Back to Journals » Infection and Drug Resistance » Volume 13
Antagonistic Activities of Cell-Free Supernatants of Lactobacilli Against Extended-Spectrum β-Lactamase Producing Klebsiella pneumoniae and Pseudomonas aeruginosa
Authors El-Mokhtar MA , Hassanein KM, Ahmed AS, Gad GFM, Amin MM, Hassanein OFE
Received 22 October 2019
Accepted for publication 23 January 2020
Published 17 February 2020 Volume 2020:13 Pages 543—552
DOI https://doi.org/10.2147/IDR.S235603
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Suresh Antony
Mohamed A El-Mokhtar, 1 Khaled M Hassanein, 1 Ahmed S Ahmed, 1 Gamal FM Gad, 2 Mohamed M Amin, 3 Osama FE Hassanein 4
1Department of Medical Microbiology and Immunology, Faculty of Medicine, Assiut University, Assiut, Egypt; 2Department of Microbiology and Immunology, Faculty of Pharmacy, Minia University, Minia, Egypt; 3Department of Microbiology and Immunology, Faculty of Medicine, Aswan University, Aswan, Egypt; 4Drug Research Center, Assiut University, Assiut, Egypt
Correspondence: Mohamed A El-Mokhtar
Department of Medical Microbiology and Immunology, Faculty of Medicine, Assiut University, Assiut 71515, Egypt
Tel +20 122 111 53 13
Email [email protected]
Aim: This study aimed to describe the inhibitory activity of cell-free supernatants (CFS) of lactobacilli against extended-spectrum β-lactamase (ESBL)-producing Klebsiella pneumoniae (K pneumoniae) and Pseudomonas aeruginosa (P aeruginosa).
Material and Methods: Pathogenic clinical strains of K pneumoniae and P aeruginosa were isolated from urine samples and selected for investigation. Anti-bacterial activities of the CFS of lactobacilli were assessed by agar well diffusion, MTT assay, as well as time-kill tests. In addition, the antibiofilm characteristics were analyzed by the microplate method against fresh and 24 h-old biofilms. The ability of CFS to interfere with bacterial invasion was analyzed by flow cytometry.
Results: Although all tested strains were ESBL producers and showed a multidrug-resistant phenotype, the CFS displayed a high anti-ESBL activity with inhibition zone diameters greater than 13 mm in the agar well diffusion assays against both pathogens. The growth kinetics of K pneumoniae and P aeruginosa were dramatically decreased in the presence of the CFS. The CFS not only inhibited the biofilm formation by these pathogens but also was able to remove the 24-h formed biofilms. The invasion abilities of FITC-labelled K pneumoniae decreased from 30.3%± 7 to 15.4%± 5 and invasion of FITC-labelled P aeruginosa was reduced from 36.9%± 7 to 25.2%± 5.
Conclusion: CFS of lactobacilli exhibit anti-ESBL activities, which suggests its potential application for controlling or preventing colonization of infections caused by ESBL-producing bacteria.
Keywords: lactobacilli, antibiofilm, ESBL, K pneumoniae, P aeruginosa
Introduction
Klebsiella pneumoniae (K pneumoniae) and Pseudomonas aeruginosa (P aeruginosa) are Gram-negative opportunistic pathogens that can cause severe nosocomial infections such as bacteremia, pneumonia, urinary tract infections and soft tissue infections, particularly in immune-compromised individuals.1 These pathogens are well-known for their ability to develop and transfer antibiotic resistance determinants such as the production of extended-spectrum β-lactamase (ESBL), which confers resistance to β-lactam antibiotics, particularly to third-generation cephalosporins.2 The spread of ESBL producing Gram-negative bacilli has increase critically worldwide and is one of the most growing problems of antibiotic resistance and leaves only limited treatment options for clinicians.3 Moreover, treatment of serious infections with these bacteria is extremely difficult due to co-resistance to multiple antibiotics.4 Their pathogenicity is multifactorial, including LPS, capsule, adherence factors and exotoxins, and till now, no effective vaccines are developed for protection from these pathogens.5 A common virulence strategy for both pathogens is the ability to form biofilms. Bacteria in biofilms are not only resistant to immune defense mechanisms but also to many antibiotics due to the production of a protecting extracellular polymer matrix.6,7 Therefore, there is an urgent need for new treatment strategies for these critical groups of pathogenic bacteria.
Lactobacilli is one of the most common probiotics that is generally recognized as safe (GRAS) biological therapeutic agent and is also used to boost the host immune responses. There are different mechanisms by which lactobacilli can exert their antimicrobial activity, including the production of inhibitory compounds, immune stimulation, competition with pathogenic bacteria for the receptor binding, and competition on nutrients. The inhibitory compounds produced by lactobacilli include organic acids such as lactic acid, acetic acid, and formic acid or bacteriocins.8,9 Through these antimicrobial mechanisms, lactobacilli have demonstrated antagonistic activates against different pathogenic bacteria, including carbapenem-resistant Enterobacteriaceae,10 Escherichia coli,11 Helicobacter pylori,12 Salmonella,13 Shigella,14 P aeruginosa,15 and Staphylococcus aureus.16
However, no previous studies have assessed the antimicrobial activity of lactobacillus against ESBL-producing K pneumoniae or P aeruginosa. Thus, in this study, we aimed to describe the potential antagonistic activities of lactobacilli’s CFS on ESBL-producing K pneumoniae and P aeruginosa.
