Yokohama, Japan, 2020. Euro Surveill. 2020;25(10). doi:10.2807/1560-7917.ES.2020.25.10.2000180
22. Sahu KK, Lal A, Mishra AK. Latest updates on COVID-2019: a changing paradigm shift. J Med Virol. 2020;92(6):533–535. doi:10.1002/jmv.25760
23. Drosten C, Gunther S, Preiser W, et al. Identification of a novel coronavirus in patients with severe acute respiratory syndrome. New Engl J Med. 2003;348(20):1967–1976. doi:10.1056/NEJMoa030747
24. Ksiazek TG, Erdman D, Goldsmith CS, et al. A novel coronavirus associated with severe acute respiratory syndrome. New Engl J Med. 2003;348(20):1953–1966. doi:10.1056/NEJMoa030781
25. Peiris JSM, Lai ST, Poon LLM, et al. Coronavirus as a possible cause of severe acute respiratory syndrome. Lancet. 2003;361(9366):1319–1325. doi:10.1016/S0140-6736(03)13077-2
26. Zhong NS, Zheng BJ, Li YM, et al. Epidemiology and cause of severe acute respiratory syndrome (SARS) in Guangdong, People’s Republic of China, in February, 2003. Lancet. 2003;362(9393):1353–1358. doi:10.1016/S0140-6736(03)14630-2
27. Guan Y, Zheng BJ, He YQ, et al. Isolation and characterization of viruses related to the SARS coronavirus from animals in Southern China. Science. 2003;302(5643):276–278. doi:10.1126/science.1087139
28. Lau SKP, Woo PCY, Li KSM, et al. Severe acute respiratory syndrome coronavirus-like virus in Chinese horseshoe bats. Proc Natl Acad Sci U S A. 2005;102(39):14040–14045. doi:10.1073/pnas.0506735102
29. Hamming I, Timens W, Bulthuis MLC, Lely AT, Navis GJ, van Goor H. Tissue distribution of ACE2 protein, the functional receptor for SARS coronavirus. A first step in understanding SARS pathogenesis. J Pathol. 2004;203(2):631–637. doi:10.1002/path.1570
30. Bertram S, Glowacka I, Muller MA, et al. Cleavage and activation of the severe acute respiratory syndrome coronavirus spike protein by human airway trypsin-like protease. J Virol. 2011;85(24):13363–13372. doi:10.1128/JVI.05300-11
31. Iwata-Yoshikawa N, Okamura T, Shimizu Y, Hasegawa H, Takeda M, Nagata N. TMPRSS2 contributes to virus spread and immunopathology in the airways of murine models after Coronavirus infection. J Virol. 2019;93(6). doi:10.1128/JVI.01815-18
32. Seto WH, Tsang D, Yung RWH, et al. Effectiveness of precautions against droplets and contact in prevention of nosocomial transmission of severe acute respiratory syndrome (SARS). Lancet. 2003;361(9368):1519–1520. doi:10.1016/S0140-6736(03)13168-6
33. Sui JH, Li WH, Murakami A, et al. Potent neutralization of severe acute respiratory syndrome (SARS) coronavirus by a human mAb to S1 protein that blocks receptor association. Proc Natl Acad Sci U S A. 2004;101(8):2536–2541. doi:10.1073/pnas.0307140101
34. Li BJ, Tang QQ, Cheng D, et al. Using siRNA in prophylactic and therapeutic regimens against SARS coronavirus in rhesus macaque. Nat Med. 2005;11(9):944–951. doi:10.1038/nm1280
35. Ni B, Shi XF, Li Y, Gao WD, Wang XL, Wu YZ. Inhibition of replication and infection of severe acute respiratory syndrome-associated coronavirus with plasmid-mediated interference RNA. Antivir Ther. 2005;10(4):527–533.
36. Zhao P, Qin ZL, Ke JS, et al. Small interfering RNA inhibits SARS-CoV nucleocapsid gene expression in cultured cells and mouse muscles. FEBS Lett. 2005;579(11):2404–2410. doi:10.1016/j.febslet.2005.02.080
37. Shi Y, Yang DH, Xiong J, Jia H, Huang B, Jin YX. Inhibition of genes expression of SARS coronavirus by synthetic small interfering RNAs. Cell Res. 2005;15(3):193–200. doi:10.1038/sj.cr.7290286
38. Wu CJ, Huang HW, Liu CY, Hong CF, Chan YL. Inhibition of SARS-CoV replication by siRNA. Antivir Res. 2005;65(1):45–48. doi:10.1016/j.antiviral.2004.09.005
39. Qin ZL, Zhao P, Zhang XL, et al. Silencing of SARS-CoV spike gene by small interfering RNA in HEK 293T cells. Biochem Biophys Res Commun. 2004;324(4):1186–1193. doi:10.1016/j.bbrc.2004.09.180
40. Zheng BJ, Guan Y, Tang Q, et al. Prophylactic and therapeutic effects of small interfering RNA targeting SARS-coronavirus. Antivir Ther. 2004;9(3):365–374.
41. Wang Z, Ren L, Zhao X, et al. Inhibition of severe acute respiratory syndrome virus replication by small interfering RNAs in mammalian cells. J Virol. 2004;78(14):7523–7527. doi:10.1128/JVI.78.14.7523-7527.2004
42. Zhang Y, Li T, Fu L, et al. Silencing SARS-CoV Spike protein expression in cultured cells by RNA interference. FEBS Lett. 2004;560(1–3):141–146. doi:10.1016/S0014-5793(04)00087-0
43. Li T, Zhang Y, Fu L, et al. siRNA targeting the leader sequence of SARS-CoV inhibits virus replication. Gene Ther. 2005;12(9):751–761. doi:10.1038/sj.gt.3302479
44. Ter Meulen J, van den Brink EN, Poon LL, et al. Human monoclonal antibody combination against SARS coronavirus: synergy and coverage of escape mutants. PLoS Med. 2006;3(7):e237. doi:10.1371/journal.pmed.0030237
45. Sui J, Li W, Roberts A, et al. Evaluation of human monoclonal antibody 80R for immunoprophylaxis of severe acute respiratory syndrome by an animal study, epitope mapping, and analysis of spike variants. J Virol. 2005;79(10):5900–5906. doi:10.1128/JVI.79.10.5900-5906.2005
46. Zhu Z, Chakraborti S, He Y, et al. Potent cross-reactive neutralization of SARS coronavirus isolates by human monoclonal antibodies. Proc Natl Acad Sci U S A. 2007;104(29):12123–12128. doi:10.1073/pnas.0701000104
47. Booth CM, Matukas LM, Tomlinson GA, et al. Clinical features and short-term outcomes of 144 patients with SARS in the greater Toronto area. JAMA. 2003;289(21):2801–2809. doi:10.1001/jama.289.21.JOC30885
48. Leung GM, Hedley AJ, Ho LM, et al. The epidemiology of severe acute respiratory syndrome in the 2003 Hong Kong epidemic: an analysis of all 1755 patients. Ann Intern Med. 2004;141(9):662–673. doi:10.7326/0003-4819-141-9-200411020-00006
49. Peiris JSM, Chu CM, Cheng VCC, et al. Clinical progression and viral load in a community outbreak of coronavirus-associated SARS pneumonia: a prospective study. Lancet. 2003;361(9371):1767–1772. doi:10.1016/S0140-6736(03)13412-5
50. Liang W, Zhu Z, Guo J, et al. Severe acute respiratory syndrome, Beijing, 2003. Emerg Infect Dis. 2004;10(1):25–31. doi:10.3201/eid1001.030553
51. Goodwin K, Viboud C, Simonsen L. Antibody response to influenza vaccination in the elderly: a quantitative review. Vaccine. 2006;24(8):1159–1169. doi:10.1016/j.vaccine.2005.08.105
52. Haynes L, Eaton SM, Burns EM, Rincon M, Swain SL. Inflammatory cytokines overcome age-related defects in CD4 T cell responses in vivo. J Immunol. 2004;172(9):5194–5199. doi:10.4049/jimmunol.172.9.5194
53. Pulendran B, Ahmed R. Translating innate immunity into immunological memory: implications for vaccine development. Cell. 2006;124(4):849–863. doi:10.1016/j.cell.2006.02.019
54. Thompson JM, Whitmore AC, Konopka JL, et al. Mucosal and systemic adjuvant activity of alphavirus replicon particles. Proc Natl Acad Sci U S A. 2006;103(10):3722–3727. doi:10.1073/pnas.0600287103
55. Zaki AM, van Boheemen S, Bestebroer TM, Osterhaus ADME, Fouchier RAM. Isolation of a novel Coronavirus from a man with pneumonia in Saudi Arabia. New Engl J Med. 2012;367(19):1814–1820. doi:10.1056/NEJMoa1211721
56. Bialek SR, Allen D, Alvarado-Ramy F, et al. First confirmed cases of middle east respiratory syndrome Coronavirus (MERS-CoV) infection in the United States, updated information on the epidemiology of MERS-CoV infection, and guidance for the public, clinicians, and public health authorities May 2014. Am J Transplant. 2014;14(7):1693–1699.
57. Cauchemez S, Van Kerkhove MD, Riley S, Donnelly CA, Fraser C, Ferguson NM. Transmission scenarios for middle east respiratory syndrome Coronavirus (MERS-CoV) and how to tell them apart. Eurosurveillance. 2013;18(24):7–13.
