Back to Journals » Infection and Drug Resistance » Volume 13
Antibiogram and Genetic Characterization of Carbapenem-Resistant Gram-Negative Pathogens Incriminated in Healthcare-Associated Infections
Authors Makharita RR , El-kholy I, Hetta HF , Abdelaziz MH, Hagagy FI, Ahmed AA, Algammal AM
Received 14 August 2020
Accepted for publication 7 October 2020
Published 4 November 2020 Volume 2020:13 Pages 3991—4002
DOI https://doi.org/10.2147/IDR.S276975
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Suresh Antony
Rabab R Makharita,1,2 Iman El-kholy,3 Helal F Hetta,4,5 Moahmed H Abdelaziz,2 Fatma I Hagagy,2 Amera A Ahmed,2,6 Abdelazeem M Algammal7
1Biology Department, Faculty of Science and Arts, University of Jeddah, Khulais, Jeddah, Saudi Arabia; 2Botany Department, Faculty of Science, Suez Canal University, Ismailia 41522, Egypt; 3Ain Shams Specialized Hospital, Faculty of Medicine, Ain Shams University, Cairo 11556, Egypt; 4Department of Medical Microbiology and Immunology, Faculty of Medicine, Assuit University, Assuit 71515, Egypt; 5Department of Internal Medicine, University of Cincinnati, College of Medicine, Cincinnati, OH 45267-0595, USA; 6Ministry of National Guard, Health Affairs, King Abdulaziz Medical City, Jeddah, Saudi Arabia; 7Department of Bacteriology, Immunology and Mycology, Faculty of Veterinary Medicine, Suez Canal University, Ismailia 41522, Egypt
Correspondence: Rabab R Makharita
Botany Department, Faculty of Science, Suez Canal University, Ismailia 41522, Egypt
Email [email protected]
Abdelazeem M Algammal
Department of Bacteriology, Immunology and Mycology, Faculty of Veterinary Medicine, Suez Canal University, Ismailia 41522, Egypt
Tel +201010667290
Email [email protected]
Purpose: Carbapenems are considered the most efficient antibiotic used in the treatment of nosocomial infections. Carbapenem-resistant Gram-negative rods are becoming a serious hazard in hospitals threatening public health. The aim of the current study was to investigate the prevalence of carbapenem-resistant Gram-negative pathogens incriminated in healthcare-associated infections, along with antimicrobial resistance profiles, carbapenemase and metallo-β-lactamase production, and their molecular characterization.
Methods: A total of 186 clinical specimens were collected from 133 patients at various hospitals in Cairo, Egypt. The obtained specimens were subjected to bacteriological examination, antimicrobial susceptibility testing, detection of carbapenemase production using the modified Hodge test (MHT), the metallo-β-lactamase production using the EDTA combined disc test (CDT), and PCR-based detection of the blaKPC and blaGES resistance genes. The identification of the highly resistant retrieved isolates was then confirmed by 16S rRNA gene sequencing.
Results: The most common isolated Gram-negative species was Klebsiella pneumoniae (40.9%), followed by Acinetobacter baumannii (18.8%), Pseudomonas aeruginosa (17.3%), Escherichia coli (15.4%), Enterobacter aerogenes (5.3%), and Proteus mirabilis (2.4%). The prevalence of carbapenem-resistant isolates was 36.1% (n=75). However, 86.5% of the recovered clinical isolates were susceptible to colistin. The MHT revealed that 33.6% (n=70) of the tested strains were positive for carbapenemase production, while the CDT showed that 33.17% (n=69) of the examined strains were metallo-β-lactamase producers. The PCR revealed that 98.6% (74/75) of the tested strains possessed the blaKPC gene; moreover, 97.3% (73/75) of the examined strains harbored the blaGES gene.
Conclusion: This study displayed the emergence of carbapenem-resistant Gram-negative pathogens incriminated in healthcare-associated infections. The accurate identification of carbapenem-resistant bacterial pathogens is pivotal for the treatment of patients, in addition to propelling appropriate contamination control measures to restrain the fast spread of such pathogens. Colistin showed a potent in vitro antimicrobial activity against the carbapenem-resistant strains.