Materials and Methods
Collection of Samples and Enrichment
Lactobacilli were isolated from plain yogurt samples that were prepared from cow milk and purchased from a local dairy shop in Assiut, Egypt. One gram of the yogurt sample was inoculated in de Man Rogosa Sharpe (MRS) broth followed by streaking on the MRS agar (Thermo Fisher Oxoid, UK). Plates were incubated anaerobically at 37°C for 48 hrs and the resulting colonies were purified by repeated subculture and preliminarily identified as Lactobacillus spp. based on Gram staining, morphology, sporulation and biochemical test results.17,18
Identification of Lactobacillus Spp. And Preparation of Cell-Free Supernatants
To identify the species of the isolated bacteria, we sequenced the 16S ribosomal RNA gene. Bacterial genomic DNA was purified from 5 mL overnight MRS broth cultures according to the manufacturer’s instructions (PureLink® Genomic DNA, Life Technologies Corp., USA). The 16S ribosomal RNA gene was amplified using Lac16f (5ʹ-AGAGGTTTGATCCTGGCTCAG-3ʹ) and Lac16r (5ʹ -CTACGGCTACCTTGTTACGA-3ʹ) primers.19 The PCR conditions were as follows: initial denaturation at 95°C for 2 mins, followed by 40 cycles of 2 mins at 95°C, 20 S at 49°C and 1 min at 72°C, and a final extension at 72°C for 5 min. The purified products were sequenced in Macrogen Corporation (Seoul, South Korea). Sequences were identified using the NCBI GenBank database using the BLAST search tool (http://www.ncbi.nlm.nih.gov. blast) to find the closest match.
For the preparation of cell-free supernatants of lactobacilli, 15 mL of MRS broth was inoculated with single separated colonies of lactobacilli and incubated at 37°C for 24 h. After incubation, samples were centrifuged at 6000 xg for 15 mins, and supernatants were filter-sterilized through a 0.22 um filter (Millipore Inc., Billerica, USA) and used freshly.
Bacterial Strains and Culture Conditions
To exclude the possibility that the effect of CFS was strain-dependent, 15 different strains of K pneumoniae and another 15 different strains of P aeruginosa were tested. K pneumoniae and P aeruginosa were isolated from urine samples of patients suffering from UTI admitted to the Urology Unit, Assiut University hospitals. The identity of these isolates was determined using the API 20E and API20NE identification system (biomerieux, France). Testing the production of ESBL is described in the next section. For preparation of bacterial suspensions, separate fresh colonies were inoculated into Muller Hinton Broth (MHB; Thermo Fisher Oxoid, UK) and cultured overnight at 37°C. Cell density was determined by measuring the optical density at 600 nm (OD600) using a spectrophotometer (Epoch, USA). These clinical isolates were used to test the antimicrobial and antibiofilm activities of the probiotic supernatants.
Antibiotic Susceptibility Testing
Susceptibility of K pneumoniae and P aeruginosa isolates to different antibiotics including, Ampicillin, Amoxycillin, Aztereonam, Cefepime, Cefotaxime, Cefoperazone, Ceftazidime, Ceftriaxone, Imipenem, Meropenem, Gentamicin, Amikacin, Amoxycillin/clavulanic acid, Trimethoprim/sulphamethoxazole were investigated by using Kirby-Bauer disk diffusion method and diameters of inhibition zones were measured and compared with the zones reported by CLSI.20 In addition, bacteria were tested for ESBL production by initially screening the isolates for reduced susceptibility to ceftazidime, ceftriaxone, and cefotaxime. Then ESBL production was confirmed using the combined disc synergy testing between ceftazidime versus ceftazidime-clavulanate and cefotaxime versus cefotaxime-clavulanate where ESBL production was indicated by a ≥5 mm increase in the inhibition zone diameter for the antimicrobial agent tested in combination with β-lactamase inhibitor versus its zone when tested alone.21
Assessment of the Antibacterial Activity Using the Well-Diffusion Method
The antimicrobial activity of supernatants isolated from lactobacilli was evaluated initially according to the agar well diffusion assay. Mueller Hinton agar plates (Oxoid, USA) were swabbed on the surface with cultures of 15 different pathogenic ESBL-producing K pneumoniae or P aeruginosa strains adjusted to approximately 105 CFU/mL. Then, 5 mm diameter wells were prepared and CFS (100 ul) was added in the wells. After incubation at 37°C for 24 h, the diameter of the inhibition zone around the well was measured.22 A negative control that consisted of MRS broth without added CFS was included.
