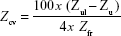Back to Journals » International Journal of Chronic Obstructive Pulmonary Disease » Volume 12
Alpha-1 antitrypsin Pi*Z gene frequency and Pi*ZZ genotype numbers worldwide: an update
Authors Blanco I , Bueno P, Diego I , Pérez-Holanda S, Casas-Maldonado F , Esquinas C, Miravitlles M
Received 21 October 2016
Accepted for publication 10 December 2016
Published 13 February 2017 Volume 2017:12 Pages 561—569
DOI https://doi.org/10.2147/COPD.S125389
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Richard Russell
Ignacio Blanco,1 Patricia Bueno,2 Isidro Diego,3 Sergio Pérez-Holanda,4 Francisco Casas-Maldonado,5 Cristina Esquinas,6 Marc Miravitlles6,7
1Alpha1-Antitrypsin Deficiency Spanish Registry (REDAAT), Fundación Respira, Spanish Society of Pneumology and Thoracic Surgery (SEPAR), Barcelona, 2Internal Medicine Department, County Hospital of Jarrio, 3Materials and Energy Department, School of Mining Engineering, Oviedo University, 4Surgical Department, University Central Hospital of Asturias (HUCA), Oviedo, Principality of Asturias, 5Pneumology Department, Complejo Hospitalario Universitario de Granada, Granada, 6Pneumology Department, Hospital Universitari Vall d’Hebron, 7CIBER de Enfermedades Respiratorias (CIBERES), Barcelona, Spain
Abstract: In alpha-1 antitrypsin deficiency (AATD), the Z allele is present in 98% of cases with severe disease, and knowledge of the frequency of this allele is essential from a public health perspective. However, there is a remarkable lack of epidemiological data on AATD worldwide, and many of the data currently used are outdated. Therefore, the objective of this study was to update the knowledge of the frequency of the Z allele to achieve accurate estimates of the prevalence and number of Pi*ZZ genotypes worldwide based on studies performed according to the following criteria: 1) samples representative of the general population, 2) AAT phenotyping characterized by adequate methods, and 3) measurements performed using a coefficient of variation calculated from the sample size and 95% confidence intervals. Studies fulfilling these criteria were used to develop maps with an inverse distance weighted (IDW)-interpolation method, providing numerical and graphical information of Pi*Z distribution worldwide. A total of 224 cohorts from 65 countries were included in the study. With the data provided by these cohorts, a total of 253,404 Pi*ZZ were estimated worldwide: 119,594 in Europe, 91,490 in America and Caribbean, 3,824 in Africa, 32,154 in Asia, 4,126 in Australia, and 2,216 in New Zealand. In addition, the IDW-interpolation maps predicted Pi*Z frequencies throughout the world even in some areas that lack real data. In conclusion, the inclusion of new well-designed studies and the exclusion of the low-quality ones have significantly improved the reliability of results, which may be useful to plan strategies for future research and diagnosis and to rationalize the therapeutic resources available.
Keywords: SERPINA1, alpha-1 antitrypsin deficiency, protease inhibitor, genetic epidemiology, inverse distance weighted interpolation, geographic information system
Introduction
Human alpha-1 antitrypsin (AAT), also named α1 proteinase inhibitor (α1-Pi) and SERPINA1 (serine protease inhibitor, group A, member 1), is a water-soluble, tissue-diffusible, medium-sized circulating glycoprotein, with a molecular weight of 52 kDa, and a blood half-life of around 5 days. AAT is mainly synthesized by hepatocytes, reaching plasma concentrations of 1–2 g L−1. From the plasma, 80% of AAT diffuses to interstitial tissues, and up to 10% reaches biological fluids. AAT acts as a major inhibitor of leukocyte elastase, proteinase-3, and bacterial serine proteases. In addition, it has anti-inflammatory and immunomodulatory properties.1 Alpha-1 antitrypsin deficiency (AATD) is a rare hereditary condition that predisposes to premature onset of COPD especially in smokers, liver cirrhosis in children and adults, neutrophilic panniculitis, systemic vasculitis, as well as other possible inflammatory, autoimmune, and neoplastic diseases.2–4
The AAT gene has two alleles, which are transmitted by autosomal codominant Mendelian inheritance. Normal alleles are present in 85%–90% of individuals and are designated as M, and the most prevalent deficiency alleles are S and Z. Consequently, the vast majority of genotypes described are the result of combinations of M, S, and Z, that is: MM (normal genotype present in approximately 90%–95% of people, expressing 100% of serum AAT); MS, SS, MZ, SZ, and ZZ (five deficiency genotypes present in the remaining 5%–10% of the population, expressing ~80%, 60%, 55%, 40%, and 15% of AAT, respectively). The Pi*Z mutation is a single-nucleotide polymorphism (SNP), which results in an amino acid substitution – glutamic acid by lysine at position 342 (Glu342Lys) – causing a substantial conformational change in the molecule causing hepatic polymerization and decreased serum concentrations. Individuals who are homozygous for the Z allele undergo significant intracellular polymerization of AAT (85%–90%), with a great reduction of plasma levels (10%–15%). AAT-Z polymers retained within the endoplasmic reticulum of hepatocytes can cause liver damage, whereas the lack of circulating protein promotes the development of COPD.2