Keywords: gram-negative rods, antibiotic resistance, carbapenem-resistance, blaKPC, blaGES, 16S rRNA gene
Introduction
The ongoing global rise of resistance to carbapenems in Gram-negative rods is considered a public health threat. Infections caused by carbapenem-resistant bacteria (CR) are extremely hard to treat and have been related to a high death rate.1 Multidrug-resistant Gram-negative pathogens are turning into an inexorably troublesome issue in hospitals and semi-permanent care services.2–4 Resistance to broad-spectrum antimicrobials such as cephalosporins is a noteworthy issue among Gram-negative rods, such as Enterobacterales, Pseudomonas sp. and Acinetobacter sp. On the other hand, carbapenems are considered as the last line of treatment for nosocomial infections and community-acquired diseases.5
Enterobacterales are significant opportunistic pathogens; they are ordinary occupants of the intestinal microflora but can cause severe illness in humans; including meningitis, pneumonia, pyelonephritis, septicemia, peritonitis, and device-related contaminations. They are among the major etiologic causes that lead to healthcare-associated infections (HAIs).6 In addition, the existence of multidrug-resistant Gram-negative bacterial pathogens from animal origins was reported by previous studies.7–10
β-Lactams for instance cephalosporins, carbapenems, penicillins, and monobactams represent 60% of all antimicrobial agents utilized.11,12 They are favored due to their safety and adequacy and since their activity may be extended or renovated by chemical manipulation, nonetheless, their significant usage led to resistance. Such antimicrobial agents are essential for controlling and curing the grave hospital-acquired infections such as those connected with medical procedures, transplantation and hospitalizations in escalated care-units, and the development of carbapenem-resistant microorganisms leaves the healthcare facilities with no powerful medications.11,13
Carbapenem-resistance mechanisms comprise the production of β-lactamases as well as the genetic mutations that affect the expression of Penicillin-Binding Proteins (PBPs) and porins.14,15 The synergism of these mechanisms could cause a remarkable carbapenem-resistance in numerous pathogens, including; Pseudomonas aeruginosa, Klebsiella pneumonia, and Acinetobacter baumannii.12,16–18 Carbapenemases are placed into three Ambler classes; Class-A (penicillinases), Class-B (metallo-β-lactamases), and class-D (oxacillinases).19,20 The combination of carbapenems with the extended-spectrum beta-lactams was considered as powerful antimicrobial agents for the treatment of infections caused by multidrug-resistant Gram-negative rods.21
Various kinds of carbapenem antibiotics including ertapenem, imipenem, meropenem, doripenem, and faropenem are most frequently used for treating severe nosocomial infections yielded by β-lactamase producing Enterobacteriaceae.22 Resistance to carbapenems are usually intervened through β-lactamases and in addition due to carbapenemases enzymes (KPC, IMP, VIM, NDM, and OXA-48), encoded by the bla genes typically situated on transmissible-components “plasmid-mediated”, encouraging the transmission of resistance among various gram-negative pathogens.23 Accurate investigation of carbapenem-resistance is essential for directing the line of treatment and additionally establishing a reliable infection control.24,25
Molecular techniques that are used for precise detection of carbapenemases genes remain the gold standard for specificity and sensitivity. Consequently, PCR-based assays are usually used to avoid the problems of phenotypic identifications.24 Amongst class-A β-lactamases, the most clinically important enzymes are KPC enzymes which encoded by K. pneumoniae carbapenemase-gene (KPC).26–29 The Guiana extended-spectrum β-lactamase (encoded by GES gene) involved up to 20 variants.30 The blaGES genes are generally carried on integrons found in different members of family Enterobacteriaceae. All the GES type variations can break-down the cephalosporins, however, certain variants have amino acid substitutions which amplify their broad effect against carbapenems.29,31,32
The present study aims to determine the prevalence, antimicrobial resistance profiles, carbapenemases and metallo-β-lactamase production, and molecular characterization of carbapenem-resistant Gram-negative pathogens incriminated in healthcare-associated infections.
Materials and Methods
Specimens Collection
All methods were carried out in accordance with relevant guidelines and regulations. Sampling and all the experimental protocols were carried out by well-trained scientists and were approved by the Ethics committee, Specialist Hospital, Ain Shams University, Cairo, Egypt. The collected specimens were a part of the routine hospital laboratory procedures. A total of 186 clinical specimens: sputum (n=44), urine (37), blood (n=33), pus (n=27), wound (n=17), endotracheal tube (n=8), stool (n=7), drains (n=5), 3 central line (n=3), cerebrospinal fluid (n=3), bronchial alveolar lavage (n=1), and 1 pleural fluid (n=1) were collected from 133 different patients admitted in governmental and private hospitals in Cairo, Egypt. Those patients were suffering from various diseases: 27 performed liver transplantation surgery, 21 were suffering from Diabetes mellitus (D.M), 18 chronic obstructive pulmonary disease (C.O.P.D), 18 urinary tract infection (UTI), 14 bronchogenic carcinoma (B.C), 12 chest infection, 8 Liver Cell Failure (L.C.F), 8 respiratory failure, and 7 were suffering from pneumonia. The samples were collected during the period from June to October 2017. All samples were collected in a clean and sterilized container under complete aseptic conditions and transported immediately to the Microbiology laboratory of Suez Canal University. Each specimen was labeled with the patient’s age, gender, site of infection, and the source of specimen, and then transported to the laboratory.33
Isolation and Phenotypic Identification of Gram-Negative Rods
The obtained samples were inoculated on MacConkey’s agar, blood agar, cetrimide agar, nutrient agar, trypticase soy agar, and eosin methylene blue agar plates (Oxoid, UK) and incubated at 37оC for 24 h. A loopful of each pure colony was inoculated in trypticase soy broth (Oxoid, UK) and incubated on a shaker at 37оC for 24 h., 0.8 mL of the obtained culture was mixed with 0.2 mL of sterile glycerol in sterile vials and immediately placed in –20о C for later use. The suspected colonies were identified according to Tille,34 Bergey, Holt35 upon Gram’s stain, colonial characteristics, motility, and biochemical reactions (oxidase, catalase, lactose, sorbitol, mannitol, and sucrose fermentation, indole, methyl red, Voges-Proskauer, citrate utilization, H2S production, gelatin hydrolysis, urease production, and nitrate reduction tests).