58. Drosten C, Seilmaier M, Corman VM, et al. Clinical features and virological analysis of a case of Middle East respiratory syndrome coronavirus infection. Lancet Infect Dis. 2013;13(9):745–751. doi:10.1016/S1473-3099(13)70154-3
59. Mailles A, Blanckaert K, Chaud P, et al. First cases of Middle East respiratory syndrome Coronavirus (MERS-CoV) infections in France, investigations and implications for the prevention of human-to-human transmission, France, May 2013. Eurosurveillance. 2013;18(24):2–6.
60. Tahir M, Gajraj R, Bardhan M, et al. Evidence of person-to-person transmission within a family cluster of novel coronavirus infections, United Kingdom, February 2013. Eurosurveillance. 2013;18(11):4–10.
61. Puzelli S, Azzi A, Santini MG, et al. Investigation of an imported case of Middle East respiratory syndrome Coronavirus (MERS-CoV) infection in Florence, Italy, May to June 2013. Eurosurveillance. 2013;18(34):2–5. doi:10.2807/1560-7917.ES2013.18.34.20564
62. Tsiodras S, Baka A, Mentis A, et al. A case of imported Middle East respiratory syndrome coronavirus infection and public health response, Greece, April 2014. Eurosurveillance. 2014;19(16):5–10. doi:10.2807/1560-7917.ES2014.19.16.20782
63. de Groot RJ, Baker SC, Baric RS, et al. Middle East respiratory syndrome coronavirus (MERS-CoV): announcement of the Coronavirus Study Group. J Virol. 2013;87(14):7790–7792. doi:10.1128/JVI.01244-13
64. Raj VS, Mou HH, Smits SL, et al. Dipeptidyl peptidase 4 is a functional receptor for the emerging human coronavirus-EMC. Nature. 2013;495(7440):251–254. doi:10.1038/nature12005
65. Barlan A, Zhao JC, Sarkar MK, et al. Receptor variation and susceptibility to middle east respiratory syndrome Coronavirus infection. J Virol. 2014;88(9):4953–4961. doi:10.1128/JVI.00161-14
66. Millet JK, Whittaker GR. Host cell entry of Middle East respiratory syndrome coronavirus after two-step, furin-mediated activation of the spike protein. Proc Natl Acad Sci U S A. 2014;111(42):15214–15219. doi:10.1073/pnas.1407087111
67. Chowell G, Blumberg S, Simonsen L, Miller MA, Viboud C. Synthesizing data and models for the spread of MERS-CoV, 2013: key role of index cases and hospital transmission. Epidemics. 2014;9:40–51. doi:10.1016/j.epidem.2014.09.011
68. Assiri A, Al-Tawfiq JA, Al-Rabeeah AA, et al. Epidemiological, demographic, and clinical characteristics of 47 cases of Middle East respiratory syndrome coronavirus disease from Saudi Arabia: a descriptive study. Lancet Infect Dis. 2013;13(9):752–761. doi:10.1016/S1473-3099(13)70204-4
69. Memish ZA, Al-Tawfiq JA, Makhdoom HQ, et al. Screening for Middle East respiratory syndrome coronavirus infection in hospital patients and their healthcare worker and family contacts: a prospective descriptive study. Clin Microbiol Infect. 2014;20(5):469–474. doi:10.1111/1469-0691.12562
70. Saad M, Omrani AS, Baig K, et al. Clinical aspects and outcomes of 70 patients with Middle East respiratory syndrome coronavirus infection: a single-center experience in Saudi Arabia. Int J Infect Dis. 2014;29:301–306. doi:10.1016/j.ijid.2014.09.003
71. Hayden FG, Farrar J, Peiris JS. Towards improving clinical management of Middle East respiratory syndrome coronavirus infection. Lancet Infect Dis. 2014;14(7):544–546. doi:10.1016/S1473-3099(14)70793-5
72. Guery B, Poissy J, El Mansouf L, et al. Clinical features and viral diagnosis of two cases of infection with Middle East respiratory syndrome coronavirus: a report of nosocomial transmission. Lancet. 2013;381(9885):2265–2272. doi:10.1016/S0140-6736(13)60982-4
73. Arabi YM, Arifi AA, Balkhy HH, et al. Clinical course and outcomes of critically ill patients with Middle East respiratory syndrome coronavirus infection. Ann Intern Med. 2014;160(6):389–397. doi:10.7326/M13-2486
74. Perera RA, Wang P, Gomaa MR, et al. Seroepidemiology for MERS coronavirus using microneutralisation and pseudoparticle virus neutralisation assays reveal a high prevalence of antibody in dromedary camels in Egypt, June 2013. Eurosurveillance. 2013;18(36):8–14. doi:10.2807/1560-7917.ES2013.18.36.20574
75. Reusken CBEM, Messadi L, Feyisa A, et al. Geographic distribution of MERS Coronavirus among Dromedary Camels, Africa. Emerg Infect Dis. 2014;20(8):1370–1374. doi:10.3201/eid2008.140590
76. Corman VM, Jores J, Meyer B, et al. Antibodies against MERS Coronavirus in Dromedary Camels, Kenya, 1992–2013. Emerg Infect Dis. 2014;20(8):1319–1322. doi:10.3201/eid2008.140596
77. Reusken CB, Ababneh M, Raj VS, et al. Middle East Respiratory Syndrome coronavirus (MERS-CoV) serology in major livestock species in an affected region in Jordan, June to September 2013. Eurosurveillance. 2013;18(50):14–20. doi:10.2807/1560-7917.ES2013.18.50.20662
78. Reusken CBEM, Haagmans BL, Muller MA, et al. Middle East respiratory syndrome coronavirus neutralising serum antibodies in dromedary camels: a comparative serological study. Lancet Infect Dis. 2013;13(10):859–866. doi:10.1016/S1473-3099(13)70164-6
79. Alagaili AN, Briese T, Mishra N, et al. Middle East respiratory syndrome Coronavirus infection in Dromedary Camels in Saudi Arabia. Mbio. 2014;5(2). doi:10.1128/mBio.01002-14.
80. Hemida MG, Chu DKW, Poon LLM, et al. MERS Coronavirus in Dromedary Camel Herd, Saudi Arabia. Emerg Infect Dis. 2014;20(7):1231–1234. doi:10.3201/eid2007.140571
81. Muller MA, Corman VM, Jores J, et al. MERS Coronavirus neutralizing antibodies in Camels, Eastern Africa, 1983–1997. Emerg Infect Dis. 2014;20(12):2093–2095. doi:10.3201/eid2012.141026
82. Alexandersen S, Kobinger GP, Soule G, Wernery U. Middle East Respiratory syndrome coronavirus antibody reactors among Camels in Dubai, United Arab Emirates, in 2005. Transbound Emerg Dis. 2014;61(2):105–108. doi:10.1111/tbed.12212
83. Meyer B, Muller MA, Corman VM, et al. Antibodies against MERS coronavirus in dromedary camels, United Arab Emirates, 2003 and 2013. Emerg Infect Dis. 2014;20(4):552–559. doi:10.3201/eid2004.131746
84. Peck KM, Cockrell AS, Yount BL, Scobey T, Baric RS, Heise MT. Glycosylation of mouse DPP4 plays a role in inhibiting Middle East respiratory syndrome Coronavirus infection. J Virol. 2015;89(8):4696–4699. doi:10.1128/JVI.03445-14
85. Cockrell AS, Yount BL, Scobey T, et al. A mouse model for MERS coronavirus-induced acute respiratory distress syndrome. Nat Microbiol. 2017;2(2). doi:10.1038/nmicrobiol.2016.226.