Effect of CFS on the Viability of the Pathogenic Bacteria
The impact of CFS on the viability of ESBL-producing K pneumoniae and P aeruginosa was evaluated using the MTT assay (Promega, USA). Briefly, K pneumoniae and P aeruginosa (15 strains each) were sub-cultured in LB medium at 37°C overnight. After reaching confluence (OD600=0.5), 100 µL of each culture was added per well in a 96-well plate, followed by the addition of 100 µL lactobacilli CFS or MRS medium alone (negative control). The cultures were incubated at 37°C for 6 h then 5 µL MTT solution (CellTiter 96 Non-Radioactive Cell Proliferation Assay, Promega, USA) was added and incubated for 1 h at 37°C in the dark. Supernatants were carefully discarded, and wells were washed with 250 µL PBS then the formed crystals were dissolved using 100 µL of a solubilizing agent. The developed color was measured at 570 nm using a microplate reader (Epoch, USA). The absorbance values were expressed as the mean percentage of viability relative to control untreated samples.
Kinetic Growth of Pathogenic Bacteria in Lactobacilli CFS-Containing Media
Suspensions of K pneumoniae or P aeruginosa (15 strains each) in MHB of bacteria (100 ul) with OD 600 =0.2 were added to wells of 96-well plate and incubated at 37°C for 1–3 h with shaking at 200 r/min. CSF was added at a concentration of 100% and the number of viable bacteria was evaluated by plating out the broth-containing bacteria on the surface of MH agar at 0, 4, 8, 12, 16 and 24 h after incubation and counting the number of CFUs.
Biofilm Removal Activity of Lactobacillus CFS
Testing the biofilm formation was performed using a 96-well microtiter plate as previously described with some modification.23 Briefly, overnight cultures of ESBL-producing K pneumoniae and P aeruginosa (15 strains each) were diluted 1:100 into 15 mL of MHB supplemented with 2% w/v sucrose (Sigma-Aldrich, USA), in presence or absence of different concentrations of probiotic supernatants. Cultures were incubated at 37C for 24 h. After incubation, the developed biofilm was washed 3 times with 200 μL of distilled water and allowed to air dry. Then, 100 μL of 0.2% crystal violet (Merck KGaA, Germany) were added to each well, and the plate was incubated for 20 min to allow for biofilm staining. Wells were washed three times with distilled water, air-dried, and treated with 200 μL of 95% ethanol (Sigma-Aldrich, USA) to dissolve the crystal violet crystals. The plate was incubated for 30 mins and the intensity of the crystal violet was measured at 570 nm using a microplate reader (Epoch, USA). The ability of CFS to affect biofilm formation was measured by comparing the absorbance values of the CFS treated wells versus untreated control wells. To test the ability of CSF to remove the formed biofilms, the same procedures described above were employed, but before the staining step, different aliquots of CFS were gently added to the overnight incubated cultures and then further incubated for 30 min at room temperature then the strength of the biofilm was re-evaluated. Each sample was analyzed in triplicate. To estimate the reduction percent of CFS, the below formula was applied:
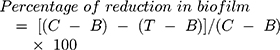
Where C is OD570nm of wells containing bacterial cells without CFS treatment, B is OD570nm of negative controls and T is OD570nm of wells treated with CFS.
Invasion Assay by Flow Cytometry
ESBL-producing K pneumoniae and P aeruginosa were first FITC labeled. To do that, overnight K pneumoniae and P aeruginosa cultures in MHB were prepared. Cells were pelleted, washed in sterile saline and then resuspended in 0.1M NaHCO3 buffer (pH 9), supplemented with 100 μg/mL FITC (Sigma Aldrich, Germany) and incubated for 30 min with constant agitation in the dark. Bacteria were then washed with saline, fixed using 0.5% formaldehyde in PBS for 30 mins, washed and resuspended in PBS supplemented with 1% bovine serum albumin (BSA). 293 cells (kindly provided by tissue culture facility of VACERA, Cairo) were seeded to 24-well plates at a density of 5x104 cells/well. Before the experiment, cells were washed with PBS and maintained in 1 mL of DMEM medium (Gibco), supplemented with 10 mM HEPES and 5% BSA. Cells were treated with FITC-labelled bacteria at an MOI of 10 and incubated at 37°C for 6 h. To test the effect of CFS of lactobacilli, 1 mL of CFS was added to test wells and the broth was added to control cells. Gentamycin (20 mg/mL) was added 20 min before the end of the incubation period to lyse any remaining extracellular bacteria. At the end of the incubation, cells were washed with PBS, trypsinized, and analyzed on a FacsCALIBUR (Becton Dickinson) flow cytometer to determine the percentage of the intracellular bacteria. The effect of CFS on the invasion of K pneumoniae and P aeruginosa to 293 cells was assessed by comparing the invasive abilities of the microbes in the presence or absence of CFS.
Statistical Analysis
The Student’s t-test was used to evaluate the effect of CFS treatment. The significance level was set at P<0.05 for all comparisons. All statistical analyses were performed using SPSS version 16.