Antimicrobial Susceptibility Testing (AST) and Minimal Inhibitory Concentration (MIC)
The susceptibility of the retrieved isolates to various commercial antimicrobial agents: imipenem (10µg), meropenem (10µg), amoxicillin-clavulanic acid (20/10µg), piperacillin-tazobactam (100/10µg), amikacin (30µg), tetracycline (30µg), ciprofloxacin (5µg), levofloxacin (5µg), trimethoprim-sulfamethoxazole (1.25/23.75µg), cefepime (30µg), ceftazidime (30µg), ceftriaxone (30µg), aztreonam (30µg), gentamycin (10µg), and colistin (10µg) (Oxoid, UK) were investigated using the disc diffusion method (Modified Kirby-Bauer method). Additionally, the Minimal Inhibitory Concentration (MIC) for imipenem was performed using the broth dilution technique. Both tests were applied as described by CLSI.36
Phenotypic Detection of Carbapenemases Production
In order to determine if strains produce carbapenemases, the retrieved Carbapenem-resistant isolates were subjected to Modified Hodge Test (MHT) as described by Clinical and Laboratory Standards Institute.36 E. coli ATCC 25922 was used as an indicator strain. The turbidity of an overnight broth culture of the indicator strain was adjusted to 0.5 McFarland standards then diluted to 1:10 broth. Using a sterile cotton swab the indicator strain was inoculated on a Mueller Hinton agar plate (Oxoid, UK), as with the routine disc diffusion technique. Then the ertapenem disc (10 μg) was positioned in the center of the agar plates. Three to five colonies of the tested organism from overnight blood agar culture were picked by sterile swab and inoculated in straight-line, from the edge of the disc, up to a distance of at least 25 mm, then plates were incubated at 37 оC overnight then were examined. The Positive result is indicated by a clover-leaf-like indentation zone of inhibition from the indicator strain and the streak of inoculum.37
Phenotypic Detection of Metallo-β-Lactamase Production
The isolated species were tested for metallo-β-lactamase (MBL) production using imipenem/EDTA combined disc test (CDT) as previously described by Yong et al38 imipenem/EDTA disc was prepared by adding 750 µg of a sterile 0.5 M EDTA solution to an imipenem disc-10 μg, then discs were dried immediately in a 37оC incubator. A 0.5 McFarland-suspension of the tested isolate was inoculated on the surface of the Muller Hinton agar plate then a disc of imipenem and another disc of imipenem/EDTA were placed at suitable distances on the inoculated plate. Then plates were incubated for 24 h at 37оC. The positive result is indicated by the increase of the inhibition zone diameter (≥17 mm) of imipenem/EDTA disc compared to the imipenem disc alone. The negative result is indicated by no difference in the diameter of the zones around the two discs.
Genotypic Detection of Class-A Carbapenemases Genes
The extraction of genomic DNA was conducted using Bacterial DNA Preparation Kit (Jena Bioscience, GmbH, Germany) according to the manufacturer’s instructions. The used oligonucleotide primers (Metabion International AG, Germany) were previously reported in the detection of Class-A carbapenemases.39 Primer-sequences, cycling conditions, and the specific amplicon size are listed in Table 1.
 |
Table 1 Oligonucleotide Primers and Their Recycling Conditions Used for Detection of Class-A Carbapenemases Genes |
PCR reaction mixtures of 25 μL (12.5 μL of Master Mix (Promega, W1, USA), 0.5 μL of each primer forward and reverse, 2.5 μL of the DNA, and up to 25 µL deionized water) were amplified in the MJ MiniTM Gradient Thermocycler apparatus (Biometra Göttingen, Germany). The PCR protocol was carried out as described by.28 A reaction with no DNA-template was used as a negative control, while reference strains, kindly supplied by the Microbiology Department, Faculty of Science, Suez Canal University, Egypt, were used as positive controls. PCR products were screened using 1.5% (w/v) agarose gel electrophoresis (Applichem GmbH, Darmstadt, Germany) for 45 m at 100 V in 1× TAE buffer (0.04 M Tris, 0.0001 M EDTA, pH 8.0), visualized using 15µL of DNA gel stain and photographed under UV transilluminator. A 100 bp ladder (Thermo Scientific, Darmstadt, Germany) was used.39
Genotypic Identification of the Retrieved Carbapenem-Resistant Isolates by 16S rRNA-Sequencing
The Molecular identification was performed by 16S rRNA sequencing for Carbapenem-resistant isolates which recorded the highest MIC value for imipenem and were positive for blaKPC and blaGES genes (n=13) including: K. pneumoniae (n=7), A. baumannii (n=2), E. coli (n=2), E. aerogenes (n=1), and one carbapenem susceptible P. mirabilis (n=1).
The universal 16S rRNA specific primers 27F-YM (5ʹ-AGAGTTTGATYMTGGCTCAG-3ʹ) and 1492R (5ʹ-TACCTTGTTACGACTT-3ʹ) were used in the PCR as previously described.40 1500 bp amplicons were produced and DNA sequenced between position 27 and 1492 of bacterial 16S rRNA genes numbered according to E. coli rRNA. PCR products were purified and sequenced by Macrogen Inc., (Seoul, Korea).
The sequences obtained were compared with those deposited in the public databases using the Basic Local Alignment Search Tool (BLAST) provided by the National Center for Biotechnology Information (NCBI) at (http://www.ncbi.nlm.nih.gov/BLAST). Sequences were submitted to NCBI and GenBank nucleotide sequence database and deposited under accession numbers KY856923 to KY856935.
Phylogenetic Analysis
The newly generated sequences of the current study as well as other related sequences downloaded from the GenBank were downloaded and concatenated into a balanced dataset for the phylogenetic analysis. The nucleotide sequences of the 16SrRNA genes of the retrieved isolates were aligned using ClustalW from MEGA software, version 5.05.41
The Juke and Cantor model of sequence similarity was conducted to estimate sequence relatedness.42 The dendrogram was constructed using Neighbor-Joining (NJ) method. Maximum Parsimony (MP) was applied to infer the phylogenetic relationships with bootstrapping of 100 replicates to get the branch support and testing the tree stability.
Statistical Analysis
The Chi-square was conducted to analyze the results using the statistical analysis software (SAS software version 9.4, SAS Institute, Cary, NC, USA). The significance level was (P < 0.05).