86. Zhu N, Zhang DY, Wang WL, et al. A novel Coronavirus from patients with pneumonia in China, 2019. New Engl J Med. 2020;382(8):727–733. doi:10.1056/NEJMoa2001017
87. Xu XT, Chen P, Wang JF, et al. Evolution of the novel coronavirus from the ongoing Wuhan outbreak and modeling of its spike protein for risk of human transmission. Sci China Life Sci. 2020;63(3):457–460. doi:10.1007/s11427-020-1637-5
88. Ji W, Wang W, Zhao XF, Zai JJ, Li XG. Cross-species transmission of the newly identified coronavirus 2019-nCoV. J Med Virol. 2020;92(4):433–440. doi:10.1002/jmv.25682
89. Lam TTY, Jia N, Zhang YW, et al. Identifying SARS-CoV-2-related coronaviruses in Malayan pangolins. Nature. 2020;583(7815):282. doi:10.1038/s41586-020-2169-0
90. Walls AC, Park YJ, Tortorici MA, Wall A, McGuire AT, Veesler D. Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell. 2020;181(2):281. doi:10.1016/j.cell.2020.02.058
91. Yan RH, Zhang YY, Li YN, Xia L, Guo YY, Zhou Q. Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science. 2020;367(6485):1444. doi:10.1126/science.abb2762
92. Letko M, Marzi A, Munster V. Functional assessment of cell entry and receptor usage for SARS-CoV-2 and other lineage B betacoronaviruses. Nat Microbiol. 2020;5(4):562. doi:10.1038/s41564-020-0688-y
93. Bestle D, Heindl MR, Limburg H, et al. TMPRSS2 and furin are both essential for proteolytic activation of SARS-CoV-2 in human airway cells. Life Sci Alliance. 2020;3(9):e202000786. doi:10.26508/lsa.202000786
94. Leung JM, Yang CX, Tam A, et al. ACE-2 expression in the small airway epithelia of smokers and COPD patients: implications for COVID-19. Eur Respir J. 2020;55(5):2000688. doi:10.1183/13993003.00688-2020
95. Chen L, Li XJ, Chen MQ, Feng Y, Xiong CL. The ACE2 expression in human heart indicates new potential mechanism of heart injury among patients infected with SARS-CoV-2. Cardiovasc Res. 2020;116(6):1097–1100. doi:10.1093/cvr/cvaa078
96. Rothe C, Schunk M, Sothmann P, et al. Transmission of 2019-nCoV infection from an asymptomatic contact in Germany. New Engl J Med. 2020;382(10):970–971. doi:10.1056/NEJMc2001468
97. Kampf G, Todt D, Pfaender S, Steinmann E. Persistence of coronaviruses on inanimate surfaces and their inactivation with biocidal agents (vol 104, pg 246, 2020). J Hosp Infect. 2020;105(3):587. doi:10.1016/j.jhin.2020.06.001
98. van Doremalen N, Bushmaker T, Morris DH, et al. Aerosol and surface stability of SARS-CoV-2 as compared with SARS-CoV-1. New Engl J Med. 2020;382(16):1564–1567. doi:10.1056/NEJMc2004973
99. Huang C, Wang Y, Li X. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China (vol 395, pg 497, 2020). Lancet. 2020;395(10223):496. doi:10.1016/S0140-6736(20)30183-5
100. Shi HS, Han XY, Jiang NC, et al. Radiological findings from 81 patients with COVID-19 pneumonia in Wuhan, China: a descriptive study. Lancet Infect Dis. 2020;20(4):425–434. doi:10.1016/S1473-3099(20)30086-4
101. Lu XX, Zhang LQ, Du H, et al. SARS-CoV-2 Infection in Children. New Engl J Med. 2020;382(17):1663–1665. doi:10.1056/NEJMc2005073
102. Qiu HY, Wu JH, Hong L, Luo YL, Song QF, Chen D. Clinical and epidemiological features of 36 children with coronavirus disease 2019 (COVID-19) in Zhejiang, China: an observational cohort study. Lancet Infect Dis. 2020;20(6):689–696. doi:10.1016/S1473-3099(20)30198-5
103. Zhou F, Yu T, Du RH, et al. Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: a retrospective cohort study. Lancet. 2020;395(10229):1054–1062. doi:10.1016/S0140-6736(20)30566-3
104. Blanco-Melo D, Nilsson-Payant BE, Liu WC, et al. Imbalanced host response to SARS-CoV-2 drives development of COVID-19. Cell. 2020;181(5):1036. doi:10.1016/j.cell.2020.04.026
105. Mercatelli D, Giorgi FM. Geographic and genomic distribution of SARS-CoV-2 mutations. Front Microbiol. 2020;11:1800. doi:10.3389/fmicb.2020.01800
106. Becerra-Flores M, Cardozo T. SARS-CoV-2 viral spike G614 mutation exhibits higher case fatality rate. Int J Clin Pract. 2020;74(8):e13525. doi:10.1111/ijcp.13525
107. Hu J, He C-L, Gao Q-Z, et al. D614G mutation of SARS-CoV-2 spike protein enhances viral infectivity. bioRxiv. 2020.
108. Wang ML, Cao RY, Zhang LK, et al. Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. Cell Res. 2020;30(3):269–271. doi:10.1038/s41422-020-0282-0
109. Choy KT, Wong AYL, Kaewpreedee P, et al. Remdesivir, lopinavir, emetine, and homoharringtonine inhibit SARS-CoV-2 replication in vitro. Antivir Res. 2020;178:104786. doi:10.1016/j.antiviral.2020.104786
110. Grein J, Ohmagari N, Shin D, et al. Compassionate use of remdesivir for patients with severe Covid-19. New Engl J Med. 2020;382(24):2327–2336. doi:10.1056/NEJMoa2007016
111. Wang YM, Zhang DY, Du GH, et al. Remdesivir in adults with severe COVID-19: a randomised, double-blind, placebo-controlled, multicentre trial. Lancet. 2020;395(10236):1569–1578. doi:10.1016/S0140-6736(20)31022-9
112. Yao X, Ye F, Zhang M, et al. In vitro antiviral activity and projection of optimized dosing design of hydroxychloroquine for the treatment of severe acute respiratory syndrome Coronavirus 2 (SARS-CoV-2). Clin Infect Dis. 2020;71(15):732–739.
113. Chen J, Liu D, Liu L, et al. [A pilot study of hydroxychloroquine in treatment of patients with moderate COVID-19]. Zhejiang Da Xue Xue Bao Yi Xue Ban. 2020;49(2):215–219. Chinese. doi:10.3785/j.issn.1008-9292.2020.03.03
114. Borba MGS, Val FFA, Sampaio VS, et al. Effect of high vs low doses of chloroquine diphosphate as adjunctive therapy for patients hospitalized with severe acute respiratory syndrome Coronavirus 2 (SARS-CoV-2) infection a randomized clinical trial. JAMA Netw Open. 2020;3(4). doi:10.1001/jamanetworkopen.2020.8857.
115. Chu CM, Cheng VCC, Hung IFN, et al. Role of lopinavir/ritonavir in the treatment of SARS: initial virological and clinical findings. Thorax. 2004;59(3):252–256. doi:10.1136/thorax.2003.012658
116. Xia S, Zhu Y, Liu MQ, et al. Fusion mechanism of 2019-nCoV and fusion inhibitors targeting HR1 domain in spike protein. Cell Mol Immunol. 2020;17(7):765–767. doi:10.1038/s41423-020-0374-2
117. Wrapp D, De Vlieger D, Corbett KS, et al. Structural basis for potent neutralization of betacoronaviruses by single-domain camelid antibodies. Cell. 2020;181(5):1004. doi:10.1016/j.cell.2020.04.031
118. Wang CY, Li WT, Drabek D, et al. A human monoclonal antibody blocking SARS-CoV-2 infection. Nat Commun. 2020;11(1).
119. Pinto D, Park YJ, Beltramello M, et al. Cross-neutralization of SARS-CoV-2 by a human monoclonal SARS-CoV antibody. Nature. 2020;583(7815):290–295. doi:10.1038/s41586-020-2349-y
120. Tanaka T, Narazaki M, Kishimoto T. Immunotherapeutic implications of IL-6 blockade for cytokine storm. Immunotherapy. 2016;8(8):959–970. doi:10.2217/imt-2016-0020
121. Schrezenmeier E, Dorner T. Mechanisms of action of hydroxychloroquine and chloroquine: implications for rheumatology. Nat Rev Rheumatol. 2020;16(3):155–166. doi:10.1038/s41584-020-0372-x
122. Della-Torre E, Campochiaro C, Cavalli G, et al. Interleukin-6 blockade with sarilumab in severe COVID-19 pneumonia with systemic hyperinflammation: an open-label cohort study. Ann Rheum Dis. 2020;79(10):1277–1285. doi:10.1136/annrheumdis-2020-218122
123. Zhai P, Ding YB, Wu X, Long JK, Zhong YJ, Li YM. The epidemiology, diagnosis and treatment of COVID-19. Int J Antimicrob Agents. 2020;55(5):105955. doi:10.1016/j.ijantimicag.2020.105955
124. Zhou P, Yang XL, Wang XG, et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020;579(7798):270. doi:10.1038/s41586-020-2012-7
125. Shen CG, Wang ZQ, Zhao F, et al. Treatment of 5 critically Ill patients with COVID-19 with convalescent plasma. JAMA. 2020;323(16):1582–1589. doi:10.1001/jama.2020.4783
126. Wrapp D, Wang NS, Corbett KS, et al. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science. 2020;367(6483):1260. doi:10.1126/science.abb2507
127. Andersen KG, Rambaut A, Lipkin WI, Holmes EC, Garry RF. The proximal origin of SARS-CoV-2. Nat Med. 2020;26(4):450–452. doi:10.1038/s41591-020-0820-9
128. Benvenuto D, Giovanetti M, Ciccozzi A, Spoto S, Angeletti S, Ciccozzi M. The 2019-new coronavirus epidemic: evidence for virus evolution. J Med Virol. 2020;92(4):455–459. doi:10.1002/jmv.25688
129. Yuan M, Wu NC, Zhu XY, et al. A highly conserved cryptic epitope in the receptor binding domains of SARS-CoV-2 and SARS-CoV. Science. 2020;368(6491):630. doi:10.1126/science.abb7269
130. Enjuanes L, Zuniga S, Castano-Rodriguez C, Gutierrez-Alvarez J, Canton J, Sola I. Molecular basis of coronavirus virulence and vaccine development. Adv Virus Res. 2016;96:245–286.
131. Song ZQ, Xu YF, Bao LL, et al. From SARS to MERS, thrusting coronaviruses into the spotlight. Viruses Basel. 2019;11(1):59. doi:10.3390/v11010059
132. Thao TTN, Labroussaa F, Ebert N, et al. Rapid reconstruction of SARS-CoV-2 using a synthetic genomics platform. Nature. 2020.