Apart from the most common AAT-S- and Z-deficient alleles, ~50 other extremely uncommon so-called rare and null deficiency alleles have been characterized. However, in clinical practice, 96% of individuals with diseases associated with AATD exhibit Pi*ZZ genotypes, whereas only 4% present combinations of the “rare” or “null” Z or S, alleles.5–7 Thus, the Pi*Z allele is present in 98% of the subjects expressing diseases related to AATD. Taking this into account as well as the current lack of data on the frequency, prevalence, and geographical distribution of these uncommon alleles, this study was limited to the epidemiology of the Pi*Z allele.2
Knowledge of the prevalence of AATD in every community is essential for health authorities, patients, and physicians as well as the pharmaceutical companies involved in the manufacture of products for augmentation therapy. Patients with AATD-related diseases are very often misdiagnosed or undiagnosed. The physician facing a case of suspected AATD should request a serum level of AAT and, if low, two strategies for laboratory testing can be used: 1) plasma AAT phenotyping by isoelectric focusing (IEF), which allows identification of the most common AAT variants or 2) genotyping by allele-specific polymerase chain reaction method, which allows identification of the most common AATD alleles: Pi*S and Pi*Z. Molecular analysis with direct sequencing of the whole SERPINA1 gene (genotyping) is the reference method for identifying rare and null deficiency allelic variants. In addition, testing of first-degree family members of subjects diagnosed with AATD should be performed in order to take appropriate preventive and therapeutic measures including counseling related to genetic alterations, the risk of tobacco smoke, occupational exposure to pollutants and hepatotoxic substances, and information on clinical management, including intravenous augmentation therapy.3
Since Laurell and Eriksson discovered AATD in 1963,8 a large number of original studies and reviews on the genetic epidemiology of the AAT gene have been published.9–22 However, most were conducted in Europe, the US, and Canada, with a remarkable lack of data from more than half of the 193 countries in the world. Moreover, there is a great lack of uniformity in the methodology used to achieve the results of some of these studies leading to the presentation of inaccurate data and the subsequent need to consider these results with caution. It is of note that in recent years, the results of other well-designed studies have substantially changed previous findings.23–25 All of the above justify the need for an update of the current data available, and therefore, the aim of this study was to develop an updated database of valid epidemiological studies on Pi*Z frequency to make accurate estimates on the prevalence and number of Pi*ZZ genotypes throughout the world.
Methods
Study selection criteria
- Samples representative of the general population.
- AAT phenotyping characterized by adequate methods, including IEF, or alternatively, crossed antigen-antibody electrophoresis or polymerase chain reaction.
- To assess the statistical validity of the results, a coefficient of variation (CV) of the Pi*Z frequencies in each cohort was calculated. This CV is a measure of the precision of results in terms of dispersion of the data around the mean. Its value depends on the number of alleles studied and on the frequencies of Pi*Z actually found, according to the formula:
|
where Zul is the 95% confidence interval (95% CI) upper limit of Z, Zll is the 95% CI lower limit of Z, and Zfr is the frequency of Z. The value of this variable was adjusted for each country, according to the methodology used in several previous peer-reviewed publications by our group.18–20
The following studies were excluded from the analysis:
- Cohort studies of patients with AATD-related diseases (eg, COPD, liver cirrhosis, etc.).
- Screening studies in which phenotypes were determined only in samples with AAT concentrations below any given cut-off point.
Calculation of Pi*Z allele frequency
The Z allelic frequency was obtained by adding the number of Z alleles and expressing this total as a fraction of the total number of Pi alleles in the population (expressed as alleles per 1,000 of all Pi types).
