Back to Archived Journals » Advances in Genomics and Genetics » Volume 5
A molecular toolbox for genetic manipulation of zebrafish
Received 22 October 2014
Accepted for publication 1 January 2015
Published 27 March 2015 Volume 2015:5 Pages 151—163
DOI https://doi.org/10.2147/AGG.S57585
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 7
Editor who approved publication: Dr John Martignetti
Wiebke A Sassen, Reinhard W Köster
Division of Cellular and Molecular Neurobiology, Zoological Institute, Technical University Braunschweig, Braunschweig, Germany
Abstract: Zebrafish is an emerging model system in biomedical research. The high degree of evolutionary conservation in genetics as well as in development allows deciphering the molecular mechanisms underlying vertebrate development and human diseases. Zebrafish offers important advantages as a genetic model organism such as a completely sequenced genome, fast external embryonic development accompanied by nearly perfect transparency, a large number of offspring year round, and molecular accessibility for genetic manipulation. This review discusses established and newly developed approaches for genetic manipulation of zebrafish and their usefulness to understand gene function in various human disorders like heart defects or Alzheimer's disease.
Keywords: zebrafish, genetic manipulation, disease models, transgenes, forward genetics, reverse genetic
Introduction
More than 30 years ago, the small freshwater fish Danio rerio attracted the attention of scientists who searched for an easy-to-handle but yet complex vertebrate to analyze conserved gene functions.1,2 Today, D. rerio is better known under its common name “zebrafish” for its striped appearance, and it is a well-established model for challenging issues in various research fields of biomedical relevance such as cardiovascular or neurodegenerative diseases.
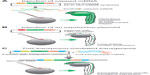
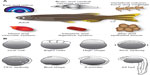
Zebrafish is a bony fish belonging to the infraclass of teleosts. Its anatomical structure as well as the function of most organs resemble the respective features of mammals, making zebrafish an excellent disease model (Figure 1A). For example, studying zebrafish heart development contributed significantly to the understanding of human cardiovascular diseases.3 The zebrafish liver can be used as a model for liver tumorigenesis or alcoholic fatty liver.4,5
Zebrafish embryos are transparent and develop rapidly outside the mother6 (Figure 1B), allowing exceptionally detailed observations of developmental processes down to the subcellular level. Because an adult female produces up to 200 eggs per week and zebrafish husbandry is relatively cost-efficient, zebrafish experiments can be easily performed with high n-numbers or even as high-throughput approaches.7 Zebrafish embryos and larvae are easily accessible for mechanical manipulations such as injection or transplantation. Chemical manipulations can be carried out by simply diluting the compound of choice in the water in which embryos or larvae are raised. However, the two key features of the zebrafish are i) that all these advantages can be combined with its genetical tractability and a broad range of manipulation methods, most of them relatively easy to perform, and ii) consequences of molecular manipulations can be followed over time through in vivo microscopy approaches thus delivering data in the context of a living vertebrate organism.8–10 This review aims to present an overview over the wide and constantly expanding spectrum of genetic manipulation approaches in zebrafish.
The zebrafish genome
In the haploid state, the zebrafish genome has a size of about 1,412 Gb distributed among 25 chromosomes.11,12 Based on data obtained by the zebrafish genome project initiated by the Sanger Institute, Howe et al presented in 2013 a sequence assembly of outstanding quality, meeting the high standards of the mouse and human genome. This so-called “reference genome” uses the widespread standard fish strain Tübingen as a genetic model for all other common standard strains.11 A follow-up version of this sequence assembly was published online just recently (http://www.ncbi.nlm.nih.gov/genome/annotation_euk/Danio_rerio/104/). However, it is important to note that the genetic heterogeneity between different zebrafish laboratory strains is higher than that in other model organisms with considerable variations even within one distinct strain and that inbreeding possibilities are limited in zebrafish.13–16
About 71% of the 20,479 protein-coding genes of Homo sapiens have orthologs in the zebrafish genome. In turn, 69% of the 26,206 protein-coding zebrafish genes have human counterparts.11 The increased gene number in the zebrafish genome – as compared to the human genome – is accepted to originate mainly from a whole-genome duplication during teleost evolution that was followed by a loss of many ohnologs.17 Hence, many but not all human genes have more than one zebrafish ortholog.11 The existence of two orthologs often comes with an advantage though. Mutations in the two ohnologs have been accumulated mostly in regulative regions. While both ohnologs together show a spatio-temporal expression pattern similar to the respective human ortholog, each individual ohnolog may display a more restricted expression pattern compared to its human ortholog. As a result, early lethal phenotypes result less often from loss-of-function experiments, allowing deeper insights into the tissue-specific functions of a gene of interest. However, it is important to keep in mind that this advantage can also come with the pitfall of redundant functionality.18,19
Howe et al also described 4,556 RNA genes.11 Although these genes do not encode a protein, many of them seem to have essential functions. One important class of such RNAs is microRNAs (miRNAs), which play a key role in the regulation of protein-coding genes. Among most vertebrates, miRNA sequences and functions are largely conserved.20,21
The close genetic similarities between zebrafish and human document the great potential of zebrafish as a model system for biomedical research. This is supported by the identification of many zebrafish genes orthologous to human disease genes listed in the Online Mendelian Inheritance in Man database.11
Zebrafish as model to elucidate vertebrate gene function
In 1996, zebrafish had its major breakthrough as a model for vertebrate genetics. Presented as a stand-alone issue of the journal Development, an international team coordinated by Christiane Nüsslein-Volhard, Wolfgang Driever, Friedrich Bonhoeffer, and other pioneering researchers published the results of several large-scale forward genetic screens. By utilizing the point mutation-inducing chemical N-ethylnitrosourea (ENU), they generated zebrafish males with randomly mutagenized sperm and crossed these fish with nontreated wild-type fish. The descendants of these potential mutation carriers were screened by visual inspection with regular stereomicroscopes for specific phenotypes during embryonic development. Of 4,264 isolated mutants, 1,163 were confirmed and subsequently analyzed. Doing so led to the identification of approximately 400 genes generating visible phenotypes when mutated that could be assigned to distinct developmental or physiological processes. Most importantly, many of these genes were found to have orthologs in higher vertebrates, thus documenting the outstanding suitability of zebrafish for human developmental forward genetics. Thus, the established zebrafish mutant library covered almost all aspects of vertebrate development, including cell migration in the early embryo, notochord formation, muscle development, and development of the nervous system and the heart and circulatory system.22
