Back to Journals » Infection and Drug Resistance » Volume 13
An NDM-1-Producing Acinetobacter towneri Isolate from Hospital Sewage in China
Authors Wang K, Li P, Li J, Hu X, Lin Y, Yang L, Qiu S, Ma H, Li P, Song H
Received 20 January 2020
Accepted for publication 26 March 2020
Published 16 April 2020 Volume 2020:13 Pages 1105—1110
DOI https://doi.org/10.2147/IDR.S246697
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Joachim Wink
Kaiying Wang,1,2,* Peihan Li,1,2,* Jinhui Li,2,* Xiaofeng Hu,2,* Yanfeng Lin,1,2 Lang Yang,1,2 Shaofu Qiu,2 Hui Ma,*,3 Peng Li,2 Hongbin Song2
1College of Military Medicine, Academy of Military Sciences, Beijing, People’s Republic of China; 2Center for Disease Control and Prevention of PLA, Beijing, People’s Republic of China; 3The Sixth Medical Center of PLA General Hospital, Beijing, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Peng Li; Hongbin Song Tel/ Fax +86-10-66948475
Email [email protected]; [email protected]
Background: The New Delhi metallo-β-lactamase-1 (NDM-1)-positive plasmid and its variants pose daunting threats to public health. Hospital sewage was considered as an important reservoir of antibiotic genes. Numerous and diverse taxa of multidrug-resistant (MDR) bacteria carrying NDM-1-positive plasmids have been identified during routine surveillance of hospital sewage. We herein report a carbapenem-resistant Acinetobacter towneri strain AeBJ009 with an NDM-1-positive plasmid isolated from hospital sewage.
Materials and Methods: Bacteria were isolated from cultures of hospital sewage and identified by using the Vitek 2 compact system and 16S rRNA sequencing. The blaNDM-1 gene was amplified and confirmed by sequencing. Antimicrobial susceptibility testing was performed using AST-GN14 on the Vitek2 compact system. In addition, the blaNDM-1 gene was located by Southern blotting. Conjugation experiment and whole-genome sequencing were performed for further analysis.
Results: Strain AeBJ009 was isolated from hospital sewage and identified as A. towneri. Antimicrobial susceptibility testing revealed an MDR phenotype. Pulsed-field gel electrophoresis and Southern blotting showed that strain AeBJ009 carries three plasmids and that blaNDM-1 is located on the 47kb plasmid pNDM-AeBJ009. However, the conjugation experiment to transfer pNDM-AeBJ009 to Escherichia coli strain J53 was unsuccessful. Whole-genome sequencing found that pNDM-AeBJ009 contains a Tn 125 element carrying blaNDM-1. The ble gene downstream of blaNDM-1 displayed a single-nucleotide polymorphism compared to its homologue on plasmid pM131_NDM1. BLAST analysis using the Comprehensive Antibiotic Resistance Database identified no gene polymorphisms with 100% identity to our ble variant.
Conclusion: The A. towneri strain AeBJ009 exhibiting an extended spectrum of antibiotic resistance was isolated from hospital sewage and may potentially exacerbate the risk of MDR bacterial infections. The prevention of nosocomial infections due to drug-resistant bacteria will require enhanced monitoring and control of MDR pathogens in environmental reservoirs.
Keywords: NDM-1, ble, Acinetobacter towneri, drug resistance
Introduction
Multidrug-resistant (MDR) pathogens and residual antibiotics have been detected in hospital sewage consequent to their discharge into wastewater during the therapy of patients with infectious diseases.1 Hospital sewage is an ideal environment for the selection of MDR pathogens,2–4 for the interspecies exchange of drug-resistance genes on mobile genetic elements, and may serve as an environmental reservoir for their further spread.5 Surveillance of hospital sewage has shown that many MDR bacteria belong to New Delhi metallo-β-lactamase-1 (NDM-1)-positive strains.6
NDM-1 can hydrolyze nearly all beta-lactam antibiotics, including carbapenems.7 The blaNDM-1 gene was widely distributed among gram-negative bacteria,8,9 most of which are found on conjugative plasmids enhancing the spread of resistance.10 Strains carrying blaNDM-1 are most often members of either the Enterobacteriaceae family11,12 or the Acinetobacter genus.13,14 Acinetobacter spp. can be isolated from multiple environmental reservoirs,15 and are considered among the most formidable nosocomial pathogens.16 NDM-1-positive plasmids have been detected with increasing frequency in Acinetobacter spp.17 Studies from China have reported that Acinetobacter towneri isolates from both clinical specimens18 and hospital sewage19 carry multiple resistance genes.
