Back to Journals » OncoTargets and Therapy » Volume 13
Adequate Lymph Node Assessments and Investigation of Gut Microorganisms and Microbial Metabolites in Colorectal Cancer
Authors Han S , Wu W , Da M , Xu J, Zhuang J, Zhang L, Zhang X, Yang X
Received 11 December 2019
Accepted for publication 18 February 2020
Published 4 March 2020 Volume 2020:13 Pages 1893—1906
DOI https://doi.org/10.2147/OTT.S242017
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Prof. Dr. Nicola Silvestris
Shuwen Han,1 Wei Wu,2 Miao Da,3 Jiamin Xu,4 Jing Zhuang,4 Longqi Zhang,5 Xiaoxiang Zhang,6 Xi Yang7
1Department of Oncology, Huzhou Central Hospital, Affiliated Central Hospital Huzhou University, Huzhou, Zhejiang, People’s Republic of China; 2Department of Gastroenterology, Huzhou Central Hospital, Huzhou, Zhejiang, People’s Republic of China; 3Department of Nursing, Huzhou Third Municipal Hospital, Huzhou, Zhejiang, People’s Republic of China; 4Graduate School of Nursing, Huzhou University, Huzhou, Zhejiang, People’s Republic of China; 5Department of Infectious Disease, Huzhou Central Hospital, Huzhou, Zhejiang, People’s Republic of China; 6Department of Clinical Examination, Huzhou Central Hospital, Huzhou, Zhejiang, People’s Republic of China; 7Department of Oncology, Huzhou Central Hospital, Huzhou, Zhejiang, People’s Republic of China
Correspondence: Xi Yang
Department of Oncology, Huzhou Central Hospital, Huzhou, Zhejiang, People’s Republic of China
Tel +86 57 2202 3301
Email [email protected]
Purpose: To analyze the lymph node metastasis status and prognosis in CRCs and to investigate the gut microorganisms and microbial metabolites at different lymph node stages.
Methods: The Surveillance, Epidemiology, and End Results (SEER) database and STAT software were used to analyze the clinical features and lymph node metastasis. Bacterial 16S V3-V4 and fungal ITS V3-V4 ribosomal RNA genes were sequenced in 53 stool samples and gas chromatography/mass spectrometry (GS/MS) was performed to detect the microbial metabolites in 48 stool samples from CRC patients.
Results: A higher number of lymph node metastases predicted a poor prognosis. Inadequate evaluation of lymph nodes affects the accuracy of prognostic assessments. We constructed a nomogram model for the assessment of prognostic factors. There were multiple characteristic bacteria identified, including Akkermansia, Megamonas, Dialister, etc., and fungi, including Penicillium, Filobasidium, Debaryomyces, etc. A total of 27 characteristic microbial metabolites in different lymph node metastasis status were also identified.
Conclusion: Gut microorganisms and microbial metabolites may provide reference and guidance for the adequate lymph node assessments (ALNA) in CRC.
Keywords: microbiome, metabolites, lymph node, metastasis, colorectal cancer
Introduction
Colorectal cancer (CRC) is one of the most common malignancies, and its morbidity and mortality rates are increasing rapidly.1,2 Although some decisions, such as multidisciplinary comprehensive treatment (MDT), molecular targeting treatment, and immunological therapy, have been applied in the treatment of CRCs, the prognosis of CRCs, especially advanced colorectal cancer, is of concern.3–6 The number of lymph nodes obtained from resected specimens is an independent risk factor for mortality in CRC patients.7 A high positive/total lymph node ratio is positively correlated with poor prognosis of CRC.8 The American Joint Committee on Cancer (AJCC)/Union for International Cancer Control (UICC) TNM staging system recommends collecting at least 12 lymph nodes for lymph node staging of CRCs.9 Adequate lymph node assessments (ALNA) are critical for accurate cancer staging, prognostic assessment, and optimal treatment selection10 For example, postoperative adjuvant chemotherapy is highly recommended for CRC patients with lymph node metastasis, while for patients without lymph node metastasis, whether to accept postoperative adjuvant chemotherapy is determined according to the comprehensive assessment of tumor size and high-risk factors.11–13 The Surveillance, Epidemiology, and End Results (SEER) Program provides a large volume of cancer-related data and statistics to reduce the cancer burden among the US population.14,15 Through retrieving and analyzing the lymph node metastasis status and outcome of CRCs in the SEER database, we confirmed the effect of lymph node metastasis status on the prognosis of CRC and tried to explain the influence of ALNA on the prognosis of CRC.