133. Xie XP, Muruato A, Lokugamage KG, et al. An infectious cDNA clone of SARS-CoV-2. Cell Host Microbe. 2020;27(5):841.
134. Chen WH, Strych U, Hotez PJ, Bottazzi ME. The SARS-CoV-2 vaccine pipeline: an overview. Curr Trop Med Rep. 2020;7(2):61–64. doi:10.1007/s40475-020-00201-6
135. Bull JJ, Nuismer SL, Antia R. Recombinant vector vaccine evolution. PLoS Comput Biol. 2019;15(7):e1006857. doi:10.1371/journal.pcbi.1006857
136. Le TT, Andreadakis Z, Kumar A, et al. The COVID-19 vaccine development landscape. Nat Rev Drug Discov. 2020;19(5):305–306. doi:10.1038/d41573-020-00151-8
137. Gomez CE, Najera JL, Krupa M, Perdiguero B, Esteban M. MVA and NYVAC as vaccines against emergent infectious diseases and cancer. Curr Gene Ther. 2011;11(3):189–217. doi:10.2174/156652311795684731
138. Sutter G, Staib C. Vaccinia vectors as candidate vaccines: the development of modified vaccinia virus Ankara for antigen delivery. Curr Drug Targets Infect Disord. 2003;3(3):263–271. doi:10.2174/1568005033481123
139. Frey SE, Winokur PL, Salata RA, et al. Safety and immunogenicity of IMVAMUNE (R) smallpox vaccine using different strategies for a post event scenario. Vaccine. 2013;31(29):3025–3033. doi:10.1016/j.vaccine.2013.04.050
140. Sullivan NJ, Sanchez A, Rollin PE, Yang ZY, Nabel GJ. Development of a preventive vaccine for Ebola virus infection in primates. Nature. 2000;408(6812):605–609. doi:10.1038/35046108
141. Shiver JW, Fu TM, Chen L, et al. Replication-incompetent adenoviral vaccine vector elicits effective anti-immunodeficiency-virus immunity. Nature. 2002;415(6869):331–335. doi:10.1038/415331a
142. Urabe M, Ding CT, Kotin RM. Insect cells as a factory to produce adeno-associated virus type 2 vectors. Hum Gene Ther. 2002;13(16):1935–1943. doi:10.1089/10430340260355347
143. Kotin RM. Large-scale recombinant adeno-associated virus production. Hum Mol Genet. 2011;20:R2–R6. doi:10.1093/hmg/ddr141
144. Flotte TR. Gene therapy progress and prospects: recombinant adeno-associated virus (rAAV) vectors. Gene Ther. 2004;11(10):805–810. doi:10.1038/sj.gt.3302233
145. Monahan PE, Samulski RJ. AAV vectors: is clinical success on the horizon? Gene Ther. 2000;7(1):24–30. doi:10.1038/sj.gt.3301109
146. Choi VW, McCarty DM, Samulski RJ. AAV hybrid serotypes: improved vectors for gene delivery. Curr Gene Ther. 2005;5(3):299–310. doi:10.2174/1566523054064968
147. Li HJ, Malani N, Hamilton SR, et al. Assessing the potential for AAV vector genotoxicity in a murine model. Blood. 2011;117(12):3311–3319. doi:10.1182/blood-2010-08-302729
148. Folegatti PM, Ewer KJ, Aley PK, et al. Safety and immunogenicity of the ChAdOx1 nCoV-19 vaccine against SARS-CoV-2: a preliminary report of a Phase 1/2, single-blind, randomised controlled trial. Lancet. 2020;396(10249):467–478. doi:10.1016/S0140-6736(20)31604-4
149. Yin H, Kanasty RL, Eltoukhy AA, Vegas AJ, Dorkin JR, Anderson DG. Non-viral vectors for gene-based therapy. Nat Rev Genet. 2014;15(8):541–555. doi:10.1038/nrg3763
150. Vincent M, de Lazaro I, Kostarelos K. Graphene materials as 2D non-viral gene transfer vector platforms. Gene Ther. 2017;24(3):123–132. doi:10.1038/gt.2016.79
151. Gregory AE, Titball R, Williamson D. Vaccine delivery using nanoparticles. Front Cell Infect Micrbiol. 2013;3.
152. Panyam J, Labhasetwar V. Biodegradable nanoparticles for drug and gene delivery to cells and tissue. Adv Drug Deliver Rev. 2003;55(3):329–347. doi:10.1016/S0169-409X(02)00228-4
153. Shen YB, Hao TY, Ou SY, Hu CR, Chen L. Applications and perspectives of nanomaterials in novel vaccine development. Med Chem Comm. 2018;9(2):226–238. doi:10.1039/C7MD00158D
154. Qi F, Wu J, Li H, Ma GH. Recent research and development of PLGA/PLA microspheres/nanoparticles: a review in scientific and industrial aspects. Front Chem Sci Eng. 2019;13(1):14–27. doi:10.1007/s11705-018-1729-4
155. Sainz V, Conniot J, Matos AI, et al. Regulatory aspects on nanomedicines. Biochem Biophys Res Commun. 2015;468(3):504–510. doi:10.1016/j.bbrc.2015.08.023
156. Mograo J, da Costa CA, Gaspar R, Florindo HF. Modulation of dendritic cells by nanotechnology-based immunotherapeutic strategies. J Biomed Nanotechnol. 2016;12(3):405–434. doi:10.1166/jbn.2016.2157
157. Bonam SR, Kotla NG, Bohara RA, Rochev Y, Webster TJ, Bayry J. Potential immuno-nanomedicine strategies to fight COVID-19 like pulmonary infections. Nano Today. 2021;36:101051. doi:10.1016/j.nantod.2020.101051
158. Barik S. New treatments for influenza. BMC Med. 2012;10. doi:10.1186/1741-7015-10-104
159. Levina AS, Repkova MN, Mazurkova NA, Zarytova VF. Nanoparticle-mediated nonviral DNA delivery for effective inhibition of influenza a viruses in cells. Ieee T Nanotechnol. 2016;15(2):248–254. doi:10.1109/TNANO.2016.2516561
160. Hendricks GL, Weirich KL, Viswanathan K, et al. Sialylneolacto-N-tetraose c (LSTc)-bearing liposomal decoys capture influenza A virus. J Biol Chem. 2013;288(12):8061–8073. doi:10.1074/jbc.M112.437202
161. Dunning J, Sahr F, Rojek A, et al. Experimental treatment of ebola virus disease with TKM-130803: a single-arm Phase 2 clinical trial. PLoS Med. 2016;13(4):e1001997. doi:10.1371/journal.pmed.1001997
162. Pereira de Oliveira M, Garcion E, Venisse N, BenoIt JP, Couet W, Olivier JC. Tissue distribution of indinavir administered as solid lipid nanocapsule formulation in mdr1a (+/+) and mdr1a (-/-) CF-1 mice. Pharm Res. 2005;22(11):1898–1905. doi:10.1007/s11095-005-7147-6
163. Rodriguez B, Asmuth DM, Matining RM, et al. Safety, tolerability, and immunogenicity of repeated doses of DermaVir, a candidate therapeutic HIV vaccine, in HIV-infected patients receiving combination antiretroviral therapy: results of the ACTG 5176 trial. J Acquir Immune Defic. 2013;64(4):351–359. doi:10.1097/QAI.0b013e3182a99590
164. Orkin C, Squires KE, Molina JM, et al. Doravirine/Lamivudine/Tenofovir disoproxil fumarate is non-inferior to Efavirenz/Emtricitabine/Tenofovir Disoproxil fumarate in treatment-naive adults with human immunodeficiency Virus-1 infection: week 48 results of the DRIVE-AHEAD trial. Clin Infect Dis. 2019;68(4):535–544. doi:10.1093/cid/ciy540
165. Price CF, Tyssen D, Sonza S, et al. SPL7013 gel (VivaGel (R)) retains potent HIV-1 and HSV-2 inhibitory activity following vaginal administration in humans. PLoS One. 2011;6(9):e24095. doi:10.1371/journal.pone.0024095
166. Cavalli R, Donalisio M, Bisazza A, et al. Enhanced antiviral activity of acyclovir loaded into nanoparticles. Nanomedicine. 2012;509:1–19.
167. Lembo D, Swaminathan S, Donalisio M, et al. Encapsulation of Acyclovir in new carboxylated cyclodextrin- based nanosponges improves the agent’s antiviral efficacy. Int J Pharm. 2013;443(1–2):262–272. doi:10.1016/j.ijpharm.2012.12.031
168. Hu RL, Li SR, Kong FJ, Hou RJ, Guan XL, Guo F. Inhibition effect of silver nanoparticles on herpes simplex virus 2. Genet Mol Res. 2014;13(3):7022–7028. doi:10.4238/2014.March.19.2
169. Rupp R, Rosenthal SL, Stanberry LR. VivaGel (TM) (SPL7013 Gel): a candidate dendrimer microbicide for the prevention of HIV and HSV infection. Int J Nanomed. 2007;2(4):561–566.