Results
Prevalence Gram-Negative Rods
The bacteriological examination revealed that 208 isolates clinical isolates were retrieved from different examined specimens. The most predominant isolated pathogens were K. pneumoniae (40.9%), followed by A. baumannii (18.8%), P. aeruginosa (17.3%), E. coli (15.4%), E. aerogenes (5.3%), and P. mirabilis (2.4%) (Table 2). Statistically, there is a significant difference in the prevalence of different bacterial pathogens recovered from various types of samples (p < 0.05).
 |
Table 2 The Prevalence of Different Bacterial Pathogens Isolated from Various Types of Samples |
Antimicrobial Resistance Profiles of the Retrieved Strains
In the present study, 86.5% of the recovered clinical isolates (n=180) exhibited remarkable sensitivity to colistin, while 13.5% of the obtained strains (n= 28) were resistant to colistin. On the other hand, the retrieved K. pneumoniae strains exhibited remarkable resistance to ceftazidime and levofloxacin (89.4%), ceftriaxone and trimethoprim-sulfamethoxazole (87.1%). Furthermore, the isolated A. baumannii strains were resistant to trimethoprim-sulfamethoxazole (89.7%), levofloxacin (87.2%), aztreonam (84.6%), tetracycline, amoxicillin-clavulanic acid, ciprofloxacin, and cefepime (82.1%).
Moreover, the recovered P. aeruginosa strains exhibited resistance to cefepime (72.2%), and trimethoprim-sulfamethoxazole and ceftriaxone (66.9%). Besides, the isolated E. coli strains exhibited resistance to ceftriaxone and ceftazidime (90.6%), levofloxacin and aztreonam (87.5%). The retrieved E. aerogenes strains exhibited resistance to ciprofloxacin (72.7%), aztreonam, cefepime, piperacillin-tazobactam, trimethoprim-sulfamethoxazole, ceftriaxone, gentamycin, ceftazidime, and levofloxacin (63.5% for each). Furthermore, the isolated P. mirabilis were resistant to ceftazidime, ciprofloxacin, cefepime, and trimethoprim-sulfamethoxazole (80% for each). Statistically, there is a significant difference in the resistance of different recovered bacterial pathogens to the tested antimicrobial agents (p < 0.05). The isolated E. aerogenes strains exhibited non-significant differences for resistance to various tested antimicrobial agents (p > 0.05) (Table 3).
 |
Table 3 The Antimicrobial Resistance Profiles of the Retrieved Gram-Negative Rods |
Concerning the carbapenem-resistance, 36% of the recovered strains (75/208) were carbapenem-resistant (resistant to both imipenem and meropenem), including: K. pneumoniae (n=36), A. baumannii (n=19), P. aeruginosa (n=10), E. coli (n=7), and E. aerogenes (n=3) (Table 3). MIC values (using imipenem) for the 75 CR-strains ranged from 2 to >128 µg mL−1 (Table 4).
 |
Table 4 The MIC Value of Carbapenem-Resistant Species Against Different Concentrations of Imipenem (µg ml−1) |
Phenotypic Detection of Carbapenem-Resistant Gram-Negative Bacterial Pathogens
Prevalence of Class-A Carbapenemases-Producing Bacterial Pathogens Using the Modified Hodge Test (MHT)
The MHT revealed that 70/208 tested strains (33.6%) were class A-Carbapenemases producing bacterial pathogens (were resistant to imipenem and meropenem) including: K. pneumoniae (n=36), A. baumannii (n=19), P. aeruginosa (n=7), E. coli (n=7), and E. aerogenes (n=1) (Table 5 and Figure 1A).
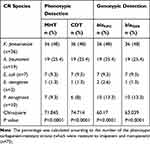 |
Table 5 The Phenotypic and Genotypic Detection of Carbapenem-Resistant Strains |
Prevalence of Metallo-β-Lactamase-Producing Bacterial Pathogens Using Combined Disk Test (CDT)
The CDT proved that 69/208 tested strains (33.17%) were metallo-β-lactamase-producing bacterial pathogens (were resistant to imipenem and meropenem) including: K. pneumoniae (n=36), A. baumannii (n=19), P. aeruginosa (n=6), E. coli (n=7), and E. aerogenes (n=1) (Table 5 and Figure 1 B). Statistically, there is a remarkably significant difference in the prevalence of class A-Carbapenemases and metallo-β-lactamase producing bacterial pathogens (p < 0.0001).
Genotypic Detection of Class-A Carbapenemases-Producing Carbapenem-Resistant Gram-Negative Bacterial Pathogens
The genotypic investigation of the presence of Class-A carbapenemases blaKPC and blaGES genes were performed by PCR for the carbapenem-resistant strains (n=75). The PCR revealed that 98.6% (74/75) of the tested strains possessed the blaKPC gene including: K. pneumonia (36/75), A. baumannii (n=19), P. aeruginosa (n=10), E. coli (n=7), and E. aerogenes (n=2). In addition, 97.3% (73/75) of the examined strains harbored the blaGES gene including: K. pneumonia (36/75), A. baumannii (n=19), P. aeruginosa (n=10), E. coli (n=7), and E. aerogenes (n=1) (Table 5 and Figure 2). Statistically, there is a highly significant difference in the prevalence of blaKPC and blaGES genes among tested strains (p < 0.0001).
16S rRNA Gene Sequencing and Phylogenetic Analysis
Thirteen isolates were identified with 16S rRNA sequencing. Twelve of these isolates were CR: K. pneumoniae (n=7), A. baumannii (n=2), E. coli (n=2), E. aerogenes (n=1), and one carbapenem susceptible P. mirabilis (n=1). Sequences were submitted to NCBI and Gene Bank nucleotide sequence database and were deposited under accession numbers (KY856923 to KY856935).
The phylogenetic relationships and placement among the isolated CR species were retrieved based on the obtained 16S rRNA sequences. The obtained phylogenetic tree has a well-supported phylogeny, providing a maximum and decisive resolution of the relationships among the examined species.