The occurrence of colorectal cancer is a multi-stage and multi-factor process.16 The colorectum, as a storage site for feces, contains the largest number of microorganisms in the human body, and these microbes affect human health.17,18 Large numbers of studies have shown that CRC patients have characteristic microbes, including Fusobacteria, Streptococcus, and Clostridium in their intestines.19–22 Gut microorganisms are involved in the development and progression of CRCs through many pathways such as through the induction of inflammation and biosynthesis of genotoxins.23,24 Recent studies have suggested that the occurrence of CRCs is not only the function of a single microorganism, but also the result of the disruption of dynamic equilibrium of the gut microecology.25,26 Microbial metabolites, such as short-chain fatty acids and secondary bile acids, act as an important part of gut microecosystem and are also involved in the occurrence of colorectal cancer.27–29 Thus, produced novel hypothesis that microorganisms and their metabolites could be used as auxiliary indexes of ALNA or risk factors of lymph node metastasis status in CRCs.
To assess the feasibility of this hypothesis, we explored the relationship between the lymph node metastasis status of CRCs and gut microorganisms as well as microbial metabolites in three aspects in the present study. First, the effect of different lymph node metastases and assessments on the prognosis of CRC was demonstrated by retrieval and analysis of relevant data in the SEER database. Secondly, fecal samples from CRC patients with different lymph node metastasis statuses were collected, and the community structure and diversity differences of gut microorganisms, including bacteria and fungi, were analyzed. Thirdly, microbial metabolites in fecal samples in CRCs with different lymph node metastasis status were quantitatively detected and analyzed to further clarify the role of the gut microecological environment in the lymph node metastasis status of CRCs. This may provide a fresh approach to the assistant reference and guidance for ALNA in CRCs.
Materials and Methods
Study Design
The study design is shown in Supplement material Figure S1. The SEER database was used to analyze the effect of lymph node metastasis status in different regions on the prognosis of CRCs. The bacterial 16S V3-V4 and fungal ITS V3–V4 ribosomal RNA genes were sequenced and GS/MS analysis was performed to detect microbial metabolites in CRC patients with different lymph node stages. Bioinformatics analysis was carried out to examine the microbial diversity, community structure, and microbial metabolite differences associated with lymph node stages.
SEER Database Retrieval
SEER*Stat software (seer.cancer.gov/seerstat) was used to access the SEER data after signing a research data agreement. All patients diagnosed with colon and rectal adenocarcinoma from 2006 to 2015 were included. The SEER database retrieval strategy is shown in Supplement material Figure S2. A total of 353,420 CRC patients were included.
Stool Sample Collection
The clinical protocols involving the patients and the informed consent forms were approved by the Chinese Clinical Trial Registry (http://www.chictr.org.cn, No. ChiCTR1800018908) and the Ethics Committee of Huzhou Central Hospital (No. 201601023).And all experiments were performed in accordance with relevant guidelines and regulations. The CRC patients confirmed by pathologic diagnosis at Huzhou Central Hospital from January 2017 to August 2018 were recruited in the present study. The inclusion criteria were as follows: ① CRC was confirmed by pathological diagnosis, and the patients volunteered to participate in the study, ② All stool samples were able to be collected before treatment (chemotherapy, radiotherapy, or surgery), ③ No CRC patients had evidence of organ metastasis before specimen collection. The exclusion criteria were as follows: ① Patients with diarrhea pre-collection, ② Patients with other complicating gut diseases, such as ulcerative colitis, Crohn’s disease, multiple primary tumors, or known primary organ failure, ③ Patients with a history of the use of oral microbial agents within 1 month and with the use of a purgative or lubricant within one week precollection. Lymph node staging was assessed 1 month after CRC surgery and determined by collaboration with pathologists, surgeons, and oncologists according to the AJCC/UICC TNM staging system. Basic information and serological indicators were obtained from the medical record management system of Huzhou Central Hospital. Approximately 30–50 g stool sample was collected and stored in a −80°C refrigerator within half an hour. The stool samples were analyzed within 3 months.