The heart is the first organ formed during vertebrate embryonic development and its formation is highly conserved among species. During early development, two bilateral heart primordia are established. They migrate toward the midline where they fuse to give rise to a fully functional heart. In zebrafish, heart formation can be observed during the first two days of development with heart contraction and subsequent circulation starting approximately after 1 day of development.23,24
One striking phenotype in heart organogenesis found during the mutagenesis screens is cardia bifida or “two hearts”. In these mutants, the two heart primordia fail to fuse preventing the formation of a functional heart. Instead, “two hearts” are found, one on each side of the midline indicating a fundamental problem with cell migration. Such mutants provide fascinating insights into the molecular mechanisms underlying vertebrate organogenesis. In the case of cardia bifida, the zebrafish mutant miles apart was mapped to a gene encoding a Sphingosine-1-phosphate receptor. Signaling induced by this receptor was demonstrated to regulate the interaction of zebrafish cells with the extracellular matrix protein Fibronectin, which marks the migration path of myocardial precursors in the embryo. In congruence with this, another cardia bifida mutant named natter was mapped to the gene locus of fibronectin.25,26
Other generated mutants also allow the analysis of organ function. For example, the pickwick mutant shows reduced heart contractility with dilations and was mapped to the zebrafish ortholog encoding the large sarcomeric protein Titin. Interestingly, mutations in the human Titin gene have been identified as cause for dilated cardiomyopathy resulting in a phenotype comparable to the pickwick mutant. Therefore, the pickwick zebrafish mutant is a promising model to study dilated cardiomyopathy.24,27
The findings summarized earlier are only a small glimpse of the huge amount of data obtained by Nüsslein-Volhard, Driever, Bonhoeffer, and their colleagues. It is particularly noteworthy that mutant screening was realized without comprehensive knowledge of the zebrafish genome. Taken together, these forward genetic screens allowed successful analysis of gene function in zebrafish and readily contributed to the understanding of vertebrate development and human diseases.
Molecular tools for the genetic manipulation of zebrafish
Random gene disruption
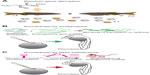
ENU is an alkylating agent that modifies single DNA bases resulting in randomly distributed point mutations (Figure 2A). Depending on the position within an open reading frame (ORF) or at splice sites, this can result in a nonsense mutation leading to protein truncation or a missense mutation changing the meaning of the affected codon. Those mutations mostly result in hypomorphic or null alleles; the generation of hypermorphic and neomorphic alleles is also known. However, silent mutations in the coding sequence or the protein are possible and small ORFs are unlikely targeted.28
ENU-derived mutagenesis of zebrafish was exploited in the past to map readily observable phenotypes to gene functions.29 Today, the almost fully annotated zebrafish genome is available for everybody by different online resources allowing the precise selection of any genomic sequence of interest (Table 1). This allows one to identify the targeted identification of ENU-induced point mutations in a selected locus by TILLING (targeting-induced local lesions in genomes). TILLING combines the use of an endonuclease specifically detecting mismatches within a DNA heteroduplex with resequencing.33
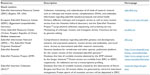  | Table 1 Resources for zebrafish researchers |
Recently, a point mutation was identified by TILLING in the zebrafish ortholog of psen1 potentially allowing one to study the function of an Alzheimer’s disease (AD) candidate gene in an animal model. PSEN1 is a highly conserved protein involved in the γ-secretase complex. γ-Secretase function is crucial for the cleavage of the amyloid precursor protein. Amyloid precursor protein miscleavage is accepted to be one possible cause for progressive dementia in AD, and psen1 mutations have been found in many patients with autosomal dominant familial AD. Various attempts were made to study psen1 function in the mouse system; however, psen1 null mice are not viable. The zebrafish psen1 allele hu2547 encodes a truncated protein variant without the proteolytic domain, and psen1hu2547/hu2547 zebrafish mutants develop almost normally and can be raised to adulthood allowing long-term analysis of impaired psen1 function in a vertebrate brain. These fish displayed significant alteration in histaminergic neuron number presumably linked to the neuronal histamine deficit observed in AD patients.34–36
The targeted identification of mutations is accelerated by combining the now available knowledge of the zebrafish genome sequence with up-to-date high-throughput sequencing. Kettleborough et al designed short biotinylated RNA probes complementary to all protein-coding sequences of zebrafish. By hybridizing these probes with genomic DNA of ENU-mutagenized zebrafish, they were able to enrich exonic sequences by pull down with streptavidin beads. Subsequently utilizing exome sequencing, their ultimate goal is to identify mutations in all protein-coding genes of zebrafish.30
ENU-based mutagenesis is highly efficient; however, the identification of a point mutation requires intensive screening by sequencing techniques. An alternative approach is the random integration of exogenous DNA. This so-called insertional mutagenesis relies on the integration mechanisms of transposable elements or retroviral vectors, which introduce a disrupting cassette encoding an easy-to-detect reporter such as a fluorescent protein (eg, a minimal promotor and an ORF encoding green fluorescent protein [GFP]) into the zebrafish genome. Successful integration can be monitored by fluorescence and by using this construct as molecular landmark, the flanking genomic regions can be easily identified allowing a reliable mapping of the hit locus. This fundamental principle is also designated as trapping.37
Utilizing a pseudotyped retrovirus for random insertion into the zebrafish genome, the research group of Nancy Hopkins created a wide range of different mutant alleles by the disruption of endogenous ORFs. Many of the identified mutants showed early developmental phenotypes thus complementing the Nüsslein-Volhard screens. However, in contrast to ENU-induced mutations, the integrated viral DNA sequence allowed fast positional cloning and identification of the disrupted genes. This trapping approach is termed gene trapping.38–41 Enhancer trapping is more directed to the identification of novel regulative elements such as enhancers and promoters. Once characterized, these stable integrates can serve as tissue- or cell type-specific markers. The most ambitious approach is protein trapping by which the ORF of an endogenous gene is fused with the ORF of the fluorescent reporter. This allows the characterization of the expression pattern of the hit gene as well as the subcellular localization of the respective protein in parallel.37,42,43
However, random mutagenesis is work and cost intensive and requires the maintenance and screening of thousands of embryos and fish. Hence, such extensive mutagenesis projects with the potential aim of genome or phenome saturation are mostly conducted by larger research teams who then kindly make the created mutant library available for the community (Table 1). However, many small-scale screens with an easy-to-score read-out and aiming for a particular phenotype have been performed in the meantime.