Results
Isolation and Identification of Lactobacilli
Different yogurt samples were collected from the milk vendors. For presumptive phenotypic identification, samples were cultured on MRS agar and after culturing for 48 h, separate while colonies on MRS agar were selected and identified as Lactobacillus spp. based on their Gran-positive rod-shaped morphology, catalase and oxidase activity, sporulation, and cell motility. Lactobacillus spp. were the predominantly isolated organisms from the total bacterial isolates. The Lactobacillus spp. were further identified to the species level by using 16S ribosomal RNA sequencing analysis. A total of 13 Lactobacillus isolates, including L. acidophilus (n=6), L. plantarum (n = 3), L. paracasei (n = 2), L. fermentum (n = 1), L. bugaricus (n = 1) were isolated in this study. In our experiments, L. acidophilus was considered since L. acidophilus was the most predominantly isolated organism. To ensure consistency and reproducibility, CFS was prepared from a single pure culture of L. acidophilus in all experiments.
Antimicrobial Susceptibility of Tested Pathogens
The antibiotics susceptibility testing of K pneumoniae and P aeruginosa isolates indicated that the tested strains were highly resistant to most tested antibiotics as ampicillin, amoxicillin, ciprofloxacin, trimethoprim/sulfamethoxazole, third-generation cephalosporins in a range of 66.6–100%. The antibiotics which showed good activity included imipenem, meropenem, gentamycin, and amikacin. All isolates were ESBL producers (Table 1).
 |
Table 1 Antibiotic Resistant Pattern of K pneumoniae and P aeruginosa Isolates |
Antibacterial Activity of CFS
CFS of lactobacilli exhibited antibacterial activity against all tested ESBL-producing strains but with variable degrees. The effect of CFS was greater on K pneumoniae than on P aeruginosa. The mean diameter of the inhibition zone was larger in case of K pneumoniae compared to P aeruginosa isolates (17±2.4 mm and 13±1.3 mm respectively; p < 0.05) as shown in Table 2. To confirm these results, we tested the effect of CFS on the viability of K pneumoniae and P aeruginosa using the MTT assay. CFS was able to reduce the viability of K pneumoniae and P aeruginosa strains by 65%±13 and 53%±15 respectively after only 6 h of incubation compared to control samples incubated in the absence of CFS.
 |
Table 2 Inhibition Zone Diameters Induced by CSF on K pneumoniae and P aeruginosa |
Kinetics of Bacterial Growth in Lactobacilli CFS-Containing Media
Figure 1 shows the results of the time-killing test and assessment of the ability of CSF to suppress the growth of ESBL-producing K pneumoniae and P aeruginosa. The growth of K pneumoniae and P aeruginosa was significantly inhibited after culturing with CFS in all tested time points compared to control cultures incubated without CFS (all p <0.05 except P aeruginosa at 4 h). The inhibitory effect was more prominent on K pneumoniae cultures, as indicated by the lower CFU, particularly after 24 h of incubation. CFS induced about 2.1 log reduction in the growth of K pneumoniae and about 1.5 log reduction of P aeruginosa growth compared to control cultures incubated in the absence of CFS.
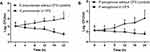 |
Figure 1 CFS of L acidophilus decreased the growth of both K pneumoniae (A) and P aeruginosa (B) in a time-dependent manner. |
Assessment of the Antibiofilm Activity
All tested strains produced an OD570 > 0.24 and were classified as potent biofilm producers. A dose-dependent reduction in biofilm formation was observed when different concentrations of CFS were added to the growing biofilms of K pneumoniae and P aeruginosa. Moreover, when 24 h biofilms were challenged with CFS, the formed biofilms were disrupted and removed by 52%±12 and 41%±15, respectively (Figure 2).
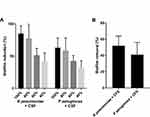 |
Figure 2 Anti-biofilm activity of different concentrations of L acidophilus-CFS against K pneumoniae and P aeruginosa (A), and against 24 h-old biofilms (B) of K pneumoniae and P aeruginosa. |
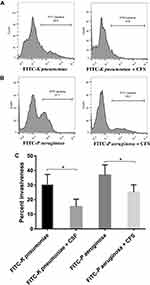
Effect of CFS on Bacterial Invasion Ability
Virulent bacteria can invade target cells and initiate their damage. We, therefore, tested the invasiveness of K pneumoniae and P aeruginosa strains in the presence of CFS. Based on the flow cytometry histogram (Figure 3), FITC-labelled K pneumoniae and FITC-labelled P aeruginosa strains displayed a similar degree of invasiveness to the 293 cells. Interestingly, CFS was able to decrease the invasive ability of these pathogens. The mean percent of intracellular FITC- K pneumoniae was reduced from 30.3%±7 to 15.4%±5 (14.9% reduction of K pneumoniae invasion) while the mean percent of FITC- P aeruginosa decreased from 36.9%±7 to 25.2%±5 (11.7% reduction of P aeruginosa invasion).