Gut Microorganism Detection
After screening according to the inclusion and exclusion criteria and eliminating the ineligible specimens, 53 stool samples were ultimately used to detect gut microorganisms. Bacterial 16S V3–V4 rRNA and fungal ITS V3–V4 rRNA genes were sequenced by using the Illumina MiSeq platform after DNA extraction from stool samples and PCR amplification. The steps involved in microorganism detection include library construction and sequencing, followed by bioinformatics analysis. The specific research methods were as previously published.22 Supplementary material 1 includes the specific research methods used for gut microorganism detection.
Microbial Metabolite Detection
Sample Preparation
After screening according to the inclusion and exclusion criteria and eliminating the ineligible specimens, 48 stool samples were ultimately used to detect gut microbial metabolites. Approximately 10 mg of lyophilized fecal samples was homogenized with 300 µL NaOH solution by using a homogenizer (BB24, Next Advance, Inc., Averill Park, NY, USA), then centrifuged at 4°C and 16,000 RPM for 20 min (Microfuge 20R,Beckman Coulter, Inc., Indianapolis, IN, USA). Next, 200 µL of the supernatant at a time was transferred to an auto-sampler vial. The residue was homogenized with 200 µL of cold methanol, then centrifuged and 167 µL of supernatant transferred into an auto-sampler vial. The mixtures in the auto-sampler vials were sealed and used for automated sample derivatization with a robotic multipurpose sample MPS2 with dual heads (Gerstel, Muehlheim, Germany). In summary, 20 µL of methyl chloroformate (MCF) was added to the mixture, then shaken for 30 s. Another 20 µL of MCF was added for the second derivatization. Next, 400 mL of CHCl3 followed by 400 mL of Na2CO3 solution (50 mmol) was added and centrifuged at 4°C and 4000 RPM for 20 min. The robotic preparation station transferred the CHCl3 layer at the bottom to an auto-sampler vial preloaded with approximately 25 mg of anhydrous Na2SO4. Samples were then shaken at 4°C and 1500 RPM for 20 min and further transferred to a capped empty auto-sampler vial for injection.
GC/TOFMS Analysis
An Agilent 6890N gas chromatography coupled to time-of-flight mass spectrometry (GC-TOFMS) system (Pegasus HT, Leco Corp., St. Joseph, MO, USA) operated in electron ionization (EI) mode was used to quantitatively analyze the microbial metabolites. One µL of each derivatized sample was injected into a Rxi-5MS capillary column ((Crossbond ® 5% diphenyl/95% dimethyl polysiloxane) 30 m (length) x 250 µm I.D., 0.25 µm film thickness). The carrier gas was helium (99.9999%), the flow rate was 1.0 mL/min and the optimized temperature gradient was the following: 45°C (1 min), 45–260°C (20°C/min), 260–320°C (40°C/min) and 320°C (2 min). Electron impact ionization (−70 eV) at a mass range of 38−550 Da was used. The source temperature was 220°C and the acquisition rate was 20 spectra per second.
Quality Control and Data Analysis
The sample control procedure and data control procedure refer to criterion (ISO9001, QAIC/CN/170149). Raw data from GC/TOFMS analysis were processed using proprietary software XploreMET (v2.0, Metabo-Profile, Shanghai, China) for automatic baseline correction, smoothing, peak picking, peak signal, library searching, and area calculation. Identification of differential metabolites was as follows: GC/MS workstation software automatically compared the fragment mass to charge ratio and abundance of characteristic ion fragmentation patterns with the available reference standards in our lab, the NISI II standard mass spectral databases, and the Fiehn databases linked to the LECO ChromaTOF software. A similarity of more than 70% was used as the reference standard.
Statistical Analysis
The Kaplan–Meier method was used to describe the analysis of survival curves. The survival curves were used to describe the effect of lymph node metastasis status on the outcome of CRCs. The variables were described with mean ± standard deviation or median value and analyzed using the Student’s t-test or SNK test, as appropriate. All statistical analyses were conducted using SPSS Statistics version 16.0 and Microsoft Excel. Bioinformatic analyses were conducted using the software packages in R studio (http://cran.r-project.org/). A two-tailed p< 0.05 was considered statistically significant.