170. Wieland SF, Vega RG, Muller R, et al. Searching for interferon-induced genes that inhibit hepatitis B virus replication in transgenic mouse hepatocytes. J Virol. 2003;77(2):1227–1236. doi:10.1128/JVI.77.2.1227-1236.2003
171. Liang TJ, Block TM, McMahon BJ, et al. Present and future therapies of Hepatitis B: from discovery to cure. Hepatology. 2015;62(6):1893–1908. doi:10.1002/hep.28025
172. Lingala S, Lau DTY, Koh C, Auh S, Ghany MG, Hoofnagle JH. Long-term lamivudine therapy in chronic hepatitis B. Aliment Pharm Ther. 2016;44(4):380–389. doi:10.1111/apt.13707
173. Marcellin P, Chang TT, Lim SG, et al. Adefovir dipivoxil for the treatment of hepatitis B e antigen-positive chronic hepatitis B. N Engl J Med. 2003;348(9):808–816. doi:10.1056/NEJMoa020681
174. Dimou E, Papadimitropoulos V, Hadziyannis SJ. The role of entecavir in the treatment of chronic hepatitis B. Ther Clin Risk Manag. 2007;3(6):1077–1086.
175. Matthews SJ. Telbivudine for the management of chronic hepatitis B virus infection. Clin Ther. 2007;29(12):2635–2653. doi:10.1016/j.clinthera.2007.12.032
176. Manzoor S, Saalim M, Imran M, Resham S, Ashraf J. Hepatitis B virus therapy: what’s the future holding for us? World J Gastroenterol. 2015;21(44):12558–12575. doi:10.3748/wjg.v21.i44.12558
177. Szunerits S, Barras A, Khanal M, Pagneux Q, Boukherroub R. Nanostructures for the inhibition of viral infections. Molecules. 2015;20(8):14051–14081. doi:10.3390/molecules200814051
178. Broglie JJ, Alston B, Yang C, et al. Antiviral Activity of Gold/Copper Sulfide Core/Shell nanoparticles against human Norovirus virus-like particles. PLoS One. 2015;10(10):e0141050. doi:10.1371/journal.pone.0141050
179. Lammers T, Sofias AM, van der Meel R, et al. Dexamethasone nanomedicines for COVID-19. Nat Nanotechnol. 2020;15(8):622–624.
180. Alshweiat A, Ambrus R, Csoka I. Intranasal nanoparticulate systems as alternative route of drug delivery. Curr Med Chem. 2019;26(35):6459–6492.
181. Costantino HR, Illum L, Brandt G, Johnson PH, Quay SC. Intranasal delivery: physicochemical and therapeutic aspects. Int J Pharm. 2007;337(1–2):1–24. doi:10.1016/j.ijpharm.2007.03.025
182. Marasini N, Kaminskas LM. Subunit-based mucosal vaccine delivery systems for pulmonary delivery - are they feasible? Drug Dev Ind Pharm. 2019;45(6):882–894. doi:10.1080/03639045.2019.1583758
183. Kumar A, Pandey AN, Jain SK. Nasal-nanotechnology: revolution for efficient therapeutics delivery. Drug Deliv. 2016;23(3):681–693. doi:10.3109/10717544.2014.920431
184. Alyane M, Barratt G, Lahouel M. Remote loading of doxorubicin into liposomes by transmembrane pH gradient to reduce toxicity toward H9c2 cells. Saudi Pharm J. 2016;24(2):165–175. doi:10.1016/j.jsps.2015.02.014
185. Bozzuto G, Molinari A. Liposomes as nanomedical devices. Int J Nanomed. 2015;10:975–999. doi:10.2147/IJN.S68861
186. Rodrigues C, Gameiro P, Prieto M, de Castro B. Interaction of rifampicin and isoniazid with large unilamellar liposomes: spectroscopic location studies. Bba Gen Subj. 2003;1620(1–3):151–159. doi:10.1016/S0304-4165(02)00528-7
187. Zou Y, Lee HY, Seo YC, Ahn J. Enhanced antimicrobial activity of nisin-loaded liposomal nanoparticles against foodborne pathogens. J Food Sci. 2012;77(3):M165–M170. doi:10.1111/j.1750-3841.2011.02580.x
188. Del Amo EM, Rimpela AK, Heikkinen E, et al. Pharmacokinetic aspects of retinal drug delivery. Prog Retin Eye Res. 2017;57:134–185. doi:10.1016/j.preteyeres.2016.12.001
189. Pattni BS, Chupin VV, Torchilin VP. New developments in liposomal drug delivery. Chem Rev. 2015;115(19):10938–10966. doi:10.1021/acs.chemrev.5b00046
190. Zuidam NJ, Lee SS, Crommelin DJ. Sterilization of liposomes by heat treatment. Pharm Res. 1993;10(11):1591–1596. doi:10.1023/A:1018916518515
191. Grit M, Crommelin JA. Chemical-stability of liposomes - implications for their physical stability. Chem Phys Lipids. 1993;64(1–3):3–18. doi:10.1016/0009-3084(93)90053-6
192. Ishida T, Harashima H, Kiwada H. Liposome clearance. Biosci Rep. 2002;22(2):197–224. doi:10.1023/A:1020134521778
193. LaBauve AE, Rinker TE, Noureddine A, et al. Lipid-coated mesoporous silica nanoparticles for the delivery of the ML336 antiviral to inhibit encephalitic alphavirus infection. Sci Rep. 2018;8.
194. Draz MS, Fang BA, Zhang PF, et al. Nanoparticle-mediated systemic delivery of siRNA for treatment of cancers and viral infections. Theranostics. 2014;4(9):872–892. doi:10.7150/thno.9404
195. Davis ME, Zuckerman JE, Choi CHJ, et al. Evidence of RNAi in humans from systemically administered siRNA via targeted nanoparticles. Nature. 2010;464(7291):1067–U1140. doi:10.1038/nature08956
196. Ku SH, Jo SD, Lee YK, Kim K, Kim SH. Chemical and structural modifications of RNAi therapeutics. Adv Drug Deliver Rev. 2016;104:16–28. doi:10.1016/j.addr.2015.10.015
197. Adams D, Gonzalez-Duarte A, O’Riordan WD, et al. Patisiran, an RNAi therapeutic, for hereditary transthyretin amyloidosis. New Engl J Med. 2018;379(1):11–21. doi:10.1056/NEJMoa1716153
198. Sonaje K, Chuang EY, Lin KJ, et al. Opening of epithelial tight junctions and enhancement of paracellular permeation by chitosan: microscopic, ultrastructural, and computed-tomographic observations. Mol Pharm. 2012;9(5):1271–1279. doi:10.1021/mp200572t
199. Chua BY, Al Kobaisi M, Zeng WG, Mainwaring D, Jackson DC. Chitosan microparticles and nanoparticles as biocompatible delivery vehicles for peptide and protein-based immunocontraceptive vaccines. Mol Pharm. 2012;9(1):81–90. doi:10.1021/mp200264m
200. Qi RG, Wang YH, Bruno PM, et al. Nanoparticle conjugates of a highly potent toxin enhance safety and circumvent platinum resistance in ovarian cancer. Nat Commun. 2017;8. doi:10.1038/s41467-017-02390-7
201. Ashton S, Song YH, Nolan J, et al. Aurora kinase inhibitor nanoparticles target tumors with favorable therapeutic index in vivo. Sci Transl Med. 2016;8(325):325ra17. doi:10.1126/scitranslmed.aad2355
202. Kim Y, Park EJ, Na DH. Recent progress in dendrimer-based nanomedicine development. Arch Pharm Res. 2018;41(6):571–582. doi:10.1007/s12272-018-1008-4
203. Ahmad Z, Shah A, Siddiq M, Kraatz HB. Polymeric micelles as drug delivery vehicles. RSC Adv. 2014;4(33):17028–17038. doi:10.1039/C3RA47370H
204. Bao G, Mitragotri S, Tong S. Multifunctional nanoparticles for drug delivery and molecular imaging. Annu Rev Biomed Eng. 2013;15:253–282. doi:10.1146/annurev-bioeng-071812-152409
205. Vauthier C, Bouchemal K. Methods for the preparation and manufacture of polymeric nanoparticles. Pharm Res. 2009;26(5):1025–1058.
206. Rao JP, Geckeler KE. Polymer nanoparticles: preparation techniques and size-control parameters. Prog Polym Sci. 2011;36(7):887–913. doi:10.1016/j.progpolymsci.2011.01.001
207. Torchilin VP. Multifunctional, stimuli-sensitive nanoparticulate systems for drug delivery. Nat Rev Drug Discov. 2014;13(11):813–827. doi:10.1038/nrd4333
208. He J, Perez MT, Zhang P, et al. A general approach to synthesize asymmetric hybrid nanoparticles by interfacial reactions. J Am Chem Soc. 2012;134(8):3639–3642. doi:10.1021/ja210844h
209. Lee CC, MacKay JA, Frechet JMJ, Szoka FC. Designing dendrimers for biological applications. Nat Biotechnol. 2005;23(12):1517–1526. doi:10.1038/nbt1171
210. Singh I, Rehni AK, Kalra R, Joshi G, Kumar M. Dendrimers and their pharmaceutical applications–a review. Pharmazie. 2008;63(7):491–496.