We here reported an MDR A. towneri strain AeBJ009 with a blaNDM-1-harboring plasmid. Further experiments and whole-genome sequencing revealed the plasmid shared nearly the same sequence with a transferable plasmid of a clinical sample but was non-conjugative, intriguing us to explore the genetic characteristics of the blaNDM-1 gene. Our results underscore the potential threat of the transmission of bla genes in environmental reservoirs, and highlight the roles of enhanced environmental surveillance of antimicrobial resistance and treatment of hospital sewage as risk reduction strategies.
Materials and Methods
Bacterial Isolation and Identification
Hospital sewage was collected and concentrated using centrifuge GR22G II (HITACHI, Japan) in Beijing in 2011. Bacteria was isolated by incubating the resuspended sediment on MacConkey agar plates containing 4μg/mL meropenem. Strain AeBJ009 was recovered and identified using the Vitek2 compact system (BioMérieux, France). The 16S rRNA gene was amplified with the universal primers (27F: 5ʹ-AGAGTTTGATCCTGGCTCAG-3ʹ and 1492R: 5ʹ-TACGGCTACCTTGTTACGACTT-3ʹ).20 The blaNDM-1 gene was detected by PCR and sequencing with primers NDM-F-38 (5ʹ-GGCGGAATGGCTCATCACGA-3ʹ) and NDM-R-344 (5ʹ-CGCAACACAGCCTGACTTTC-3′).21 An NDM-1-carrying K. pneumoniae strain KP1400322 from our lab was used as positive control.
Antimicrobial Susceptibility Testing
The strain was cultured on LB agar plates. One hundred and forty-five microliter bacterial suspension of a 0.5-McFarland turbidity was mixed with 3mL 0.45% NaCl solution. The AST-GN14 card filled with the mixture was used. The minimum inhibitory concentrations (MICs) of ampicillin, amoxicillin/clavulanic acid, piperacillin, cefazolin, ceftazidime, ceftriaxone, cefepime, aztreonam, imipenem, meropenem, amikacin, gentamicin, ciprofloxacin, levofloxacin, tetracycline, nitrofurantoin and sulfamethoxazole/trimethoprim were determined by the Vitek2 compact system (BioMérieux, France) following the manufacturer’s instructions. Antimicrobial sensitivity results were interpreted according to the Clinical and Laboratory Standards Institute guidelines (M100-S24).23 The E. coli ATCC25922 and E. coli J53 were used as quality control and negative control for antimicrobial testing, respectively.
Southern Blotting
Four hundred microliter bacterial suspension with turbidity of 3.7–4.2 McF was used for gel preparation. Genomic DNA was digested with S1 nuclease (Code No: 2410A, Takara). The linearized DNA fragments were separated by pulsed-field gel electrophoresis (PFGE) (Bio-Rad, Hercules, CA, USA). DNA on gel was dyed with ethidium bromide. Plasmid DNA was transferred to a Hybond N+® membrane (Sigma-Aldrich, St. Louis, MO, USA) and hybridized with a blaNDM-1 probe that was labeled with digoxin. The experiment was performed according to the manufacturer’s manual of the DIG High Prime DNA Labeling and Detection Start Kit I (Cat. No: 11745832910, Roche).
Conjugation Experiment
Conjugation was carried out by broth and filter mating using azide-resistant Escherichia coli strain J53 as the recipient and the AeBJ009 strain as the donor. The donor/recipient LB suspensions in logarithmic phase were mixed at a 4:1 ratio. The mixture was incubated at 37°C for 18 hours. Transconjugants were selected on MacConkey agar plates containing 200μg/mL sodium azide and 4μg/mL meropenem.