Morpholinos
Instead of chemical mutagenesis, another possibility in zebrafish to interfere with a gene’s function is to inject short antisense oligonucleotides named morpholinos into early embryos to specifically knockdown the expression of the gene of interest generating a hypomorphic phenotype.44 Depending on the target sequence, morpholino-binding blocks splicing or translation of a specific messenger RNA (mRNA) resulting in reduced or even lost expression of the respective gene product (Figure 2B). This reverse genetics approach is often used to analyze the function of genes without available mutant allele or to produce intermediate phenotypes of different strengths.45
Although morpholinos are commonly used in the zebrafish community, concerns were raised regarding the specificity of this technique. Robu et al showed that morpholinos can upregulate the activity of the apoptosis factor p53. Thereby, induced cell death frequently masks the actual phenotype. Co-injection of a p53-specific morpholino is accepted to compensate this off-target effect; however, uncertainties remain.46,47 For example, a recent study revealed the fundamental differences between the morpholino-induced knockdown of the serine-threonine protein kinase PAK4 and a pak4 null allele. pak4−/− zebrafish embryos and adults develop normally, whereas the knockdown shows a severe developmental phenotype.48 Nevertheless, morpholinos may still be a valuable tool to analyze gene function if the specificity is characterized well.49,50
RNA interference
In many model organisms, RNA interference (RNAi) is a standard technique for specific expression knockdown. Exogenous double-strand RNA is processed to short single-strand RNA fragments that bind to an mRNA with complementary sequence which is then degraded. All this is carried out by a highly conserved protein machinery.51 Although all components of this pathway are found in zebrafish, most attempts to establish a well-functioning RNAi approach struggled with technical problems or off-target effects. However, it was demonstrated that the processing of endogenous genetically encoded miRNAs by the zebrafish RNAi machinery is conserved to other vertebrates and essential for development.52 Therefore, the typical miRNA structure of zebrafish was recently used as template for a novel RNAi tool by replacing the miRNA stem loop by a short target sequence matching the protein-coding mRNA of interest. Expression of such constructs reduced the expression levels of the respective genes and caused phenotypes comparable to corresponding loss-of-function mutants and integration of these constructs into the genome produced stable transgenic “RNAi lines”. By the combination of modified miRNAs with a tissue-specific expression system, which is also inducible, cell type-specific gene knockdowns have been already achieved.53–55 These encouraging approaches raise the expectation that zebrafish finally has a well-functioning and adjustable RNAi tool and that this technique may see a revival in zebrafish in the future (Figure 2C).
Ectopic expression by mRNA
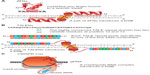
Injection of synthetic capped mRNA encoding the protein of interest into early embryo stages (eg, one cell embryos) is a standard method within the zebrafish community. Injected mRNA is distributed more or less evenly to every newly born cell of the embryo (Figure 3A). Thus, mRNA-derived overexpression although not being cell type specific is a fast and valuable tool for functional gene analysis by global expression of gain- and loss-of-function variants of a gene of interest. Because the injected mRNA is only stable during early stages, this method is limited to characterize early processes of embryogenesis. Also, mRNA injection is often used to verify a loss-of-function phenotype by rescue.44,56
Transgenesis
Instead of injecting mRNA, it is also possible to inject an expression plasmid into zebrafish embryos. Common promoters such as the CMV promoter work well in zebrafish and allow the expression of the protein of interest.57 To achieve cell type-specific expression, many well-characterized enhancer elements for zebrafish are available, which can be combined, for example with an ORF encoding a fluorescent protein. Moreover, a plasmid can carry a multicistron expression cassette allowing the parallel and even equimolar expression of several components at once. Unsurprisingly, fluorescent reporters play a major role in zebrafish research due to the transparent nature of the zebrafish embryo throughout development; therefore, a broad variety of advanced genetically encoded fluorescent microscopy tools have been developed.58 In contrast to injected mRNA, plasmid DNA distributes in a mosaic manner and hence only few cells in the multicellular embryo will express the protein of interest (Figure 3B). Therefore, plasmid injection is common for the analysis of the behavior of specially selected small cell populations or even single cells and to address the cell autonomy of gain-of-function and loss-of-function phenotypes.59 These transgenesis effects are long lasting in contrast to mRNA-mediated transgene expression, but transgene expression is rarely transmitted through the germ line.