211. Pasut G, Scaramuzza S, Schiavon O, Mendichi R, Veronese FM. PEG-epirubicin conjugates with high drug loading. J Bioact Compat Polym. 2005;20(3):213–230. doi:10.1177/0883911505053377
212. Tomalia DA, Reyna LA, Svenson S. Dendrimers as multi-purpose nanodevices for oncology drug delivery and diagnostic imaging. Biochem Soc T. 2007;35:61–67. doi:10.1042/BST0350061
213. Saraf S. Process optimization for the production of nanoparticles for drug delivery applications. Expert Opin Drug Deliv. 2009;6(2):187–196. doi:10.1517/17425240902735806
214. Jesus S, Schmutz M, Som C, Borchard G, Wick P, Borges O. Hazard assessment of polymeric nanobiomaterials for drug delivery: what can we learn from literature so far. Front Bioeng Biotech. 2019;7. doi:10.3389/fbioe.2019.00261
215. Stiriba SE, Frey H, Haag R. Dendritic polymers in biomedical applications: from potential to clinical use in diagnostics and therapy. Angew Chem Int Ed Engl. 2002;41(8):1329–1334. doi:10.1002/1521-3773(20020415)41:8<1329::AID-ANIE1329>3.0.CO;2-P
216. Salazar-Gonzalez JA, Gonzalez-Ortega O, Rosales-Mendoza S. Gold nanoparticles and vaccine development. Expert Rev Vaccines. 2015;14(9):1197–1211. doi:10.1586/14760584.2015.1064772
217. Neto LMM, Kipnis A, Junqueira-Kipnis AP. Role of metallic nanoparticles in vaccinology: implications for infectious disease vaccine development. Front Immunol. 2017;8.
218. Ding Y, Jiang Z, Saha K, et al. Gold nanoparticles for nucleic acid delivery. Mol Ther. 2014;22(6):1075–1083. doi:10.1038/mt.2014.30
219. Roca M, Haes AJ. Probing cells with noble metal nanoparticle aggregates. Nanomedicine (Lond). 2008;3(4):555–565. doi:10.2217/17435889.3.4.555
220. Gustafson HH, Holt-Casper D, Grainger DW, Ghandehari H. Nanoparticle uptake: the phagocyte problem. Nano Today. 2015;10(4):487–510. doi:10.1016/j.nantod.2015.06.006
221. Buzea C, Pacheco II, Robbie K. Nanomaterials and nanoparticles: sources and toxicity. Biointerphases. 2007;2(4):Mr17–Mr71. doi:10.1116/1.2815690
222. Bansal SA, Kumar V, Karimi J, Singh AP, Kumar S. Role of gold nanoparticles in advanced biomedical applications. Nanoscale Adv. 2020;2(9):3764–3787.
223. Kato T, Takami Y, Deo VK, Park EY. Preparation of virus-like particle mimetic nanovesicles displaying the S protein of Middle East respiratory syndrome coronavirus using insect cells. J Biotechnol. 2019;306:177–184. doi:10.1016/j.jbiotec.2019.10.007
224. Murata K, Lechmann M, Qiao M, Gunji T, Alter HJ, Liang TJ. Immunization with hepatitis C virus-like particles protects mice from recombinant hepatitis C virus-vaccinia infection. Proc Natl Acad Sci U S A. 2003;100(11):6753–6758. doi:10.1073/pnas.1131929100
225. Quan FS, Compans RW, Nguyen HH, Kang SM. Induction of heterosubtypic immunity to influenza virus by intranasal immunization. J Virol. 2008;82(3):1350–1359. doi:10.1128/JVI.01615-07
226. Lee YT, Ko EJ, Lee Y, et al. Intranasal vaccination with M2e5x virus-like particles induces humoral and cellular immune responses conferring cross-protection against heterosubtypic influenza viruses. PLoS One. 2018;13(1).
227. Bundy BC, Swartz JR. Efficient disulfide bond formation in virus-like particles. J Biotechnol. 2011;154(4):230–239. doi:10.1016/j.jbiotec.2011.04.011
228. Lu Y, Chan W, Ko BY, VanLang CC, Swartz JR. Assessing sequence plasticity of a virus-like nanoparticle by evolution toward a versatile scaffold for vaccines and drug delivery. Proc Natl Acad Sci U S A. 2015;112(40):12360–12365. doi:10.1073/pnas.1510533112
229. Bundy BC, Franciszkowicz MJ, Swartz JR. Escherichia coli-based cell-free synthesis of virus-like particles. Biotechnol Bioeng. 2008;100(1):28–37. doi:10.1002/bit.21716
230. Ashley CE, Carnes EC, Phillips GK, et al. Cell-Specific delivery of diverse cargos by bacteriophage MS2 virus-like particles. ACS Nano. 2011;5(7):5729–5745. doi:10.1021/nn201397z
231. Hovlid ML, Lau JL, Breitenkamp K, et al. Encapsidated atom-transfer radical polymerization in Q beta virus-like nanoparticles. ACS Nano. 2014;8(8):8003–8014. doi:10.1021/nn502043d
232. Peyret H, Gehin A, Thuenemann EC, et al. Tandem fusion of hepatitis B core antigen allows assembly of virus-like particles in bacteria and plants with enhanced capacity to accommodate foreign proteins. PLoS One. 2015;10(4):e0120751. doi:10.1371/journal.pone.0120751
233. Patel KG, Swartz JR. Surface functionalization of virus-like particles by direct conjugation using azide-alkyne click chemistry. Bioconjug Chem. 2011;22(3):376–387. doi:10.1021/bc100367u
234. Zepeda-Cervantes J, Ramirez-Jarquin JO, Vaca L. Interaction between virus-like particles (VLPs) and pattern recognition receptors (PRRs) from dendritic cells (DCs): toward better engineering of VLPs. Front Immunol. 2020;11.
235. Champion JA, Katare YK, Mitragotri S. Particle shape: a new design parameter for micro- and nanoscale drug delivery carriers. J Control Release. 2007;121(1–2):3–9. doi:10.1016/j.jconrel.2007.03.022
236. Henry BD, Neill DR, Becker KA, et al. Engineered liposomes sequester bacterial exotoxins and protect from severe invasive infections in mice. Nat Biotechnol. 2015;33(1):81–88. doi:10.1038/nbt.3037
237. Hu CMJ, Fang RH, Copp J, Luk BT, Zhang LF. A biomimetic nanosponge that absorbs pore-forming toxins. Nat Nanotechnol. 2013;8(5):336–340. doi:10.1038/nnano.2013.54
238. Keller MD, Ching KL, Liang FX, et al. Decoy exosomes provide protection against bacterial toxins. Nature. 2020;579(7798):260. doi:10.1038/s41586-020-2066-6
239. Zhang PF, Chen YX, Zeng Y, et al. Virus-mimetic nanovesicles as a versatile antigen-delivery system. Proc Natl Acad Sci U S A. 2015;112(45):E6129–E6138. doi:10.1073/pnas.1505799112
240. Wang J, Li P, Yu Y, et al. Pulmonary surfactant-biomimetic nanoparticles potentiate heterosubtypic influenza immunity. Science. 2020;367(6480):eaau0810. doi:10.1126/science.aau0810
241. Zhang PF, Zhang L, Qin ZE, et al. Genetically engineered liposome-like nanovesicles as active targeted transport platform. Adv Mater. 2018;30(7).
242. Rao L, Xia S, Xu W, et al. Decoy nanoparticles protect against COVID-19 by concurrently adsorbing viruses and inflammatory cytokines. Proc Natl Acad Sci U S A. 2020;117(44):27141–27147. doi:10.1073/pnas.2014352117
243. Zhang Q, Honko A, Zhou J, et al. Cellular nanosponges inhibit SARS-CoV-2 infectivity. Nano Lett. 2020;20(7):5570–5574. doi:10.1021/acs.nanolett.0c02278
244. Patil AA, Rhee WJ. Exosomes: biogenesis, composition, functions, and their role in pre-metastatic niche formation. Biotechnol Bioproc E. 2019;24(5):689–701. doi:10.1007/s12257-019-0170-y
245. Kuate S, Cinatl J, Doerr HW, Uberla K. Exosomal vaccines containing the S protein of the SARS coronavirus induce high levels of neutralizing antibodies. Virology. 2007;362(1):26–37. doi:10.1016/j.virol.2006.12.011
246. Thirabanjasak D, Tantiwongse K, Thorner PS. Angiomyeloproliferative lesions following autologous stem cell therapy. J Am Soc Nephrol. 2010;21(7):1218–1222. doi:10.1681/ASN.2009111156
247. Buyanovskaya OA, Kuleshov NP, Nikitina VA, Voronina ES, Katosova LD, Bochkov NP. Spontaneous aneuploidy and clone formation in adipose tissue stem cells during different periods of culturing. Bull Exp Biol Med. 2009;148(1):109–112. doi:10.1007/s10517-009-0647-3
248. Bruno S, Grange C, Deregibus MC, et al. Mesenchymal stem cell-derived microvesicles protect against acute tubular injury. J Am Soc Nephrol. 2009;20(5):1053–1067. doi:10.1681/ASN.2008070798
249. Baj-Krzyworzeka M, Majka M, Pratico D, et al. Platelet-derived microparticles stimulate proliferation, survival, adhesion, and chemotaxis of hematopoietic cells. Exp Hematol. 2002;30(5):450–459. doi:10.1016/S0301-472X(02)00791-9
250. Gasser O, Hess C, Miot S, Deon C, Sanchez JC, Schifferli JA. Characterisation and properties of ectosomes released by human polymorphonuclear neutrophils. Exp Cell Res. 2003;285(2):243–257. doi:10.1016/S0014-4827(03)00055-7
251. Horstman LL, Jy W, Jimenez JJ, Bidot C, Ahn YS. New horizons in the analysis of circulating cell-derived microparticles. Keio J Med. 2004;53(4):210–230. doi:10.2302/kjm.53.210
252. Hotary KB, Allen ED, Brooks PC, Datta NS, Long MW, Weiss SJ. Membrane type I matrix metalloproteinase usurps tumor growth control imposed by the three-dimensional extracellular matrix. Cell. 2003;114(1):33–45. doi:10.1016/S0092-8674(03)00513-0
253. Safaei R, Larson BJ, Cheng TC, et al. Abnormal lysosomal trafficking and enhanced exosomal export of cisplatin in drug-resistant human ovarian carcinoma cells. Mol Cancer Ther. 2005;4(10):1595–1604. doi:10.1158/1535-7163.MCT-05-0102
254. Shedden K, Xie XT, Chandaroy P, Chang YT, Rosania GR. Expulsion of small molecules in vesicles shed by cancer cells: association with gene expression and chemosensitivity profiles. Cancer Res. 2003;63(15):4331–4337.