Results
Retrieval and Analysis of Lymph Node Metastasis Status and Outcome in the SEER Database
The classification criteria of lymph node metastasis status and pathological stage refer to the AJCC/UICC TNM staging system. N0, N1, N2 and Nx represent no regional lymph node metastasis status, 1–3 regional lymph node metastases or tumor deposit, more than 4 regional lymph node metastases and regional lymph nodes could not be evaluated, respectively. After retrieving and screening the SEER database, 353420 CRC cases were included in the study. Table 1 includes lymph node metastasis status including N0, N1, N2, and Nx, and univariate analyses for multiple factors such as age, gender, race, histologic subtype, and histologic grade. In addition, adequate lymph node examination of the included cases was described.
 |
Table 1 Lymph Node Metastasis Status and Univariate Analysis in Patients with CRC Between 2006 and 2015 |
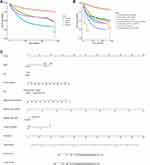
The survival curve shown in panel A of Figure 1 revealed that the 10-year survival rate varied significantly among the different lymph node metastasis statuses in CRC patients. A high number of lymph node metastases predicts a poor prognosis. The curve of the Nx group overlaps that of N1 group, indicating a failure to accurately assess lymph node metastasis status is not conducive to accurate prognosis evaluation.
To further illustrate the important value of ANLA, more possibilities for lymph node assessment were considered and mapped in the survival curve in panel B of Figure 1. In the evaluation process of lymph node metastasis status, multiple conditions including “All nodes examined are negative,” “Exact number of nodes positive,” “90 or more nodes are positive,” “Positive aspiration of lymph node(s) was performed,” “Positive nodes are documented, but number is unspecified,” “No nodes were examined,” and “Unknown whether nodes are positive: not applicable: not stated in patient record” were included in the 10-year survival analysis. The overall trend is that the overall survival time of CRC patients with lymph node metastasis was shorter than that of patients without lymph node metastasis. However, different methods of lymph node evaluation or unclear number of lymph nodes have an impact on the accuracy of prognosis.
Moreover, as shown in panel C of Figure 1, based on prognostic factors including race, gender, age, site, regional nodes examined, regional nodes positive, regional node type, and lymph node metastasis status, a nomogram model was constructed and used for predicting 5- and 10-year overall survival (OS) and cancer-specific survival (CSS).
Association of Gut Microbiota with Lymph Node Metastasis Status in CRCs
After screening and identification, overall, 53 CRC patients in the Huzhou Central Hospital from January 2016 to August 2018 were enrolled in the microbiology study. The clinical characteristics are shown in Table 2. There was no significant difference between the patients with different lymph node metastasis status in regards to site, age, sex, smoking, drinking history, and serological indicators such as glutamic oxaloacetic transaminase, alanine transaminase, albumin, creatinine, fasting blood glucose (FBG), triglycerides, total cholesterol, carbohydrate antigen 153, carbohydrate antigen 742, carbohydrate antigen 199, and carcinoembryonic antigen.
 |
Table 2 Characteristics of the Study Participants with Regard to the Association of Gut Microbe with Lymph Node Metastasis |
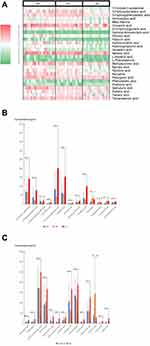
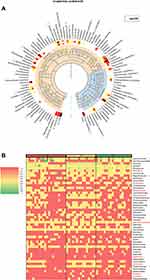
Figure 2 shows the association of gut bacteria with lymph node metastasis status in CRCs. The taxonomic tree heatmap in panel A of Figure 2 displays the composition and proportion of gut bacteria at different taxonomic levels. The innermost layer shows the taxonomic tree. The circle from the inside to the outside represents different taxon levels from phylum to genus. Orange circles represent the statistical difference among the three groups (Kruskal algorithm). The intermediate layer represents the average abundance in the form of a heatmap. The deeper the color, the higher was the species abundance. The outermost layer indicates an annotated genus. The heatmap in panel B of Figure 2 showed the relative abundance of gut bacteria from each sample in the top 50 at the genus level. The bacterial annotations marked red indicated statistical difference (SNK test). A comprehensive analysis of panel A and panel B of Figure 4 showed that there were statistically significant differences in the relative abundance of gut bacteria including Akkermansia, Megamonas, Dialister, Enterobacteriaceae, Erysipelotrichaceae incertae sedis, Porphyromonas, Prevotellaceae, Butyricicoccus, Desulfovibrio, Lactobacillus, and Porphyromonadaceae among the different lymph node metastasis status groups in CRCs.