To efficiently create stable transgenes, the respective construct needs to be integrated actively into the zebrafish genome. For this, several transposon-based tools are available for zebrafish such as Sleeping Beauty, Tol1, and Tol2 of which the latter is the most commonly used one due to its detailed characterized function and efficiency in mediating genome integrations. Minimal Tol2 cis sequences are only ~200 bp long; thus, they can be easily cloned to the flanking sites of any desired transgene to be integrated into the zebrafish genome. Co-injected with synthetic capped mRNA encoding for Tol2 transposase in the early embryo, the Tol2-flanked transgene integrates with a high chance into the germ line where its integration becomes stable due to fast Tol2 transposase-encoding mRNA and protein turnover and thus the transgene is transmitted to the progeny of founder fish (Figure 3C). However, problems such as position effects and silencing of the integrated transgene can occur as usually found with random transgene integration methods in other model organisms.60–62
Up to now, many different transgenic lines have been generated and are available from the large international stock centers (Table 1) or are shared between zebrafish labs by simply shipping eggs in a Falcon tube via regular surface mail. Especially popular are transgenic reporter lines, which carry a cell type-specific enhancer to drive the expression of a fluorescent protein.58 Thus, the development of an organ or the migration of a cell population can be analyzed in an intact living vertebrate embryo throughout development. A widely used reporter for neurobiological research is the Tg(elavl3:EGFP) line (formerly known as HuC:GFP). The elavl3 enhancer element drives expression specifically in postmitotic neuronal progenitors and expression is maintained in differentiated neurons. Hence, the development, migration and morphology of those neurons can be easily observed by fluorescence by confocal microscopy. However, this line was created before the development of the Tol2 system by just injecting a linearized plasmid construct. Random integration of such constructs in the genome is possible, but happens with rather low efficiency.63
To make the use of cell type-specific enhancer elements and driven transgenes more flexible, combinatorial genetics have been introduced into the zebrafish system. One approach is the targeted expression of an ORF driven by a so-called upstream activating sequence (UAS) upon the specific binding of the transcription activator factor Gal4.64–66 In turn, the expression of the Gal4 protein is driven by a cell type-specific enhancer such as elavl3. By the combination of an enhancer:Gal4 transgene with an UAS:ORF of choice transgene by the intercross of respective stable transgenic fish lines, one achieves the specific spatial and temporal expression of the protein of interest. Due to the combinatorial nature of this system, the number of potential matings rises exponentially with the number of generated driver and effector lines.58 This can be used to identify distinct cell populations by the targeted expression of a fluorescent protein and also to genetically model human diseases in zebrafish in particular with early lethal phenotypes, because driver and effector fish can be propagated separately. For example, the neuronal elavl3 enhancer was used to specifically drive the expression of a human pathogenic variant of the protein Tau by Gal4/UAS-mediated transcription activation. Tau is a microtubule-associated protein known to be a risk factor for AD, and stable transgenic zebrafish expressing the human pathogen showed AD-related phenotypes such as delayed neuronal differentiation, behavioral abnormalities and AD-like histopathology.67
Gal4-activated UAS elements can drive the expression of several ORFs to produce the respective gene products in parallel58 and also the generation of triple and quadruple transgenic fish is possible.68 The Gal4/UAS system is commonly used and new Gal4 tools are constantly being further developed as for example tamoxifen-inducible Gal4 expression;69 however, alternative strategies exist such as the chemical-inducible LexPR system or tetracycline-controlled transcriptional activation.70,71
Genome editing
Currently, the zebrafish field witnesses another grant splash in the development of fast and efficient targeted mutagenesis approaches. Genome editing utilizes specifically designed endonucleases for sequence-specific manipulation. The first-generation technique was based on zinc finger nucleases (ZFNs). ZFNs are composed of a FokI endonuclease domain and a DNA-binding domain consisting of zinc fingers, small protein motifs binding each to a specific DNA triplet. Zinc fingers with different binding properties can be combined to form a sequence-specific domain targeting the genomic sequence of interest. Two ZFNs bind to neighboring target sites, thus constituting a functional FokI dimer, which subsequently introduces a double-strand break (DSB) between the two ZFN-binding sites (Figure 4A). DSBs can be repaired by nonhomologous end joining (NHEJ); however, this mechanism is prone to error resulting in small deletions or insertions. Hence, a ZFN-induced DSB can lead to a disruption of gene function. DSB repair by homologous recombination (HR) and thereby site-directed integration of foreign sequences is also possible adding another even more precise method of mutagenesis.72
Developed over the last decade, ZFNs were demonstrated to work in zebrafish and many other model organisms. However, zinc fingers can influence each other’s binding properties making the specificity of a zinc finger array not entirely predictable.73 Verification of binding properties takes time and can be very expensive, which prevented the establishment of genome editing by ZFNs as a standard method in the past. Recently, this situation has changed dramatically. During the last 4 years, two novel related techniques took the scientific community by storm: transcription activator-like effector nucleases (TALENs) and clustered regularly interspaced short palindromic repeats (CRISPR)-Cas.
Similar to ZFNs, TALENs consist of an endonuclease domain of FokI and a site-specific DNA-binding domain. The DNA-binding domain is composed of short repeats whose sequence is highly conserved except two adjacent residues. These repeat variable diresidues determine the binding specificity of a single repeat to one of the four DNA bases with repeat variable diresidues not influencing each other in their binding properties. Thus, a target DNA sequence can simply be translated into a sequence of linear repeats with different repeat variable diresidues (Figure 4B).74,75 Today, a TALEN targeting the DNA sequence of interest can be generated in only a few steps by standard cloning procedures making this technique affordable and manageable to everyone.76–78
Zebrafish was the first vertebrate in which TALENs have been tested. Indeed, genomic DSBs induced by a pair of TALENs followed by incorrect NHEJ resulted in heritable mutations.79 HR-directed repair of a DSB by the integration of a donor construct with homologous region was also already successful.80
The latest tool for targeted genome editing is of remarkable simple elegance: CRISPR-Cas does not even require the design and synthesis of a DNA targeting domain – the method simply relies on classic RNA injection protocols in zebrafish. A short guide RNA complementary to the genomic target sequence of interest is co-delivered with synthetic capped mRNA encoding Cas9 endonuclease. The guide RNA mediates Cas9 endonuclease binding to the target site followed by the generation of a DSB (Figure 4C).81 Up to now, CRISPR-Cas proved to be just as efficient as TALENs and allows gene disruption by incorrect NHEJ or by targeted DNA integration.82,83 Respective vectors optimized for zebrafish or other organisms are already available requiring only the individual design of the guide RNA.84,85
In conclusion, targeted genome editing led to a dramatic change in the zebrafish community. Instead of searching the databases for mutants generated by random gene disruption or arguing about the specificity of morpholinos, one can generate a stable mutant allele in line with their own preferences within weeks. Up to now, there are many possibilities: DSBs directed to the start of an ORF followed by incorrect NHEJ can lead to a frameshift or a premature stop codon resulting in a genuine null allele. Directed to another region of a gene, truncated or alternative splice variants of the transcript can be generated. HR-based repair of a DSB can be exploited to introduce a wide variety of DNA sequences, as, for example, a simple gene-disrupting cassette or a more complex fluorescence reporter, which would be later driven by the promoter of the targeted gene. By using the sequence of the targeted locus as HR template for a DSB, one could introduce before built-in point mutations or small protein tags.37,83,86–90 It is equally conceivable to use the wild-type sequence for repairing an existing mutation to perform a rescue experiment.