Whole-Genome Sequencing and Analysis
Genomic DNA was extracted by TIANamp Bacteria DNA Kit (Cat. No: DP302, TIANGEN BIOTECH) from cultured bacterium. A total of 700ng genomic DNA was used for library preparation using NEBNext® Ultra™ II DNA Library Prep Kit (Cat. No: E7645S, NEB) according to the manufacturer’s manual. Whole-genome sequencing was performed by Novogene Company (Beijing, China) on the Illumina HiSeq X platform using Dual Flow Cell with 150bp pair-end reads. Reads were assembled de novo by using Spades (v3.6.2) with k-mer sizes of 21, 33, 77, 99 and 127.24 Gene annotation of plasmids was performed on the RAST webserver using the RASTtk pipeline with default parameters.25 Plasmid replicon types were identified using PlasmidFinder 1.3.26
Results
Isolation, Identification, and Antimicrobial Susceptibility Testing of Strain AeBJ009
Strain AeBJ009 was identified as A. towneri by using the Vitek2 compact system, and confirmed by blasting the 16S rRNA sequence against the nt database of NCBI (Figure S1). A. towneri AeBJ009 showed resistance to cefazolin, ceftazidime, ceftriaxone, meropenem, ciprofloxacin, nitrofurantoin, and trimethoprim/sulfamethoxazole, but was sensitive to amoxicillin/clavulanic acid, piperacillin, cefepime, aztreonam, amikacin, levofloxacin and tetracycline (Table 1). The results of S1 PFGE revealed that AeBJ009 carried three plasmids (~47kb, ~76kb and ~300kb), and Southern blotting indicated that blaNDM-1 was located on the ~47kb plasmid (Figure S2). Conjugation using E. coli J53 as the recipient was unsuccessful.
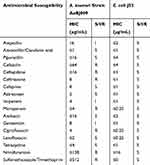 |
Table 1 Antibiotic Susceptibilities of A. towneri Strain AeBJ009 |
Genetic Features and Plasmid Characteristics
Genome sequencing and analysis revealed that the A. towneri AeBJ009 chromosome contains approximately 3.02Mb. There were 18 resistance genes (Table 2) including blaPER-1, blaNDM-1, blaBRP, blaOXA-58, blaCTX-M-55 and blaTEM-30. Three resistant genes (blaNDM-1, blaBRP and aph (3ʹ)-VI) were located on the same plasmid, designated as pNDM-AeBJ009. This plasmid has a length of 47,271 bp and 40.8% guanine-cytosine content, and contains 55 open reading frames. The plasmid contains three sections that encode plasmid transfer and replication, a type IV secretion system, and Tn125, respectively (Figure 1).
 |
Table 2 Resistance Genes of A. towneri Strain AeBJ009 |
pNDM-AeBJ009 could not be matched to any plasmid replicon type by PlasmidFinder. An NCBI BLAST analysis showed that the identity between plasmids pNDM-AeBJ009 and pM131_NDM1 (JX072963.1)27 is over 99% (Figure 2). A composite transposon Tn125 structure of pNDM-AeBJ009 is located downstream of the aphA6 gene carrying blaNDM-1. The second copy of ISAba125 in blaNDM-1 downstream is reversed from that of plasmid pNDM-JAVP01.28 Compared with plasmid pNDM229,29 there is no ISAba14 copy between the downstream Tn125 element and tnpR in plasmid pNDM-AeBJ009 (Figure 2)
Plasmid pNDM-AeBJ009 has 41 single-nucleotide polymorphism (SNPs) compared to plasmid pM131_NDM1. Tn125 has 14 SNPs, while the upstream and downstream IS have two and eleven SNPs, respectively. The only SNP in ble lead to the change of the 17th base from cytosine to thymidine. This SNP substitutes threonine for isoleucine during translation. Of the remaining SNP loci, four were in orfA and one in VirB10.