Inverse distance weighted multivariate interpolation maps
Pi*Z gene frequencies were used to develop maps using a geographic information system (GIS) of an inverse distance weighted (IDW) interpolation method. IDW interpolation is based on the assumption that the attributed value of an unsampled point is the weighted average of known values within the neighborhood, and the weights are inversely related to the distances between the prediction location and the locations sampled.26 With this approach, Pi*Z frequencies were represented by scales of 10 colors (with red representing the maximum frequencies; yellow, green, and grey intermediate frequencies, and blue minimum frequencies). The colors of these maps can be converted into quantitative data, providing both graphical and numerical information on their geographical distribution worldwide.
Results
Cohorts selected and number of subjects studied
A total of 224 cohorts (90 from Europe, 42 from America, 17 from Africa, 66 from Asia, 8 from Australia, and 1 from New Zealand; Supplementary material) were selected. The database comprised 65 of the 193 countries in the world: 25 from Europe, 23 from America, 15 from Asia, 1 from Australia, and 1 from New Zealand.
These cohorts included a total of 193,444 subjects, distributed as follows: 75,603 in Europe; 85,118 in America; 5,031 in Africa; 21,353 in Asia; 5,536 in Australia; and 1,303 in New Zealand.
Pi*Z frequency and Pi*ZZ prevalence
Tables 1–5 show the estimated frequency of Pi*Z (expressed per 1,000, with 95% CI), the prevalence of Pi*ZZ (expressed as 1:x), and the calculated number of Pi*ZZ genotypes (with 95% CI) in the countries included in the study.
  | Table 1 Estimated Pi*Z allele frequency (per 1,000), Pi*ZZ gene prevalence (1:x), and number of Pi*ZZ genotypes among 25 European countries |
  | Table 3 Estimated Pi*Z allele frequency (per 1,000), Pi*ZZ gene prevalence (1:x), and number of Pi*ZZ genotypes in 10 African countries |
  | Table 4 Estimated Pi*Z allele frequency (per 1,000), Pi*ZZ gene prevalence (1:x), and number of Pi*ZZ genotypes in 15 Asian countries |
  | Table 5 Estimated Pi*Z allele frequency (per 1,000), Pi*ZZ gene prevalence (1:x), and number of Pi*ZZ genotypes in Australia and New Zealand |
The Atlantic and sea coast regions of Europe showed the highest frequency and prevalence of Pi*Z and Pi*ZZ (Figure 1), which gradually fell in eastern Europe and practically disappeared in Asia, except in Pakistan and Thailand, which presented moderate values (Figure 2). Some regions of eastern Africa had moderate Pi*Z values, being very low or absent in the remainder of Africa (Figure 3).
  | Figure 1 Distribution of Pi*Z frequencies in Europe (×1,000). |
  | Figure 2 Distribution of Pi*Z frequencies in the world (×1,000). |
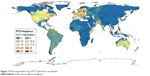  | Figure 3 IDW-interpolation map of Pi*Z distribution worldwide. |
In North America, the frequency of Pi*Z is high in the east of Canada and gradually decreases to the west and the north of the country. The same was found in the US with high values in the eastern states, which progressively decreased in the western and south-eastern areas of the country. Pi*Z frequencies were found to be low in Mexico, Central America, and in the Caribbean and South America countries.
South-eastern Australia showed moderate Pi*Z frequencies, being very high in New Zealand.
IDW-interpolation maps of Pi*Z frequencies
Figures 1–3 show the IDW-interpolation maps with the Pi*Z values found in Europe (Figure 1) and worldwide (Figures 2 and 3). Figure 3 shows the distribution of Pi*Z frequencies in the world. The largest number of studies was found in Europe, being scarce and scattered throughout the remainder of the world, making the results from areas with limited data less reliable.
Estimated numbers of Pi*ZZ genotypes
With the data provided by the cohorts selected from the 65 countries studied, a total of 253,404 ZZ genotypes were estimated and distributed as follows: Europe (119,594; Table 1); America (91,490; Table 2); Africa (3,824; Table 3); Asia (32,154; Table 4); and Australia (4,126) and New Zealand (2,216) (Table 5).
Of the almost 120,000 subjects identified as carrying Pi*ZZ genotype in Europe, most may be found in Germany (20,611), France (17,191), the Iberian Peninsula (14,522 in Spain and 4,944 in Portugal), the UK (13,044), Italy (10,652), Poland (6,791), the Netherlands (5,353), Denmark (4,090), Latvia (4,005), Belgium (3,193), and lower numbers in other European countries (Table 1).