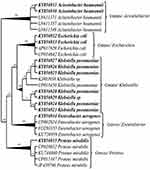
The 16S rRNA gene sequences of the bacterial isolates diverged into five phylogenetic clades: Acinetobacter clade, composed of five accessions, two of which were new to GenBank (KY856930 and KY856931); Escherichia coli clade, composed of four accessions, two new to GenBank (KY856932, and KY856933); Klebsiella clade, of nine accessions, seven new to GenBank (KY856923, KY856924, KY856925, KY856926, KY856927, KY856928, and KY856929); Enterobacter clade of four accessions, one new to GenBank (KY856934); and Proteus clade, of five accessions, only one new to GenBank (KY856935) (Figure 3).
Discussion
The existence of carbapenem-resistant bacterial pathogens is mainly accompanied by failure of medication, leading to significant morbidity and mortality. The on-going overall rise of carbapenem-resistant Gram-negative rods causes expanding public health threats.4 The laboratory detection of carbapenemases-producers among clinical isolates is crucial to limit its rapid spread. In the present study, the bacteriological examination revealed that 208 isolates were retrieved from different examined specimens. The most predominant recovered pathogen was K. pneumoniae (40.9%), followed by A. baumannii (18.8%), P. aeruginosa (17.3%), E. coli (15.4%), E. aerogenes. (5.3%), and P. mirabilis (2.4%) (Table 2). Statistically, there is a significant difference in the prevalence of different bacterial pathogens recovered from various types of samples (p < 0.05). These findings are in agreement with those obtained by Talaat et al43 who illustrated that the most predominant Gram-rods in hospital settings were K. pneumoniae (28.7%) and Acinetobacter sp. (13.7%).
Concerning the carbapenem-resistance, in the present study, 36% (75/208) of recovered strains were carbapenem-resistant (resistant to both imipenem and meropenem) including: K. pneumoniae (n=36), A. baumannii (n=19), P. aeruginosa (n=10), E. coli (n=7), and E. aerogenes (n=3). These findings agree with those obtained by Fattouh et al44,and Ramana et al45 who reported that among the tested members of Enterobacteriaceae, K. pneumoniae is the most predominant carbapenem-resistant pathogen (44.1%). Centers for Disease Control Prevention, CDC 46 reported that the prevalence of carbapenem-resistant pathogens elevated from 1.2% (2001) to 4.2% (2011) in intensive-care hospitals in the United States.
Furthermore, in the current study, the retrieved isolates exhibited remarkable multidrug resistance to most tested antimicrobial classes including cephalosporins, monobactams, tetracycline, quinolones, aminoglycosides, and beta-lactam-beta-lactamase inhibitor combinations, while showed high sensitivity to lipopeptides (colistin) as described in (Table 3). Colistin is considered the most effective antimicrobial against carbapenemases-producing bacterial pathogens. The standard therapeutic strategy of the carbapenem-resistant strains has yet to be identified. Nevertheless, the current strategy depends upon the results of the in vitro antibiogram: tigecycline, polymyxins, and combinations of carbapenems with other antibiotics were involved.47,48
In the current study, 13.5% (28/208) of the retrieved clinical isolates were resistant to colistin. The emergence of such strains is considered a public health threat. Earlier study reported that prevalence of colistin resistance among the KPC-producers was 22.4% among the recovered species.49 The emergence of multidrug-resistant Gram-negative rods is also reported by previous studies.44,50 The occurrence of such strains is always accompanied with bad prognosis as well as medication-failure.51
Regarding the prevalence of class A-Carbapenemases and metallo-β-lactamase producing bacterial pathogens, the MHT revealed that 33.6% (n=70) of the tested strains were positive for carbapenemases production, while the CDT proved that 33.17% (n=69) of the examined strains were metallo-β-lactamase production. This prevalence is similar to that reported by Fattouh et al.44 Although, the MHT is considered a simple, effective, reliable, and cheap diagnostic tool that could be used in monitoring for carbapenemases-production in KPC, it exhibited a lower specificity and sensitivity in other species such as Enterobacter.52
The combination of MHT, CDT, and the antibiotic sensitivity test is an effective tests for the phenotypic screening of carbapenem-resistant strains.37,53 The class-A-β-lactamases (especially GES and KPC enzymes) are the most prevalent type of the β-lactamase-group; which may attribute to the frequent occurrence of their specific encoding genes in various pathogenic bacterial species.54 These enzymes break down the β-lactam antibiotics and are stable to clavulanate, sulbactam and tazobactam.27
PCR is considered the most significant specific technique used for the determination of carbapenem-resistance isolates in clinical specimens. In the current study, the genotypic investigation of the presence of Class-A carbapenemases blaKPC and blaGES genes was performed by PCR for the carbapenem-resistant strains (n=75). The PCR revealed that 98.6% (74/75) of the tested strains possessed the blaKPC gene; moreover, 97.3% (73/75) of the examined strains harbored the blaGES gene. The results of the blaKPC gene are similar to those obtained by Raghunathan et al,55 who reported that prevalence of the blaKPC gene was 82% in examined strains. Jeong et al56 reported the first K. pneumoniae expressing GES-5 in Korea. Nordmann et al24 and Yigit et al26 reported the emergence of carbapenem-resistant K. pneumoniae that produce the new carbapenemase, KPC, as the first report globally. Since then, KPC-producing isolates have become more widespread globally.
In the current study, the identification of 13 isolates; K. pneumoniae (n=7), A. baumannii (n=2), E. coli (n=2), E. aerogenes (n=1), and P. mirabilis (n=1) was confirmed by 16S rRNA sequencing. The obtained phylogenetic tree has a very well supporting phylogeny, providing a maximum and decisive resolution of the relationships among the examined species. It was divergent into five Genera; Acinetobacter, Escherichia, Klebsiella, Enterobacter, and Proteus as illustrated in Figure 3.