Figure 3 shows the association of gut fungi with lymph node metastasis status in CRCs. The drawing principles and statistical analysis are the same as in Figure 4. The taxonomic tree heatmap in panel A of Figure 3 displays the composition and proportion of gut fungi at different taxonomic levels. The heatmap in panel B of Figure 3 shows the relative abundance of gut fungi from each sample in the top 50 at the genus level. A comprehensive analysis of panel A and panel B of Figure 5 showed that there were statistically significant differences in the relative abundance of gut fungi including Penicillium, Filobasidium, Debaryomyces, Dipodascaceae, Saccharomycetales, and Ochroconis among the different lymph node metastasis status groups in CRCs.
Association of Microbial Metabolites with Lymph Node Metastasis Status in CRCs
After screening and identification, 48 CRC patients in Huzhou Central Hospital from January 2016 to August 2018 were enrolled in the microbial metabolomics study. The clinical characteristics are shown in Table 3. There was no significant difference between the patients with different lymph node metastasis status in regards to site, age, sex, smoking, drinking history or serological indicators such as glutamic oxaloacetic transaminase, alanine transaminase, albumin, creatinine, FBG, triglycerides, total cholesterol, carbohydrate antigen 153, carbohydrate antigen 742, carbohydrate antigen 199, or carcinoembryonic antigen.
 |
Table 3 Characteristics of the Study Participants with Regard to the Association of Microbial Metabolites with Lymph Node Metastasis |
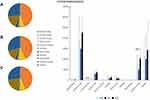
A total of 124 microbial related metabolites in 10 classes were included in the present study. Figure 4 shows the association of microbial metabolites with lymph node metastasis in CRCs at the class level. The bar chart shows the concentration of microbial metabolites in the samples of each group. There were statistical differences between the microbial metabolites including cinnamic acids and organic acids (SNK test). Panels A, B and C of Figure 4 represent the N0, N1 and N2 groups, respectively. The pie chart shows the composition and proportion of microbial metabolites in the samples of each group. The results showed that there were no significant differences in the concentration of microbial metabolites at the class level, in which amino acids accounted for the largest proportion.
The concentrations of all 124 metabolites of stool samples from the CRCs with different lymph node metastasis stages are shown in Supplementary material Figure S3. Figure 5 shows the microbial metabolites with statistical differences (SNK test). The heatmap in panel A of Figure 5 shows the relative concentrations of 27 microbial metabolites with statistically significant differences. The bar chart (panel B and C) shows the difference information of pairwise comparisons between each group and the absolute concentration measurement data with regard to the 27 microbial metabolites among the three groups.
Discussion
In the present study, we demonstrated the important value of ALNA for the prognosis of CRCs and further explored the feasibility of a hypothesis that microorganisms and their metabolites as auxiliary reference indicators of ALNA. Through the analysis of data from the SEER database with regard to the lymph node metastasis in CRCs, we found that a higher number of lymph node metastases predicted a poor prognosis. Inadequate evaluation of lymph nodes affects the accuracy of prognostic assessment. Numerous studies suggest there is an association between the occurrence and prognosis of CRC with gut microbiota. Gut bacteria, fungi, and microbial metabolites in CRCs with different lymph node metastasis status were detected and compared. The results showed that there were multiple characteristic bacteria including Akkermansia, Megamonas, Dialister, Enterobacteriaceae, Erysipelotrichaceae incertae sedis, Porphyromonas, Prevotellaceae, Butyricicoccus, Desulfovibrio, Lactobacillus, and Porphyromonadaceae and fungi including Penicillium, Filobasidium, Debaryomyces, Dipodascaceae, Saccharomycetales, and Ochroconis, as well as 27 characteristic microbial metabolites in different lymph node metastasis status groups. This study provides a preliminary exploration for verification of the hypothesis that the microorganism and its metabolites can be used as auxiliary indexes of ALNA or risk factors of lymph node metastasis status in CRCs.