In North, Central, and South America and the Caribbean islands ~90,000 individuals carry Pi*ZZ genotypes (Table 2), most being found in the US (~70,000). In addition, it has been estimated that there may be around 7,000 ZZ genotypes in Canada; 4,000 in Mexico; 6,000 in Brazil; 2,000 in Colombia, Venezuela, and Argentina; 1,000 in Peru; 700 in Chile; 600 in Cuba; and lower numbers in other countries from the American continents.
Table 3 shows that among the >1.2 billion inhabitant of Africa, there may be only around 4,000 Pi*ZZ, being mainly distributed in Nigeria and Somalia.
It has been estimated that among the >4 billion people from Asia, ~30,000 have the Pi*ZZ, most of whom may be found in Pakistan, Thailand, and South Korea (Table 4).
In Australia and New Zealand, Pi*ZZ has been estimated to be present in approximately 6,000 inhabitants, most being descendants of Anglo Saxon settlers (Table 5).
Discussion
A recent study analyzing the database of studies on the genetic epidemiology of AATD estimated that a total of 181,894 Pi*ZZ genotypes could be found in 97 countries in which data were available.21 In parallel, the first maps of IDW interpolation in AATD were created to graphically express the frequency of the distribution of the S and Z alleles in countries with real data and make estimates for countries in which no data were available.22 However, subsequent well-designed studies such as that by an Irish group23 which analyzed 1,100 individuals randomly selected from the general Irish population and a more recent study in 2,482 Finnish individuals24 reported much higher frequencies of Pi*Z, which were comparable to the highest frequencies described in neighboring countries.
However, this study followed strict scientific rigor in the selection of studies to evaluate with the application of the established selection and exclusion criteria. Indeed, numerous studies carried out in small samples of minority ethnic groups (eg, Australian aboriginal; Maoris of New Zealand; Asian, Amerindian, Alaska natives; Hawaiian and other Pacific Islander natives from the US, etc.) were excluded from the analysis because they are not representative of the inhabitants of a specific country, despite their unquestionable anthropological interest. In addition, several pioneering studies using starch gel electrophoresis for lab diagnosis were ruled out, because this technique has some difficulty for detecting the Z bands of the MZ phenotypes, thereby underestimating the true frequency of the Z allele.13 Screening studies in which phenotypes were determined in samples with AAT concentrations below a low cut-off point were also discarded, since normal phenotypes are omitted with this methodology while deficiency types are overrepresented.10–12 For example, the Sveger et al10 study in 200,000 newborns used an AAT cut-off of <40%, and although this strategy was useful to detect all the Pi*Z phenotypes, it is not able to detect all the MM and MS, most MZ and SS, and a considerable number of SZ. Finally, several studies conducted with small sample cohorts and very broad limits of 95% CI (such as those performed in the Greek islands, the Balkans, Romania, etc.) were discarded due to the inaccuracy of their results.
Taking all of the above into account, the number of cohorts included was reduced to 224 and the number of countries studied to 65. Nonetheless, the number of estimated ZZ patients in the world (>250,000) substantially increased and the quality of the IDW-interpolation maps significantly improved. This new analysis confirms that the vast majority of Pi*ZZ individuals are found in Europe (~120,000), and in Caucasians of European descent living in the US (~70,000). We also found that there may be a smaller, albeit significant, number of homozygous Z in Canada, Australia, New Zealand, and in some South American countries with populations of European origin.
Despite careful selection of the studies finally included, our estimates may be biased because of the composition of the samples studied, mainly due to newborns as well as several of the inclusion of subjects labeled under the generic title of “healthy unrelated people”, who may include blood donors, school or college students, soldiers, hospital staff, persons undergoing pre-employment or routine medical examinations, active workers, indigenous subjects, and so on. Indeed, only a few samples from a minority of studies were representative of the general population such as a study carried out in a randomly sampled Spanish population of 1,116 subjects enrolled in the municipal population register,27 or the aforementioned studies conducted in Ireland and Finland 23,24. Nevertheless, with our approach, some of the classically reported results notably improved, providing values of Pi*Z frequencies which are significantly higher than previously estimated for Caucasian populations living in the US, Australia, and in northern, western, and central European countries.
It was of note that high Pi*Z frequencies were obtained for several regions in which AATD has not previously been studied (eg, the Canary Islands), and these regions might be considered as targets for further targeted high-risk case finding screening studies, to identify and provide adequate clinical management to individuals at risk for AATD-associated diseases.