The conventional phenotypic identification of bacterial pathogens is not 100% exact. The 16S rRNA gene sequencing has elicited a specific, reliable molecular technique; which is useful in the accurate identification of scarcely isolated strains as well as phenotypically eccentric ones. Furthermore, the 16S rRNA gene is the most conserved gene in various bacterial pathogens; beneficial in phylogenetic analyses. The 16S rRNA sequencing gives a complete conceive of whether the tested strains belonged to distantly or closely related bacterial lineages.37
In conclusion, the present study emphasized that the most predominant carbapenem-resistant bacterial pathogens incriminated in healthcare-associated infections were K. pneumoniae, A. baumannii, P. aeruginosa, and E. coli. The retrieved clinical isolates exhibited remarkable multidrug resistance to cephalosporins, monobactams, tetracycline, quinolones, aminoglycosides, and beta-lactam-beta-lactamase inhibitor combinations that is considered a public health threat. The carbapenem-resistance is commonly associated with class-A carbapenemases and metallo-β-lactamases production as well as the presence of both blaKPC and blaGES resistance-genes. The combination of phenotypic and genotypic detection of carbapenem-resistant strains is a reliable effective epidemiological tool. The continuous application of antimicrobial susceptibility testing is essential to detect multidrug-resistant strains. The accurate identification of carbapenem-resistant bacterial pathogens is pivotal for the treatment of patients, in addition to propelling appropriate contamination control measures to restrain the fast spread of these pathogens.
Acknowledgments
The authors gratefully acknowledge Dr. Ahmed EL-Banhawy, Suez Canal University, Faculty of Science, Botany Department, Ismailia, Egypt for his help in the phylogenetic analysis. We also appreciate the help of Dr. Oliver Harry Ellingham, Fungiscape, UK, for the help in improving the manuscript language.
Author Contributions
All authors contributed to data analysis, drafting or revising the article, have agreed on the journal to which the article will be submitted, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Disclosure
The authors declare no conflict of interest.
References
1. Aguilera-Alonso D, Escosa-García L, Saavedra-Lozano J, Cercenado E, Baquero-Artigao F. Carbapenem-Resistant Gram-Negative Bacterial Infections in Children. Antimicrob Agents Chemother. 2020;64:3.
2. Makharita RRM, El-kholy EMA, Abdelaziz MH, Ahmed AMO. Antibiotic Resistance Patterns of Bacterial Pathogens Isolated from Urinary Tract Infections in Hospitalized Patients in Ismailia City, Egypt. 6th International Conference of Biotechnology and Its Applications in Botany and Microbiology. Menoufia University; 2016.
3. Centers for Disease Control and Prevention. CRE toolkit—guidance for control of carbapenem-resistant Enterobacteriaceae (CRE). Atlanta, GA: CDC Web site. http://www.infectologia.edu.uy/images/stories/pdf/4_guias_clinicas/control_infecciones/estrategias_control/guias_control_BGNCarbapenemasa_CDC_2012.pdf.
4. World Health Organization. Report on the Burden of Endemic Health Care-Associated Infection Worldwide; 2011.
5. Orsi GB, Falcone M, Venditti M. Surveillance and management of multidrug-resistant microorganisms. Expert Rev Anti Infect Ther. 2011;9(8):653–679.
6. Jacob JT, Klein GE, Laxminarayan R, et al. Vital signs: carbapenem-resistant Enterobacteriaceae. Morbidity Mortality Weekly Rep. 2013;62(9):165–169.
7. Enany ME, Algammal AM, Nasef SA, et al. The occurrence of the multidrug resistance (MDR) and the prevalence of virulence genes and QACs resistance genes in E. coli isolated from environmental and avian sources. AMB Express. 2019;9(1):192. doi:10.1186/s13568-019-0920-4
8. Algammal AM, El-Kholy AW, Riad EM, et al. Genes Encoding the Virulence and the Antimicrobial Resistance in Enterotoxigenic and Shiga-Toxigenic E. coli Isolated from Diarrheic Calves. Toxins. 2020;12(6):383. doi:10.3390/toxins12060383
9. Algammal AM, Enany ME, EL-Tarabili RM, Gobashy MOI, Helmy YA. Prevalence, antimicrobial resistance profiles, virulence and enterotoxin-determinant genes of MRSA isolated from subclinical bovine mastitis in Egypt. Pathogens. 2020;9:1–11. doi:10.3390/pathogenes9050362.
10. El-Sayed AM, Algammal AM, Aboel-Atta ME, Mabrok M, Emam AM. Pathogenicity, genetic typing, and antibiotic sensetivity of Vibrio alginolyticus isolated from Oreochromis niloticus and Tilapia zillii. Revue de Medecine Veterinaire journal. 2019;170:80–86.
11. Algammal AM, Mabrok M, Sivaramasamy E, et al. Emerging MDR-Pseudomonas aeruginosa in fish commonly harbor oprL and toxA virulence genes and blaTEM, blaCTX-M, and tetA antibiotic-resistance genes. Sci Rep. 2020;10(1):15961. doi:10.1038/s41598-020-72264-4
12. Algammal AM, Mohamed MF, Tawfiek BA, et al. Molecular typing, antibiogram and PCR-RFLP based detection of Aeromonas hydrophila complex isolated from Oreochromis niloticus. Pathogens. 2020;9(3):238. doi:10.3390/pathogens9030238.