Discussion
In this study, we described the inhibitory activities of lactobacilli’s CFS on ESBL producing K. pneumoniae and P. aeruginosa. Lactobacilli are well-known probiotics that are commonly used in dairy products.24 These probiotic strains exhibit antibacterial activities through different mechanisms, including the production of organic acids, hydrogen peroxide, antimicrobial peptides, and bacteriocins.25,26 CFS of Lactobacillus Casei had anti-Shigella activities in vitro. This antibacterial activity was attributed to the production of organic acids since neutralization of the supernatants abolished their antibacterial effects.27 Similarly, lactobacillus CFS exhibited antibacterial activities against a wide range of pathogens, including Listeria monocytogens, Staphylococcus aureus, Staphylococcus epidermis, Bacillus subtilis, Salmonella typhimurium and E coli O157:H7, S. typhimurium, E. cloacae, P. aeruginosa, E. faecalis and Clostridium difficile, H. pylori, Campylobacter jejuni.26,28–31 CFS preparations for 46 lactic acid bacteria isolated from different raw and fermented milk products exhibited antibacterial activities against Enterococcus faecalis, E. coli, Salmonella spp, Shigella sonnei, Staphylococcus aureus, MRSA, and Listeria monocytogenes.32 Interestingly, the antimicrobial activity of probiotics is not confined to bacteria. Yasui, Kiyoshima, Ushijima33 showed that probiotic administrated provided protection against rotavirus-induced diarrhea. Moreover, it was demonstrated that the inhibitory activity of lactobacilli supernatants was not affected by environmental factors and storage conditions29 which is an advantage if CFS is used commercially as a therapeutic agent.
In addition to their antimicrobial activities, consumption of lactic acid bacteria has many health benefits, including enhancement of the immune system, prevention of intestinal infections, increasing lactose metabolism, decreased levels of blood ammonia and cholesterol, and strong tolerance to gastric acid and bile.34,35 In addition, CFS of Lactobacillus spp. derived from Malaysian kefir had a strong antioxidant activities.36 Some lactic acid bacteria have shown promising results in controlling the pathogenic bacteria in poultry, which in turn supports their use in improving the poultry performance and prevents transmission of poultry pathogens to humans.37
We employed different strategies to confirm the inhibitory activity of the CFS. First, the classical well diffusion test was used as a screening tool. The ability of CFS to induce remarkable inhibition zones in all tested strains shows that the inhibitory activity is not strain-dependent. Given the fact that all tested K pneumoniae and P aeruginosa strains were ESBL producers, it could be concluded that CFS is an excellent strategy to get rid of these difficult-to-treat pathogens. Similar to our results, Chen et al10 showed that some Lactobacillus strains are able to inhibit carbapenem-resistant Enterobacteriaceae.
Different tests were used to explore the antimicrobial activities of lactobacilli’s CFS. In the time-kill studies, the presence of CFS decreased the growth rate of both K pneumoniae and P aeruginosa by at least 1.5 logs after only 24 h. Additionally, CFS was able to reduce the viability of the strains, as evidenced by the MTT assay pointing to a decrease in the metabolic activity of these cells. Consistent with our results, Lactobacillus acidophilus isolated from the stool of an Iranian child induced a 90% reduction of P aeruginosa growth after 72 h of incubation.38
Virulent bacteria are able to invade their target cells and initiate their damage. We, therefore, tested the invasiveness of K pneumoniae and P aeruginosa strains in the presence of CFS. According to our flow cytometry results, CFS induced a partial inhibition of the invasion of the Gram-negative pathogens. Our results provide another mechanism by which lactobacilli can prevent the pathogenesis of these bacteria. Previous reports have shown that lactobacilli have adhesive properties to the uroepithelial cells, which prevents colonization by uropathogens.39,40 Inhibition of the adherence and colonization of uropathogens to the uroepithelial cells is carried out by cell wall fragments of the lactobacilli, lipoteichoic acid, competitive exclusion, or steric hindrance.41 Healthy women who received encapsulated Lactobacillus rhamnosus GR-1 plus Lactobacillus fermentum RC-14 for 28 days, had correlated with a healthy vaginal flora.42 Taken together, the ability of probiotics to inhibit the adherence and invasive characters of uropathogens underlines their potential use in the prevention of genitourinary tract infections such as UTI or vaginitis.