Discussion
The blaNDM-1 gene has been identified in multiple strains since first reported in Klebsiella pneumoniae.7 Many studies report that blaNDM-1 is not only found in clinical isolates, but also in environmental bacteria from sources such as hospital sewage.3 In this study, we isolated an A. towneri strain with an NDM-1-carrying plasmid from hospital sewage. The blaNDM-1 gene was located on the common transposon Tn125, which is a primary carrier in the spread of blaNDM-1 in Acinetobacter spp.30,31
Compared to other whole plasmid sequences in the NCBI database, pNDM-AeBJ009 was similar only to pM131_NDM1, and quite different from others. Similar to pM131_NDM, pNDM-AeBJ009 did not match any plasmid replicon type by PlasmidFinder. According to plasmid typing principles, the pNDM-AeBJ009 might lack a typing sequence, or its typing sequence is not yet in the database. Further studies are needed for the typing of pNDM-AeBJ009.
In the conjugation experiment, horizontal transfer of pNDM-AeBJ009 to E. coli strain J53 failed. However, the similar plasmid pM131_NDM1 can be effectively transferred to non-pathogenic Acinetobacter spp.27 An attempt to transfer plasmid pNDM22929 to different recipient strains also failed because of truncation or absence of conjugation-related genes. The plasmid pNDM-JAVP0128 was transferred to the recipient strain A. baumannii ATCC 17978 but failed to transfer to E. coli MC1061. Compared to pM131_NDM1, pNDM-AeBJ009 has 14 SNPs in Tn125 and one SNP in the conjugation-related gene virB10. These mutations may have caused the failed conjugation. Another possible explanation for failed conjugation may be that the E. coli J53 strain may not be a suitable recipient for pNDM-AeBJ009. For example, morphologic features of the strain may affect cell wall permeability and consequently impair conjugation. We cannot conclude that pNDM-AeBJ009 has no potential for horizontal transfer; further studies are needed to clarify this issue. The different transferability of these two highly similar plasmids indicates that further understanding of the mechanisms promoting plasmid transfer is therefore of importance.
Our genotypic analysis disclosed a SNP in ble in which the 17th base is changed from cytosine to thymidine, causing an amino acid substitution during translation. This result suggests that the ble SNP may be a new drug-resistance mutation that increases the risk of plasmid-mediated high-level drug resistance. This novel ble variant may have emerged under selection pressure from biologically relevant concentrations of multiple antibiotics in hospital sewage. Though studies on the NDM variants showed that the amino acid changes were associated with different hydrolysis activity,32 the effect of the amino acid substitution on the ble gene needs further study.
Moreover, other genes encoding OXA-58, CTX-M-55, and TEM-30 were also detected in our AeBJ009 strain. A previous study revealed that the prevalence of these beta-lactamase genes in hospital effluent was very high.3 The challenge of resistance genes and novel mutants in hospital sewage is worthy of further study. As a reservoir of multiple resistance genes, hospital sewage may have an especially important clinical and epidemiologic relevance.
Conclusions
We report an A. towneri strain AeBJ009 that was isolated from hospital sewage and characterized by bioinformatic analysis. This strain carried multiple bla genes and a novel ble variant. The co-existence of these genes may confer increased carbapenem resistance. The findings of resistant bacteria and novel resistance genes in hospital sewage highlight the potential role of hospital sewage as an environmental reservoir of MDR pathogens that deserves further attention for surveillance and risk management.
Accession Number
The whole-genome sequence of Acinetobacter towneri strain AeBJ009 has been deposited at DDBJ/ENA/GenBank under accession NZ_SIST01000000. The complete sequences of plasmid pNDM-AeBJ009 and the new ble variant have been deposited in GenBank under accession numbers CM016430 and MN886241, respectively.
Acknowledgments
The present study was funded by grants from the Mega-projects of Science and Technology Research (No. 2018ZX10712001-002-002), the Beijing Natural Science Foundation (No. 5172029) and the Beijing Noval program (No. Z181100006218110).
Disclosure
The authors report no conflicts of interest in this work.