The parallel targeting of two TALEN pairs or two guide RNAs plus Cas9 endonuclease, respectively, to the same chromosome was demonstrated to cause very large deletions or even chromosomal inversions in zebrafish, which might be useful to disrupt whole gene clusters.91 In addition, the combination of Cas9 endonuclease with several guide RNAs matching different loci can be used to generate multiple specific mutations within one approach.84 As in some of these animals, both alleles are affected, a first insight into the phenotype to expect can already be observed a few days after injection. Other potential applications are the targeted manipulation of gene expression by using the TALEN repeat domain to guide effector domains to a promoter region or inactive yet binding Cas9 endonuclease to block transcription at specific genomic sites.92,93
However, all novel techniques have their drawbacks. In the case of genome editing, these are off-target effects leading to undesired background mutations. Thus, the target sequences must be chosen carefully: to reduce off-target effects to the minimum, they must be as unique as possible in the whole genome. A further possibility at least for the CRISPR-Cas system is the use optimized Cas9 enzymes. One example is the Cas9 nickase, which does not introduce a DSB but only a single-strand break and seems to reduce off-target effects dramatically.94
Taken together, TALENs and CRISPR-Cas take reverse genetics of zebrafish and other genetically tractable organisms to an entirely new level as they enable targeted mutagenesis by inducing small mutations and even in frame knock-in to any chromosomal locus of choice. While such an approach was previously available only for mouse, it now offers these same genetic possibilities for zebrafish in an easy, cheap, and fast manner.
Zebrafish as a tool for drug development
The diverse methods for genetic manipulation of zebrafish make it possible to study a wide variety of biomedical research issues in a living vertebrate. Such research projects could contribute to understanding cellular and molecular mechanisms underlying genetic human diseases. But how could zebrafish be used for the next step of developing therapeutic strategies for disease intervention? One possibility is using zebrafish as a screening or validation tool for new drugs.
During the first days of development, zebrafish embryos can be kept in multiwell plates without the need for feeding them thus allowing the analysis of a great number of embryos in parallel. Automated systems can support manual effort, eg, by pipetting, sorting, and even searching for phenotypes.95 Therefore, a whole compound library can be tested fast and efficiently directly in the context of a living vertebrate.96
Zebrafish chemical screening can be used to assay developmental toxicity of different substances by phenotype and lethal dosing.97 Also the identification of specific modulators of developmental pathways is feasible. For example, dorsomorphin was identified in an in vivo screen searching for antagonists of the bone morphogenetic protein signaling pathway, which is essential for dorsoventral patterning during early embryonic development. When applied during early development, Dorsomorphin was shown to specifically inhibit bone morphogenetic protein type I receptors resulting in a dorsalized phenotype with reduced ventral tissues.98 To identify modulators of the development of distinct organs and tissues, fluorescence reporter strains can be used for quantitative analysis in microplate readers.99 Ultimately, compounds can be screened for their efficacy to alter or even rescue a disease phenotype. gridlock is a zebrafish mutant with a constricted aorta preventing proper blood flow. This phenotype resembles aortic coarctation in humans, a congenital heart disease. Chemical screening identified two compounds having the potential to suppress the gridlock phenotype by stimulating pathway components downstream of the gridlock gene product.100
Taken together, chemical screening utilizing zebrafish is a conclusive method to screen libraries of drug candidates for toxicity, bioavailability, and dosing or to identify compounds with specific metabolic functions. However, zebrafish does not substitute for either cell-based assays or drug testing on mammals such as mice. But by providing a large-scale platform for compound screening as “hinge” between compound screening using cultured cells and the costly and laborious drug validation on mammalian disease models, zebrafish drug screening can significantly help to narrow down potentially active substances with low toxicity and good tissue availability and is expected to speed up drug discovery significantly.
One successful identified substance with high biomedical relevance is prostaglandin E2 (PGE2). North et al101 showed in a chemical screen that PGE2-regulating compounds affected the number of hematopoietic stem cells in zebrafish and that a stable PGE2 derivative improved the repopulation of murine bone marrow with hematopoietic stem cells. Thus, PGE2 is a high potential candidate for hematopoietic stem cell therapies of human patients; as a consequence, a PGE2-derived drug was tested successfully in a clinical Phase I trial proving that the drug was not harmful when applied to healthy individuals. Phase II is already underway to test efficacy.102,103 This encouraging example raises the hope that biomedical zebrafish research may be directly transferred to humans.
Conclusion and future perspectives
The powerful combination of established molecular tools like cell type-specific expression and exiting new technologies such as genome editing provides almost limitless perspectives for the use of zebrafish as a biomedical model. Equipped with the comprehensive knowledge of the zebrafish genome, researchers have just begun to understand the full significance of the close relation between fish and human. By taking the full advantage of the molecular toolbox for genetic manipulation, significant progress has been already made; however, there is even more to expect for the near future when zebrafish genetics will have its full impact on next-generation sequencing or system biology.104,105
Acknowledgments
WAS and RWK thank all members of the Köster group for helpful discussions during the process of writing and Tobias Kruse for valuable comments on the manuscript. WAS is funded by a grant of the Deutsche Forschungsgemeinschaft (KO 1949/5-1). RWK acknowledges funding by the Federal State of Lower Saxony, Niedersächsisches Vorab (VWZN2889).
Disclosure
The authors declare that they have no conflicts of interest.