255. Yang W, Peters JI, Williams RO
256. Patton JS, Byron PR. Inhaling medicines: delivering drugs to the body through the lungs. Nat Rev Drug Discov. 2007;6(1):67–74. doi:10.1038/nrd2153
257. Cavalcanti IDL, de Fatima Ramos Dos Santos Medeiros SM, Dos Santos Macedo DC, Ferro Cavalcanti IM, de Britto Lira Nogueira MC. Nanocarriers in the delivery of hydroxychloroquine to the respiratory system: an alternative to COVID-19? Curr Drug Deliv. 2020;17. doi:10.2174/1567201817666200827110445
258. Zeng XM, Martin GP, Marriott C. The controlled delivery of drugs to the lung. Int J Pharm. 1995;124(2):149–164. doi:10.1016/0378-5173(95)00104-Q
259. Justo OR, Moraes AM. Incorporation of antibiotics in liposomes designed for tuberculosis therapy by inhalation. Drug Deliv. 2003;10(3):201–207. doi:10.1080/713840401
260. Schreier H, Gonzalezrothi RJ, Stecenko AA. Pulmonary delivery of liposomes. J Control Release. 1993;24(1–3):209–223. doi:10.1016/0168-3659(93)90180-D
261. Taylor KMG, Taylor G, Kellaway IW, Stevens J. The stability of liposomes to nebulization. Int J Pharm. 1990;58(1):57–61. doi:10.1016/0378-5173(90)90287-E
262. Bi R, Shao W, Wang Q, Zhang N. Spray-freeze-dried dry powder inhalation of insulin-loaded liposomes for enhanced pulmonary delivery. J Drug Target. 2008;16(9):639–648. doi:10.1080/10611860802201134
263. White S, Bennett DB, Cheu S, et al. EXUBERA: pharmaceutical development of a novel product for pulmonary delivery of insulin. Diabetes Technol Ther. 2005;7(6):896–906. doi:10.1089/dia.2005.7.896
264. Joshi M, Misra AN. Pulmonary disposition of budesonide from liposomal dry powder inhaler. Methods Find Exp Clin Pharmacol. 2001;23(10):531–536. doi:10.1358/mf.2001.23.10.677118
265. Cryan SA, Devocelle M, Moran PJ, Hickey AJ, Kelly JG. Increased intracellular targeting to airway cells using octaarginine-coated liposomes: in vitro assessment of their suitability for inhalation. Mol Pharm. 2006;3(2):104–112. doi:10.1021/mp050070i
266. Almeida AJ, Souto E. Solid lipid nanoparticles as a drug delivery system for peptides and proteins. Adv Drug Deliver Rev. 2007;59(6):478–490. doi:10.1016/j.addr.2007.04.007
267. Densmore CL. Advances in noninvasive pulmonary gene therapy. Curr Drug Deliv. 2006;3(1):55–63. doi:10.2174/156720106775197547
268. De Smedt SC, Demeester J, Hennink WE. Cationic polymer based gene delivery systems. Pharm Res. 2000;17(2):113–126. doi:10.1023/A:1007548826495
269. Boas U, Heegaard PMH. Dendrimers in drug research. Chem Soc Rev. 2004;33(1):43–63. doi:10.1039/b309043b
270. Bai SH, Thomas C, Ahsan F. Dendrimers as a carrier for pulmonary delivery of enoxaparin, a low-molecular weight heparin. J Pharm Sci. 2007;96(8):2090–2106. doi:10.1002/jps.20849
271. Rudolph C, Lausier J, Naundorf S, Muller RH, Rosenecker J. In vivo gene delivery to the lung using polyethylenimine and fractured polyamidoamine dendrimers. J Gene Med. 2000;2(4):269–278. doi:10.1002/1521-2254(200007/08)2:4<269::AID-JGM112>3.0.CO;2-F
272. Weiss C, Carriere M, Fusco L, et al. Toward nanotechnology-enabled approaches against the COVID-19 pandemic. ACS Nano. 2020;14(6):6383–6406. doi:10.1021/acsnano.0c03697
273. Cavalcanti IDL, Nogueira MCDL. Pharmaceutical nanotechnology: which products are been designed against COVID-19? J Nanopart Res. 2020;22(9). doi:10.1007/s11051-020-05010-6
274. Ivanov YD, Malsagova KA, Pleshakova TO, et al. Ultrasensitive detection of 2,4-dinitrophenol using nanowire biosensor. J Nanotechnol. 2018;2018.
275. Seo G, Lee G, Kim MJ, et al. Rapid detection of COVID-19 causative virus (SARS-CoV-2) in human nasopharyngeal swab specimens using field-effect transistor-based biosensor (vol 14, pg 5135, 2020). ACS Nano. 2020;14(9):12257–12258. doi:10.1021/acsnano.0c06726
276. Moitra P, Alafeef M, Dighe K, Frieman MB, Pan D. Selective naked-eye detection of SARS-CoV-2 mediated by N gene targeted antisense oligonucleotide capped plasmonic nanoparticles. ACS Nano. 2020;14(6):7617–7627. doi:10.1021/acsnano.0c03822
277. Baker AN, Richards SJ, Guy CS, et al. The SARS-COV-2 spike protein binds sialic acids and enables rapid detection in a lateral flow point of care diagnostic device. ACS Cent Sci. 2020;6(11):2046–2052. doi:10.1021/acscentsci.0c00855
278. Zhang JY, Zeng H, Gu J, Li HB, Zheng LX, Zou QM. Progress and prospects on vaccine development against SARS-CoV-2. Vaccines-Basel. 2020;8(2).
279. Jiang SB, Bottazzi ME, Du LY, et al. Roadmap to developing a recombinant coronavirus S protein receptor-binding domain vaccine for severe acute respiratory syndrome. Expert Rev Vaccines. 2012;11(12):1405–1413. doi:10.1586/erv.12.126
280. Lu RJ, Zhao X, Li J, et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet. 2020;395(10224):565–574. doi:10.1016/S0140-6736(20)30251-8
281. Coleman CM, Liu YV, Mu HY, et al. Purified coronavirus spike protein nanoparticles induce coronavirus neutralizing antibodies in mice. Vaccine. 2014;32(26):3169–3174. doi:10.1016/j.vaccine.2014.04.016
282. Takashima Y, Osaki M, Ishimaru Y, Yamaguchi H, Harada A. Artificial molecular clamp: a novel device for synthetic polymerases. Angew Chem Int Ed Engl. 2011;50(33):7524–7528. doi:10.1002/anie.201102834
283. Tai WB, He L, Zhang XJ, et al. Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: implication for development of RBD protein as a viral attachment inhibitor and vaccine. Cell Mol Immunol. 2020;17(6):613–620. doi:10.1038/s41423-020-0400-4
284. Liu C, Zhou QQ, Li YZ, et al. Research and development on therapeutic agents and vaccines for COVID-19 and related human coronavirus diseases. ACS Cent Sci. 2020;6(3):315–331. doi:10.1021/acscentsci.0c00272
285. Tu YF, Chien CS, Yarmishyn AA, et al. A review of SARS-CoV-2 and the ongoing clinical trials. Int J Mol Sci. 2020;21(7):2657. doi:10.3390/ijms21072657
286. Pardi N, Hogan MJ, Porter FW, Weissman D. mRNA vaccines - a new era in vaccinology. Nat Rev Drug Discov. 2018;17(4):261–279.
287. Zeng C, Hou X, Yan J, et al. Leveraging mRNAs sequences to express SARS-CoV-2 antigens in vivo. bioRxiv. 2020. doi:10.1101/2020.04.01.019877
288. Mulligan MJ, Lyke KE, Kitchin N, et al. Phase 1/2 study of COVID-19 RNA vaccine BNT162b1 in adults. Nature. 2020;586(7830):589–593. doi:10.1038/s41586-020-2639-4
289. Lurie N, Saville M, Hatchett R, Halton J. Developing Covid-19 vaccines at pandemic speed. New Engl J Med. 2020;382(21):1969–1973. doi:10.1056/NEJMp2005630
290. Sheahan TP, Sims AC, Leist SR, et al. Comparative therapeutic efficacy of remdesivir and combination lopinavir, ritonavir, and interferon beta against MERS-CoV. Nat Commun. 2020;11(1). doi:10.1038/s41467-019-13940-6.