13. Sievert DM, Ricks P, Edwards JR, et al. Antimicrobial-resistant pathogens associated with healthcare-associated infections summary of data reported to the National Healthcare Safety Network at the Centers for Disease Control and Prevention, 2009–2010. Infect Control Hospital Epidemiology. 2013;34(1):1–14. doi:10.1086/668770
14. Martínez-Martínez L. Extended-spectrum β-lactamases and the permeability barrier. Clin Microbiol Infect. 2008;14(s1):82–89. doi:10.1111/j.1469-0691.2007.01860.x
15. Sawa T, Kooguchi K, Moriyama K. Molecular diversity of extended-spectrum β-lactamases and carbapenemases, and antimicrobial resistance. J Intensive Care. 2020;8(1):13. doi:10.1186/s40560-020-0429-6
16. Limansky AS, Mussi MA, Viale AM. Loss of a 29-kilodalton outer membrane protein in Acinetobacter baumannii is associated with imipenem resistance. J Clin Microbiol. 2002;40(12):4776–4778.
17. Mena A, Plasencia V, García L, et al. Characterization of a large outbreak by CTX-M-1-producing Klebsiella pneumoniae and mechanisms leading to in vivo carbapenem resistance development. J Clin Microbiol. 2006;44(8):2831–2837. doi:10.1128/JCM.00418-06
18. Rodríguez-Martínez J-M, Poirel L, Nordmann P. Molecular Epidemiology and Mechanisms of Carbapenem Resistance in Pseudomonas aeruginosa. Pseudomonas Aeruginosa. Antimicrobial Agents and Chemotherapy. 2009;53(11):4783–4788. doi:10.1128/AAC.00574-09
19. Walsh TR. Emerging carbapenemases: a global perspective. Int J Antimicrob Agents. 2010;36:S8–S14. doi:10.1016/S0924-8579(10)70004-2
20. Ambler RP, Coulson AFW, Frère JM, et al. A standard numbering scheme for the class A β-lactamases. Biochem J. 1991;276(1):269–270. doi:10.1042/bj2760269
21. Yong D, Toleman MA, Giske CG, et al. Characterization of a new metallo-β-lactamase gene, blaNDM-1, and a novel erythromycin esterase gene carried on a unique genetic structure in Klebsiella pneumoniae sequence type 14 from India. Antimicrob Agents Chemother. 2009;53(12):5046–5054. doi:10.1128/AAC.00774-09
22. Papp-Wallace KM, Endimiani A, Taracila MA, Bonomo RA. Carbapenems: past, present, and future. Antimicrob Agents Chemother. 2011;55(11):4943–4960.
23. Poirel L, Potron A, Nordmann P. OXA-48-like carbapenemases: the phantom menace. J Antimicrobial Chem. 2012;67(7):1597–1606. doi:10.1093/jac/dks121
24. Nordmann P, Dortet L, Poirel L. Carbapenem resistance in Enterobacteriaceae: here is the storm! Trends Mol Med. 2012;18(5):263–272.
25. Humphries RM, Hindler JA, Epson E, et al. Carbapenem-resistant Enterobacteriaceae detection practices in California: what are we missing? Clin Infect Dis. 2018;66(7):1061–1067. doi:10.1093/cid/cix942
26. Yigit H, Queenan AM, Anderson GJ, et al. Novel Carbapenem-Hydrolyzing β-Lactamase, KPC-1, from a Carbapenem-Resistant Strain of Klebsiella pneumoniae. Klebsiella Pneumoniae. Antimicrobial Agents and Chemotherapy. 2001;45(4):1151–1161. doi:10.1128/AAC.45.4.1151-1161.2001
27. Nordmann P, Cuzon G, Naas T. The real threat of Klebsiella pneumoniae carbapenemase-producing bacteria. Lancet Infect Dis. 2009;9(4):228–236.
28. Weisburg WG, Barns SM, Pelletier DA, Lane DJ. 16S ribosomal DNA amplification for phylogenetic study.. J Bacteriol. 1991;173(2):697–703. doi:10.1128/JB.173.2.697-703.1991
29. Golkar Z, Bagasra O, Pace DG. Bacteriophage therapy: a potential solution for the antibiotic resistance crisis. J Infect Dev Countries. 2014;8(02):129–136. doi:10.3855/jidc.3573
30. Sankar MJ, Agarwal R, Deorari AK, Paul VK. Sepsis in the newborn. The Indian Journal of Pediatrics. 2008;75(3):261–266. doi:10.1007/s12098-008-0056-z
31. Gardner SL. Sepsis in the neonate. Crit Care Nurs Clin North Am. 2009;21(1):121–141. doi:10.1016/j.ccell.2008.11.002
32. Hashizume T, Ishino F, Nakagawa J-I, Tamaki S, Matsuhashi M. Studies on the mechanism of action of imipenem (N-formimidoylthienamycin) in vitro: binding to the penicillin-binding proteins (PBPs) in Escherichia coli and Pseudomonas aeruginosa, and inhibition of enzyme activities due to the PBPs in E. coli.. The Journal of Antibiotics. 1984;37(4):394–400. doi:10.7164/antibiotics.37.394
33. Linscott AJ. Collection, Transport, and Manipulation of Clinical Specimens and Initial Laboratory Concerns. In: Clinical Microbiology Procedures Handbook.