Several experimental and clinical studies have assessed the potential use of lactobacilli for the prevention or treatment of infections caused by bacterial biofilms. CFS was able to reduce biofilm formation in both bacterial strains. The antibiofilm activity of lactobacilli is due to their ability to inhibit the adherence of pathogenic bacteria to biological surfaces. The ability of CFS to inhibit and remove bacterial biofilms is linked to the presence of exopolysaccharides and bio-surfactants. Kim et al43 reported that 1 mg/mL of exopolysaccharide of L. acidophilus could remove 87% and 94% of biofilms formed by enterohemorrhagic E coli on polystyrene and polyvinyl chloride microplates, respectively. Lactobacillus strain RC-14 produced a biosurfactant that has been reported to significantly inhibit the infection and adherence of S. aureus to surgical implants.44
Generally, the CFS of the L acidophilus showed stronger antibacterial and antibiofilm activities against K pneumoniae than P aeruginosa. These results are in agreement with previously reported findings that a strain of L acidophilus isolated from a commercial vaginal product was not able to inhibit the growth of resistant clinical isolates of P aeruginosa. One mechanism proposed for the reduced anti-pseudomonas activity is the resistance of P aeruginosa to the antimicrobial substances in the CFS of the lactobacilli (hydrogen peroxide, lactic acid, and bacteriocin-like molecules).38 P aeruginosa is intrinsically resistant to different antibiotics and can easily develop resistance to many other classes of antimicrobials such as those in the CFS of lactobacilli. Moreover, different virulence factors were observed in P aeruginosa, such as their elastolytic activity and dense biofilm formation, which may contribute to their ability to partially inhibit the effect of CFS.45 P. aeruginosa has a low outer membrane permeability and expresses efflux pumps that can expel antimicrobial agents out of the cell leading to intrinsic resistance to these antimicrobial agents.46 In addition to the intrinsic ability to resist antimicrobial agents, P. aeruginosa can gain resistance to antimicrobials through mutations or acquisition of resistance genes, which increases the challenges in the eradication of this pathogen and may lead to persistent infections.47
For many years different investigators have elucidated the effective role of lactobacilli in the treatment of different pathological conditions. This work supports the therapeutic efficacy of lactobacilli in treating multi-drug resistant infections, particularly those caused by K pneumoniae and P aeruginosa, and elucidates some of the mechanisms by which CFS of lactobacilli can counteract these pathogens. Based on our results, we suggest the application of CFS of lactobacilli for controlling or preventing ESBL colonization or infection caused by the Gram-negative pathogens K pneumoniae and P aeruginosa.
Abbreviations
ESBL, extended-spectrum β-lactamase; CFS, cell-free supernatants.
Acknowledgment
We would like to thank the Drug Research Xenter and the Medical Research Center at Assiut University for providing the necessary research equipment.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Chung PY. The emerging problems of Klebsiella pneumoniae infections: carbapenem resistance and biofilm formation. FEMS Microbiol Lett. 2016;363(20):fnw219. doi:10.1093/femsle/fnw219
2. Ahmed SH, Daef EA, Badary MS, Mahmoud MA, Abd-Elsayed AA. Nosocomial blood stream infection in intensive care units at Assiut University Hospitals (Upper Egypt) with special reference to extended spectrum beta-lactamase producing organisms. BMC Res Notes. 2009;2:76. doi:10.1186/1756-0500-2-76
3. Sheu CC, Lin SY, Chang YT, Lee CY, Chen YH, Hsueh PR. Management of infections caused by extended-spectrum beta-lactamase-producing Enterobacteriaceae: current evidence and future prospects. Expert Rev Anti Infect Ther. 2018;16(3):205–218. doi:10.1080/14787210.2018.1436966
4. Elgendy SG, Abdel Hameed MR, El-Mokhtar MA. Tigecycline resistance among Klebsiella pneumoniae isolated from febrile neutropenic patients. J Med Microbiol. 2018;67(7):972–975. doi:10.1099/jmm.0.000770
5. Wu M, Li X. Chapter 87 - Klebsiella pneumoniae and Pseudomonas aeruginosa. In: Tang Y-W, Sussman M, Liu D, Poxton I, Schwartzman J, editors. Molecular Medical Microbiology.
6. Ramos-Vivas J, Chapartegui-Gonzalez I, Fernandez-Martinez M, et al. Biofilm formation by multidrug resistant Enterobacteriaceae strains isolated from solid organ transplant recipients. Sci Rep. 2019;9(1):8928. doi:10.1038/s41598-019-45060-y
7. Rasamiravaka T, Labtani Q, Duez P, El Jaziri M. The formation of biofilms by Pseudomonas aeruginosa: a review of the natural and synthetic compounds interfering with control mechanisms. Biomed Res Int. 2015;2015:759348. doi:10.1155/2015/759348