References
1. Lamba M, Graham DW, Ahammad SZ. Hospital wastewater releases of carbapenem-resistance pathogens and genes in Urban India. Environ Sci Technol. 2017;51(23):13906–13912. doi:10.1021/acs.est.7b03380
2. Walsh TR, Weeks J, Livermore DM, Toleman MA. Dissemination of NDM-1 positive bacteria in the New Delhi environment and its implications for human health: an environmental point prevalence study. Lancet Infect Dis. 2011;11(5):355–362. doi:10.1016/S1473-3099(11)70059-7
3. Marathe NP, Berglund F, Razavi M, et al. Sewage effluent from an Indian hospital harbors novel carbapenemases and integron-borne antibiotic resistance genes. Microbiome. 2019;7(1):97. doi:10.1186/s40168-019-0710-x
4. Bengtsson-Palme J, Larsson DGJ. Concentrations of antibiotics predicted to select for resistant bacteria: proposed limits for environmental regulation. Environ Int. 2016;86:140–149. doi:10.1016/j.envint.2015.10.015
5. Zhang C, Qiu S, Wang Y, et al. Higher isolation of NDM-1 producing Acinetobacter baumannii from the sewage of the hospitals in Beijing. PLoS One. 2013;8(6):e64857. doi:10.1371/journal.pone.0064857
6. Moges F, Endris M, Belyhun Y, Worku W. Isolation and characterization of multiple drug resistance bacterial pathogens from waste water in hospital and non-hospital environments, Northwest Ethiopia. BMC Res Notes. 2014;7(1):215. doi:10.1186/1756-0500-7-215
7. Yong D, Toleman MA, Giske CG, et al. Characterization of a new metallo-β-lactamase gene, blaNDM-1, and a novel erythromycin esterase gene carried on a unique genetic structure in Klebsiella pneumoniae sequence type 14 from India. Antimicrob Agents Chemother. 2009;53(12):5046–5054. doi:10.1128/AAC.00774-09
8. Bhattacharya D, Thamizhmani R, Bhattacharya H, et al. Emergence of New Delhi metallo-β-lactamase 1 (NDM-1) producing and multidrug resistant uropathogens causing urinary tract infections in Andaman Islands, India. Microbial Drug Resist. 2013;19(6):457–462. doi:10.1089/mdr.2013.0070
9. Dortet L, Poirel L, Nordmann P. Worldwide dissemination of the NDM-type carbapenemases in Gram-negative bacteria. Biomed Res Int. 2014;2014:1–12. doi:10.1155/2014/249856
10. Kumarasamy KK, Toleman MA, Walsh TR, et al. Emergence of a new antibiotic resistance mechanism in India, Pakistan, and the UK: a molecular, biological, and epidemiological study. Lancet Infect Dis. 2010;10(9):597–602. doi:10.1016/S1473-3099(10)70143-2
11. Samuelsen Ø, Thilesen CM, Heggelund L, Vada AN, Kümmel A, Sundsfjord A. Identification of NDM-1-producing Enterobacteriaceae in Norway. J Antimicrob Chemother. 2011;66(3):670–672. doi:10.1093/jac/dkq483
12. Deshpande P, Rodrigues C, Shetty A, Kapadia F, Hedge A, Soman R. New Delhi Metallo-beta lactamase (NDM-1) in Enterobacteriaceae: treatment options with carbapenems compromised. J Assoc Physicians India. 2010;58:147–149.