34. Tille PM. Bloodstream Infections. Bailey and Scott’s Diagnostic Microbiology.
35. Bergey D, Holt J. Bergey’s Manual of Determinative Microbiology.
36. Clinical and Laboratory Standards Institute. M100S Performance Standards for Antimicrobial Susceptibility Testing.
37. Girlich D, Poirel L, Nordmann P. Value of the modified Hodge test for detection of emerging carbapenemases in Enterobacteriaceae. J Clin Microbiol. 2012;50(2):477–479. doi:10.1128/JCM.05247-11
38. Yong D, Lee K, Yum JH, Shin HB, Rossolini GM, Chong Y. Imipenem-EDTA Disk Method for Differentiation of Metallo- -Lactamase-Producing Clinical Isolates of Pseudomonas spp. and Acinetobacter spp.. J Clin Microbiol. 2002;40(10):3798–3801. doi:10.1128/JCM.40.10.3798-3801.2002
39. Hong SS, Kim K, Huh JY, Jung B, Kang MS, Hong SG. Multiplex PCR for Rapid Detection of Genes Multiplex PCR for Rapid Detection of Genes Encoding Class A Carbapenemases. Ann Lab Med. 2012;32(5):359–361. doi:10.3343/alm.2012.32.5.359
40. Frank JA, Reich CI, Sharma S, Weisbaum JS, Wilson BA, Olsen GJ. Critical evaluation of two primers commonly used for amplification of bacterial 16S rRNA genes. Appl Environ Microbiol. 2008;74(8):2461–2470. doi:10.1128/AEM.02272-07
41. Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. MEGA5: molecular Evolutionary Genetics Analysis Using Maximum Likelihood, Evolutionary Distance, and Maximum Parsimony Methods. Mol Biol Evol. 2011;28(10):2731–2739. doi:10.1093/molbev/msr121
42. Juke T, Cantor C. Evolution of Protein Molecules. Mammalian Protein Metabolism. Vol. 3. New York: Academic Press, Inc; 1969.
43. Talaat M, El-Shokry M, El-Kholy J, et al. National surveillance of health care–associated infections in Egypt: developing a sustainable program in a resource-limited country. Am J Infect Control. 2016;44(11):1296–1301. doi:10.1016/j.ajic.2016.04.212
44. Fattouh M, El-din AN, Omar MA. Detection of Klebsiella pneumoniae Carbapenemase (KPC) Producing Gram Negative Superbugs: an Emerging Cause of Multidrug-Resistant Infections in General Surgery Department of Sohag University Hospital, Egypt. Int J Curr Microbiol App Sci. 2015;4(5):1–15.
45. Ramana KV, Ratna Rao R, Sharada CV, Kareem MA, Ratna Mani MS. Modified Hodge test: a useful and the low-cost phenotypic method for detection of carbapenemase producers in Enterobacteriaceae members. Journal of Natural Science, Biology and Medicine. 2013;4(2):346. doi:10.4103/0976-9668.117009
46. Centers for Disease Control Prevention. Office of Infectious Disease Antibiotic Resistance Threats in the United States. Atlanta, GA: Centers for Disease Control and Prevention Web site; 2013. Available from: https://www.cdc.gov/drugresistance/threat-report-2013/pdf/ar-threats-2013-508.pdf.
47. Drusano G, Liu W, Fregeau C, Kulawy R, Louie A. Differing effects of combination chemotherapy with meropenem and tobramycin on cell kill and suppression of resistance of wild-type Pseudomonas aeruginosa PAO1 and its isogenic MexAB efflux pump-overexpressed mutant. Antimicrob Agents Chemother. 2009;53(6):2266–2273. doi:10.1128/AAC.01680-08
48. Arnold RS, Thom KA, Sharma S, Phillips M, Johnson JK, Morgan DJ. Emergence of Klebsiella pneumoniae Carbapenemase-Producing Bacteria. South Med J. 2011;104(1):40. doi:10.1097/SMJ.0b013e3181fd7d5a
49. Monaco M, Giani T, Raffone M, et al. Colistin resistance superimposed to endemic carbapenem-resistant Klebsiella pneumoniae: a rapidly evolving problem in Italy, November 2013 to April 2014. Euro Surveill. 2014;19(42):20939. doi:10.2807/1560-7917.ES2014.19.42.20939
50. Endimiani A, Hujer AM, Perez F, et al. Characterization of blaKPC-containing Klebsiella pneumoniae isolates detected in different institutions in the Eastern USA. J Antimicrobial Chem. 2009;63(3):427–437. doi:10.1093/jac/dkn547
51. Tumbarello M, Viale P, Viscoli C, et al. Predictors of Mortality in Bloodstream Infections Caused by Klebsiella pneumoniae Carbapenemase-Producing K. pneumoniae: importance of Combination Therapy. Clin Infect Dis. 2012;55(7):943–950. doi:10.1093/cid/cis588
52. Centers for Disease Control and Prevention. Facility Guidance for Control of Carbapenem-Resistant Enterobacteriaceae (Cre)—november 2015 Update CRE Toolkit. Centers for Disease Control and Prevention. Atlanta, GA; 2015. https://www.cdc.gov/hai/pdfs/cre/cre-guidance-508.pdf.
53. Pournaras S, Zarkotou O, Poulou A, et al. A combined disk test for direct differentiation of carbapenemase-producing Enterobacteriaceae in surveillance rectal swabs. J Clin Microbiol. 2013;51(9):2986–2990. doi:10.1128/JCM.00901-13
54. Walther-Rasmussen J, Høiby HN. Class A carbapenemases. J Antimicrobial Chem. 2007;60(3):470–482. doi:10.1093/jac/dkm226
55. Raghunathan A, Samuel L, Tibbetts RJ. Evaluation of a real-time PCR assay for the detection of the Klebsiella pneumoniae carbapenemase genes in microbiological samples in comparison with the modified Hodge test. Am J Clin Pathol. 2011;135(4):566–571.
56. Jeong SH, Bae IK, Kim D, et al. First outbreak of Klebsiella pneumoniae clinical isolates producing GES-5 and SHV-12 extended-spectrum β-lactamases in Korea. Antimicrob Agents Chemother. 2005;49(11):4809–4810. doi:10.1128/AAC.49.11.4809-4810.2005
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.