8. Rushdy AA, Gomaa EZ. Antimicrobial compounds produced by probiotic Lactobacillus brevis isolated from dairy products. Ann Microbiol. 2013;63(1):81–90. doi:10.1007/s13213-012-0447-2
9. Inglin RC, Stevens MJ, Meile L, Lacroix C, Meile L. High-throughput screening assays for antibacterial and antifungal activities of Lactobacillus species. J Microbiol Methods. 2015;114:26–29. doi:10.1016/j.mimet.2015.04.011
10. Chen CC, Lai CC, Huang HL, et al. Antimicrobial activity of lactobacillus species against carbapenem-resistant enterobacteriaceae. Front Microbiol. 2019;10:789. doi:10.3389/fmicb.2019.00789
11. Kumar M, Dhaka P, Vijay D, et al. Antimicrobial effects of Lactobacillus plantarum and Lactobacillus acidophilus against multidrug-resistant enteroaggregative Escherichia coli. Int J Antimicrob Agents. 2016;48(3):265–270. doi:10.1016/j.ijantimicag.2016.05.014
12. Aiba Y, Suzuki N, Kabir AM, Takagi A, Koga Y. Lactic acid-mediated suppression of Helicobacter pylori by the oral administration of Lactobacillus salivarius as a probiotic in a gnotobiotic murine model. Am J Gastroenterol. 1998;93(11):2097–2101. doi:10.1111/j.1572-0241.1998.00600.x
13. Hudault S, Lievin V, Bernet-Camard MF, Servin AL. Antagonistic activity exerted in vitro and in vivo by Lactobacillus casei (strain GG) against Salmonella typhimurium C5 infection. Appl Environ Microbiol. 1997;63(2):513–518. doi:10.1128/AEM.63.2.513-518.1997
14. Zhang Y, Shi X, Hao S, et al. Inhibition of Shigella sonnei-induced epithelial barrier disruption by surface-layer associated proteins of lactobacilli from Chinese fermented food. J Dairy Sci. 2018;101(3):1834–1842. doi:10.3168/jds.2017-13417
15. Shokri D, Khorasgani MR, Mohkam M, Fatemi SM, Ghasemi Y, Taheri-Kafrani A. The inhibition effect of lactobacilli against growth and biofilm formation of Pseudomonas aeruginosa. Probiotics Antimicrob Proteins. 2018;10(1):34–42. doi:10.1007/s12602-017-9267-9
16. Kang MS, Lim HS, Oh JS, et al. Antimicrobial activity of Lactobacillus salivarius and Lactobacillus fermentum against Staphylococcus aureus. Pathog Dis. 2017;75(2). doi:10.1093/femspd/ftx009
17. Bergey DH, Holt JG. Bergey’s Manual of Determinative Bacteriology. Williams & Wilkins; 1994.
18. Schillinger U. Isolation and identification of lactobacilli from novel-type probiotic and mild yoghurts and their stability during refrigerated storage. Int J Food Microbiol. 1999;47(1–2):79–87. doi:10.1016/S0168-1605(99)00014-8
19. Liu W, Bao Q, Qing M, et al. Isolation and identification of lactic acid bacteria from tarag in eastern inner Mongolia of China by 16S rRNA sequences and DGGE analysis. Microbiol Res. 2012;167(2):110–115. doi:10.1016/j.micres.2011.05.001
20. CLSI. Performance standards for antimicrobial susceptibility testing: CLSI document M100-S24. Wayne, PA: Clinical and Laboratory Standards Institute; 2014.
21. Jarlier V, Nicolas MH, Fournier G, Philippon A. Extended broad-spectrum beta-lactamases conferring transferable resistance to newer beta-lactam agents in Enterobacteriaceae: hospital prevalence and susceptibility patterns. Rev Infect Dis. 1988;10(4):867–878. doi:10.1093/clinids/10.4.867
22. de Carvalho AA, de Paula RA, Mantovani HC, de Moraes CA. Inhibition of Listeria monocytogenes by a lactic acid bacterium isolated from Italian salami. Food Microbiol. 2006;23(3):213–219. doi:10.1016/j.fm.2005.05.009
23. Sancineto L, Piccioni M, De Marco S, et al. Diphenyl diselenide derivatives inhibit microbial biofilm formation involved in wound infection. BMC Microbiol. 2016;16(1):220. doi:10.1186/s12866-016-0837-x
24. Vijaya Kumar B, Vijayendra SV, Reddy OV. Trends in dairy and non-dairy probiotic products - a review. J Food Sci Technol. 2015;52(10):6112–6124. doi:10.1007/s13197-015-1795-2
25. Weichselbaum E. Potential benefits of probiotics–main findings of an in-depth review. Br J Community Nurs. 2010;15(3):110,112,114. doi:10.12968/bjcn.2010.15.3.46897
26. Muhammad Z, Ramzan R, Abdelazez A, et al. Assessment of the antimicrobial potentiality and functionality of Lactobacillus plantarum strains isolated from the conventional inner Mongolian fermented cheese against foodborne pathogens. Pathogens. 2019;8(2):71. doi:10.3390/pathogens8020071
27. Mirnejad R, Vahdati AR, Rashidiani J, Erfani M, Piranfar V. The antimicrobial effect of lactobacillus casei culture supernatant against multiple drug resistant clinical isolates of Shigella sonnei and Shigella flexneri in vitro. Iran Red Crescent Med J. 2013;15(2):122–126. doi:10.5812/ircmj
28. Abedi D, Feizizadeh S, Akbari V, Jafarian-Dehkordi A. In vitro anti-bacterial and anti-adherence effects of Lactobacillus delbrueckii subsp bulgaricus on Escherichia coli. Res Pharm Sci. 2013;8(4):260–268.