References
Streisinger G, Walker C, Dower N, Knauber D, Singer F. Production of clones of homozygous diploid zebra fish (Brachydanio rerio). Nature. 1981;291(5813):293–296. | |
Nawrocki L, BreMiller R, Streisinger G, Kaplan M. Larval and adult visual pigments of the zebrafish, Brachydanio rerio. Vision Res. 1985;25(11):1569–1576. | |
Nguyen CT, Lu Q, Wang Y, Chen J-N. Zebrafish as a model for cardiovascular development and disease. Drug Discov Today Dis Models. 2008;5(3):135–140. | |
Lam SH, Gong Z. Modeling liver cancer using zebrafish: a comparative oncogenomics approach. Cell Cycle. 2006;5(6):573–577. | |
Jang ZH, Chung HC, Ahn YG, et al. Metabolic profiling of an alcoholic fatty liver in zebrafish (Danio rerio). Mol Biosyst. 2012;8(7):2001–2009. | |
Kimmel CB, Ballard WW, Kimmel SR, Ullmann B, Schilling TF. Stages of embryonic development of the zebrafish. Dev Dyn. 1995;203(3):253–310. | |
Adatto I, Lawrence C, Thompson M, Zon LI. A new system for the rapid collection of large numbers of developmentally staged zebrafish embryos. PLoS One. 2011;6(6):e21715. | |
Cha YR, Weinstein BM. Visualization and experimental analysis of blood vessel formation using transgenic zebrafish. Birth Defects Res C Embryo Today. 2007;81(4):286–296. | |
Rieger S, Wang F, Sagasti A. Time-lapse imaging of neural development: zebrafish lead the way into the fourth dimension. Genesis. 2011;49(7):534–545. | |
Hocking JC, Distel M, Köster RW. Studying cellular and subcellular dynamics in the developing zebrafish nervous system. Exp Neurol. 2013;242:1–10. | |
Howe K, Clark MD, Torroja CF, et al. The zebrafish reference genome sequence and its relationship to the human genome. Nature. 2013;496(7446):498–503. | |
Daga RR, Thode G, Amores A. Chromosome complement, C-banding, Ag-NOR and replication banding in the zebrafish Danio rerio. Chromosome Res. 1996;4(1):29–32. | |
Guryev V, Koudijs MJ, Berezikov E, et al. Genetic variation in the zebrafish. Genome Res. 2006;16(4):491–497. | |
Shinya M, Sakai N. Generation of highly homogeneous strains of zebrafish through full sib-pair mating. G3 (Bethesda). 2011;1(5):377–386. | |
Patowary A, Purkanti R, Singh M, et al. A sequence-based variation map of zebrafish. Zebrafish. 2013;10(1):15–20. | |
LaFave MC, Varshney GK, Vemulapalli M, Mullikin JC, Burgess SM. A defined zebrafish line for high-throughput genetics and genomics: NHGRI-1. Genetics. 2014;198(1):167–170. | |
Taylor JS, Braasch I, Frickey T, Meyer A, Van de Peer Y. Genome duplication, a trait shared by 22000 species of ray-finned fish. Genome Res. 2003;13(3):382–390. | |
Chiang C, Litingtung Y, Lee E, et al. Cyclopia and defective axial patterning in mice lacking Sonic hedgehog gene function. Nature. 1996;383(6599):407–413. | |
Schauerte HE, van Eeden FJ, Fricke C, Odenthal J, Strähle U, Haffter P. Sonic hedgehog is not required for the induction of medial floor plate cells in the zebrafish. Development. 1998;125(15):2983–2993. | |
Heimberg AM, Sempere LF, Moy VN, Donoghue PCJ, Peterson KJ. MicroRNAs and the advent of vertebrate morphological complexity. Proc Natl Acad Sci U S A. 2008;105(8):2946–2950. | |
Nachtigall P, Dias M, Pinhal D. Evolution and genomic organization of muscle microRNAs in fish genomes. BMC Evol Biol. 2014;14(1):196. | |
Nüsslein-Volhard C. The zebrafish issue of development. Development. 2012;139(22):4099–4103. | |
Rohr S, Otten C, Abdelilah-Seyfried S. Asymmetric involution of the myocardial field drives heart tube formation in zebrafish. Circ Res. 2008;102(2):e12–e19. | |
Bakkers J. Zebrafish as a model to study cardiac development and human cardiac disease. Cardiovasc Res. 2011;91(2):279–288. | |
Chen JN, Haffter P, Odenthal J, et al. Mutations affecting the cardiovascular system and other internal organs in zebrafish. Development. 1996;123:293–302. | |
Kawahara A. Genetic dissection of cardiac progenitor migration. Inflamm Regen. 2009;29(5):324–328. | |
Stainier DY, Fouquet B, Chen JN, et al. Mutations affecting the formation and function of the cardiovascular system in the zebrafish embryo. Development. 1996;123:285–292. | |
Coghill EL, Hugill A, Parkinson N, et al. A gene-driven approach to the identification of ENU mutants in the mouse. Nat Genet. 2002;30(3):255–256. | |
Driever W, Solnica-Krezel L, Schier AF, et al. A genetic screen for mutations affecting embryogenesis in zebrafish. Development. 1996;123:37–46. | |
Kettleborough RN, Busch-Nentwich EM, Harvey SA, et al. A systematic genome-wide analysis of zebrafish protein-coding gene function. Nature. 2013;496(7446):494–497. | |
Westerfield M, Doerry E, Kirkpatrick AE, Douglas SA. Zebrafish informatics and the ZFIN database. Methods Cell Biol. 1999;60:339–355. | |
Varshney GK, Huang H, Zhang S, et al. The Zebrafish Insertion Collection (ZInC): a web based, searchable collection of zebrafish mutations generated by DNA insertion. Nucleic Acids Res. 2013;41:D861–D864. | |
Moens CB, Donn TM, Wolf-Saxon ER, Ma TP. Reverse genetics in zebrafish by TILLING. Brief Funct Genomic Proteomic. 2008;7(6):454–459. | |
Shen J, Bronson RT, Chen DF, Xia W, Selkoe DJ, Tonegawa S. Skeletal and CNS defects in Presenilin-1-deficient mice. Cell. 1997;89(4):629–639. | |
Sundvik M, Chen Y-C, Panula P. Presenilin1 regulates histamine neuron development and behavior in zebrafish, Danio rerio. J Neurosci. 2013;33(4):1589–1597. | |
Bateman RJ, Aisen PS, De Strooper B, et al. Autosomal-dominant Alzheimer’s disease: a review and proposal for the prevention of Alzheimer’s disease. Alzheimers Res Ther. 2011;3(1):1. | |