By retrospectively analyzing the prognosis of CRCs with different lymph node metastases in the SEER database, we found that the accuracy of lymph node assessments and the difference in the number of lymph node metastasis status both affect the prognosis of CRCs. ALNA requires the cooperation of surgeons and pathologists. Doctors’ subjectivity is difficult to overcome, as is the existence of accidental errors. In addition, some other factors are also great obstacles to ALNA. For instance, the detection of lymph nodes with skipping metastases tests preoperative imaging technology, and the acquisition of paravascular lymph nodes tests the surgeon’s skills. To summarize, it is necessary to find auxiliary means of ALNA from a new perspective.
It is of great significance to study the association between lymph node metastases and gut microorganisms in CRC. On the one hand, it provides a novel direction for translational medicine study in the stage of tumor and prognostic evaluation from the perspective of microbiology. On the other hand, interventions, such as fecal transplantation, oral administration of microorganisms or microbial metabolites, and dietary choices may provide alternative treatments and auxiliary treatment for patients with lymph node metastases. Screening and dynamic monitoring of characteristic microorganisms and microbial metabolites can support effective interventions. Thirdly, dynamic monitoring of characteristic microorganisms and microbial metabolites may provide a simple and rapid early warning and screening method for tumor recurrence and metastasis.
The intersection of microbial epidemiology and pathology may lead to a new interdisciplinary discipline to establish a basis for individualized cancer prevention and treatment. It may provide new microbiological criteria for subgroups of CRC and etiologic heterogeneity to investigate the association of environmental factors with CRC in strata of characteristic microorganisms and microbial metabolites. It may also provide new insights into the mechanism of tumor recurrence and metastasis in the future.
Microorganisms can adjust and modify their surroundings due to their metabolic products, enzymes, and proteins.27 The bacteria, fungi, their metabolic products, and relevant nutrients required by microorganisms make up the gut microecosystem. Analysis of characteristic microorganisms and their metabolites provides a direction for the next step in clarifying the molecular mechanism of the lymph node metastasis of colorectal cancer.
The complexity and variability of intestinal microorganisms inevitably lead to some deficiencies in this study. The investigation may be affected by a variety of factors such as geographical location, nosocomial infection, or pollution contamination in the fecal collection process. The influence of individual dietary habits on intestinal microorganisms is inevitable. Patients from the same region and ethnic group were included to minimize the influence of diet on intestinal microorganisms. The small sample size limits the applicability of the findings. A multi-center, large study will provide more powerful data to support clarification of the microbial differences in CRC patients with different lymph node metastasis status.
Some commonly used oral probiotics have been used to treat intestinal diseases, such as diarrhea and constipation. However, there is no definite benefit from these probiotics in the treatment of CRC. These microorganisms which were screened out in the present study can be used as potential probiotics for CRC treatment. However, There are still many difficulties in extraction, separation, culture and mass production of these microorganisms. Therefore, there is still a lot of work to be done in the follow-up research.
A nomogram model was constructed for the assessment of prognostic risk based on the analysis of SEER data in the present study. The next research direction will include the data of microbiology and microbial metabolomics after the validation of larger clinical samples. A more complete and scientific ALNA model or prognostic assessment model will be developed by applying machine learning models, such as neural network model or random forest model. This will provide support for the more accurate prognosis assessment and more optimal therapeutic regimen of CRC.
Abbreviations
ALNA, adequate lymph node assessments; CRC, colorectal cancer; SEER, surveillance, epidemiology, and end results; GS, gas chromatography; MS, mass spectrometer; MDT, multidisciplinary comprehensive treatment; AJCC, American Joint Committee on Cancer; NCCN, National Comprehensive Cancer Network; UICC, International Cancer Control; PCR, polymerase chain reaction; rRNA, ribosomal ribonucleic acid; MCF, methyl chloroformate; TOFMS, time-of-flight mass spectrometry; EI, electron ionization; FBG, fasting blood glucose; OTU, operational taxonomic unit.