Although some authors have suggested multiple origins of the Pi*Z mutation, it seems very likely that the mutation occurred in Sweden and was spread into the European continent through the maritime and inland waterways via the Baltic countries. The Pi*Z mutation has a relatively recent origin in Caucasians living in northern Europe, having occurred approximately 2000 years or 66 generations ago.28 Along the following centuries, emigrants from Europe distributed the gene throughout non-European countries to which they emigrated. Interestingly, the IDW-interpolation maps of Pi*Z distribution developed in this study agree with this hypothesis.
Currently, massive screenings in newborns and healthy individuals from the general population have become obsolete and are only justified in regions with a high prevalence of AATD and significant under-diagnosis. In addition, before ordering the genetic test, the physician should weigh the pros (eg, early diagnosis, preventive measures, genetic counseling, specific replacement therapy) and cons (eg, psychological trauma and family, social, and negative occupational impact, including possible stigmatization) that the detection of this inherited condition could induce in a healthy person.29
Conversely, a targeted high-risk case finding approach is highly recommended.30 The joint American Thoracic Society and European Respiratory Society statement and most scientific societies worldwide recommend targeted testing for AATD in individuals included in groups at risk.2,3,31 This initiative is a relatively feasible and cost-effective tool to increase the detection of AATD in individuals with obstructive lung diseases, chronic liver disease at any age, granulomatosis with polyangiitis, neutrophilic panniculitis, and family members of patients with confirmed AATD.
In conclusion, the updating of the data regarding the prevalence of the Pi*ZZ genotype worldwide provided in this study may be useful for future research and to develop diagnostic approaches in targeted risk groups and optimize the therapeutic resources currently available (this information is available from the authors upon request).
Authors’ contribution
This study was designed by IB, PB, and ID. IB provided the database and wrote the draft document. PB, SP-H, FC, CE, and MM collaborated in the redaction and revision of the manuscript. Professor ID designed the figures summarizing the database. All authors contributed toward data analysis, drafting and revising the paper and agree to be accountable for all aspects of the work.
Disclosure
The authors report no conflicts of interest in this work.
References
De Serres F, Blanco I. Role of alpha-1 antitrypsin in human health and disease. J Intern Med. 2014;276(4):311–335. | ||
American Thoracic Society/European Respiratory Society Statement. Standards for the diagnosis and management of individuals with alpha1-antitrypsin deficiency. Am J Respir Crit Care Med. 2003;168:818–900. | ||
Casas F, Blanco I, Martínez MT, et al. Indications for active case searches and intravenous alpha-1 antitrypsin treatment for patients with alpha-1 antitrypsin deficiency chronic pulmonary obstructive disease: an update. Arch Bronconeumol. 2015;51(4):185–192. | ||
Stone H, Pye A, Stockley RA. Disease associations in alpha-1-antitrypsin deficiency. Respir Med. 2014;108(2):338–343. | ||
Lee JH, Brantly M. Molecular mechanisms of alpha1-antitrypsin null alleles. Respir Med. 2000;94(Suppl C):S7–S11. | ||
Silva D, Oliveira MJ, Guimarães M, Lima R, Gomes S, Seixas S. Alpha-1-antitrypsin (SERPINA1) mutation spectrum: three novel variants and haplotype characterization of rare deficiency alleles identified in Portugal. Respir Med. 2016;116:8–18. | ||
Ferrarotti I, Carroll TP, Ottaviani S, et al. Identification and characterisation of eight novel SERPINA1 Null mutations. Orphanet J Rare Dis. 2014;9:172. | ||
Laurell CB, Eriksson S. The electrophoretic alpha1-globulin pattern of serum in alpha1-antitrypsin deficiency. Scand J Clin Lab Invest. 1963;15(2):132–140. | ||
Fagerhol MK, Tenfjord OW. Serum Pi types in some European, American, Asian and African populations. Acta Pathol Microbiol Scand. 1968;72(4):601–608. | ||