Trinh LA, Fraser SE. Enhancer and gene traps for molecular imaging and genetic analysis in zebrafish. Dev Growth Differ. 2013;55(4):434–445. | |
Amsterdam A, Burgess S, Golling G, et al. A large-scale insertional mutagenesis screen in zebrafish. Genes Dev. 1999;13(20):2713–2724. | |
Burgess S, Hopkins N. Use of pseudotyped retroviruses in zebrafish as genetic tags. Methods Enzymol. 2000;327:145–161. | |
Amsterdam A, Nissen RM, Sun Z, Swindell EC, Farrington S, Hopkins N. Identification of 315 genes essential for early zebrafish development. Proc Natl Acad Sci U S A. 2004;101(35):12792–12797. | |
Varshney GK, Lu J, Gildea DE, et al. A large-scale zebrafish gene knockout resource for the genome-wide study of gene function. Genome Res. 2013;23(4):727–735. | |
Amsterdam A, Varshney GK, Burgess SM. Retroviral-mediated insertional mutagenesis in zebrafish. Methods Cell Biol. 2011;104:59–82. | |
Varshney GK, Burgess SM. Mutagenesis and phenotyping resources in zebrafish for studying development and human disease. Brief Funct Genomics. 2014;13(2):82–94. | |
Rosen JN, Sweeney MF, Mably JD. Microinjection of zebrafish embryos to analyze gene function. J Vis Exp. 2009;(25):ii:1115. | |
Nasevicius A, Ekker SC. Effective targeted gene “knockdown” in zebrafish. Nat Genet. 2000;26(2):216–220. | |
Robu ME, Larson JD, Nasevicius A, et al. p53 activation by knockdown technologies. PLoS Genet. 2007;3(5):e78. | |
Gerety SS, Wilkinson DG. Morpholino artifacts provide pitfalls and reveal a novel role for pro-apoptotic genes in hindbrain boundary development. Dev Biol. 2011;350(2):279–289. | |
Law SHW, Sargent TD. The serine-threonine protein kinase PAK4 is dispensable in zebrafish: identification of a morpholino-generated pseudophenotype. PLoS One. 2014;9(6):e100268. | |
Eisen JS, Smith JC. Controlling morpholino experiments: don’t stop making antisense. Development. 2008;135(10):1735–1743. | |
Bill BR, Petzold AM, Clark KJ, Schimmenti LA, Ekker SC. A primer for morpholino use in zebrafish. Zebrafish. 2009;6(1):69–77. | |
Perrimon N, Ni J-Q, Perkins L. In vivo RNAi: today and tomorrow. Cold Spring Harb Perspect Biol. 2010;2(8):a003640. | |
Kelly A, Hurlstone AF. The use of RNAi technologies for gene knockdown in zebrafish. Brief Funct Genomics. 2011;10(4):189–196. | |
Dong M, Fu YF, Du TT, et al. Heritable and lineage-specific gene knockdown in zebrafish embryo. PLoS One. 2009;4(7):e6125. | |
De Rienzo G, Gutzman JH, Sive H. Efficient shRNA-mediated inhibition of gene expression in zebrafish. Zebrafish. 2012;9(3):97–107. | |
Dong Z, Peng J, Guo S. Stable gene silencing in zebrafish with spatiotemporally targetable RNA interference. Genetics. 2013;193(4):1065–1071. | |
Finckbeiner S, Ko PJ, Carrington B, et al. Transient knockdown and overexpression reveal a developmental role for the zebrafish enosf1b gene. Cell Biosci. 2011;1:32. | |
Rupp RA, Snider L, Weintraub H. Xenopus embryos regulate the nuclear localization of XMyoD. Genes Dev. 1994;8(11):1311–1323. | |
Weber T, Köster R. Genetic tools for multicolor imaging in zebrafish larvae. Methods. 2013;62(3):279–291. | |
Mumm JS, Williams PR, Godinho L, et al. In vivo imaging reveals dendritic targeting of laminated afferents by zebrafish retinal ganglion cells. Neuron. 2006;52(4):609–621. | |
Ni J, Clark KJ, Fahrenkrug SC, Ekker SC. Transposon tools hopping in vertebrates. Brief Funct Genomic Proteomic. 2008;7(6):444–453. | |
Kawakami K. Transposon tools and methods in zebrafish. Dev Dyn. 2005;234(2):244–254. | |
Balciunas D, Wangensteen KJ, Wilber A, et al. Harnessing a high cargo-capacity transposon for genetic applications in vertebrates. PLoS Genet. 2006;2(11):e169. | |
Park HC, Kim CH, Bae YK, et al. Analysis of upstream elements in the HuC promoter leads to the establishment of transgenic zebrafish with fluorescent neurons. Dev Biol. 2000;227(2):279–293. | |
Brand AH, Perrimon N. Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development. 1993;118(2):401–415. | |
Scheer N, Campos-Ortega JA. Use of the Gal4-UAS technique for targeted gene expression in the zebrafish. Mech Dev. 1999;80(2):153–158. | |
Scheer N, Riedl I, Warren JT, Kuwada JY, Campos-Ortega JA. A quantitative analysis of the kinetics of Gal4 activator and effector gene expression in the zebrafish. Mech Dev. 2002;112(1–2):9–14. | |
Paquet D, Bhat R, Sydow A, et al. A zebrafish model of tauopathy allows in vivo imaging of neuronal cell death and drug evaluation. J Clin Invest. 2009;119(5):1382–1395. | |
Almeida AD, Boije H, Chow RW, et al. Spectrum of Fates: a new approach to the study of the developing zebrafish retina. Development. 2014;141(9):1971–1980. | |
Gerety SS, Breau MA, Sasai N, Xu Q, Briscoe J, Wilkinson DG. An inducible transgene expression system for zebrafish and chick. Development. 2013;140(10):2235–2243. | |
Emelyanov A, Parinov S. Mifepristone-inducible LexPR system to drive and control gene expression in transgenic zebrafish. Dev Biol. 2008;320(1):113–121. | |
Knopf F, Schnabel K, Haase C, Pfeifer K, Anastassiadis K, Weidinger G. Dually inducible TetON systems for tissue-specific conditional gene expression in zebrafish. Proc Natl Acad Sci U S A. 2010;107(46):19933–19938. | |
Urnov FD, Rebar EJ, Holmes MC, Zhang HS, Gregory PD. Genome editing with engineered zinc finger nucleases. Nat Rev Genet. 2010;11(9):636–646. | |