291. Lee JW, Gupta N, Serikov V, Matthay MA. Potential application of mesenchymal stem cells in acute lung injury. Expert Opin Biol Ther. 2009;9(10):1259–1270. doi:10.1517/14712590903213651
292. Reddy ST, van der Vlies AJ, Simeoni E, et al. Exploiting lymphatic transport and complement activation in nanoparticle vaccines. Nat Biotechnol. 2007;25(10):1159–1164. doi:10.1038/nbt1332
293. Bachmann MF, Jennings GT. Vaccine delivery: a matter of size, geometry, kinetics and molecular patterns. Nat Rev Immunol. 2010;10(11):787–796. doi:10.1038/nri2868
294. Reddy ST, Berk DA, Jain RK, Swartz MA. A sensitive in vivo model for quantifying interstitial convective transport of injected macromolecules and nanoparticles. J Appl Physiol. 2006;101(4):1162–1169.
295. Heesters BA, Myers RC, Carroll MC. Follicular dendritic cells: dynamic antigen libraries. Nat Rev Immunol. 2014;14(7):495–504. doi:10.1038/nri3689
296. Amanna IJ, Raue HP, Slifka MK. Development of a new hydrogen peroxide-based vaccine platform. Nat Med. 2012;18(6):974. doi:10.1038/nm.2763
297. Hanson MC, Crespo MP, Abraham W, et al. Nanoparticulate STING agonists are potent lymph node-targeted vaccine adjuvants. J Clin Invest. 2015;125(6):2532–2546. doi:10.1172/JCI79915
298. Moon JJ, Suh H, Bershteyn A, et al. Interbilayer-crosslinked multilamellar vesicles as synthetic vaccines for potent humoral and cellular immune responses. Nat Mater. 2011;10(3):243–251.
299. Ma CQ, Wang LL, Tao XR, et al. Searching for an ideal vaccine candidate among different MERS coronavirus receptor-binding fragments-The importance of immunofocusing in subunit vaccine design. Vaccine. 2014;32(46):6170–6176. doi:10.1016/j.vaccine.2014.08.086
300. Wang LS, Shi W, Joyce MG, et al. Evaluation of candidate vaccine approaches for MERS-CoV. Nat Commun. 2015;6.
301. Gangadaran P, Hong CM, Oh JM, et al. In vivo non-invasive imaging of radio-labeled exosome-mimetics derived from red blood cells in mice. Front Pharmacol. 2018;9. doi:10.3389/fphar.2018.00817
302. Darnell MER, Subbarao K, Feinstone SM, Taylor DR. Inactivation of the coronavirus that induces severe acute respiratory syndrome, SARS-CoV. J Virol Methods. 2004;121(1):85–91. doi:10.1016/j.jviromet.2004.06.006
303. Chin AWH, Chu JTS, Perera MRA, et al. Stability of SARS-CoV-2 in different environmental conditions. Lancet Microbe. 2020;1(1):e10. doi:10.1016/S2666-5247(20)30003-3
304. Sim W, Barnard RT, Blaskovich MAT, Ziora ZM. Antimicrobial silver in medicinal and consumer applications: a patent review of the past decade (2007–2017). Antibiotics Basel. 2018;7(4). doi:10.3390/antibiotics7040093
305. Sohal IS, O’Fallon KS, Gaines P, Demokritou P, Bello D. Ingested engineered nanomaterials: state of science in nanotoxicity testing and future research needs. Part Fibre Toxicol. 2018;15. doi:10.1186/s12989-018-0265-1
306. Kaiser JP, Zuin S, Wick P. Is nanotechnology revolutionizing the paint and lacquer industry? A critical opinion. Sci Total Environ. 2013;442:282–289. doi:10.1016/j.scitotenv.2012.10.009
307. Elechiguerra JL, Burt JL, Morones JR, et al. Interaction of silver nanoparticles with HIV-1. J Nanobiotechnology. 2005;3:6. doi:10.1186/1477-3155-3-6
308. De Gusseme B, Sintubin L, Baert L, et al. Biogenic silver for disinfection of water contaminated with viruses. Appl Environ Microbiol. 2010;76(4):1082–1087. doi:10.1128/AEM.02433-09
309. Orlowski P, Tomaszewska E, Gniadek M, et al. Tannic acid modified silver nanoparticles show antiviral activity in herpes simplex virus Type 2 infection. PLoS One. 2014;9(8):e104113. doi:10.1371/journal.pone.0104113
310. Du T, Liang JG, Dong N, et al. Glutathione-capped Ag2S nanoclusters inhibit coronavirus proliferation through blockage of viral RNA synthesis and budding. ACS Appl Mater Interfaces. 2018;10(5):4369–4378. doi:10.1021/acsami.7b13811
311. Zodrow K, Brunet L, Mahendra S, et al. Polysulfone ultrafiltration membranes impregnated with silver nanoparticles show improved biofouling resistance and virus removal. Water Res. 2009;43(3):715–723. doi:10.1016/j.watres.2008.11.014
312. Joe YH, Park DH, Hwang J. Evaluation of Ag nanoparticle coated air filter against aerosolized virus: anti-viral efficiency with dust loading. J Hazard Mater. 2016;301:547–553. doi:10.1016/j.jhazmat.2015.09.017
313. Warnes SL, Little ZR, Keevil CW. Human coronavirus 229E remains infectious on common touch surface materials. Mbio. 2015;6(6). doi:10.1128/mBio.01697-15
314. Grass G, Rensing C, Solioz M. Metallic copper as an antimicrobial surface. Appl Environ Microb. 2011;77(5):1541–1547. doi:10.1128/AEM.02766-10
315. Borkow G, Zhou SS, Page T, Gabbay J. A novel anti-influenza copper oxide containing respiratory face mask. PLoS One. 2010;5(6):e11295. doi:10.1371/journal.pone.0011295
316. Tu ZX, Guday G, Adeli M, Haag R. Multivalent Interactions between 2D nanomaterials and biointerfaces. Adv Mater. 2018;30(33).
317. Song ZY, Wang XY, Zhu GX, et al. Virus capture and destruction by label-free graphene oxide for detection and disinfection applications. Small. 2015;11(9–10):1171–1176. doi:10.1002/smll.201401706
318. Xie ZX, Huang JL, Luo SS, et al. Ultrasensitive electrochemical immunoassay for avian influenza subtype H5 using nanocomposite. PLoS One. 2014;9(4).
319. Jung JH, Cheon DS, Liu F, Lee KB, Seo TS. A graphene oxide based immuno-biosensor for pathogen detection. Angew Chem Int Ed. 2010;49(33):5708–5711. doi:10.1002/anie.201001428
320. Singh R, Hong S, Jang J. Label-free detection of influenza viruses using a reduced graphene oxide-based electrochemical immunosensor integrated with a microfluidic platform. Sci Rep. 2017;7.
321. Palmieri V, Papi M. Can graphene take part in the fight against COVID-19? Nano Today. 2020;33:100883. doi:10.1016/j.nantod.2020.100883
322. Arias LS, Pessan JP, Vieira APM, Lima TMT, Delbem ACB, Monteiro DR. Iron oxide nanoparticles for biomedical applications: a perspective on synthesis, drugs, antimicrobial activity, and toxicity. Antibiotics (Basel). 2018;7(2). doi:10.3390/antibiotics7020046
323. Abo-Zeid Y, Williams GR. The potential anti-infective applications of metal oxide nanoparticles: a systematic review. Wiley Interdiscip Rev Nanomed Nanobiotechnol. 2020;12(2):e1592. doi:10.1002/wnan.1592
324. Coyne DW. Ferumoxytol for treatment of iron deficiency anemia in patients with chronic kidney disease. Expert Opin Pharmacother. 2009;10(15):2563–2568. doi:10.1517/14656560903224998
325. Abo-zeid Y, Ismail NSM, McLean GR, Hamdy NM. A molecular docking study repurposes FDA approved iron oxide nanoparticles to treat and control COVID-19 infection. Eur J Pharm Sci. 2020;153:153. doi:10.1016/j.ejps.2020.105465
326. Ahamed M, Alhadlaq HA, Alam J, Khan MA, Ali D, Alarafi S. Iron oxide nanoparticle-induced oxidative stress and genotoxicity in human skin epithelial and lung epithelial cell lines. Curr Pharm Des. 2013;19(37):6681–6690. doi:10.2174/1381612811319370011
327. Ahamed M, Alhadlaq HA, Khan MAM, Akhtar MJ. Selective killing of cancer cells by iron oxide nanoparticles mediated through reactive oxygen species via p53 pathway. J Nanopart Res. 2013;15(1). doi:10.1007/s11051-012-1225-6
328. Rab S, Afjal A, Javaid M, Haleem A, Vaishya R. An update on the global vaccine development for coronavirus. Diabetes Metab Syndr. 2020;14(6):2053–2055. doi:10.1016/j.dsx.2020.10.023
329. Yadavalli T, Shukla D. Role of metal and metal oxide nanoparticles as diagnostic and therapeutic tools for highly prevalent viral infections. Nanomed Nanotechnol. 2017;13(1):219–230. doi:10.1016/j.nano.2016.08.016
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.