Ethics Approval and Consent to Participate
The clinical protocols involving the patients and the informed consent form were approved by the Chinese Clinical Trial Registry (http://www.chictr.org.cn, No. ChiCTR1800018908) and Ethics Committee of Huzhou Central Hospital (No. 201601023). All experiments were performed in accordance with relevant guidelines and regulations. All patients provided written informed consent. This study was conducted in accordance with the Declaration of Helsinki.
Acknowledgments
The authors gratefully acknowledge the SEER database (established by the National Cancer Institute, USA), which made the data of BTCC available. We thank the patients and volunteers for their contributions to sample collection.
Author Contributions
All authors contributed to data analysis, drafting and revising the article, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Funding
This work was supported by the Major Science and Technology Projects for Medical and Health Care of Zhejiang Province (No. WKJ-ZJ-2013), and Public Welfare Technology Application Research Program of Huzhou (No. 2019GZB01).
Disclosure
The authors report no conflicts of interest in this work.
References
1. Ferlay J, Colombet M, Soerjomataram I, et al. Cancer incidence and mortality patterns in Europe: estimates for 40 countries and 25 major cancers in 2018. Eur J Cancer. 2018;103:356–387. doi:10.1016/j.ejca.2018.07.005
2. Arnold M, Sierra MS, Laversanne M, et al. Global patterns and trends in colorectal cancer incidence and mortality. Gut. 2017;66(4):683–691. doi:10.1136/gutjnl-2015-310912
3. Lin W-L, Sun J-L, Chang S-C, et al. Effectiveness of the multidisciplinary team model in treating colorectal cancer. Gastroenterol Nurs. 2018;41(6):491–496. doi:10.1097/SGA.0000000000000348
4. Fan J, Shang D, Han B, et al. Adoptive cell transfer: is it a promising immunotherapy for colorectal cancer? Theranostics. 2018;8(20):5784–5800. doi:10.7150/thno.29035
5. Battaglin F, Naseem M, Lenz H-J, et al. Microsatellite instability in colorectal cancer: overview of its clinical significance and novel perspectives. Clin Adv Hematol Oncol. 2018;16(11):735–745.
6. Heinemann V, von Weikersthal LF, Decker T, et al. FOLFIRI plus cetuximab versus FOLFIRI plus bevacizumab as first-line treatment for patients with metastatic colorectal cancer (FIRE-3): a randomised, open-label, Phase 3 trial. Lancet Oncol. 2014;15(10):1065–1075. doi:10.1016/S1470-2045(14)70330-4
7. Emre A, Akbulut S, Sertkaya M, et al. Assessment of risk factors affecting mortality in patients with colorectal cancer. Prz Gastroenterol. 2018;13(2):109–117. doi:10.5114/pg.2018.73348
8. Fortea-Sanchis C, Martinez-Ramos D, Escrig-Sos J. The lymph node status as a prognostic factor in colon cancer: comparative population study of classifications using the logarithm of the ratio between metastatic and nonmetastatic nodes (LODDS) versus the pN-TNM classification and ganglion ratio systems. BMC Cancer. 2018;18(1):1208. doi:10.1186/s12885-018-5048-4
9. Shia J, Wang H, Nash GM, Klimstra DS. Lymph node staging in colorectal cancer: revisiting the benchmark of at least 12 lymph nodes in R0 resection. J Am Coll Surg. 2012;214(3):348–355. doi:10.1016/j.jamcollsurg.2011.11.010
10. Reha J, Mukkamalla SKR, Rathore R, et al. Adequate lymph node evaluation in the elderly is associated with improved survival in patients with stage I-III colon cancer: a validation study using the National Cancer Data Base. Eur J Surg Oncol. 2018;44(1):148–156. doi:10.1016/j.ejso.2017.11.005