Sveger T. Liver disease in alpha1-antitrypsin deficiency detected by screening of 200,000 infants. N Engl J Med. 1976;294(24):1316–1321. | ||
O’Brien ML, Buist NR, Murphey WH. Neonatal screening for alpha1-antitrypsin deficiency. J Pediatr. 1978;92(6):1006–1010. | ||
Silverman EK, Miletich JP, Pierce JA, et al. Alpha-1-antitrypsin deficiency. High prevalence in the St Louis area determined by direct population screening. Am Rev Respir Dis. 1989;140(4):961–966. | ||
Hutchison DC. Alpha 1-antitrypsin deficiency in Europe: geographical distribution of Pi types S and Z. Respir Med. 1998;92(3):367–377. | ||
Blanco I, E Fernández, EF Bustillo. Alpha-1-antitrypsin PI phenotypes S and Z in Europe: an analysis of the published surveys. Clin Genet. 2001;60(1):31–41. | ||
Blanco I, Bustillo EF, Rodríguez MC. Distribution of a1-antitrypsin PI S and PI Z frequencies in countries outside Europe: a meta-analysis. Clin Genet. 2001;60(6):431–444. | ||
De Serres FJ. Worldwide racial and ethnic distribution of alpha1-antitrypsin deficiency: summary of an analysis of published genetic epidemiologic surveys. Chest. 2002;122(5):1818–1829. | ||
Luisetti M, Seersholm N. Alpha1-antitrypsin deficiency. 1: epidemiology of alpha1-antitrypsin deficiency. Thorax. 2004;59(2):164–169. | ||
De Serres FJ, Blanco I, Fernández-Bustillo E. Health implications of alpha1-antitrypsin deficiency in sub-Sahara African countries and their emigrants in Europe and the New World. Genet Med. 2005;7(3):175–184. | ||
Blanco I, de Serres FJ, Fernandez-Bustillo E, Lara B, Miravitlles M. Estimated numbers and prevalence of PI*S and PI*Z alleles of alpha1-antitrypsin deficiency in European countries. Eur Respir J. 2006;27(1):77–84. | ||
De Serres FJ, Blanco I, Fernández-Bustillo E. Estimated numbers and prevalence of PI*S and PI*Z deficiency alleles of alpha1-antitrypsin deficiency in Asia. Eur Respir J. 2006;28(6):1091–1099. | ||
De Serres FJ, Blanco I. Prevalence of α1-antitrypsin deficiency alleles PI*S and PI*Z worldwide and effective screening for each of the five phenotypic classes PI*MS, PI*MZ, PI*SS, PI*SZ, and PI*ZZ: a comprehensive review. Ther Adv Respir Dis. 2012;6(5):277–295. | ||
Blanco I, de Serres FJ, Cárcaba V, Lara B, Fernández-Bustillo E. Alpha-1 antitrypsin deficiency PI*Z and PI*S gene frequency distribution using on maps of the world by an inverse distance weighting (IDW) multivariate interpolation method. Hepat Mon. 2012;12(10 HCC):e7434. | ||
Carroll TP, O’Connor CA, Floyd O, et al. The prevalence of alpha-1 antitrypsin deficiency in Ireland. Respir Res. 2011;12:91. | ||
Häggblom J, Kettunen K, Karjalainen J, Heliövaara M, Jousilahti P, Saarelainen S. Prevalence of PI*Z and PI*S alleles of alpha-1-antitrypsin deficiency in Finland. Eur Clin Respir J. 2015;2:28829. | ||
Chorostowska-Wynimko J, Struniawski R, Popławska B, Borszewska-Kornacka M. The incidence of alpha-1-antitrypsin (A1AT) deficiency alleles in population of Central Poland – preliminary results from newborn screening. Pneumonol Alergol Pol. 2012;80(5):450–453. | ||
Lu GY, Wong DW. An adaptive inverse-distance weighting spatial interpolation technique. Compute Geosci. 2008;34(9):1044–1055. | ||
Blanco I, Fernández E, Rodríguez MC, Fernández A. Allelic frequency of the gene alpha-1-antitrypsin in the general population in a county in Asturias. Med Clin (Barc). 1999;133(10):366–370. Spanish. | ||
Lace B, Sveger T, Krams A, Cernevska G, Krumina A. Age of SERPINA1 gene PI Z mutation: Swedish and Latvian population analysis. Ann Hum Genet. 2008;72(Pt 3):300–304. | ||
Stoller JK, Sandhaus RA, Turino G, Dickson R, Rodgers K, Strange C. Delay in diagnosis of alpha1-antitrypsin deficiency: a continuing problem. Chest. 2005;128(4):1989–1994. | ||
De la Roza C, Rodríguez-Frías F, Lara B, Vidal R, Jardí R, Miravitlles M. Results of a case-detection programme for alpha-1-antitrypsin deficiency in COPD patients. Eur Respir J. 2005;26(4):616–622. | ||
Alpha 1-antitrypsin deficiency: memorandum from a WHO meeting. Bull World Health Organ. 1997;75(5):397–415. |
 © 2017 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2017 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.