Ramirez CL, Foley JE, Wright DA, et al. Unexpected failure rates for modular assembly of engineered zinc fingers. Nat Methods. 2008;5(5):374–375. | |
Moscou MJ, Bogdanove AJ. A simple cipher governs DNA recognition by TAL effectors. Science. 2009;326(5959):1501. | |
Christian M, Cermak T, Doyle EL, et al. Targeting DNA double-strand breaks with TAL effector nucleases. Genetics. 2010;186(2):757–761. | |
Cermak T, Doyle EL, Christian M, et al. Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting. Nucleic Acids Res. 2011;39(12):e82. | |
Ding Q, Lee YK, Schaefer EA, et al. A TALEN genome-editing system for generating human stem cell-based disease models. Cell Stem Cell. 2013;12(2):238–251. | |
Ma AC, Lee HB, Clark KJ, Ekker SC. High efficiency In Vivo genome engineering with a simplified 15-RVD GoldyTALEN design. PLoS One. 2013;8(5):e65259. | |
Huang P, Xiao A, Zhou M, Zhu Z, Lin S, Zhang B. Heritable gene targeting in zebrafish using customized TALENs. Nat Biotechnol. 2011;29(8):699–700. | |
Zu Y, Tong X, Wang Z, et al. TALEN-mediated precise genome modification by homologous recombination in zebrafish. Nat Methods. 2013;10(4):329–331. | |
Cong L, Ran FA, Cox D, et al. Multiplex genome engineering using CRISPR/Cas systems. Science. 2013;339(6121):819–823. | |
Hwang WY, Fu Y, Reyon D, et al. Efficient genome editing in zebrafish using a CRISPR-Cas system. Nat Biotechnol. 2013;31(3):227–229. | |
Hruscha A, Krawitz P, Rechenberg A, et al. Efficient CRISPR/Cas9 genome editing with low off-target effects in zebrafish. Development. 2013;140(24):4982–4987. | |
Jao L-E, Wente SR, Chen W. Efficient multiplex biallelic zebrafish genome editing using a CRISPR nuclease system. Proc Natl Acad Sci U S A. 2013;110(34):13904–13909. | |
Shen B, Zhang J, Wu H, et al. Generation of gene-modified mice via Cas9/RNA-mediated gene targeting. Cell Res. 2013;23(5):720–723. | |
Auer TO, Del Bene F. CRISPR/Cas9 and TALEN-mediated knock-in approaches in zebrafish. Methods. 2014;69(2):142–150. | |
Irion U, Krauss J, Nüsslein-Volhard C. Precise and efficient genome editing in zebrafish using the CRISPR/Cas9 system. Development. 2014;141(24):4827–4830. | |
Kimura Y, Hisano Y, Kawahara A, Higashijima S. Efficient generation of knock-in transgenic zebrafish carrying reporter/driver genes by CRISPR/Cas9-mediated genome engineering. Sci Rep. 2014;4:6545. | |
Campbell JM, Hartjes KA, Nelson TJ, Xu X, Ekker SC. New and TALENted genome engineering toolbox. Circ Res. 2013;113(5):571–587. | |
Blackburn PR, Campbell JM, Clark KJ, Ekker SC. The CRISPR system – keeping zebrafish gene targeting fresh. Zebrafish. 2013;10(1):116–118. | |
Xiao A, Wang Z, Hu Y, et al. Chromosomal deletions and inversions mediated by TALENs and CRISPR/Cas in zebrafish. Nucleic Acids Res. 2013;41(14):e141. | |
Zhang F, Cong L, Lodato S, Kosuri S, Church GM, Arlotta P. Efficient construction of sequence-specific TAL effectors for modulating mammalian transcription. Nat Biotechnol. 2011;29(2):149–153. | |
Qi LS, Larson MH, Gilbert LA, et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell. 2013;152(5):1173–1183. | |
Cho SW, Kim S, Kim Y, et al. Analysis of off-target effects of CRISPR/Cas-derived RNA-guided endonucleases and nickases. Genome Res. 2014;24(1):132–141. | |
Vogt A, Cholewinski A, Shen X, et al. Automated image-based phenotypic analysis in zebrafish embryos. Dev Dyn. 2009;238(3):656–663. | |
Yeh J-RJ, Munson KM, Elagib KE, Goldfarb AN, Sweetser DA, Peterson RT. Discovering chemical modifiers of oncogene-regulated hematopoietic differentiation. Nat Chem Biol. 2009;5(4):236–243. | |
Selderslaghs IWT, Van Rompay AR, De Coen W, Witters HE. Development of a screening assay to identify teratogenic and embryotoxic chemicals using the zebrafish embryo. Reprod Toxicol. 2009;28(3):308–320. | |
Yu PB, Hong CC, Sachidanandan C, et al. Dorsomorphin inhibits BMP signals required for embryogenesis and iron metabolism. Nat Chem Biol. 2008;4(1):33–41. | |
Walker SL, Ariga J, Mathias JR, et al. Automated reporter quantification in vivo: high-throughput screening method for reporter-based assays in zebrafish. PLoS One. 2012;7(1):e29916. | |
Peterson RT, Shaw SY, Peterson TA, et al. Chemical suppression of a genetic mutation in a zebrafish model of aortic coarctation. Nat Biotechnol. 2004;22(5):595–599. | |
North TE, Goessling W, Walkley CR, et al. Prostaglandin E2 regulates vertebrate haematopoietic stem cell homeostasis. Nature. 2007;447(7147):1007–1011. | |
Stachura DL, Svoboda O, Campbell CA, et al. The zebrafish granulocyte colony-stimulating factors (Gcsfs): 2 paralogous cytokines and their roles in hematopoietic development and maintenance. Blood. 2013;122(24):3918–3928. | |
Ablain J, Zon LI. Of fish and men: using zebrafish to fight human diseases. Trends Cell Biol. 2013;23(12):584–586. | |
Deo RC, MacRea CA. The zebrafish: scalable in vivo modeling for systems biology. Wiley Interdiscip Rev Syst Biol Med. 2011;3(3):335–346. | |
Henke K, Bowen ME, Harris MP. Perspectives for identification of mutations in the zebrafish: making use of next-generation sequencing technologies for forward genetic approaches. Methods. 2013;62(3):185–196. |
 © 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms
and incorporate the Creative Commons Attribution
- Non Commercial (unported, 3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms
and incorporate the Creative Commons Attribution
- Non Commercial (unported, 3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.