11. Ajani JA, D’Amico TA, Almhanna K, et al. Gastric cancer, Version 3.2016, NCCN clinical practice guidelines in oncology. J Natl Compr Canc Netw. 2016;14(10):1286–1312. doi:10.6004/jnccn.2016.0137
12. Tournigand C, André T, Bonnetain F, et al. Adjuvant therapy with fluorouracil and oxaliplatin in stage II and elderly patients (between ages 70 and 75 years) with colon cancer: subgroup analyses of the Multicenter International Study of Oxaliplatin, Fluorouracil, and Leucovorin in the adjuvant treatment of colon cancer trial. J Clin Oncol. 2012;30(27):3353–3360. doi:10.1200/JCO.2012.42.5645
13. Sargent DJ, Marsoni S, Monges G, et al. Defective mismatch repair as a predictive marker for lack of efficacy of fluorouracil-based adjuvant therapy in colon cancer. J Clin Oncol. 2010;28(20):3219–3226. doi:10.1200/JCO.2009.27.1825
14. Wang X-J, Chi P, Zhang -Y-Y, et al. Survival outcome of adjuvant radiotherapy after local excision for T2 early rectal cancer: an analysis based on the surveillance, epidemiology, and end result registry database. Eur J Surg Oncol. 2018;44(12):1865–1872. doi:10.1016/j.ejso.2018.08.024
15. Borstlap WA, Tanis PJ, Koedam TWA, et al. A multi-centred randomised trial of radical surgery versus adjuvant chemoradiotherapy after local excision for early rectal cancer. BMC Cancer. 2016;16:513. doi:10.1186/s12885-016-2557-x
16. Calvert PM, Frucht H. The genetics of colorectal cancer. Ann Intern Med. 2002;137(7):603–612. doi:10.7326/0003-4819-137-7-200210010-00012
17. Feng Q, Chen WD, Wang YD. Gut microbiota: an integral moderator in health and disease. Front Microbiol. 2018;9:151. doi:10.3389/fmicb.2018.00151
18. Schippa S, Conte MP. Dysbiotic events in gut microbiota: impact on human health. Nutrients. 2014;6(12):5786–5805. doi:10.3390/nu6125786
19. Brennan CA, Garrett WS. Fusobacterium nucleatum - symbiont, opportunist and oncobacterium. Nat Rev Microbiol. 2019;17(3):156–166. doi:10.1038/s41579-018-0129-6
20. Kelly D, Yang L, Pei Z, Microbiota G. Fusobacteria, and colorectal cancer. Diseases. 2018;6(4).
21. Zheng Y, Luo Y, Lv Y, et al. Clostridium difficile colonization in preoperative colorectal cancer patients. Oncotarget. 2017;8(7):11877–11886. doi:10.18632/oncotarget.14424
22. Han S, Pan Y, Yang X, et al. Intestinal microorganisms involved in colorectal cancer complicated with dyslipidosis. Cancer Biol Ther. 2019;20(1):81–89. doi:10.1080/15384047.2018.1507255
23. Lucas C, Barnich N, Nguyen HTT. Microbiota, inflammation and colorectal cancer. Int J Mol Sci. 2017;18(6):1310.
24. Dai Z, Zhang J, Wu Q, et al. The role of microbiota in the development of colorectal cancer. Int J Cancer. 2018. 145(8):2032–2041.
25. Hale VL, Jeraldo P, Chen J, et al. Distinct microbes, metabolites, and ecologies define the microbiome in deficient and proficient mismatch repair colorectal cancers. Genome Med. 2018;10(1):78. doi:10.1186/s13073-018-0586-6
26. O’Keefe SJ. Diet, microorganisms and their metabolites, and colon cancer. Nat Rev Gastroenterol Hepatol. 2016;13(12):691–706. doi:10.1038/nrgastro.2016.165
27. Han S, Gao J, Zhou Q, et al. Role of intestinal flora in colorectal cancer from the metabolite perspective: a systematic review. Cancer Manag Res. 2018;10:199–206. doi:10.2147/CMAR
28. Louis P, Hold GL, Flint HJ. The gut microbiota, bacterial metabolites and colorectal cancer. Nat Rev Microbiol. 2014;12(10):661–672. doi:10.1038/nrmicro3344
29. Ma H, Yu Y, Wang M, et al. Correlation between microbes and colorectal cancer: tumor apoptosis is induced by sitosterols through promoting gut microbiota to produce short-chain fatty acids. Apoptosis. 2019;24(1–2):168–183. doi:10.1007/s10495-018-1500-9
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.