29. Koohestani M, Moradi M, Tajik H, Badali A. Effects of cell-free supernatant of Lactobacillus acidophilus LA5 and Lactobacillus casei 431 against planktonic form and biofilm of Staphylococcus aureus. Vet Res Forum. 2018;9(4):301–306. doi:10.30466/vrf.2018.33086
30. Forestier C, De Champs C, Vatoux C, Joly B. Probiotic activities of Lactobacillus casei rhamnosus: in vitro adherence to intestinal cells and antimicrobial properties. Res Microbiol. 2001;152(2):167–173. doi:10.1016/S0923-2508(01)01188-3
31. Strus M, Pakosz K, Gosciniak H, et al. Antagonistyczne dzialanie bakterii z rodzaju Lactobacillus wobec beztlenowych i mikroaerofilnych czynnikow zakazen przewodu pokarmowego (Helicobacter pylori, Campylobacter coli, Campylobacter jejuni, Clostridium difficile) [Antagonistic activity of Lactobacillus bacteria strains against anaerobic gastrointestinal tract pathogens (Helicobacter pylori, Campylobacter coli, Campylobacter jejuni, Clostridium difficile)]. Med Dosw Mikrobiol. 2001;53(2):133–142.
32. Bin Masalam MS, Bahieldin A, Alharbi MG, et al. Isolation, molecular characterization and probiotic potential of lactic acid bacteria in Saudi raw and fermented milk. Evid Based Complement Alternat Med. 2018;2018:7970463. doi:10.1155/2018/7970463
33. Yasui H, Kiyoshima J, Ushijima H. Passive protection against rotavirus-induced diarrhea of mouse pups born to and nursed by dams fed Bifidobacterium breve YIT4064. J Infect Dis. 1995;172(2):403–409. doi:10.1093/infdis/172.2.403
34. Angmo K, Kumari A, Savitri BTC. Probiotic characterization of lactic acid bacteria isolated from fermented foods and beverage of Ladakh. LWT Food Sci Technol. 2016;66:428–435. doi:10.1016/j.lwt.2015.10.057
35. Liong MT, Shah NP. Bile salt deconjugation ability, bile salt hydrolase activity and cholesterol co-precipitation ability of lactobacilli strains. Int Dairy J. 2005;15(4):391–398. doi:10.1016/j.idairyj.2004.08.007
36. Talib N, Mohamad NE, Yeap SK, et al. Isolation and characterization of Lactobacillus spp. from kefir samples in Malaysia. Molecules. 2019;24(14):2606. doi:10.3390/molecules24142606
37. Reuben RC, Roy PC, Sarkar SL, Alam R-U, Jahid IK. Isolation, characterization, and assessment of lactic acid bacteria toward their selection as poultry probiotics. BMC Microbiol. 2019;19(1):253. doi:10.1186/s12866-019-1626-0
38. Jamalifar H, Rahimi H, Samadi N, et al. Antimicrobial activity of different Lactobacillus species against multi- drug resistant clinical isolates of Pseudomonas aeruginosa. Iran J Microbiol. 2011;3(1):21–25.
39. Bruce AW, Reid G. Intravaginal instillation of lactobacilli for prevention of recurrent urinary tract infections. Can J Microbiol. 1988;34(3):339–343. doi:10.1139/m88-062
40. Reid G, Cook RL, Bruce AW. Examination of strains of lactobacilli for properties that may influence bacterial interference in the urinary tract. J Urol. 1987;138(2):330–335. doi:10.1016/S0022-5347(17)43137-5
41. Chan RC, Reid G, Irvin RT, Bruce AW, Costerton JW. Competitive exclusion of uropathogens from human uroepithelial cells by Lactobacillus whole cells and cell wall fragments. Infect Immun. 1985;47(1):84–89. doi:10.1128/IAI.47.1.84-89.1985
42. Reid G, Beuerman D, Heinemann C, Bruce AW. Probiotic Lactobacillus dose required to restore and maintain a normal vaginal flora. FEMS Immunol Med Microbiol. 2001;32(1):37–41. doi:10.1111/j.1574-695X.2001.tb00531.x
43. Kim Y, Oh S, Kim SH. Released exopolysaccharide (r-EPS) produced from probiotic bacteria reduce biofilm formation of enterohemorrhagic Escherichia coli O157: H7. Biochem Biophys Res Commun. 2009;379(2):324–329. doi:10.1016/j.bbrc.2008.12.053
44. Gan BS, Kim J, Reid G, Cadieux P, Howard JC. Lactobacillus fermentum RC-14 inhibits Staphylococcus aureus infection of surgical implants in rats. J Infect Dis. 2002;185(9):1369–1372. doi:10.1086/jid.2002.185.issue-9
45. Alexandre Y, Le Berre R, Barbier G, Le Blay G. Screening of Lactobacillus spp. for the prevention of Pseudomonas aeruginosa pulmonary infections. BMC Microbiol. 2014;14:107. doi:10.1186/1471-2180-14-107
46. Breidenstein EB, de la Fuente-núñez C, Hancock RE. Pseudomonas aeruginosa: all roads lead to resistance. Trends Microbiol. 2011;19(8):419–426. doi:10.1016/j.tim.2011.04.005
47. Pang Z, Raudonis R, Glick BR, Lin TJ, Cheng Z. Antibiotic resistance in Pseudomonas aeruginosa: mechanisms and alternative therapeutic strategies. Biotechnol Adv. 2019;37(1):177–192. doi:10.1016/j.biotechadv.2018.11.013
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.