13. Pan C-Y, Chen J-C, Chen T-L, Wu J-L, Hui C-F, Chen J-Y. Piscidin is highly active against carbapenem-resistant acinetobacter baumannii and NDM-1-producing klebsiella pneumonia in a systemic septicaemia infection mouse model. Mar Drugs. 2015;13(4):2287–2305. doi:10.3390/md13042287
14. Huang Y-M, Zhong L, Zhang X-F, et al. NDM-1-producing citrobacter freundii, escherichia coli, and acinetobacter baumannii identified from a single patient in China. Antimicrob Agents Chemother. 2015;59(8):5073–5077. doi:10.1128/AAC.04682-14
15. Barbosa BG, Fernandez-García L, Gato E, et al. Genome sequence of airborne Acinetobacter sp. strain 5-2Ac02 in the hospital environment, close to the species of Acinetobacter towneri. Genome Announc. 2016;4(6):e01343–16. doi:10.1128/genomeA.01343-16
16. Peleg AY, Seifert H, Paterson DL. Acinetobacter baumannii: emergence of a successful pathogen. Clin Microbiol Rev. 2008;21(3):538–582. doi:10.1128/CMR.00058-07
17. Jones LS, Toleman MA, Weeks JL, Howe RA, Walsh TR, Kumarasamy KK. Plasmid carriage of blaNDM-1 in clinical Acinetobacter baumannii isolates from India. Antimicrob Agents Chemother. 2014;58(7):4211–4213. doi:10.1128/AAC.02500-14
18. Zou D, Huang Y, Liu W, et al. Complete sequences of two novel bla NDM-1-harbouring plasmids from two Acinetobacter towneri isolates in China associated with the acquisition of Tn 125. Sci Rep. 2017;7(1):9405. doi:10.1038/s41598-017-09624-0
19. Jiang N, Zhang X, Zhou Y, Zhang Z, Zheng X. Whole-genome sequencing of an NDM-1-and OXA-58-producing Acinetobacter towneri isolate from hospital sewage in Sichuan Province, China. J Glob Antimicrob Resist. 2019;16:4–5. doi:10.1016/j.jgar.2018.11.015
20. Lane DJ. 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M, editors. Nucleic Acid Techniques in Bacterial Systematics. New York: John Wiley and Sons; 1991:115–175.
21. Chen Y, Zhou Z, Jiang Y, Yu Y. Emergence of NDM-1-producing Acinetobacter baumannii in China. J Antimicrob Chemother. 2011;66(6):1255–1259. doi:10.1093/jac/dkr082
22. Li J, Hu X, Yang L, et al. New Delhi Metallo-β-Lactamase 1-Producing Klebsiella pneumoniae ST719 Isolated from a Neonate in China. Microbial Drug Resist. 2019. doi:10.1089/mdr.2019.0058
23. Performance standards for antimicrobial susceptibility testing: twenty-four informational supplement. CLSI document M100-S24; 2014.
24. Bankevich A, Nurk S, Antipov D, et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol. 2012;19(5):455–477. doi:10.1089/cmb.2012.0021
25. Aziz RK, Bartels D, Best AA, et al. The RAST Server: rapid annotations using subsystems technology. BMC Genomics. 2008;9(1):75. doi:10.1186/1471-2164-9-75
26. Carattoli A, Zankari E, García-Fernández A, et al. In silico detection and typing of plasmids using PlasmidFinder and plasmid multilocus sequence typing. Antimicrob Agents Chemother. 2014;58(7):3895–3903. doi:10.1128/AAC.02412-14
27. Huang T-W, Lauderdale T-L, Liao T-L, et al. Effective transfer of a 47 kb NDM-1-positive plasmid among Acinetobacter species. J Antimicrob Chemother. 2015;70(10):2734–2738. doi:10.1093/jac/dkv191
28. Espinal P, Mosqueda N, Telli M, et al. Identification of NDM-1 in a putatively novel Acinetobacter species (“NB14”) closely related to Acinetobacter pittii. Antimicrob Agents Chemother. 2015;59(10):6657–6660. doi:10.1128/AAC.01455-15
29. Brovedan M, Marchiaro PM, Morán-Barrio J, et al. Complete sequence of a blaNDM-1-harboring plasmid in an Acinetobacter bereziniae clinical strain isolated in Argentina. Antimicrob Agents Chemother. 2015;59(10):6667–6669. doi:10.1128/AAC.00367-15
30. Poirel L, Bonnin RA, Boulanger A, Schrenzel J, Kaase M, Nordmann P. Tn125-related acquisition of blaNDM-like genes in Acinetobacter baumannii. Antimicrob Agents Chemother. 2012;56(2):1087–1089. doi:10.1128/AAC.05620-11
31. Bontron S, Nordmann P, Poirel L. Transposition of Tn125 encoding the NDM-1 carbapenemase in Acinetobacter baumannii. Antimicrob Agents Chemother. 2016;60(12):7245–7251. doi:10.1128/AAC.01755-16
32. Liu Z, Li J, Wang X, et al. Novel variant of New Delhi metallo-β-lactamase, NDM-20, in Escherichia coli. Front Microbiol. 2018;9:248. doi:10.3389/fmicb.2018.00248
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.


